Pb2+ Effects on Growth, Lipids, and Protein and DNA Profiles of the Thermophilic Bacterium Thermus Thermophilus
Abstract
:1. Introduction
2. Materials and Methods
2.1. Chemicals
2.2. Biological System and Cultural Condition
2.3. Homogenate Preparation
2.4. Lipid and Fatty Acid Analysis
2.5. Extracellular Phase Preparation
2.6. Enzyme Activities
2.7. Protein Determination
2.8. Protein Profile
2.9. Two-Dimensional Electrophoresis
2.10. DNA Preparation and Genotyping Analysis
2.11. Statistical Analysis
3. Results
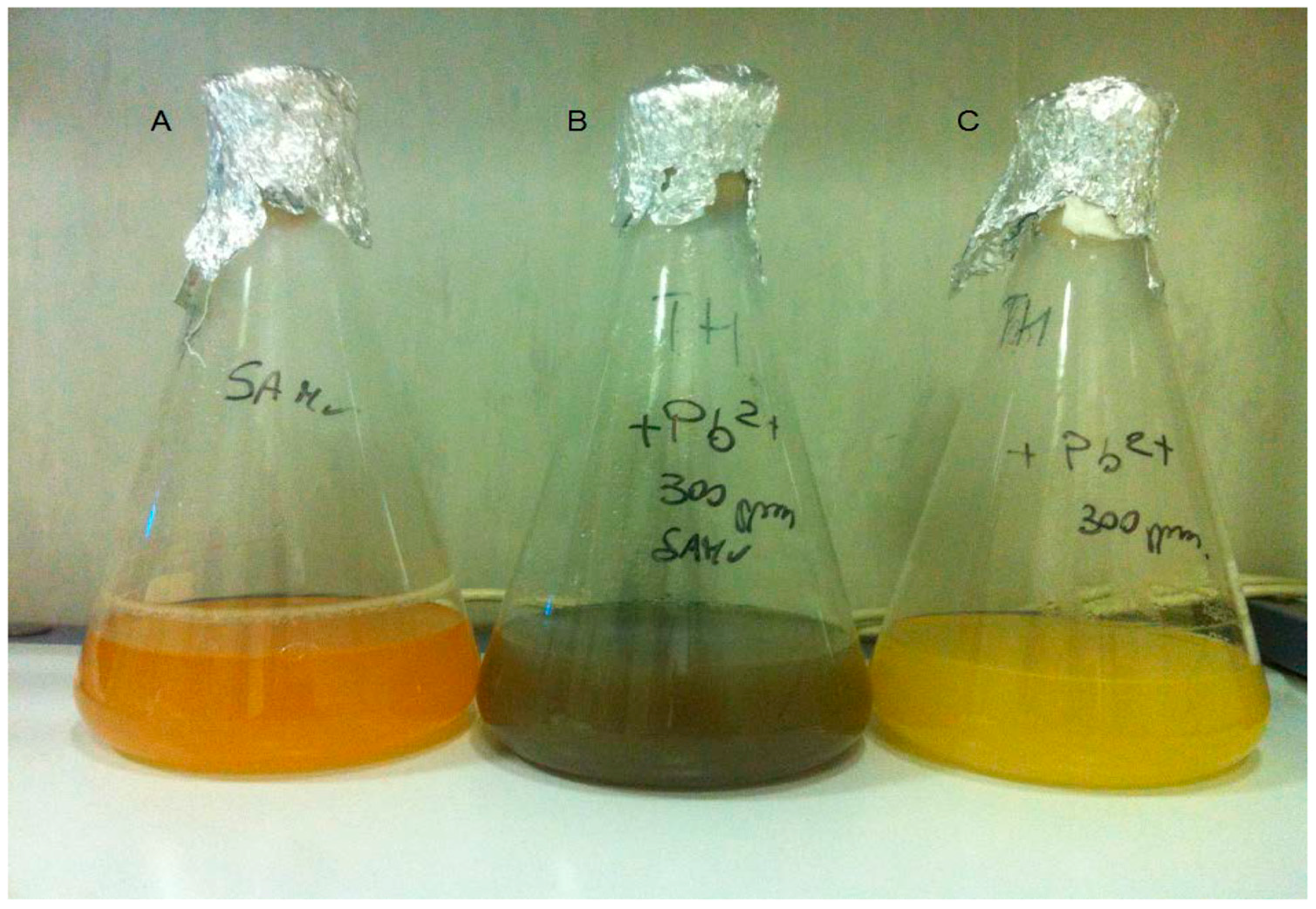
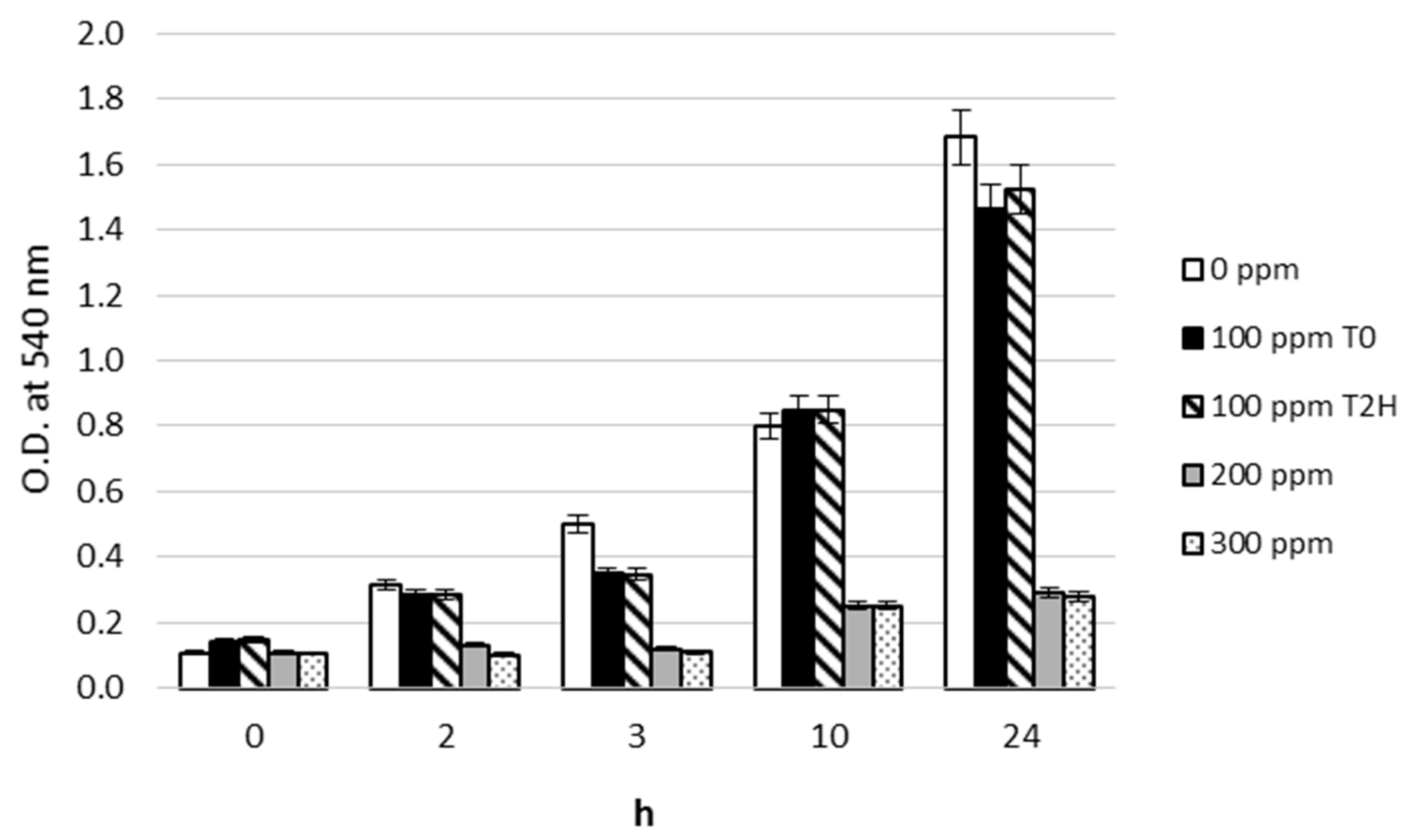
3.1. Pb2+ Effects on Growth
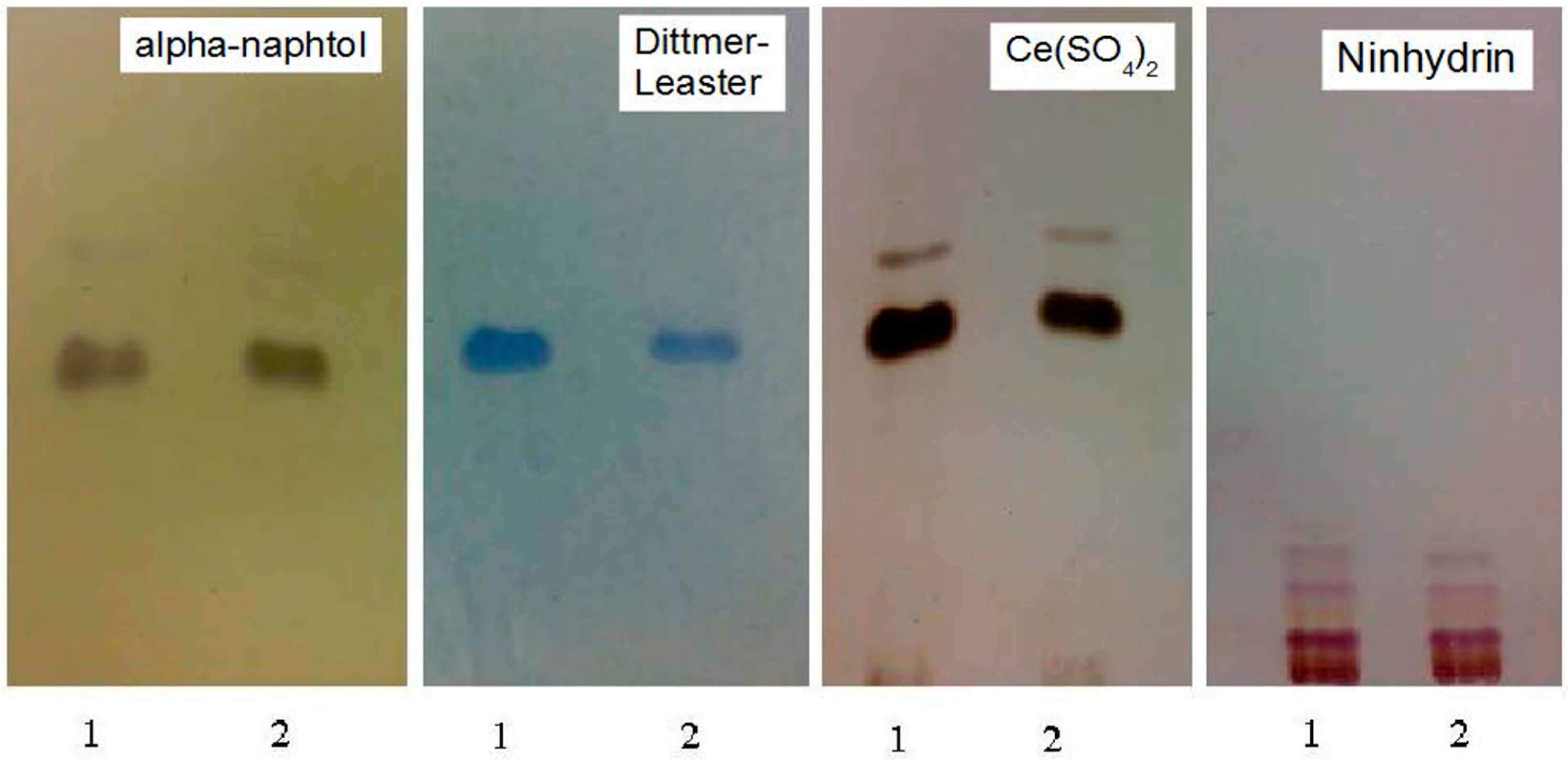
3.2. Pb2+ Effects on Lipid Pattern and Fatty Acid Composition
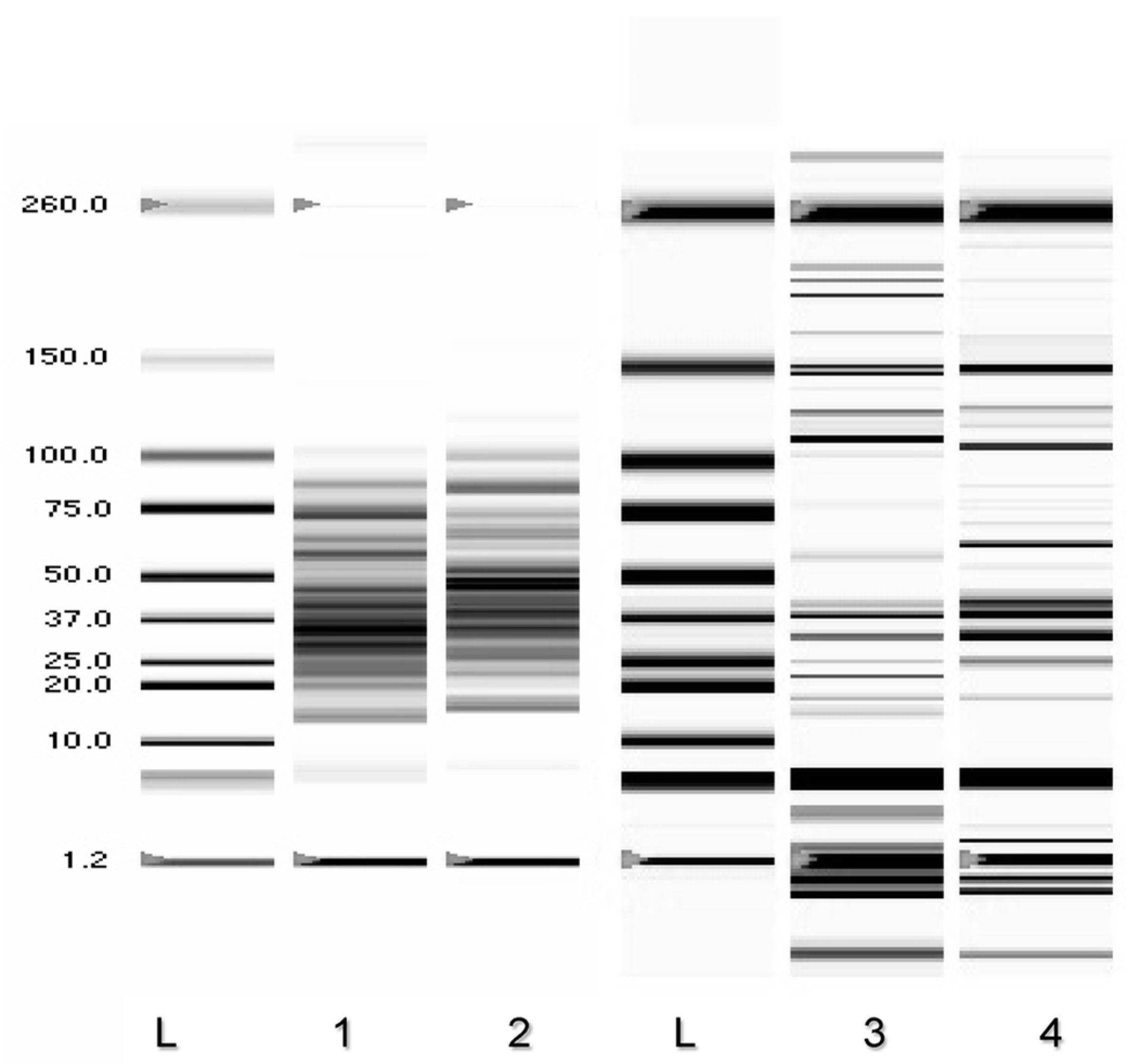
3.3. Pb2+ Effects on Cell Protein Profiles and Enzyme Activities
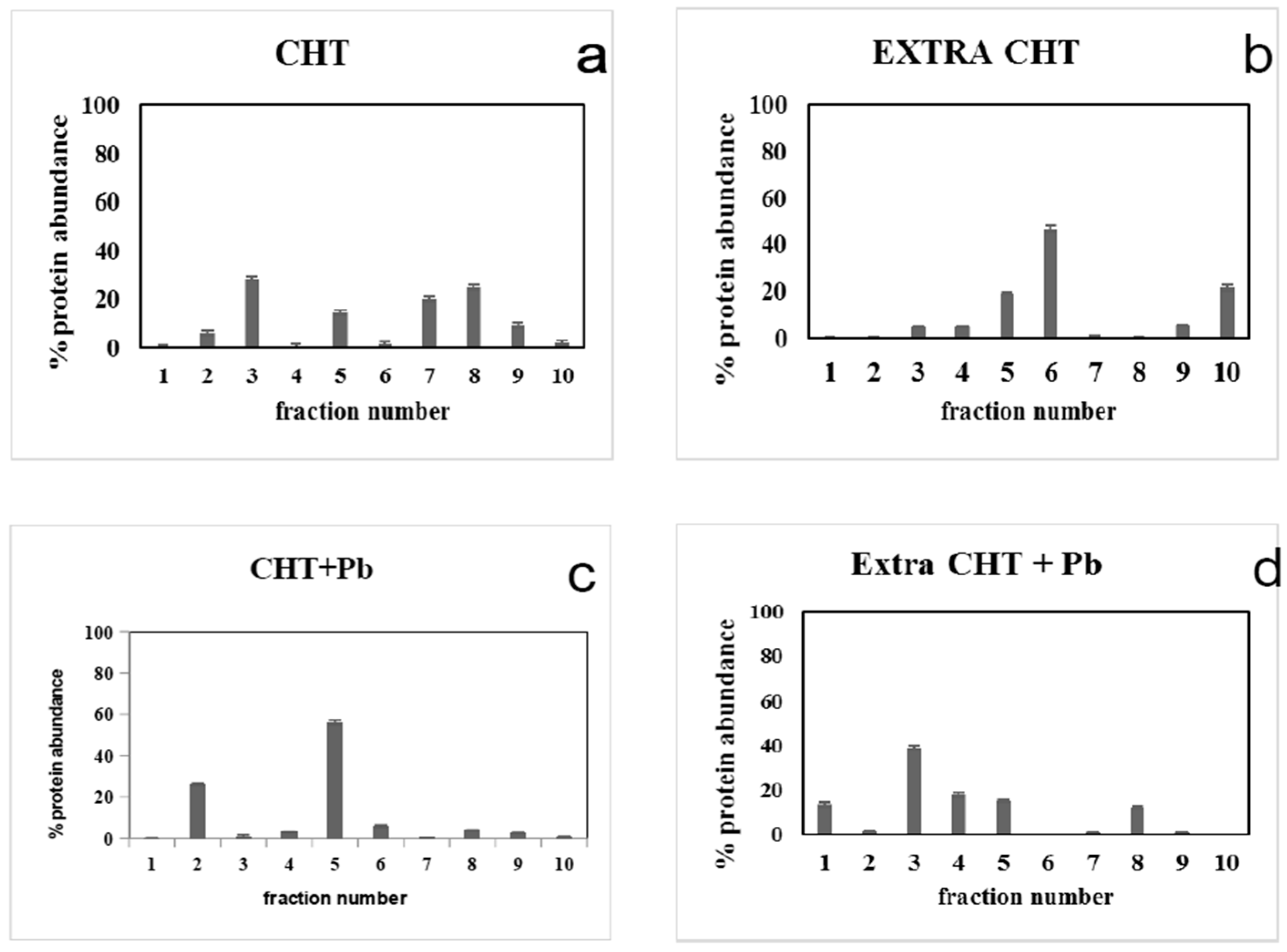
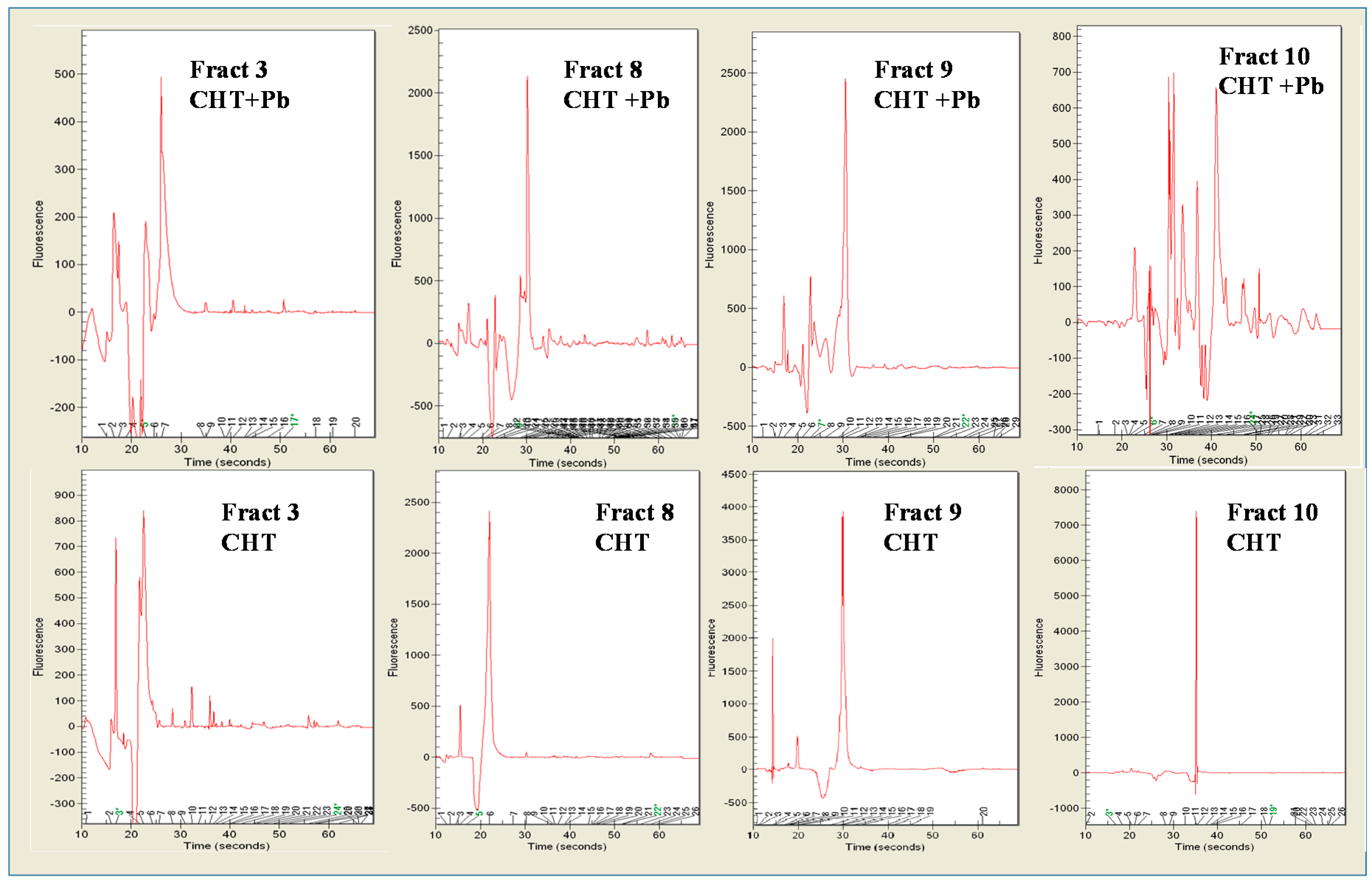
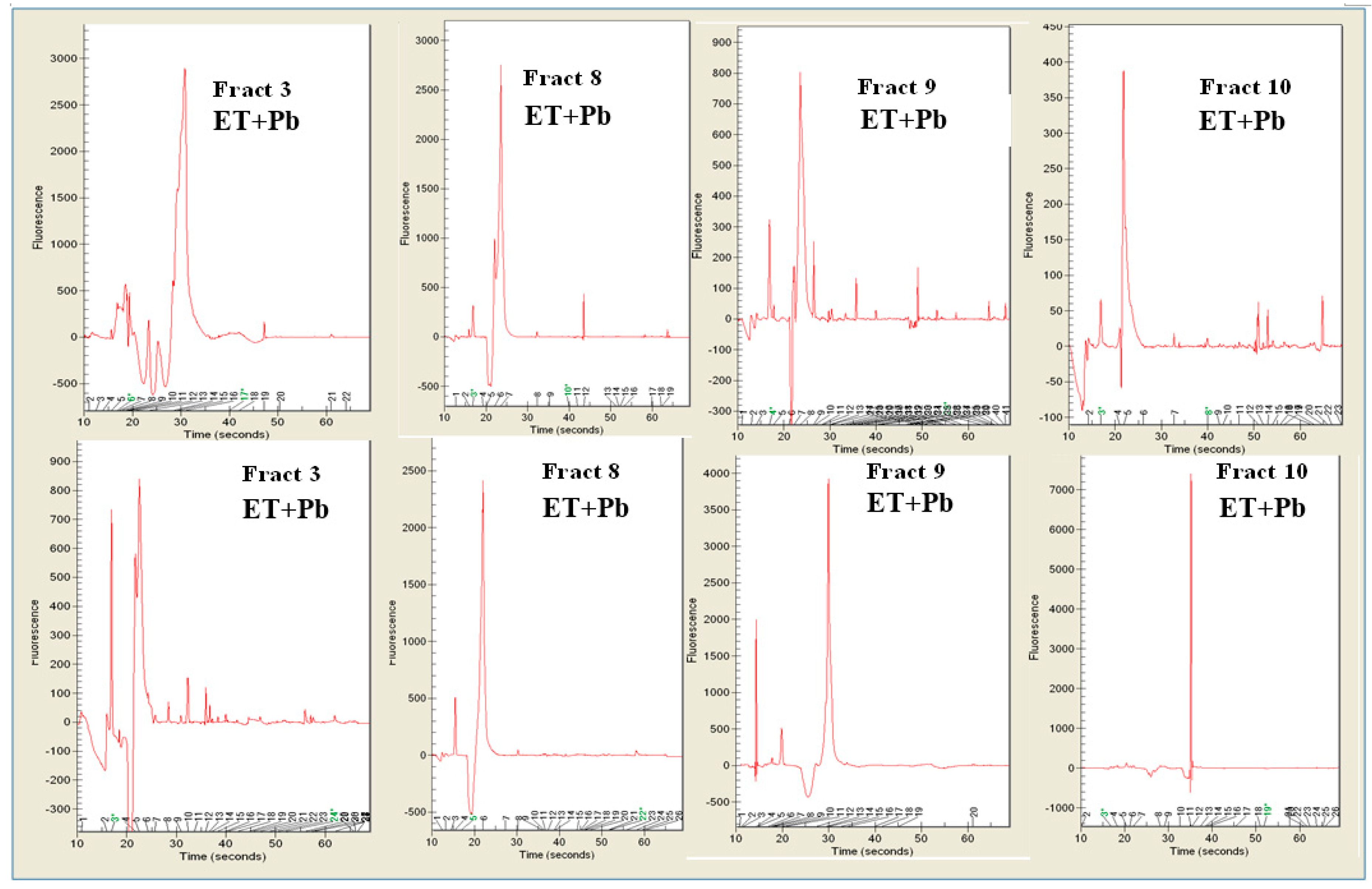
3.4. Pb2+ Effects on Protein Pattern; Two-Dimensional Electrophoresis
3.5. Determination of DNA Melting Temperature
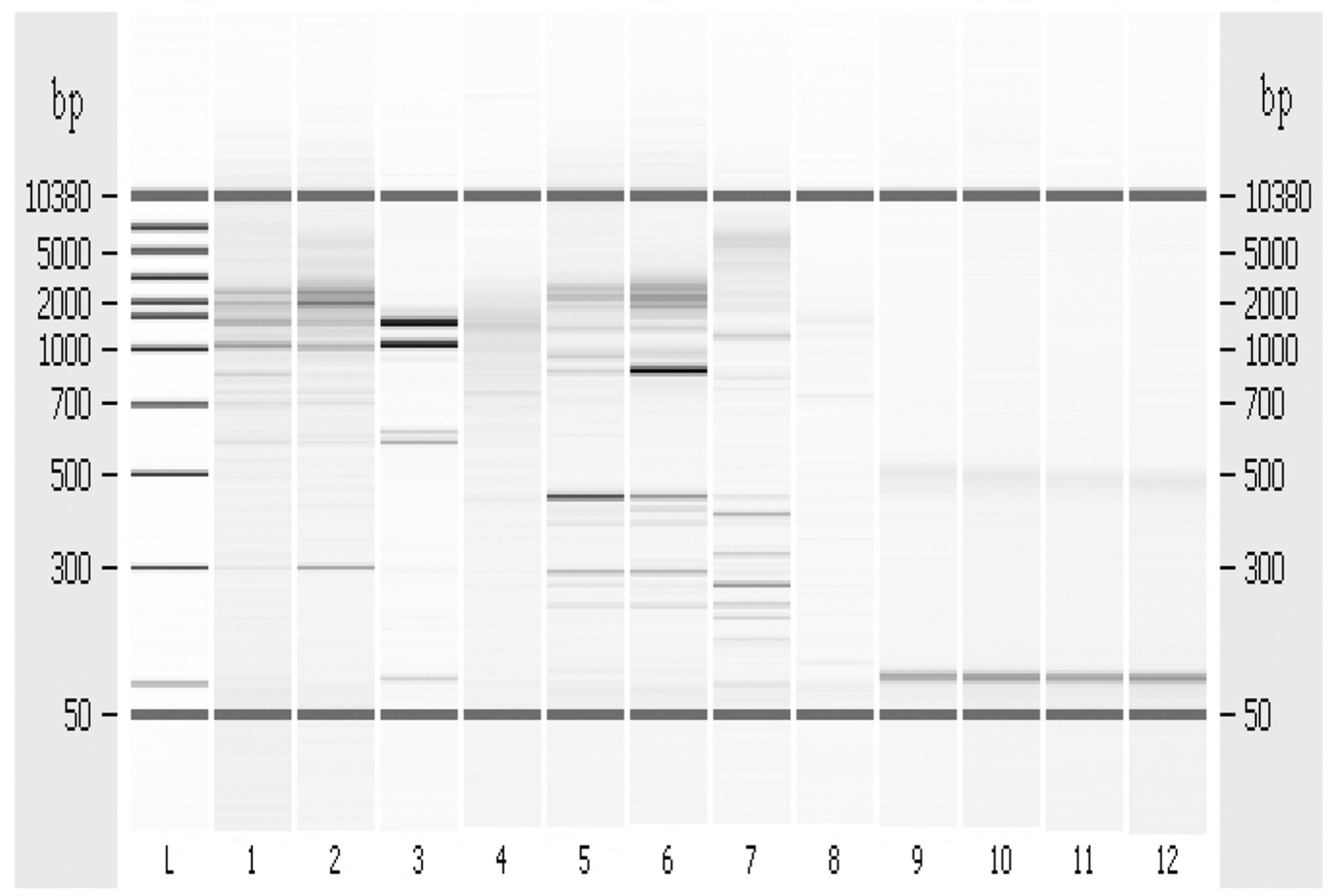
3.6. Random Amplified Polymorphic DNA-PCR
4. Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gavaghan, H. Lead, unsafe at any level. Bull. World Health Organ. 2002, 80, 82. [Google Scholar] [PubMed]
- Lanphear, B.P.; Hornung, R.; Khoury, J.; Yolton, K.; Baghurst, P.; Bellinger, D.C.; Canfield, R.L.; Dietrich, K.N.; Bornschein, R.; Greene, T.; et al. Low-level environmental lead exposure and children’s intellectual function: An International Pooled Analysis. Environ. Health Perspect. 2005, 113, 894–899. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Konopka, A.; Zakharova, T.; Bischoff, M.; Oliver, L.; Nakatsu, C.; Turco, R.F. Microbial biomass and activity in lead-contaminated soil. Appl. Environ. Microbiol. 1999, 65, 2256–2259. [Google Scholar] [PubMed]
- Zhou, Q.; Zhang, J.; Fu, J.; Shi, J.; Jiang, G. Biomonitoring: An appealing tool for assessment of metal pollution in the aquatic ecosystem. Anal. Chim. Acta 2008, 606, 135–150. [Google Scholar] [CrossRef] [PubMed]
- Hernberg, S. Lead poisoning in a historical perspective. Am. J. Ind. Med. 2000, 38, 244–254. [Google Scholar] [CrossRef]
- Hong, S.; Candelone, J.P.; Patterson, C.C.; Boutron, C.F. Greenland ice evidence of hemispheric lead pollution two millennia ago by greek and roman civilizations. Science 1994, 265, 1841–1843. [Google Scholar] [CrossRef] [PubMed]
- Flegal, A.R.; Smith, D.R. Measurements of environmental lead contamination and human exposure. Rev. Environ. Contam. Toxicol. 1995, 143, 1–45. [Google Scholar] [PubMed]
- Landsberger, S.; Iskander, F.; Basunia, S.; Barnes, D.; Kaminski, M. Lead and copper contamination of soil from industrial activities and firing ranges. Biol. Trace Elem. Res. Winter 1999, 71–72, 387–396. [Google Scholar] [CrossRef]
- De Vries, W.; Römkens, P.F.; Schütze, G. Critical soil concentrations of cadmium, lead, and mercury in view of health effects on humans and animals. Rev. Environ. Contam. Toxicol. 2007, 191, 91–130. [Google Scholar] [PubMed]
- Wierzbicka, M.; Antosiewicz, D. How lead can easily enter the food chain a study of plant roots. Sci. Total Environ. 1993, 1, 423–429. [Google Scholar] [CrossRef]
- Dudka, S.; Miller, W.P. Accumulation of potentially toxic elements in plants and their transfer to human food chain. J. Environ. Sci. Health B 1999, 34, 681–708. [Google Scholar] [CrossRef] [PubMed]
- Shi, W.; Becker, J.; Bischoff, M.; Turco, R.F.; Konopka, A.E. Association of microbial community composition and activity with lead, chromium, and hydrocarbon contamination. Appl. Environ. Microbiol. 2002, 68, 3859–3866. [Google Scholar] [CrossRef] [PubMed]
- Valls, M.; De Lorenzo, V. Exploiting the genetic and biochemical capacities of bacteria for the remediation of heavy metal pollution. FEMS Microbiol. Rev. 2002, 26, 327–338. [Google Scholar] [CrossRef] [PubMed]
- Nies, D.H. Heavy metal-resistant bacteria as extremophiles: Molecular physiology and biotechnological use of Ralstonia sp. CH34. Extremophiles 2000, 4, 77–82. [Google Scholar] [CrossRef] [PubMed]
- Hamdy, M.K.; Noyes, O.R. Formation of Methyl Mercury by Bacteria. Appl. Microbiol. 1975, 30, 424–432. [Google Scholar] [PubMed]
- Nies, D.H. Efflux-mediated heavy metal resistance in prokaryotes. FEMS Microbiol. Rev. 2003, 27, 313–339. [Google Scholar] [CrossRef]
- Özdemir, S.; Kılınç, E.; Poli, A.; Nicolaus, B. Biosorption of heavy metals (Cd2+, Cu2+, Co2+ and Mn2+) by thermophilic bacteria, Geobacillus thermantarcticus and Anoxybacillus amylolyticus: Equilibrium and kinetic studies. Bioremediat. J. 2013, 17, 86–96. [Google Scholar] [CrossRef]
- Hetzer, A.; Daughney, C.J.; Morgan, H.W. Cadmium ion biosorption by the thermophilic bacteria Geobacillus stearothermophilus and G. thermocatenulatus. Appl. Environ. Microbiol. 2006, 72, 4020–4027. [Google Scholar] [CrossRef] [PubMed]
- Burnett, P.G.G.; Daughney, C.J.; Peak, D. Cd adsorption onto Anoxybacillus flavithermus: Surface complexation modeling and spectroscopic investigations. Geochim. Cosmochim. Acta 2006, 70, 5253–5269. [Google Scholar] [CrossRef]
- Chatterjee, S.K.; Bhattacharjee, I.; Chandra, G. Biosorption of heavy metals from industrial waste water by Geobacillus thermodenitrificans. J. Hazard Mater. 2010, 175, 117–125. [Google Scholar] [CrossRef] [PubMed]
- Babak, L.; Šupinova, P.; Zichova, M.; Burdychova, R.; Vitova, E. Biosorption of Cu, Zn and Pb by thermophilic bacteria—Effect of biomass concentration on biosorption capacity. Acta Univ. Agric. Silvic. Mendel. Brun. 2012, 60, 9–18. [Google Scholar] [CrossRef]
- Spada, S.; Pembroke, T.J.; Wall, G.J. Isolation of a novel Thermus thermophilus metal efflux protein that improves Escherichia coli growth under stress conditions. Extremophiles 2002, 6, 301–308. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.H.; Hocheng, H.; Chang, H.Y.; Shih, A. Metal Removal rate of Thiobacillus thiooxidans without pre-secreted metabolite. J. Mater. Process. Technol. 2008, 201, 560–564. [Google Scholar] [CrossRef]
- Hocheng, H.; Chang, J.; Jadhav, U. Micromachining of various metals by using Acidithiobacillus ferrooxindans 13820 culture supernatant experiments. J. Clean. Prod. 2012, 20, 180–185. [Google Scholar] [CrossRef]
- Romano, I.; Lama, L.; Schiano Moriello, V.; Poli, A.; Gambacorta, A.; Nicolaus, B. Isolation of a new thermohalophilic Thermus thermophilus strain from hot spring, able to grow on a renewable source of polysaccharides. Biotechnol. Lett. 2004, 26, 45–49. [Google Scholar] [CrossRef] [PubMed]
- Nicolaus, B.; Manca, M.C.; Lama, L.; Esposito, E.; Gambacorta, A. Lipid modulation by environmental stresses in two models of extremophiles isolated from Antarctica. Polar Biol. 2001, 24, 1–8. [Google Scholar] [CrossRef]
- Poli, A.; Laezza, G.; Gul-Guven, R.; Orlando, P.; Nicolaus, B. Geobacillus galactosidasius sp. nov., a new thermophilic galactosidase-producing bacterium isolated from compost. Syst. Appl. Microbiol. 2011, 34, 419–423. [Google Scholar] [CrossRef] [PubMed]
- Lama, L.; Poli, A.; Nicolaus, B. Geobacillus thermantarcticus as source of biotechnological thermozymes and exopolysaccharides: A review. Curr. Trends Microbiol. 2012, 8, 1–12. [Google Scholar]
- Bradford, M.M. A rapid and sensitive method for quantitation of microgram quantities of protein using the principles of protein-dye binding. Anal. Biochem. 1976, 72, 248–254. [Google Scholar] [CrossRef]
- Poli, A.; Salerno, A.; Laezza, G.; Di Donato, P.; Dumontet, S.; Nicolaus, B. Heavy metal resistance of some thermophiles: Potential use of α-amylase from Anoxybacillus amylolyticus as a microbial enzymatic bioassay. Res. Microbiol. 2009, 160, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Nazzaro, F.; Fratianni, F.; Nicolaus, B.; Poli, A.; Orlando, P. The prebiotic source influences the growth, biochemical features and survival under simulated gastrointestinal conditions of the probiotic Lactobacillus acidophilus. Anaerobe 2012, 18, 280–285. [Google Scholar] [CrossRef] [PubMed]
- Poli, A.; Romano, I.; Cordella, P.; Orlando, P.; Nicolaus, B.; Ceschi Berrini, C. Anoxybacillus thermarum sp. nov., a novel thermophilic bacterium isolated from thermal mud in Euganean hot springs, Abano Terme, Italy. Extremophiles 2009, 13, 867–874. [Google Scholar] [CrossRef] [PubMed]
- Ronimus, R.S.; Parker, L.E.; Morgan, H.W. The utilization of RAPD-PCR for identifying thermophilic and mesophilic Bacillus species. FEMS Microbiol. Lett. 1997, 147, 75–79. [Google Scholar] [CrossRef] [PubMed]
- Barabote, R.D.; Xie, G.; Leu, D.H.; Normand, P.; Necsulea, A.; Daubin, V.; Me´digue, C.; Adney, W.S.; Xu, X.C.; Lapidus, A.; et al. Complete genome of the cellulolytic thermophile Acidothermus cellulolyticus 11B provides insights into its ecophysiological and evolutionary adaptations. Genome Res. 2009, 19, 1033–1043. [Google Scholar] [CrossRef] [PubMed]
- Özdemir, S.; Kilinc, E.; Poli, A.; Nicolaus, B.; Guven, K. Biosorption of Cd, Cu, Ni, Mn and Zn from aqueous solutions by thermophilic bacteria, Geobacillus toebii sub. sp. decanicus and Geobacillus thermoleovorans sub. sp. stromboliensis: Equilibrium, kinetic and thermodynamic studies. Chem. Eng. J. 2009, 152, 195–206. [Google Scholar] [CrossRef]
- Mastascusa, V.; Romano, I.; Di Donato, P.; Poli, A.; Della Corte, V.; Rotundi, A.; Bussoletti, E.; Quarto, M.; Pugliese, M.; Nicolaus, B. Extremophiles Survival to Simulated Space Conditions: An Astrobiology Model Study. Orig. Life Evol. Biosph. 2014, 44, 231–237. [Google Scholar] [CrossRef] [PubMed]
- Finore, I.; Orlando, P.; Di Donato, P.; Leone, L.; Nicolaus, B.; Poli, A. Nesterenkonia aurantiaca sp. nov., an alkaliphilic actinobacterium isolated from Antarctica. Int. J. Syst. Evol. Microbiol. 2016, 66, 1554–1560. [Google Scholar] [CrossRef] [PubMed]

| Fatty Acids | 0 ppm Pb2+ | 100 ppm Pb2+ |
|---|---|---|
| iso-C15:0 | 21.54 ± 1.1% | 16.96 ± 0.9% |
| iso-C17:0 | 47.98 ± 2.2% | 60.36 ± 3.1% |
| anteiso-C17:0 | 12.24 ± 0.4% | 10.08 ± 0.3% |
| Sample | Proteins in Crude Homogenate (mg/mL) | Proteins in Crude Homogenate (mg/g dry cell) | Proteins in Extracellular Fraction (mg/mL) | Proteins in Extracellular Fraction (mg/g dry cell) | β-Glucosidase Specific Activity (U/mg) | α-Maltosidase Specific Activity (U/mg) |
|---|---|---|---|---|---|---|
| 0 ppm Pb2+ | 36.03 ± 1.8 | 51.47 ± 2.2 | 2.07 ± 0.06 | 2.96 ± 0.07 | 4.1 ± 0.12 | 8.0 ± 0.22 |
| 100 ppm Pb2+ | 31.29 ± 1.5 | 44.7 ± 1.9 | 1.77 ± 0.05 | 2.53 ± 0.06 | 3.7 ± 0.11 | 6.6 ± 0.18 |
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Nicolaus, B.; Poli, A.; Di Donato, P.; Romano, I.; Laezza, G.; Gioiello, A.; Ulgiati, S.; Fratianni, F.; Nazzaro, F.; Orlando, P.; et al. Pb2+ Effects on Growth, Lipids, and Protein and DNA Profiles of the Thermophilic Bacterium Thermus Thermophilus. Microorganisms 2016, 4, 45. https://doi.org/10.3390/microorganisms4040045
Nicolaus B, Poli A, Di Donato P, Romano I, Laezza G, Gioiello A, Ulgiati S, Fratianni F, Nazzaro F, Orlando P, et al. Pb2+ Effects on Growth, Lipids, and Protein and DNA Profiles of the Thermophilic Bacterium Thermus Thermophilus. Microorganisms. 2016; 4(4):45. https://doi.org/10.3390/microorganisms4040045
Chicago/Turabian StyleNicolaus, Barbara, Annarita Poli, Paola Di Donato, Ida Romano, Giusi Laezza, Alessia Gioiello, Sergio Ulgiati, Florinda Fratianni, Filomena Nazzaro, Pierangelo Orlando, and et al. 2016. "Pb2+ Effects on Growth, Lipids, and Protein and DNA Profiles of the Thermophilic Bacterium Thermus Thermophilus" Microorganisms 4, no. 4: 45. https://doi.org/10.3390/microorganisms4040045











