Elements in the Development of a Production Process for Modified Vaccinia Virus Ankara
Abstract
:1. Introduction
2. Modified Vaccinia Ankara
2.1. Viral Vectored Vaccines
2.2. Properties and Challenges Associated with MVA (Modified Vaccinia Virus Ankara)
3. A New Cell Substrate
3.1. Avian Finite and Continuous Cell Substrates
3.2. Duck versus Chicken
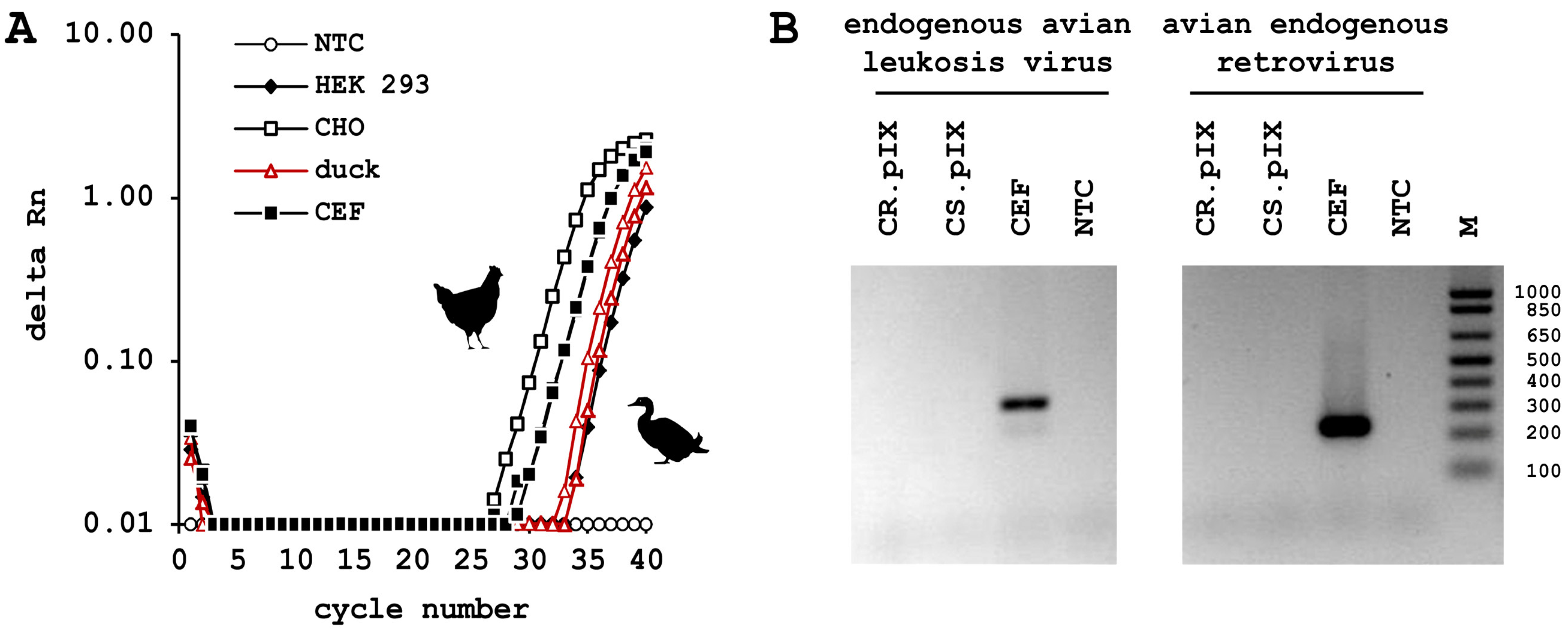
3.3. CR.pIX
- (1)
- (2)
- (3)
- Immortalization is the result of a concerted action of distinct and separate E1A and E1B gene products. This synergy necessitates concurrent events for experimental immortalization but also increases the probability against accidental tumor induction by a contamination.
- (4)
- That many of the biochemical pathways leading to immortalization by the E1 region are known may further facilitate defined-risks estimates. Briefly, by relieving inhibition of the E2F transcription factors, the E1A protein promotes cell cycle progression. However, various parallel mechanisms, including induction of DNA damage responses and augmented expression of an alternate reading frame (ARF) of the INK4a/ARF gene locus, lead into activation of apoptosis [71]. One pro-apoptotic signaling cascade is mediated by the gatekeeper of the genome, the p53 protein. This central node must be controlled by the viral E1B 55K gene product for efficient adenovirus replication [72]. Another chain of events occurs at the mitochondrial membrane. Inhibition of this pathway, that is modulated by caspase proteolysis, is exerted by the homolog to cellular Bcl2 proteins, the viral E1B 19K gene product [72,73]. The surprising property of E1A protein to sensitize cells for apoptotic stimuli and to suppress proliferation of some tumors [71,74] is investigated as therapeutic option in virotherapy [75] and E1A gene therapy for cancer patients [74,76,77].
- (5)
- For certain applications there may also be a virological advantage if E1 proteins are expressed in a host cell. Some vaccine strains are attenuated via lesions in factors that allow the virus to mask itself against the innate immunity of the cell. E1A has also been implicated in the viral defense against antiviral responses [78,79]. Although antiviral defenses within cells are activated and mediated by multiple, partially overlapping pathways [80], replication of some attenuated viruses may benefit and yields of infectious units may increase if the cellular defense is already repressed at some nodes of the redundant pathways.
4. Application
4.1. Process Development
4.2. Metabolic Properties


4.3. A Novel MVA Genotype


4.4. Current Status
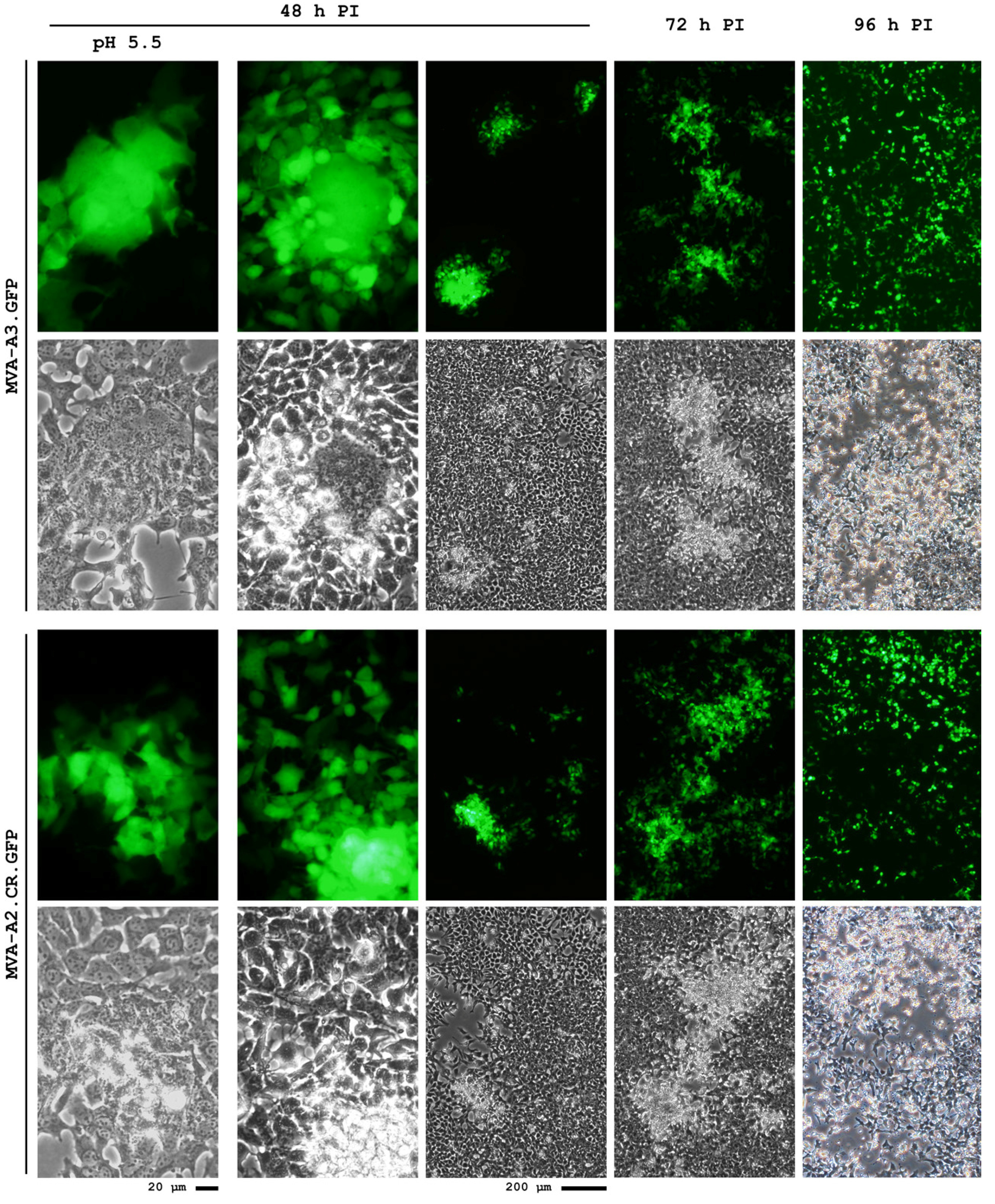
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Ehreth, J. The value of vaccination: A global perspective. Vaccine 2003, 21, 4105–4117. [Google Scholar] [CrossRef]
- Plotkin, S.A. Vaccines: The fourth century. Clin. Vaccine Immunol. 2009, 16, 1709–1719. [Google Scholar] [CrossRef]
- Lozano, R.; Naghavi, M.; Foreman, K.; Lim, S.; Shibuya, K.; Aboyans, V.; Abraham, J.; Adair, T.; Aggarwal, R.; Ahn, S.Y.; et al. Global and regional mortality from 235 causes of death for 20 age groups in 1990 and 2010: A systematic analysis for the Global Burden of Disease Study 2010. Lancet 2012, 380, 2095–2128. [Google Scholar] [CrossRef]
- Rerks-Ngarm, S.; Pitisuttithum, P.; Nitayaphan, S.; Kaewkungwal, J.; Chiu, J.; Paris, R.; Premsri, N.; Namwat, C.; de Souza, M.; Adams, E.; et al. Vaccination with ALVAC and AIDSVAX to prevent HIV-1 infection in Thailand. N. Engl. J. Med. 2009, 361, 2209–2220. [Google Scholar] [CrossRef]
- Gilbert, S.C.; Moorthy, V.S.; Andrews, L.; Pathan, A.A.; McConkey, S.J.; Vuola, J.M.; Keating, S.M.; Berthoud, T.; Webster, D.; McShane, H.; et al. Synergistic DNA-MVA prime-boost vaccination regimes for malaria and tuberculosis. Vaccine 2006, 24, 4554–4561. [Google Scholar] [CrossRef]
- Dunachie, S.J.; Hill, A.V.S. Prime-boost strategies for malaria vaccine development. J. Exp. Biol. 2003, 206, 3771–3779. [Google Scholar] [CrossRef]
- Dörsam, V.; Weimer, T.; Schmeel, A.; Hein, B.; Enssle, K.; Chumakov, K.M.; Fibi, M.R. Increased safety level of serotype 3 Sabin oral poliomyelitis vaccine lots by improved seed virus, and tissue culture and virus infection conditions. Vaccine 2000, 18, 2435–2443. [Google Scholar] [CrossRef]
- Kemper, A.R.; Davis, M.M.; Freed, G.L. Expected adverse events in a mass smallpox vaccination campaign. Eff. Clin. Pract. 2002, 5, 84–90. [Google Scholar]
- Radosević, K.; Rodriguez, A.; Lemckert, A.; Goudsmit, J. Heterologous prime-boost vaccinations for poverty-related diseases: Advantages and future prospects. Expert Rev. Vaccines 2009, 8, 577–592. [Google Scholar] [CrossRef]
- Nascimento, I.P.; Leite, L.C. Recombinant vaccines and the development of new vaccine strategies. Braz. J. Med. Biol. Res. 2012, 45, 1102–1111. [Google Scholar] [CrossRef]
- Lu, S. Heterologous prime-boost vaccination. Curr. Opin. Immunol. 2009, 21, 346–351. [Google Scholar] [CrossRef]
- Cottingham, M.G.; van Maurik, A.; Zago, M.; Newton, A.T.; Anderson, R.J.; Howard, M.K.; Schneider, J.; Skinner, M.A. Different levels of immunogenicity of two strains of Fowlpox virus as recombinant vaccine vectors eliciting T-cell responses in heterologous prime-boost vaccination strategies. Clin. Vaccine Immunol. 2006, 13, 747–757. [Google Scholar] [CrossRef]
- Lin, S.-W.; Hensley, S.E.; Tatsis, N.; Lasaro, M.O.; Ertl, H.C. Recombinant adeno-associated virus vectors induce functionally impaired transgene product-specific CD8+ T cells in mice. J. Clin. Invest. 2007, 117, 3958–3970. [Google Scholar]
- Robert-Guroff, M. Replicating and non-replicating viral vectors for vaccine development. Curr. Opin. Biotechnol. 2007, 18, 546–556. [Google Scholar] [CrossRef]
- Hodge, J.W.; Schlom, J. Comparative studies of a retrovirus versus a poxvirus vector in whole tumor-cell vaccines. Cancer Res. 1999, 59, 5106–5111. [Google Scholar]
- Coutant, F.; Frenkiel, M.-P.; Despres, P.; Charneau, P. Protective antiviral immunity conferred by a nonintegrative lentiviral vector-based vaccine. PLoS One 2008, 3, e3973. [Google Scholar]
- Cyrklaff, M.; Risco, C.; Fernández, J.J.; Jiménez, M.V.; Estéban, M.; Baumeister, W.; Carrascosa, J.L. Cryo-electron tomography of vaccinia virus. Proc. Natl. Acad. Sci. USA 2005, 102, 2772–2777. [Google Scholar] [CrossRef]
- Arif, B.M. Recent advances in the molecular biology of entomopoxviruses. J. Gen. Virol. 1995, 76, 1–13. [Google Scholar] [CrossRef]
- Gubser, C.; Hué, S.; Kellam, P.; Smith, G.L. Poxvirus genomes: A phylogenetic analysis. J. Gen. Virol. 2004, 85, 105–117. [Google Scholar] [CrossRef]
- Li, G.; Chen, N.; Feng, Z.; Buller, R.M.L.; Osborne, J.; Harms, T.; Damon, I.; Upton, C.; Esteban, D.J. Genomic sequence and analysis of a vaccinia virus isolate from a patient with a smallpox vaccine-related complication. Virol. J. 2006, 3. [Google Scholar] [CrossRef]
- Meisinger-Henschel, C.; Schmidt, M.; Lukassen, S.; Linke, B.; Krause, L.; Konietzny, S.; Goesmann, A.; Howley, P.; Chaplin, P.; Suter, M.; et al. Genomic sequence of chorioallantois vaccinia virus Ankara, the ancestor of modified vaccinia virus Ankara. J. Gen. Virol. 2007, 88, 3249–3259. [Google Scholar]
- Thézé, J.; Takatsuka, J.; Li, Z.; Gallais, J.; Doucet, D.; Arif, B.; Nakai, M.; Herniou, E.A. New insights into the evolution of Entomopoxvirinae from the complete genome sequences of four entomopoxviruses infecting Adoxophyes honmai, Choristoneura biennis, Choristoneura rosaceana, and Mythimna separa. J. Virol. 2013, 87, 7992–8003. [Google Scholar] [CrossRef]
- Jin, X.; Ramanathan, M., Jr.; Barsoum, S.; Deschenes, G.R.; Ba, L.; Binley, J.; Schiller, D.; Bauer, D.E.; Chen, D.C.; Hurley, A.; et al. Safety and immunogenicity of ALVAC vCP1452 and recombinant gp160 in newly human immunodeficiency virus type 1-infected patients treated with prolonged highly active antiretroviral therapy. J. Virol. 2002, 76, 2206–2216. [Google Scholar] [CrossRef]
- Cebere, I.; Dorrell, L.; McShane, H.; Simmons, A.; McCormack, S.; Schmidt, C.; Smith, C.; Brooks, M.; Roberts, J.E.; Darwin, S.C.; et al. Phase I clinical trial safety of DNA- and modified virus Ankara-vectored human immunodeficiency virus type 1 (HIV-1) vaccines administered alone and in a prime-boost regime to healthy HIV-1-uninfected volunteers. Vaccine 2006, 24, 417–425. [Google Scholar] [CrossRef]
- Vuola, J.M.; Keating, S.; Webster, D.P.; Berthoud, T.; Dunachie, S.; Gilbert, S.C.; Hill, A.V. Differential immunogenicity of various heterologous prime-boost vaccine regimens using DNA and viral vectors in healthy volunteers. J. Immunol. 2005, 174, 449–455. [Google Scholar]
- Webster, D.P.; Dunachie, S.; Vuola, J.M.; Berthoud, T.; Keating, S.; Laidlaw, S.M.; McConkey, S.J.; Poulton, I.; Andrews, L.; Andersen, R.F.; et al. Enhanced T cell-mediated protection against malaria in human challenges by using the recombinant poxviruses FP9 and modified vaccinia virus Ankara. Proc. Natl. Acad. Sci. USA 2005, 102, 4836–4841. [Google Scholar] [CrossRef]
- McShane, H.; Behboudi, S.; Goonetilleke, N.; Brookes, R.; Hill, A.V. Protective immunity against Mycobacterium tuberculosis induced by dendritic cells pulsed with both CD8+- and CD4+-T-cell epitopes from antigen 85A. Infect. Immun. 2002, 70, 1623–1626. [Google Scholar] [CrossRef]
- Kantoff, P.W.; Schuetz, T.J.; Blumenstein, B.A.; Glode, L.M.; Bilhartz, D.L.; Wyand, M.; Manson, K.; Panicali, D.L.; Laus, R.; Schlom, J.; et al. Overall survival analysis of a phase II randomized controlled trial of a Poxviral-based PSA-targeted immunotherapy in metastatic castration-resistant prostate cancer. J. Clin. Oncol. 2010, 28, 1099–1105. [Google Scholar] [CrossRef]
- Dangoor, A.; Lorigan, P.; Keilholz, U.; Schadendorf, D.; Harris, A.; Ottensmeier, C.; Smyth, J.; Hoffmann, K.; Anderson, R.; Cripps, M.; et al. Clinical and immunological responses in metastatic melanoma patients vaccinated with a high-dose poly-epitope vaccine. Cancer Immunol. Immunother. 2010, 59, 863–873. [Google Scholar] [CrossRef]
- Mayr, A.; Hochstein-Mintzel, V; Stickl, H. Abstammung, Eigenschaften und Verwendung des attenuierten Vaccinia-Stammes MVA. Infection 1975, 3, 6–14. (in German). [Google Scholar] [CrossRef]
- Mayr, A.; Munz, E. Changes in the vaccinia virus through continuing passages in chick embryo fibroblast cultures. Zentralbl Bakteriol Orig. 1964, 195, 24–35. (in German). [Google Scholar]
- Meyer, H.; Sutter, G.; Mayr, A. Mapping of deletions in the genome of the highly attenuated vaccinia virus MVA and their influence on virulence. J. Gen. Virol. 1991, 72, 1031–1038. [Google Scholar] [CrossRef]
- Carroll, M.W.; Moss, B. Host range and cytopathogenicity of the highly attenuated MVA strain of vaccinia virus: propagation and generation of recombinant viruses in a nonhuman mammalian cell line. Virology 1997, 238, 198–211. [Google Scholar] [CrossRef]
- Blanchard, T.J.; Alcami, A.; Andrea, P.; Smith, G.L. Modified vaccinia virus Ankara undergoes limited replication in human cells and lacks several immunomodulatory proteins: Implications for use as a human vaccine. J. Gen. Virol. 1998, 79, 1159–1167. [Google Scholar]
- Drexler, I.; Heller, K.; Wahren, B.; Erfle, V.; Sutter, G. Highly attenuated modified vaccinia virus Ankara replicates in baby hamster kidney cells, a potential host for virus propagation, but not in various human transformed and primary cells. J. Gen. Virol. 1998, 79, 347–352. [Google Scholar]
- Sutter, G.; Moss, B. Nonreplicating vaccinia vector efficiently expresses recombinant genes. Proc. Natl. Acad. Sci. USA 1992, 89, 10847–10851. [Google Scholar] [CrossRef]
- Sutter, G.; Wyatt, L.S.; Foley, P.L.; Bennink, J.R.; Moss, B. A recombinant vector derived from the host range-restricted and highly attenuated MVA strain of vaccinia virus stimulates protective immunity in mice to influenza virus. Vaccine 1994, 12, 1032–1040. [Google Scholar] [CrossRef]
- Drillien, R.; Spehner, D.; Hanau, D. Modified vaccinia virus Ankara induces moderate activation of human dendritic cells. J. Gen. Virol. 2004, 85, 2167–2175. [Google Scholar] [CrossRef]
- Liu, L.; Chavan, R.; Feinberg, M.B. Dendritic cells are preferentially targeted among hematolymphocytes by Modified Vaccinia Virus Ankara and play a key role in the induction of virus-specific T cell responses in vivo. BMC Immunol. 2008, 9, 15. [Google Scholar] [CrossRef]
- Cottingham, M.G.; Carroll, M.W. Recombinant MVA vaccines: Dispelling the myths. Vaccine 2013, 31, 4247–4251. [Google Scholar] [CrossRef]
- Kremer, M.; Volz, A.; Kreijtz, J.H.C.M.; Fux, R.; Lehmann, M.H.; Sutter, G. Easy and Efficient Protocols for Working with Recombinant Vaccinia Virus MVA. In Vaccinia Virus and Poxvirology; Humana Press: New York, NY, USA, 2012; pp. 59–92. [Google Scholar]
- Smith, G.L.; Moss, B. Infectious poxvirus vectors have capacity for at least 25,000 base pairs of foreign DNA. Gene 1983, 25, 21–28. [Google Scholar] [CrossRef]
- Stickl, H.; Hochstein-Mintzel, V.; Mayr, A.; Huber, H.C.; Schäfer, H.; Holzner, A. MVA vaccination against smallpox: clinical tests with an attenuated live vaccinia virus strain (MVA) (author’s transl). Dtsch. Med. Wochenschr. 1974, 99, 2386–2392. (in German). [Google Scholar] [CrossRef]
- Mayr, A. Smallpox vaccination and bioterrorism with pox viruses. Comp. Immunol. Microbiol. Infect. Dis. 2003, 26, 423–430. [Google Scholar] [CrossRef]
- Coulibaly, S.; Brühl, P.; Mayrhofer, J.; Schmid, K.; Gerencer, M.; Falkner, F.G. The nonreplicating smallpox candidate vaccines defective vaccinia Lister (dVV-L) and modified vaccinia Ankara (MVA) elicit robust long-term protection. Virology 2005, 341, 91–101. [Google Scholar] [CrossRef]
- Cox, H.R. Active immunization against poliomyelitis. Bull. N. Y. Acad. Med. 1953, 29, 943–960. [Google Scholar]
- Hess, R.D.; Weber, F.; Watson, K.; Schmitt, S. Regulatory, biosafety and safety challenges for novel cells as substrates for human vaccines. Vaccine 2012, 30, 2715–2727. [Google Scholar] [CrossRef]
- Jacobs, J.P.; Jones, C.M.; Baille, J.P. Characteristics of a human diploid cell designated MRC-5. Nature 1970, 227, 168–170. [Google Scholar] [CrossRef]
- Enserink, M. Influenza. Crisis underscores fragility of vaccine production system. Science 2004, 306, 385. [Google Scholar] [CrossRef]
- Uscher-Pines, L.; Barnett, D.J.; Sapsin, J.W.; Bishai, D.M.; Balicer, R.D. A systematic analysis of influenza vaccine shortage policies. Public Health 2008, 122, 183–191. [Google Scholar] [CrossRef]
- Jordan, I.; Vos, A.; Beilfuss, S.; Neubert, A.; Breul, S.; Sandig, V. An avian cell line designed for production of highly attenuated viruses. Vaccine 2009, 27, 748–756. [Google Scholar] [CrossRef]
- Böni, J.; Stalder, J.; Reigel, F.; Schüpbach, J. Detectionof reverse transcriptase activity in live attenuated virus vaccines. Clin. Diagn. Virol. 1996, 5, 43–53. [Google Scholar] [CrossRef]
- Weissmahr, R.N.; Schüpbach, J.; Böni, J. Reverse transcriptase activity in chicken embryo fibroblast culture supernatants is associated with particles containing endogenous avian retrovirus EAV-0 RNA. J. Virol. 1997, 71, 3005–3012. [Google Scholar]
- Herniou, E.; Martin, J.; Miller, K.; Cook, J.; Wilkinson, M.; Tristem, M. Retroviral diversity and distribution in vertebrates. J. Virol. 1998, 72, 5955–5966. [Google Scholar]
- Lander, E.S.; Linton, L.M.; Birren, B.; Nusbaum, C.; Zody, M.C.; Baldwin, J.; Devon, K.; Dewar, K.; Doyle, M.; FitzHugh, W.; et al. Initial sequencing and analysis of the human genome. Nature 2001, 409, 860–921. [Google Scholar] [CrossRef]
- Belshaw, R.; Pereira, V.; Katzourakis, A.; Talbot, G.; Paces, J.; Burt, A.; Tristem, M. Long-term reinfection of the human genome by endogenous retroviruses. Proc. Natl. Acad. Sci. USA 2004, 101, 4894–4899. [Google Scholar]
- Huda, A.; Polavarapu, N.; Jordan, I.K.; McDonald, J.F. Endogenous retroviruses of the chicken genome. Biol. Direct 2008, 3, 9. [Google Scholar] [CrossRef]
- Huang, Y.; Li, Y.; Burt, D.W.; Chen, H.; Zhang, Y.; Qian, W.; Kim, H.; Gan, S.; Zhao, Y.; Li, J.; et al. The duck genome and transcriptome provide insight into an avian influenza virus reservoir species. Nat. Genet. 2013, 45, 776–783. [Google Scholar] [CrossRef]
- Reverse transcriptase activity in chicken-cell derived vaccine. Wkly. Epidemiol. Rec. 1998, 73, 209–212.
- Payne, L.N.; Howes, K.; Gillespie, A.M.; Smith, L.M. Host range of Rous sarcoma virus pseudotype RSV(HPRS-103) in 12 avian species: Support for a new avian retrovirus envelope subgroup, designated J. J. Gen. Virol. 1992, 73, 2995–2997. [Google Scholar] [CrossRef]
- Shoyab, M.; Baluda, M.A. Homology between avian oncornavirus RNAs and DNA from several avian species. J. Virol. 1975, 16, 1492–1502. [Google Scholar]
- Philipp, H.-C.; Kolla, I. Laboratory host systems for extraneous agent testing in avian live virus vaccines: Problems encountered. Biol. J. Int. Assoc. Biol. Stand. 2010, 38, 350–351. [Google Scholar]
- Grachev, V.; Magrath, D.; Griffiths, E. WHO requirements for the use of animal cells as in vitro substrates for the production of biologicals (Requirements for biological susbstances no. 50). Biol. J. Int. Assoc. Biol. Stand. 1998, 26, 175–193. [Google Scholar]
- Yang, H.; Zhang, L.; Galinski, M. A probabilistic model for risk assessment of residual host cell DNA in biological products. Vaccine 2010, 28, 3308–3311. [Google Scholar] [CrossRef]
- Wierenga, D.E.; Cogan, J.; Petricciani, J.C. Administration of tumor cell chromatin to immunosuppressed and non-immunosuppressed non-human primates. Biologicals 1995, 23, 221–224. [Google Scholar] [CrossRef]
- Jordan, I.; Northoff, S.; Thiele, M.; Hartmann, S.; Horn, D.; Höwing, K.; Bernhardt, H.; Oehmke, S.; von Horsten, H.; Rebeski, D.; et al. A chemically defined production process for highly attenuated poxviruses. Biologicals 2011, 39, 50–58. [Google Scholar] [CrossRef]
- Graham, F.L.; Smiley, J.; Russell, W.C.; Nairn, R. Characteristics of a human cell line transformed by DNA from human adenovirus type 5. J. Gen. Virol. 1977, 36, 59–74. [Google Scholar] [CrossRef]
- Fallaux, F.J.; Bout, A.; van der Velde, I.; van den Wollenberg, D.J.; Hehir, K.M.; Keegan, J.; Auger, C.; Cramer, S.J.; van Ormondt, H.; van der Eb, A.J.; et al. New helper cells and matched early region 1-deleted adenovirus vectors prevent generation of replication-competent adenoviruses. Hum. Gene Ther. 1998, 9, 1909–1917. [Google Scholar] [CrossRef]
- “Designe” Cells as Substrates for the Manufacture of Viral Vaccines. Available online: http://www.fda.gov/ohrms/dockets/ac/01/briefing/3750b1_01.htm (accessed on 27 September 2013).
- Munoz, F.M.; Piedra, P.A.; Demmler, G.J. Disseminated adenovirus disease in immunocompromised and immunocompetent children. Clin. Infect. Dis. 1998, 27, 1194–1200. [Google Scholar]
- Frisch, S.M.; Mymryk, J.S. Adenovirus-5 E1A: Paradox and paradigm. Nat. Rev. Mol. Cell Biol. 2002, 3, 441–452. [Google Scholar] [CrossRef]
- Berk, A.J. Recent lessons in gene expression, cell cycle control, and cell biology from adenovirus. Oncogene 2005, 24, 7673–7685. [Google Scholar] [CrossRef]
- Henry, H.; Thomas, A.; Shen, Y.; White, E. Regulation of the mitochondrial checkpoint in p53-mediated apoptosis confers resistance to cell death. Oncogene 2002, 21, 748–760. [Google Scholar] [CrossRef]
- Frisch, S.M. E1A as a tumor suppressor gene: Commentary re S. Madhusudan et al. A multicenter Phase I gene therapy clinical trial involving intraperitoneal administration of E1A-lipid complex in patients with recurrent epithelial ovarian cancer overexpressing HER-2/neu oncogene. Clin. Cancer Res. 2004, 10, 2905–2907. [Google Scholar] [CrossRef]
- Choi, I.-K.; Yun, C.-O. Recent developments in oncolytic adenovirus-based immunotherapeutic agents for use against metastatic cancers. Cancer Gene Ther. 2013, 20, 70–76. [Google Scholar] [CrossRef]
- Madhusudan, S.; Tamir, A.; Bates, N.; Flanagan, E.; Gore, M.E.; Barton, D.P.J.; Harper, P.; Seckl, M.; Thomas, H.; Lemoine, N.R.; et al. A multicenter Phase I gene therapy clinical trial involving intraperitoneal administration of E1A-lipid complex in patients with recurrent epithelial ovarian cancer overexpressing HER-2/neu oncogene. Clin. Cancer Res. 2004, 10, 2986–2996. [Google Scholar]
- Yamaguchi, H.; Chen, C.-T.; Chou, C.-K.; Pal, A.; Bornmann, W.; Hortobagyi, G.N.; Hung, M.-C. Adenovirus 5 E1A enhances histone deacetylase inhibitors-induced apoptosis through Egr-1-mediated Bim upregulation. Oncogene 2010, 29, 5619–5629. [Google Scholar] [CrossRef]
- Anderson, K.P.; Fennie, E.H. Adenovirus early region 1A modulation of interferon antiviral activity. J. Virol. 1987, 61, 787–795. [Google Scholar]
- Leonard, G.T.; Sen, G.C. Restoration of interferon responses of adenovirus E1A-expressing HT1080 cell lines by overexpression of p48 protein. J. Virol. 1997, 71, 5095–5101. [Google Scholar]
- Unterholzner, L.; Bowie, A.G. The interplay between viruses and innate immune signaling: Recent insights and therapeutic opportunities. Biochem. Pharmacol. 2008, 75, 589–602. [Google Scholar] [CrossRef]
- Lohr, V.; Rath, A.; Genzel, Y.; Jordan, I.; Sandig, V.; Reichl, U. New avian suspension cell lines provide production of influenza virus and MVA in serum-free media: studies on growth, metabolism and virus propagation. Vaccine 2009, 27, 4975–4982. [Google Scholar] [CrossRef]
- Lohr, V.; Genzel, Y.; Jordan, I.; Katinger, D.; Mahr, S.; Sandig, V.; Reichl, U. Live attenuated influenza viruses produced in a suspension process with avian AGE1.CR.pIX cells. BMC Biotechnol. 2012, 12, 79. [Google Scholar]
- Meiser, A.; Boulanger, D.; Sutter, G.; Krijnse Locker, J. Comparison of virus production in chicken embryo fibroblasts infected with the WR, IHD-J and MVA strains of vaccinia virus: IHD-J is most efficient in trans-Golgi network wrapping and extracellular enveloped virus release. J. Gen. Virol. 2003, 84, 1383–1392. [Google Scholar] [CrossRef]
- Jordan, I.; Woods, N.; Whale, G.; Sandig, V. Production of a viral-vectored vaccine candidate against tuberculosis. BioProcess Int. 2012, 10, 46–55. [Google Scholar]
- Theiler, M.; Smith, H.H. The use of yellow fever virus modified by in vitro cultivation for human immunization. J. Exp. Med. 1937, 65, 787–800. [Google Scholar] [CrossRef]
- Harper, J.M.; Wang, M.; Galecki, A.T.; Ro, J.; Williams, J.B.; Miller, R.A. Fibroblasts from long-lived bird species are resistant to multiple forms of stress. J. Exp. Biol. 2011, 214, 1902–1910. [Google Scholar] [CrossRef]
- Wu, G.; Bazer, F.W.; Burghardt, R.C.; Johnson, G.A.; Kim, S.W.; Knabe, D.A.; Li, P.; Li, X.; McKnight, J.R.; Satterfield, M.C.; et al. Proline and hydroxyproline metabolism: Implications for animal and human nutrition. Amino Acids 2011, 40, 1053–1063. [Google Scholar] [CrossRef]
- Singer, M.A. Do mammals, birds, reptiles and fish have similar nitrogen conserving systems? Com. Biochem. Physiol. B Biochem. Mol. Biol. 2003, 134, 543–558. [Google Scholar] [CrossRef]
- Cortin, V.; Thibault, J.; Jacob, D.; Garnier, A. High-titer adenovirus vector production in 293S cell perfusion culture. Biotechnol. Prog. 2004, 20, 858–863. [Google Scholar] [CrossRef]
- Hädicke, O.; Lohr, V.; Genzel, Y.; Reichl, U.; Klamt, S. Evaluating differences of metabolic performances: Statistical methods and their application to animal cell cultivations. Biotechnol. Bioeng. 2013, 110, 2633–2642. [Google Scholar] [CrossRef]
- Frensing, T.; Heldt, F.S.; Pflugmacher, A.; Behrendt, I.; Jordan, I.; Flockerzi, D.; Genzel, Y.; Reichl, U. Continuous influenza virus production in cell culture shows a periodic accumulation of defective interfering particles. PLoS One 2013, 8, e72288. [Google Scholar] [CrossRef]
- Frey, T.K.; Hemphill, M.L. Generation of defective-interfering particles by rubella virus in Vero cells. Virology 1988, 164, 22–29. [Google Scholar] [CrossRef]
- Kilburn, D.G.; van Wezel, A.L. The effect of growth rate in continuous-flow cultures on the replication of rubella virus in BHK cells. J. Gen. Virol. 1970, 9, 1–7. [Google Scholar] [CrossRef]
- Jordan, I.; Horn, D.; John, K.; Sandig, V. A genotype of modified vaccinia Ankara (MVA) that facilitates replication in suspension cultures in chemically defined medium. Viruses 2013, 5, 321–339. [Google Scholar] [CrossRef]
- Heljasvaara, R.; Rodríguez, D.; Risco, C.; Carrascosa, J.L.; Esteban, M.; Rodríguez, J.R. The major core protein P4a (A10L gene) of vaccinia virus is essential for correct assembly of viral DNA into the nucleoprotein complex to form immature viral particles. J. Virol. 2001, 75, 5778–5795. [Google Scholar] [CrossRef]
- Byrd, C.M.; Bolken, T.C.; Hruby, D.E. The vaccinia virus I7L gene product is the core protein proteinase. J. Virol. 2002, 76, 8973–8976. [Google Scholar] [CrossRef]
- Kato, S.E.M.; Strahl, A.L.; Moussatche, N.; Condit, R.C. Temperature-sensitive mutants in the vaccinia virus 4b virion structural protein assemble malformed, transcriptionally inactive intracellular mature virions. Virology 2004, 330, 127–146. [Google Scholar] [CrossRef]
- Yeh, W.W.; Moss, B.; Wolffe, E.J. The vaccinia virus A9L gene encodes a membrane protein required for an early step in virion morphogenesis. J. Virol. 2000, 74, 9701–9711. [Google Scholar] [CrossRef]
- Harrison, S.C.; Alberts, B.; Ehrenfeld, E.; Enquist, L.; Fineberg, H.; McKnight, S.L.; Moss, B.; O’Donnell, M.; Ploegh, H.; Schmid, S.L.; et al. Discovery of antivirals against smallpox. Proc. Natl. Acad. Sci. USA 2004, 101, 11178–11192. [Google Scholar] [CrossRef]
- Moss, B. Poxvirus cell entry: how many proteins does it take? Viruses 2012, 4, 688–707. [Google Scholar] [CrossRef]
- Blasco, R.; Sisler, J.R.; Moss, B. Dissociation of progeny vaccinia virus from the cell membrane is regulated by a viral envelope glycoprotein: Effect of a point mutation in the lectin homology domain of the A34R gene. J. Virol. 1993, 67, 3319–3325. [Google Scholar]
- Katz, E.; Wolffe, E.; Moss, B. Identification of second-site mutations that enhance release and spread of vaccinia virus. J. Virol. 2002, 76, 11637–11644. [Google Scholar] [CrossRef]
- Husain, M.; Weisberg, A.S.; Moss, B. Resistance of a vaccinia virus A34R deletion mutant to spontaneous rupture of the outer membrane of progeny virions on the surface of infected cells. Virology 2007, 366, 424–432. [Google Scholar] [CrossRef]
- McIntosh, A.A.; Smith, G.L. Vaccinia virus glycoprotein A34R is required for infectivity of extracellular enveloped virus. J. Virol. 1996, 70, 272–281. [Google Scholar]
- Ward, B.M. Visualization and characterization of the intracellular movement of vaccinia virus intracellular mature virions. J. Virol. 2005, 79, 4755–4763. [Google Scholar] [CrossRef]
- Chakrabarti, S.; Sisler, J.R.; Moss, B. Compact, synthetic, vaccinia virus early/late promoter for protein expression. BioTechniques 1997, 23, 1094–1097. [Google Scholar]
- Gallego-Gómez, J.C.; Risco, C.; Rodríguez, D.; Cabezas, P.; Guerra, S.; Carrascosa, J.L.; Esteban, M. Differences in virus-induced cell morphology and in virus maturation between MVA and other strains (WR, Ankara, and NYCBH) of vaccinia virus in infected human cells. J. Virol. 2003, 77, 10606–10622. [Google Scholar] [CrossRef]
- Sancho, M.C.; Schleich, S.; Griffiths, G.; Krijnse-Locker, J. The block in assembly of modified vaccinia virus Ankara in HeLa cells reveals new insights into vaccinia virus morphogenesis. J. Virol. 2002, 76, 8318–8334. [Google Scholar] [CrossRef]
- Wolff, M.W.; Siewert, C.; Lehmann, S.; Hansen, S.P.; Djurup, R.; Faber, R.; Reichl, U. Capturing of cell culture-derived modified Vaccinia Ankara virus by ion exchange and pseudo-affinity membrane adsorbers. Biotechnol. Bioeng. 2010, 105, 761–769. [Google Scholar]
- Wolff, M.W.; Siewert, C.; Hansen, S.P.; Faber, R.; Reichl, U. Purification of cell culture-derived modified Vaccinia Ankara virus by pseudo-affinity membrane adsorbers and hydrophobic interaction chromatography. Biotechnol. Bioeng. 2010, 107, 312–320. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Jordan, I.; Lohr, V.; Genzel, Y.; Reichl, U.; Sandig, V. Elements in the Development of a Production Process for Modified Vaccinia Virus Ankara. Microorganisms 2013, 1, 100-121. https://doi.org/10.3390/microorganisms1010100
Jordan I, Lohr V, Genzel Y, Reichl U, Sandig V. Elements in the Development of a Production Process for Modified Vaccinia Virus Ankara. Microorganisms. 2013; 1(1):100-121. https://doi.org/10.3390/microorganisms1010100
Chicago/Turabian StyleJordan, Ingo, Verena Lohr, Yvonne Genzel, Udo Reichl, and Volker Sandig. 2013. "Elements in the Development of a Production Process for Modified Vaccinia Virus Ankara" Microorganisms 1, no. 1: 100-121. https://doi.org/10.3390/microorganisms1010100




