1. Introduction
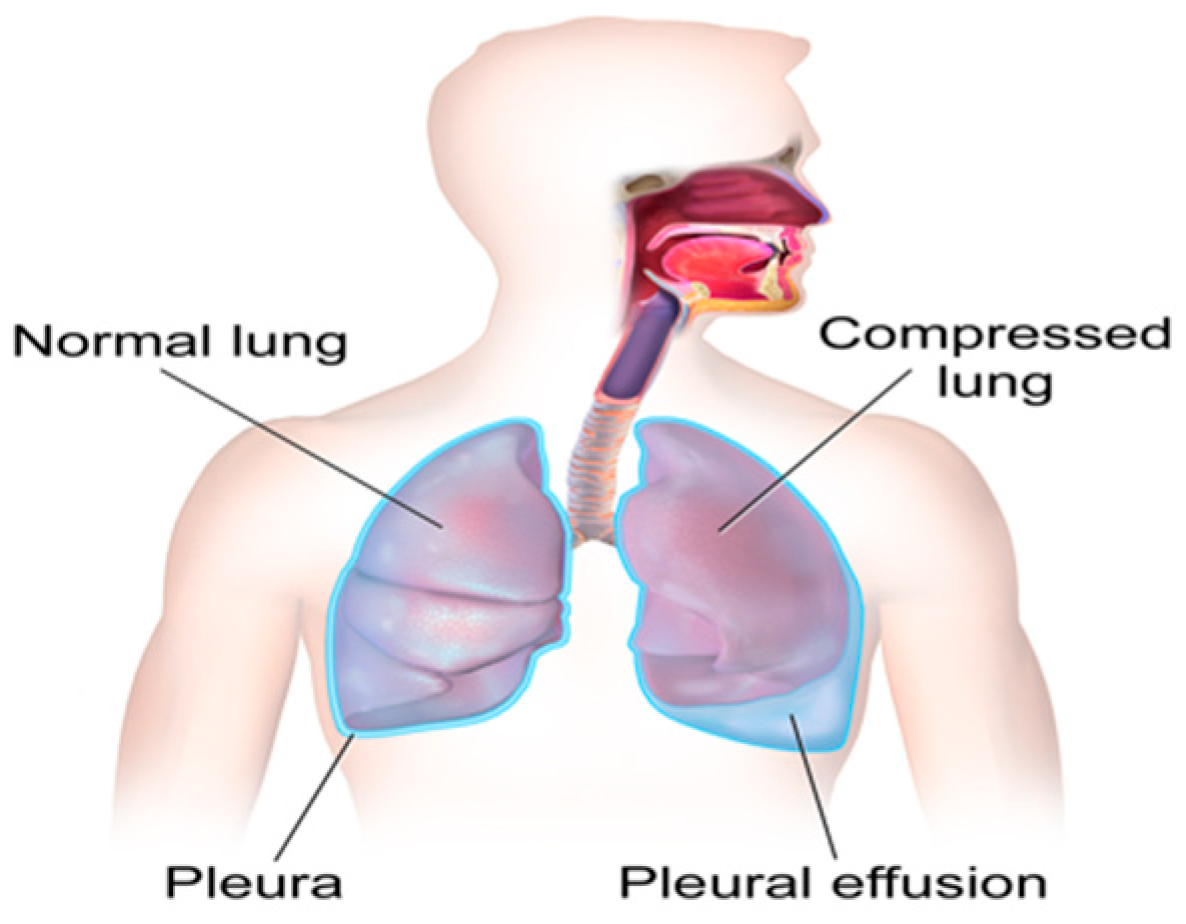
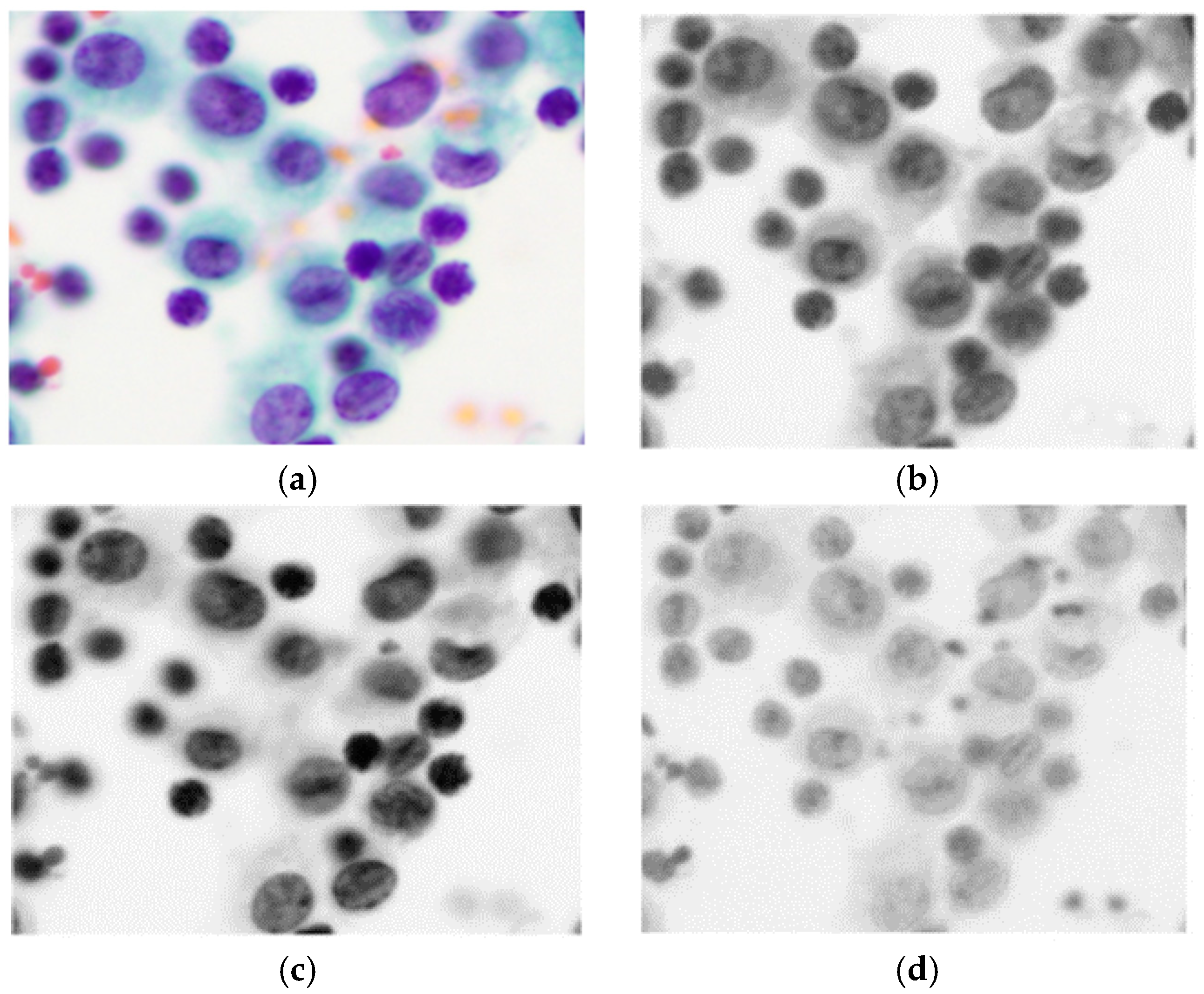
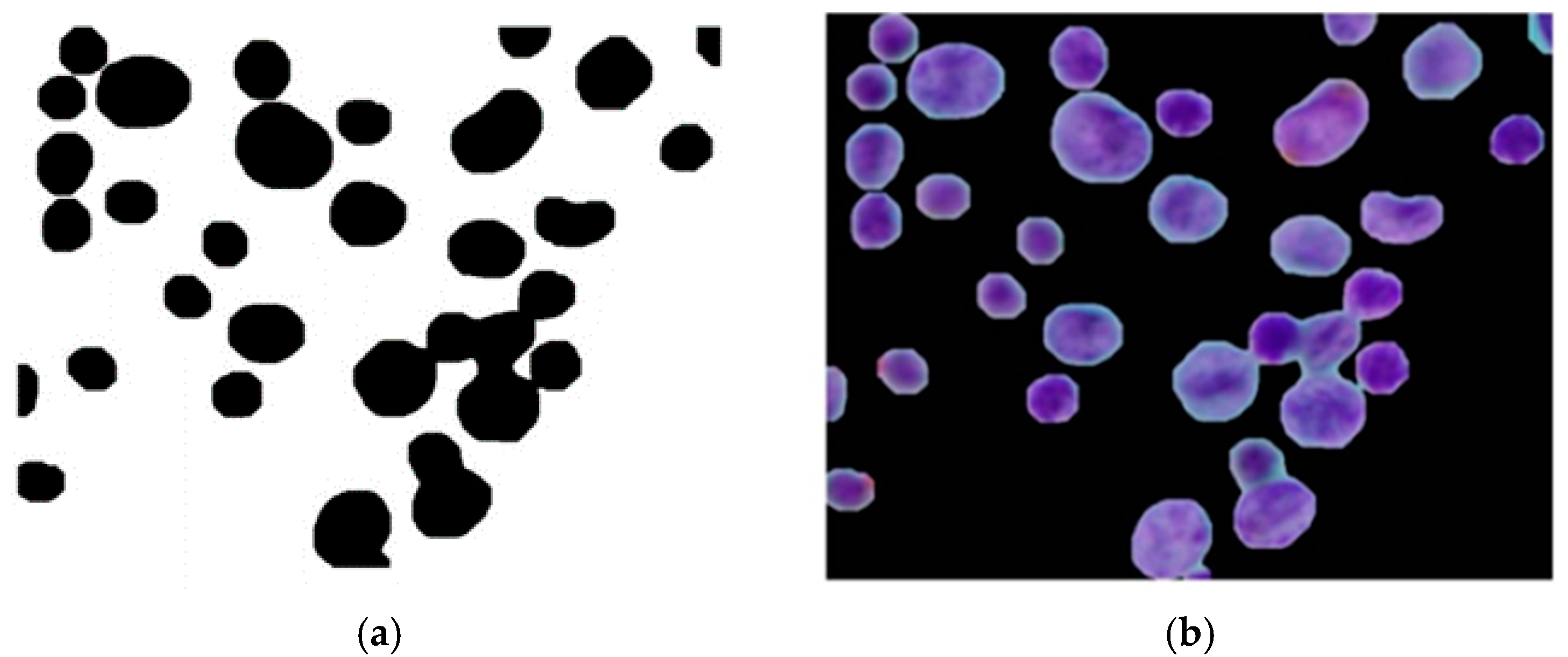
Cancer is a class of diseases characterized by malignant cells, and malignant pleural effusion (MPE) is the excessive accumulation of cancerous effusion in the pleura, as shown in
Figure 1 [
1]. MPE is one of the most aggressive cancerous effusions and a sign of an advanced stage of cancer. It is a common problem for cancer patients, and around half of cancer patients end up developing MPE. MPE can be caused by metastatic cancers or primary cancers (mesothelioma). MPE often implies an advanced stage of cancer and confers a poor prognosis [
2,
3]. Thus, fast and accurate diagnosis and prognosis of cancer cells in pleural effusion is a first priority task required so that cytologists can arrange effective treatment plans.
Cytology examination is currently considered the gold standard for diagnosing cancerous cells in pleural effusion. Cytologists take a small amount of effusion then fix and stain it on a glass slide using certain staining methods. They then visually examine the cytology slides under a microscope in order to diagnose for abnormality in every single cell [
4,
5]. However, classical cytological diagnosis is laborious, tedious, and unreasonably time-consuming. It is also prone to different diagnosis results depending on the observer. Recently, there has been growing interest in automated cell analysis systems which can serve as assistance tools to help cytologists during cytology examinations. They can provide fast, accurate and objective diagnostic results for cell analysis [
6,
7].
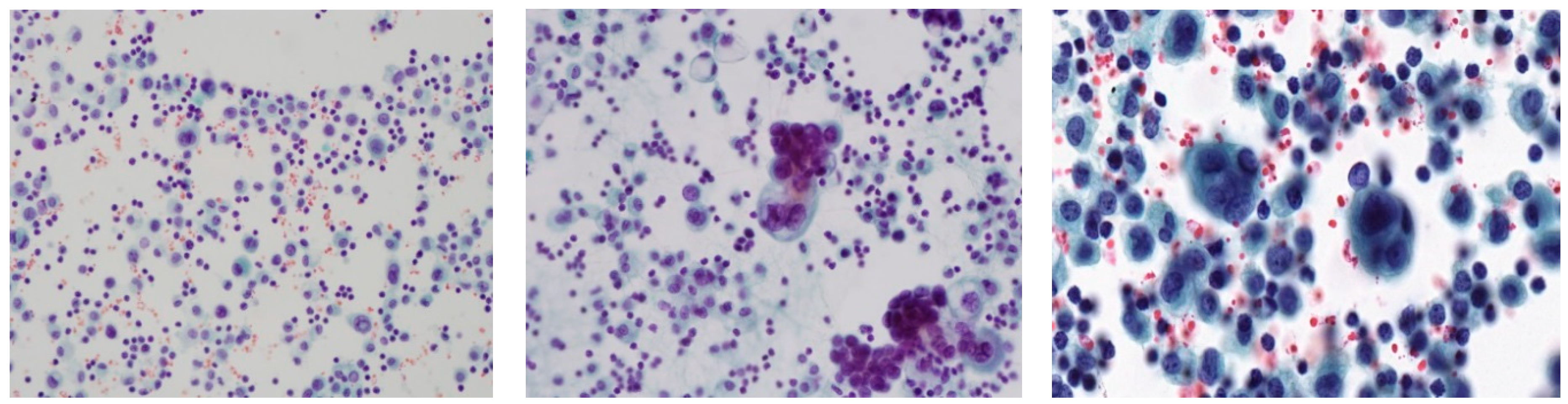
One of the difficulties found while developing such systems is the presence of overlapping nuclei. Nuclei morphology, e.g., size, shape, and density are the most important features used by cytologists in predicting cell malignancy. For instance, the excessive enlargement of nuclei and their irregular shapes are highly suggestive of malignancy. Accurate delineation of each cell contour is essential for the quantitative analysis of cell morphology. In practice, there is a great deal of overlapping nuclei occurrence in cytological pleural effusion (CPE) images. Although human experts find little difficulty in differentiating between single and overlapping nuclei, it is still a challenging task for automatic systems. Overlapping nuclei in CPE images often appear as dark purple regions, and there is a high degree of resemblance among the nuclei forming the overlapped or clustered regions. Thus, an automatic system may wrongly interpret overlapping nuclei as single nuclei. It is difficult to retrieve and quantitatively analyze features such as nucleus morphology and density if cells are touching, overlapping or clustered. Furthermore, the excessive enlargement of size and irregular shapes of overlapping nuclei regions may lead automatic systems to misclassify them as malignant cells. Thus, overlapping nuclei should be detected and distinguished from single ones prior to nuclei feature learning.
Many researchers have reported several methods for delineating the interregional contours of overlapping cell nuclei or for splitting the overlapping nuclei into individual ones. Watershed methods [
8,
9] and concavity analysis based methods [
10,
11] are the most widely used overlapping-nuclei splitting methods in microscopy image analysis. Recently, Kumar et al. reported a rule-based clump isolation method for separating overlapping nuclei [
12]. In another study, Wang et al. presented a bottleneck rule method for isolating overlapping cells [
13]. These previous studies indicate that there has been a tremendous interest in accurately delineating individual cell nuclei in cell image analysis.
Nevertheless, it is crucial to accurately determine the presence of overlapping nuclei prior to the occurrence of any splitting process. Some studies have been devoted to distinguishing overlapping nuclei from single ones. For instance, Tafavogh et al. [
14,
15] demonstrated a method for the identification of overlapping nuclei on microscopic images of neuroblastoma. Nuclei are segmented using a mean shift method, and three size-and shape-based features of cells namely (i) area, (ii) diameter equality, and (iii) concavity dominance are extracted to differentiate between single and overlapped cells using step-by-step conditional filtering. Abbas et al. [
16] proposed a method for detecting overlapping nuclei in microscopic red blood cell images prior to performing a splitting process. First, an image is binarized using an automatic thresholding approach, then three features, namely (i) convex hull, (ii) area, and (iii) elongation, are extracted. The pre-labeled value of each feature through parameter-tuning is used to determine overlapping nuclei. Wang et al. [
13] reported on a pre-determination scheme to identify overlapping nuclei using shape-based classification. Five shape features that is (i) solidity, (ii) convexity, (iii) eccentricity, (iv) area, and (v) variance are extracted for each nucleus and fed as input to an SVM classifier to classify single and overlapping cell nuclei. Four different types of image set: oil cells, yeast cells, blood cells and curvularia cells, are used to evaluate the method and obtain a classification accuracy of 86%, 90%, 88%, and 88% respectively. Guven et al. [
17] proposed an unsupervised data-clustering method to determine the presence of overlapping cell nuclei from Pap smear cervical images. The nuclei borders are firstly outlined using a morphological operation. Three shape-based features and two minima based features are extracted and used as inputs to a fuzzy clustering method to discriminate between single and overlapping nuclei. The method is evaluated using 290 nuclei and obtains an F-score of 79.1%, a recall of 67.4%, and precision of 95.7%. The methods in [
14,
15,
16] are parameter-dependent and limited to objects with a great variation of size and shape. The method proposed in [
13] is based only on shape and size features. In the case of cytology pleural effusion images, the forms of overlapping nuclei vary greatly. Thus, it can be deduced that considering only size and shape features may not be sufficient for discriminating between overlapping and single nuclei. The method presented in [
17] takes into account not only shape features but also local minima based features and judges for the presence of overlapping nuclei using an unsupervised clustering method, which yields acceptable performance. However, the method is designed specifically for cervical cells. It cannot be taken for granted that this method will provide good results with pleural effusion cells. The originators of the aforementioned method did not take into consideration the textural pattern difference between single and overlapping nuclei despite the fact that the texture pattern between single and overlapping nuclei varies greatly. Moreover, supervised learning techniques could greatly help to attain a more accurate detection rate [
18]. For our method, we extract a new combination of 16 geometric (i.e., size and shape) and 10 textural features and select the most relevant features from a total of 26 that are then used in identifying overlapping nuclei in CPE images. Using the selected features, five supervised learning methods, namely naïve Bayes (NB), support vector machine (SVM), K nearest neighborhood (KNN), decision tree (DT), and random forest (RF), are examined for the classification of single and overlapping nuclei. It should also be noted that our study objective is focused on accurately detecting overlapping nuclei to improve the extraction of each nucleus. Our proposed method is not a separation algorithm for overlapping nuclei or an extraction algorithm for interregional contours of overlapping nuclei.
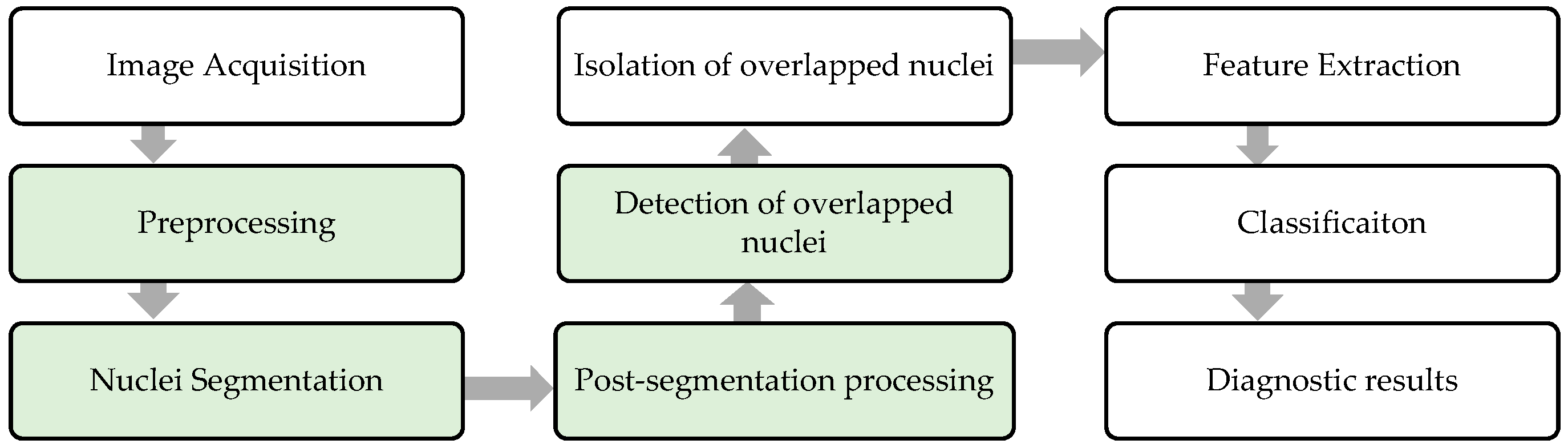
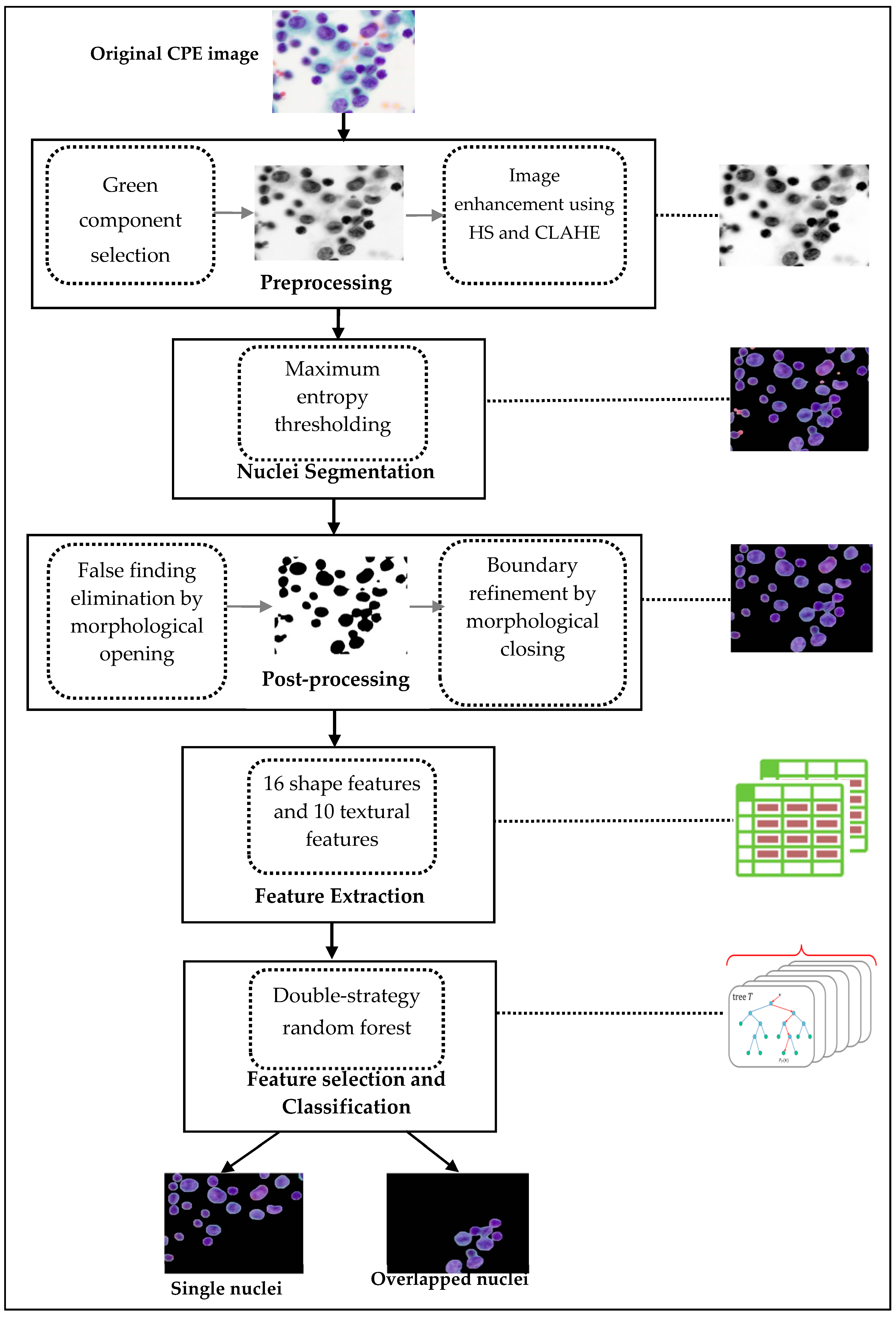
We hereby propose the following novel ideas to distinguish between single and overlapping nuclei in CPE images using three main steps: (i) nuclei segmentation: extracting candidate nuclei using maximum entropy thresholding supplemented by preprocessing and refinement; (ii) feature extraction: extracting a new combination of nuclei features, 16 geometric features and 10 textural features; and (iii) classification: selecting the most relevant features and determining whether the nucleus is single or overlapping using a double-strategy random forest algorithm. The performance of the proposed method was assessed using six evaluation metrics namely sensitivity, specificity, precision, F1 score, accuracy, and G-mean on a local dataset containing 4000 nuclei. The experiment results were acquired in various ways. Firstly, the classification accuracy of using all features and selecting them by random forest was investigated and compared. Then, the accuracy of four alternative classifiers, namely naïve NB, SVM, KNN, and DT, was further examined and compared with the results achieved by random forest. Third, the performance of previous studies was investigated and compared with the results achieved from the proposed method. In addition, the computation efficiency of nuclei segmentation, feature extraction, and classification was analyzed to prove the reliability and suitability of the proposed method for real-time use. This analysis demonstrates that the proposed method is relatively simple, computationally affordable, and yields promising results. Thus, it can serve as a feasible, reproducible and cost-effective tool in the development of an advanced system for diagnosing cancer cells in CPE images.
The rest of this paper is divided into five sections.
Section 2 presents the image acquisition and dataset description processes.
Section 3 presents the methodology proposed in this study containing preprocessing, nuclei candidate extraction, post-processing, feature extraction, and classification. The experiment results and discussion are presented in
Section 4. The conclusion is given in the last section,
Section 5.
4. Experiment Results and Discussions
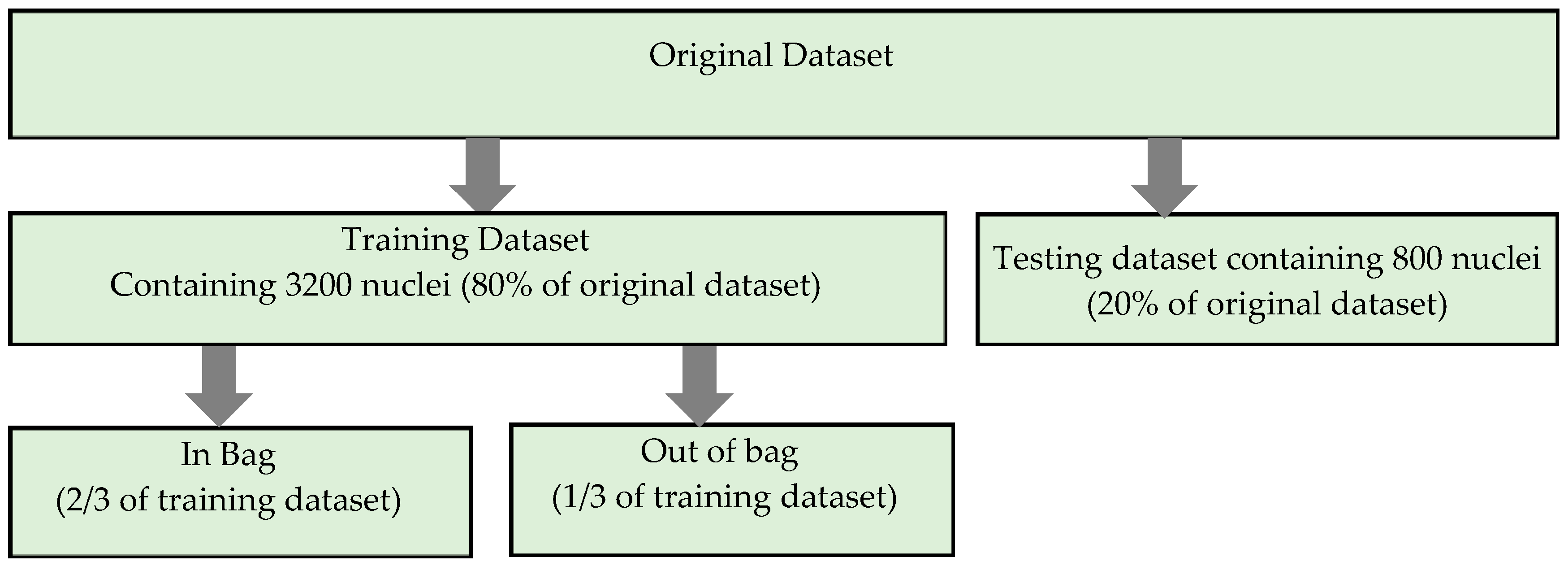
In our study, the experiments were carried out in a MATLAB_R2016b environment using an Intel(R) Core (TM) i7 CPU 3.40–3.70 GHz personal computer and Microsoft Windows 7, 64-bit operating system. The study is based on a local dataset containing 124 CPE images. The main contribution of the study is the development of an effective algorithm that can accurately determine the presence of overlapping nuclei in CPE images. The first step of the proposed algorithm was to deal with image quality. Histogram stretching and CLAHE image enhancement methods were utilized in order to enhance the contrast of cell nuclei. Then, maximum entropy thresholding based nuclei segmentation was employed to extract candidate nuclei regions from surrounding objects in the image. The segmentation performance of maximum entropy based nuclei segmentation was evaluated in the test images and yielded a 92% detection accuracy. After the nuclei were detected, an overlapping nuclei detection scheme was developed. A new combination of 16 geometrical and 10 textural features was extracted from 4000 nuclei containing single and overlapping nuclei. Thus, our dataset is made up of 4000 × 26-dimensional datasets. It is partitioned into training and testing sets in an 80/20 ratio, as given in
Table 4.
Then, double-strategy RF was utilized to select the most important features and classify single and overlapping nuclei using selected high-ranking features. One of the important parameters that we needed to adjust while constructing RF was the number of decision trees to be grown. The optimal number of decision trees was obtained through empirical tuning. OBB errors using a different number of decision trees are illustrated in
Figure 13. From the graph, it can be seen that OOB errors decrease at above 250 decision trees, and start to stabilize from 300 trees. Thus, we grew 400 decision trees in order to maintain classification stability and keep the computation cost low.
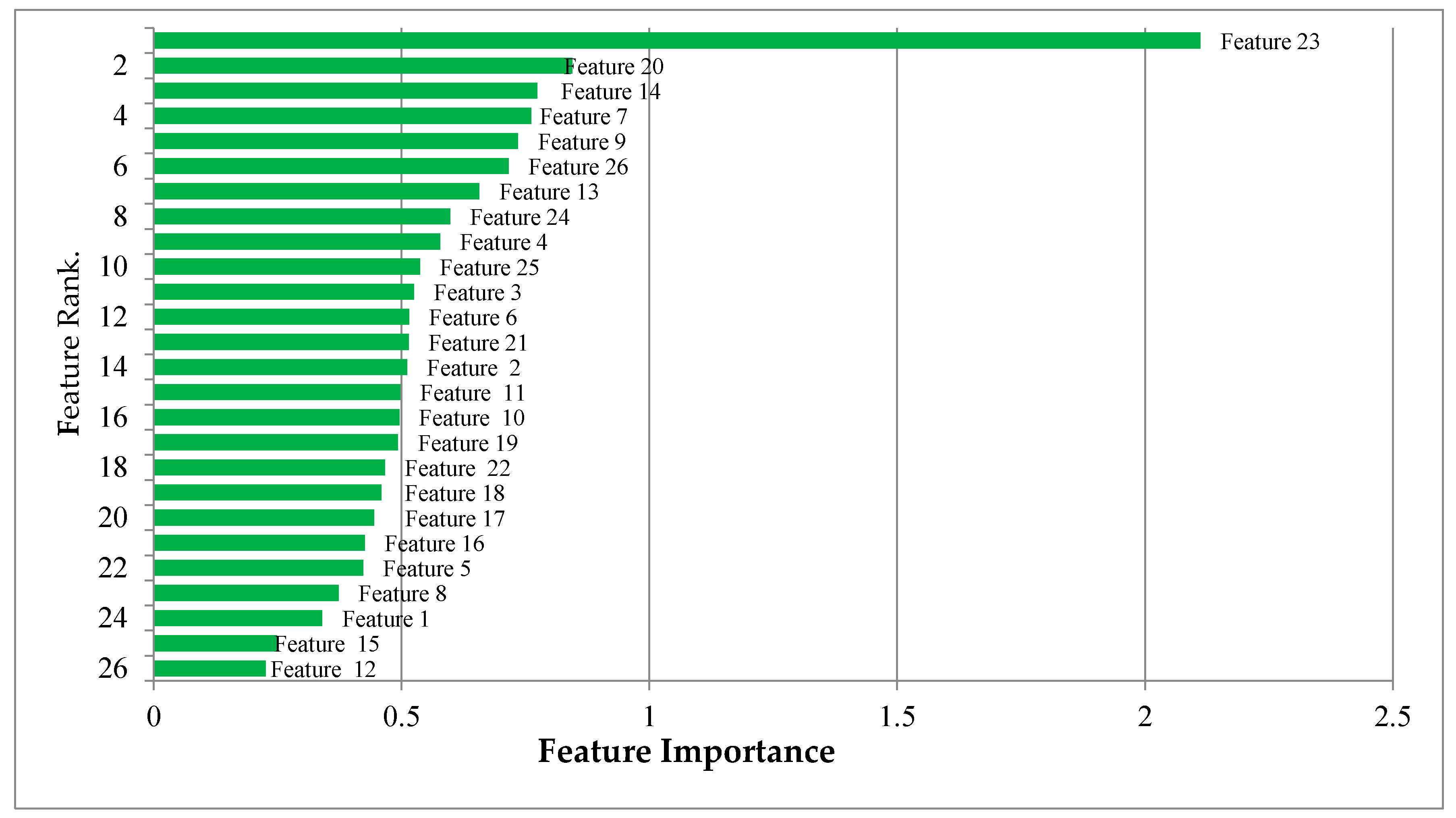
Once a random forest was constructed with 400 decision trees, feature selection was performed by scoring OOB permutation errors using each feature. The importance of features ranked by RF is given in
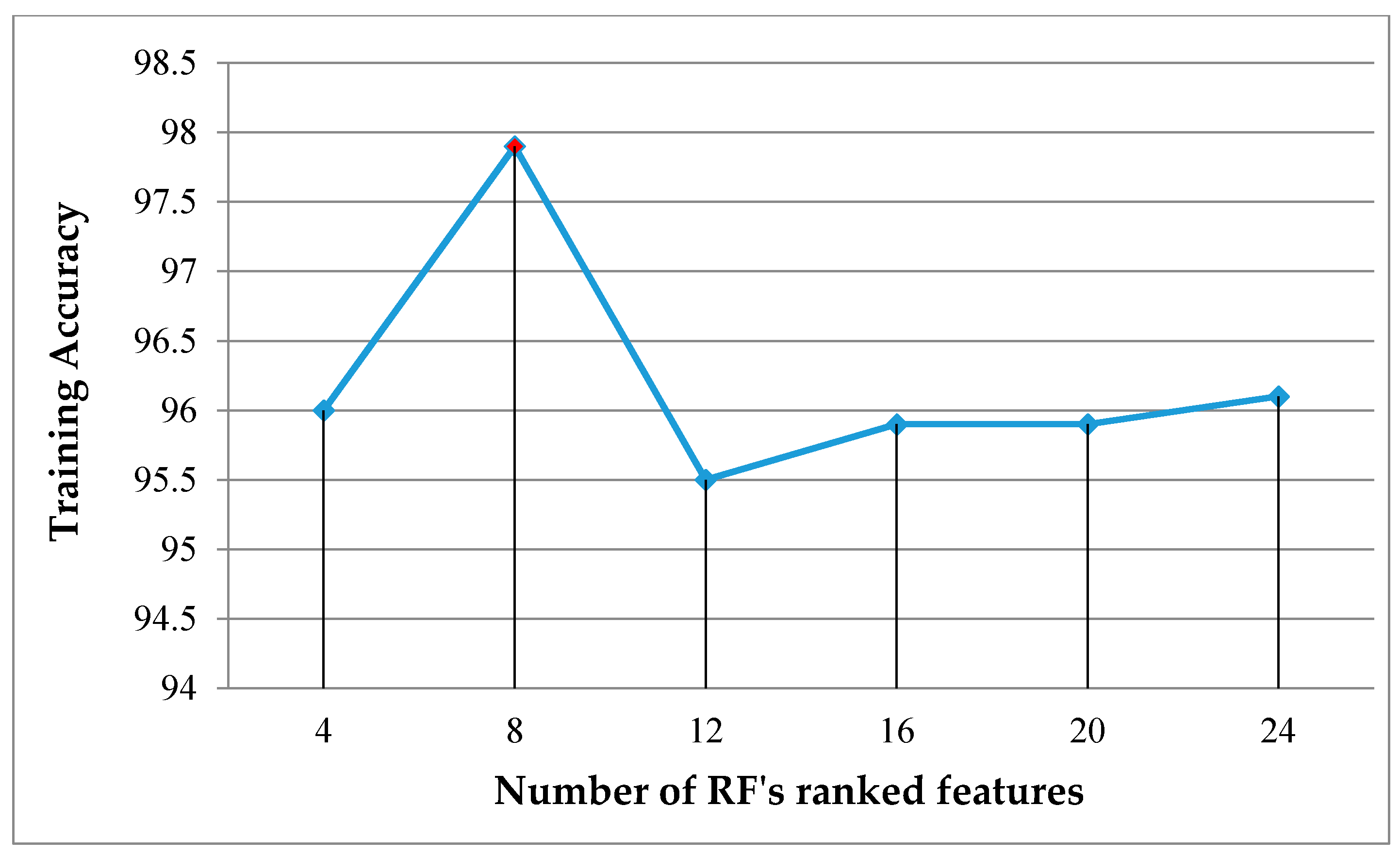
Figure 14. To select the most significant features, we experimentally tested the different feature numbers in ascending rank order and examined their training accuracy as given in
Figure 15. The chart shows that the first eight ranked features achieved the highest training accuracy, and those features are described in
Table 5. We fed those selected features as input to train the RF ensemble classifier. The trained RF classifier was used to validate the testing dataset. The classification accuracy of using RF’s selected features was compared to the accuracy of using all features. In addition, we also examined four alternative classifiers: NB [
40], SVM [
41], KNN [
42], and DT [
43] by coupling with all features and RF’s selected features. The classification accuracies of using all features and RF selected features blending with five classifiers are presented in
Table 6 and
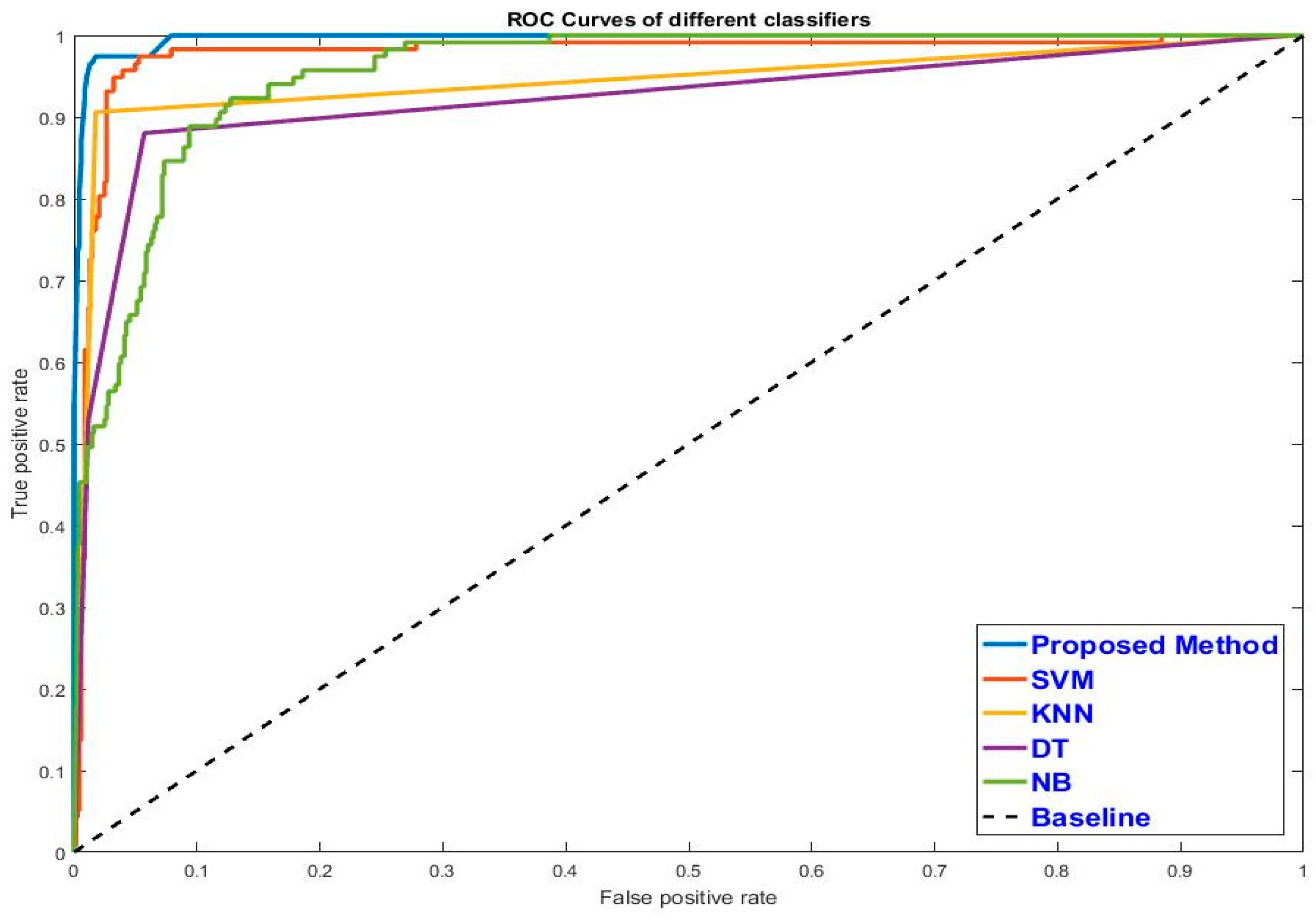
Table 7, respectively. From the experiment results, it is shown that using RF selected features provides better accuracy compared to using all features for most classifiers except NB. The results also reveal that the RF ensemble classifier yields preferable accuracy compared to NB, SVM, KNN, and DT classifiers. The synergy between RF’s selected features and the RF ensemble classifier reached the highest classification accuracy. In order to evaluate the classifiers graphically, we plotted an ROC curve for each classifier, as given in
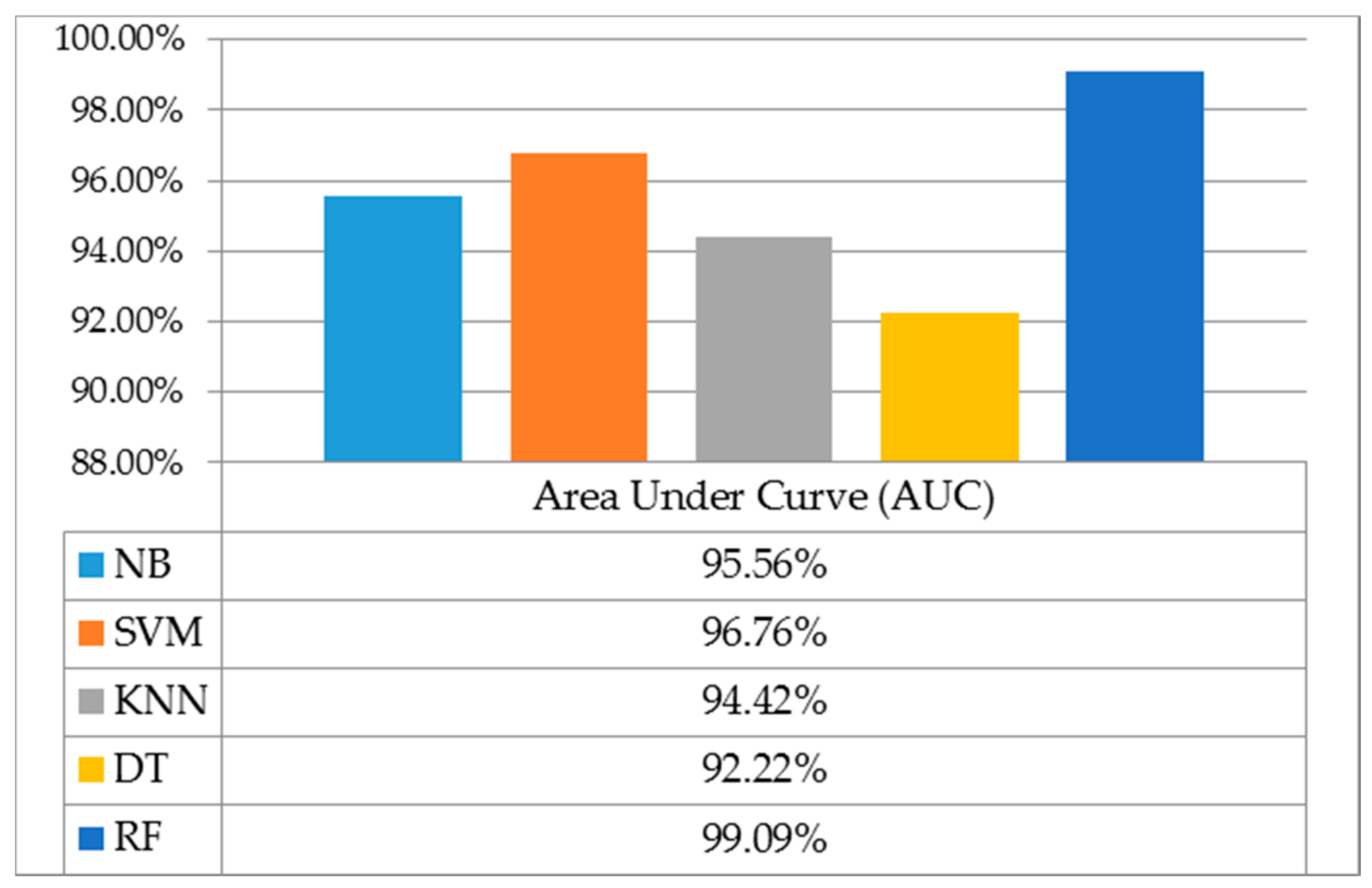
Figure 16. The curves show that the RF ensemble classifier gains higher accuracy and stability compared to others. From the ROC curves, we further computed the AUC of different classifiers as presented in
Figure 17. An RF ensemble classifier using RF-selected features reached the highest AUC by a given 99.09%.
In order to compare with previous studies, there was no common dataset, and previous studies were evaluated based on different types of images. In order to make a fair, objective comparison, we adopted the methodologies of previous studies to our application. It should be noted that all the methods in the comparison were evaluated with the same experiment settings and the same dataset used to test the proposed method. Thus, the evaluation results were compared fairly without affecting any other factors. The comparison of classification accuracy obtained along with their corresponding methodologies is presented in
Table 8. From the experiment results, it is inferred that our proposed method provides superior accuracy compared to previous works [
13,
17]. It is also reasonable to conclude that our combination of geometric and textural features is more discriminant than the features used in previous studies for classifying single and overlapping nuclei. To validate computational efficiency, we also analyzed the processing time of each processing step and the entire algorithm as given in
Table 9 and found that computational complexity is relatively simple. The visual results of detected overlapped nuclei using our proposed method are depicted in
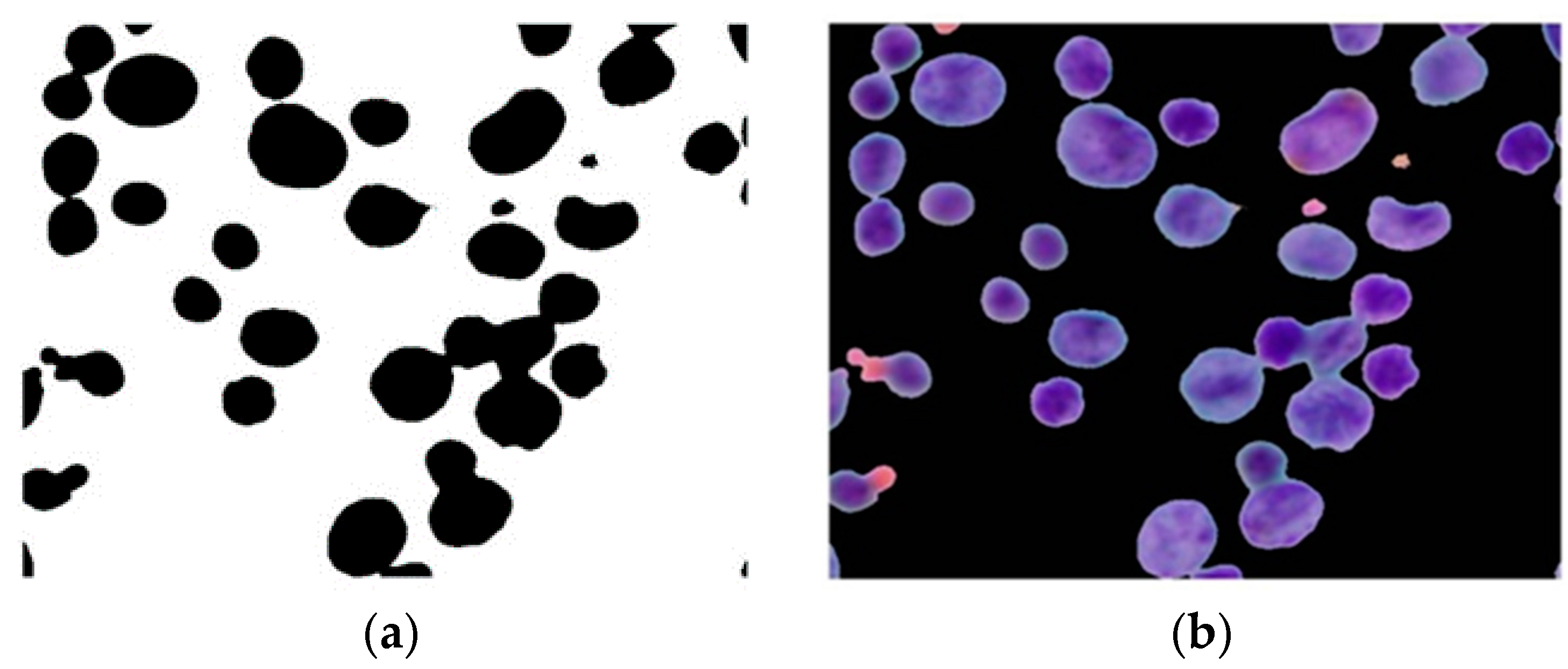
Figure 18.
The proposed algorithm can be utilized to accurately detect and classify touching, overlapping or clustering nuclei from single nuclei. Due to its high accuracy and computational simplicity, it can serve as a new supportive tool in developing new overlapping cell separation algorithms. Moreover, our method has the potential to integrate with existing overlapping-separation methods, such as watershed methods, contour concavity analysis, rule-based methods, etc., to separate overlapping nuclei. It can be deduced that accurately detecting overlapping nuclei before decomposing them into their constituent parts can help to reduce the workload of separation methods because these methods need to work only on detected overlapping nuclei instead of on all nuclei. It should also be noted that the proposed algorithm may determine the presence of overlapping nuclei even if there are no overlapping nuclei in the image. Since the aim of our study focuses on developing a determination algorithm for overlapping nuclei, isolating overlapping nuclei, or extracting the interregional contour of each nucleus has been deferred for future study.
5. Conclusions
This paper presents a method for the automated detection and classification of overlapping nuclei from CPE images using maximum entropy thresholding, new combinations of geometric and textural features, and double-strategy RF. First, the images were enhanced on their green color channel using histogram stretching and CLAHE. Then, maximum entropy thresholding was employed to segment the cell nuclei from their surrounding background (i.e., cytoplasm, blood cells, artifacts, and so on). The post-processing step was performed to eliminate any false findings and preserve the shape of the segmented nuclei using morphological operations. A new combination of 16 geometrical and 10 textural features was extracted for each extracted nucleus region. A double-strategy RF algorithm was applied to perform two tasks: ensemble feature selection to select the most relevant features, and an ensemble classifier to identify the presence of overlapped nuclei using selected features. RF ensemble feature selection selected eight features out of a total of 26 features that were used as input to the RF ensemble classifier. The proposed method was evaluated on 4000 nuclei from CPE images with respect to six performance measures and AUC. It yielded 96.6% sensitivity, 98.7% specificity, 92.62% precision, 94.6% F1 score, 98.4% accuracy, 97.6% G mean, and AUC 99.0%. Only 5.17 s of computation time was required to run the entire algorithm. The performance from using RF’s selected features was compared to the performance of all features by coupling with five different classifiers: NB, SVM, KNN, DT, and RF. The comparison revealed that RF’s selected features were better in terms of generalization capability and yielded significant improvements in accuracy for most classifiers, except NB. It is also worth noting that the RF ensemble classifier provided favorable accuracy compared to other classifiers. The synergy between the proposed features and a double-strategy RF achieved the promising results. Furthermore, the achieved results were compared with the results obtained from previous works. The results prove that the proposed algorithm yields superior results compared to previous works. It is our finding that the combination of geometric and textural features is more effective than the features used in previous studies. Due to its high accuracy and computational simplicity, the proposed algorithm can be used as a new basis in developing algorithms for separating overlapping nuclei, and can also serve as a new supportive tool in developing advanced automated cell analysis systems. Furthermore, the proposed method has the potential to integrate with the existing separation algorithms of overlapping nuclei to enhance separation accuracy by accurately locating the overlapping nuclei to be separated. Separation of overlapping nuclei or extraction of the interregional contour of each nucleus has been deferred for future study.