Quorum Sensing in Some Representative Species of Halomonadaceae
Abstract
:1. Introduction
2. Results and Discussion
2.1. Detection of AHLs in the Halomonadaceae Family
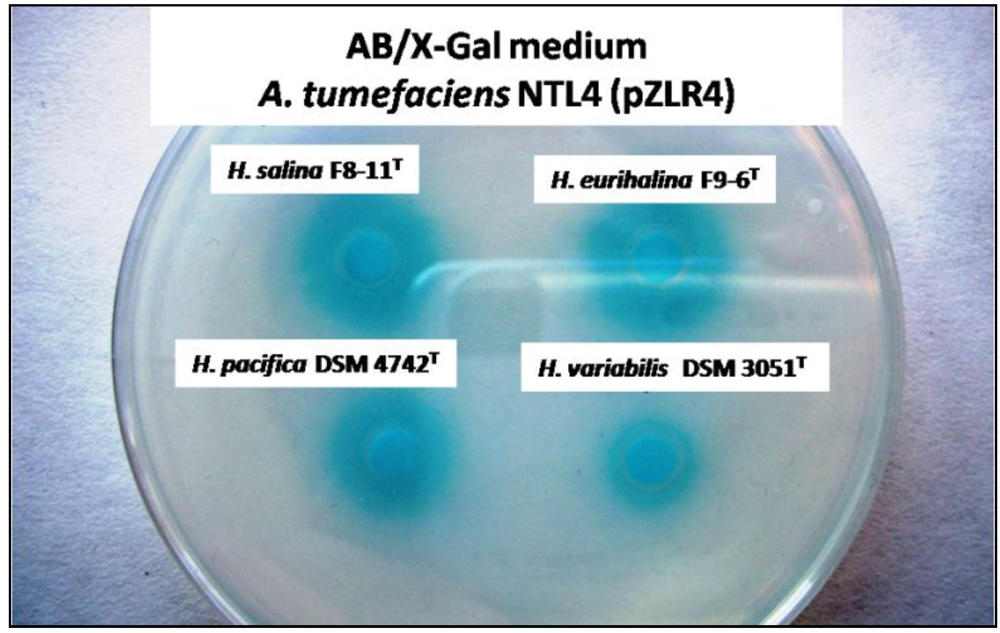
2.2. Characterization of the AHLs

2.3. Distribution of the Autoinducer Synthase Gene

3. Experimental Section
| Species | Strain | Ecological Niches |
|---|---|---|
| 1. Carnimonas nigrificans | CTCBS1T | Cured meat, Spain |
| 2. Chromohalobacter salexigens | DSM 3043T | Solar saltern, Netherlands |
| 3. Cobetia marina | 219T | Sea water, USA |
| 4. Halomonas alimentaria | YKJ-16T | Jeotgal, a traditional Korean fermented seafood, Korea |
| 5. H. almeriensis | M8T | Solar saltern, south-east Spain |
| 6. H. anticariensis | FP35T | Saline wetland, southern Spain |
| 7. H. aquamarina | 558T | Pacific ocean |
| 8. H. campaniensis | 5AGT | Mineral pool, Italy |
| 9. H. cerina | SP4T | Saline soil, Spain |
| 10. H. denitrificans | M29T | Saline water, Korea |
| 11. H. desiderata | FB2T | Municipal sewage works, Germany |
| 12. H. elongata | 1H9T | Solar saltern, Netherlands |
| 13. H. eurihalina | F9-6T | Saline soil, Spain |
| 14. H. fontilapidosi | CR-5T | Saline soil, southern Spain |
| 15. H. gudaonensis | SL014B-69T | Saline soil contaminated by crude oil, China |
| 16. H. halmophila | ACAM 71T | Dead Sea, Israel |
| 17. H. halodenitrificans | ATCC 13511T | Meat in brine |
| 18. H. halodurans | DSM 5160T | Great Bay estuary, USA |
| 19. H. koreensis | SS20T | Solar saltern, Korea |
| 20. H. magadiensis | 21 MIT | Soda lake, East-African Rift Valley |
| 21. H. maura | S-31T | Saltern, Morocco |
| 22. H. meridiana | ACAM 246T | Saline lake, Antarctic |
| 23. H. mongoliensis | Z-7009T | Soda lake, Mongolia |
| 24. H. nitroreducens | 11-ST | Solar saltern, Chile |
| 25. H. organivorans | G-16.1T | Hypersaline habitats contaminated by aromatic organic compounds, southern Spain |
| 26. H. pacifica | DSM 4742T | Pacific ocean |
| 27. H. pantelleriensis | AAPT | Hard lake sand, Pantelleria island, Italy |
| 28. H. ramblicola | RS-16T | Saline soil, south-east Spain |
| 29. H. rifensis | HK-31T | Solar saltern, Morocco |
| 30. H. saccharevitans | AJ275T | Salt lake and a subterranean saline well, China |
| 31. H. salina | F8-11T | Saline soil, Spain |
| 32. H. shengliensis | SLO14B-85T | Saline soil contaminated with crude oil, China |
| 33. H. stenophila | N12T | Saline soil, Spain |
| 34. H. subglaciescola | ACAM 12T | Antarctic hypersaline, meromictic lake |
| 35. H. variabilis | DSM 3051T | Great Salt Lake, USA |
| 36. H. ventosae | Al-12T | Saline soil, south-eastern Spain |
| 37. Halotalea alkalilenta | AW-7T | Alkaline olive-mill waste (alpechin), Greece |
| 38. Kushneria avicenniae | MW2aT | Salty leaves of Avicennia germnans trees growing near solar salterns, Puerto Rico |
| 39. K. indalinina | CG2.1T | Solar saltern, south-east Spain |
| 40. K. marisflavi | SW32T | Water from the Yellow Sea, Korea |
| 41. Modicisalibacter tunisiensis | LIT2T | Oilfield-water injection sample, southern Tunisia |
| 42. Salinicola halophilus | CG4.1T | Solar saltern, south-east Spain |
| 43. S. salarius | M27T | Saline water, Korea |
4. Conclusions
Acknowledgments
References
- Brock, T. Halophilic-blue-green algae. Arch. Microbiol. 1976, 107, 109–111. [Google Scholar] [CrossRef]
- Rodríguez-Valera, F. Characteristics and microbial ecology of hypersaline environments. In Halophilic Bacteria; Rodríguez-Valera, F., Ed.; CRC Press: Boca Raton, Florida, USA, 1988; Volume 1, pp. 3–30. [Google Scholar]
- Cifuentes, A.; Antón, J.; De Wit, R.; Rodríguez-Valera, F. Diversity of Bacteria and Archaea in sulphate-reducing enrichment cultures inoculated from serial dilution of Zostera noltii rhizosphere samples. Environ. Microbiol. 2003, 5, 754–764. [Google Scholar] [CrossRef]
- González-Toril, E.; Llobet-Brossa, E.; Casamayor, E.O.; Amann, R.; Amils, R. Microbial ecology of an extreme acidic environment, the Tinto River. Appl. Environ. Microbiol. 2003, 69, 4853–4865. [Google Scholar] [CrossRef]
- Horikoshi, K.; Grant, W.D. Extremophiles: Microbial Life in Extreme Environments; Wiley-Liss: New York, USA, 1998. [Google Scholar]
- Ventosa, A. Unusual micro-organisms from unusual habitats: Hypersaline environments. In Prokaryotic Diversity: Mechanisms and Significance; Logan, N.A., Lappin-Scott, H.M., Oyston, P.C.F., Eds.; Cambridge University Press: New York, USA, 2006; pp. 223–253. [Google Scholar]
- Franzmann, P.D.; Wehmeyer, U.; Stackebrandt, E. Halomonadaceae fam. nov., a new family of the Class Proteobacteria to accommodate the genera Halomonas and Deleya. Syst. Appl. Microbiol. 1988, 11, 16–19. [Google Scholar] [CrossRef]
- Ventosa, A.; Nieto, J.J.; Oren, A. Biology of moderately halophilic aerobic bacteria. Microbiol. Mol. Biol. Rev. 1998, 62, 504–544. [Google Scholar]
- Euzéby, J.P. List of Prokaryotic Names with Standing in Nomenclature. 2012. Available online: http://www.bacterio.cict.fr/ (accessed on 15 November 2012).
- Kushner, D.; Kamekura, M. Physiology of halophilic eubacteria. In Halophilic Bacteria; Rodríguez-Valera, F., Ed.; CRC Press: Boca Raton, Florida, USA, 1988; Volume 1, pp. 87–103. [Google Scholar]
- Ventosa, A.; Mellado, E.; Sánchez-Porro, C.; Márquez, M.C. Halophilic and halotolerant micro-organisms from soils. In Microbiology of Extreme Soils; Dion, P., Nautiyal, C., Eds.; Springer-Verlag: Heidelberg, Germany, 2008; Volume 13, pp. 87–115. [Google Scholar]
- Nieto, J.J.; Carmen, V.M. Synthesis of osmoprotectants by moderately halophilic bacteria: Genetic and applied aspects. In Recent Research and Development in Endocrinology; Transworld Research Network: Kerala, India, 2002; pp. 403–418. [Google Scholar]
- De la Haba, R.R.; Sánchez-Porro, C.; Márquez, M.C.; Ventosa, A. Taxonomy of Halophiles. In Extremophiles Handbook; Horikoshi, K., Ed.; Springer: New York, NY, USA, 2011. [Google Scholar]
- Oren, A. Industrial and environmental applications of halophilic microorganisms. Environ. Technol. 2010, 31, 825–834. [Google Scholar] [CrossRef]
- Kaye, J.Z.; Baross, J.A. High incidence of halotolerant bacteria in Pacific hydrothermal-vent and pelagic environments. FEMS Microbiol. Ecol. 2000, 32, 249–260. [Google Scholar] [CrossRef]
- Oueriaghli, N.; González-Domenech, C.M.; Martínez-Checa, F.; Muyzer, M.; Quesada, E.; Béjar, V. Estudio molecular de la diversidad del género Halomonas en Rambla Salada mediante DGGE, CARD-FISH y análisis multivariable. In Presented at the XIV Reunión del Grupo de Taxonomía Filogenia y Biodiversidad Microbiana (SEM), Granada, España, 10-11 May 2012.
- Parker, C.T.; Sperandio, V. Cell-to-cell signalling during pathogenesis. Cell. Microbiol. 2009, 11, 363–369. [Google Scholar] [CrossRef]
- Williams, P. Quorum sensing, communication and cross-kingdom signalling in the bacterial world. Microbiology 2007, 153, 3923–3938. [Google Scholar] [CrossRef]
- Jung, K.; Fried, L.; Behr, S.; Heermann, R. Histidine kinases and response regulators in networks. Curr. Opin. Microbiol. 2012, 15, 118–124. [Google Scholar]
- González, J.E.; Marketon, M.M. Quorum sensing in nitrogen-fixing rhizobia. Microbiol. Mol. Biol. Rev. 2003, 67, 574–592. [Google Scholar] [CrossRef]
- Whitehead, N.A.; Barnard, A.M.; Slater, H.; Simpson, N.J.; Salmond, G.P. Quorum-sensing in Gram-negative bacteria. FEMS Microbiol. Rev. 2001, 25, 365–404. [Google Scholar] [CrossRef]
- Eberhard, A.; Longin, T.; Widrig, C.A.; Stranick, S.J. Synthesis of the lux gene autoinducer in Vibrio fischeri is positively autoregulated. Arch. Microbiol. 1991, 155, 294–297. [Google Scholar] [CrossRef]
- Fuqua, W.C.; Winans, S.C.; Greenberg, E.P. Quorum sensing in bacteria: the LuxR-LuxI family of cell density-responsive transcriptional regulators. J. Bacteriol. 1994, 176, 269–275. [Google Scholar]
- Swift, S.; Williams, P.; Stewart, G.S.A.B. N-acyl homoserine lactones and quorum sensing in proteobacteria. In Cell-Cell Signaling in Bacteria.; Dunny, G.M., Winans, S.C., Eds.; American Society of Microbiology Press: Washington, DC, USA, 1999; pp. 291–314. [Google Scholar]
- Case, R.J.; Labbate, M.; Kjelleberg, S. AHL-driven quorum-sensing circuits: their frequency and function among the Proteobacteria. The ISME J. 2008, 2, 345–349. [Google Scholar] [CrossRef]
- Llamas, I.; Quesada, E.; Martínez-Cánovas, M.J.; Gronquist, M.; Eberhard, A.; González, J.E. Quorum sensing in halophilic bacteria: Detection of N-acyl-homoserine lactones in the exopolysaccharide-producing species of Halomonas. Extremophiles. 2005, 9, 333–341. [Google Scholar] [CrossRef]
- Tahrioui, A.; Quesada, E.; Llamas, I. The hanR/hanI quorum-sensing system of Halomonas anticariensis, a moderately halophilic bacterium. Microbiology 2011, 157, 3378–3387. [Google Scholar] [CrossRef]
- Tahrioui, A.; Quesada, E.; Llamas, I. Genetic and phenotypic analysis of the GacS/GacA system in the moderate halophile Halomonas anticariensis. Microbiology 2013, 159, 461–473. [Google Scholar]
- Amjres, H.; Béjar, V.; Quesada, E.; Abrini, J.; Llamas, I. Halomonas rifensis sp. nov., an exopolysaccharide-producing, halophilic bacterium isolated from a solar saltern. Int. J. Syst. Evol. Microbiol. 2011, 61, 2600–2605. [Google Scholar] [CrossRef]
- Dobson, S.J.; Franzmann, P.D. Unification of the genera Deleya (Baumann et al. 1983), Halomonas (Vreeland et al. 1980), and Halovibrio (Fendrich 1988) and the Species Paracoccus halodenitrificans (Robinson and Gibbons 1952) into a single genus, Halomonas, and placement of the Genus Zymobacter in the family Halomonadaceae. Int. J. Syst. Bacteriol. 1996, 46, 550–558. [Google Scholar] [CrossRef]
- González-Domenech, C.M.; Béjar, V.; Martínez-Checa, F.; Quesada, E. Halomonas nitroreducens sp. nov., a novel nitrate- and nitrite-reducing species. Int. J. Syst. Evol. Microbiol. 2008, 58, 872–876. [Google Scholar] [CrossRef]
- González-Domenech, C.M.; Martínez-Checa, F.; Quesada, E.; Béjar, V. Halomonas cerina sp. nov., a moderately halophilic, denitrifying, exopolysaccharide-producing bacterium. Int. J. Syst. Evol. Microbiol. 2008, 58, 803–809. [Google Scholar] [CrossRef]
- González-Domenech, C.M.; Martínez-Checa, F.; Quesada, E.; Béjar, V. Halomonas fontilapidosi sp. nov., a moderately halophilic, denitrifying bacterium. Int. J. Syst. Evol. Microbiol. 2009, 59, 1290–1296. [Google Scholar]
- Llamas, I.; Béjar, V.; Martínez-Checa, F.; Martínez-Cánovas, M.J.; Molina, I.; Quesada, E. Halomonas stenophila sp. nov., a halophilic bacterium that produces sulphate exopolysaccharides with biological activity. Int. J. Syst. Evol. Microbiol. 2011, 61, 2508–2514. [Google Scholar] [CrossRef]
- Luque, R.; Béjar, V.; Quesada, E.; Martínez-Checa, F.; Llamas, I. Halomonas ramblicola sp. nov., a moderately halophilic bacterium from Rambla Salada, a Mediterranean hypersaline rambla in south-east Spain. Int. J. Syst. Evol. Microbiol. 2012, 62, 2903–2909. [Google Scholar] [CrossRef]
- Martínez-Cánovas, M.J.; Béjar, V.; Martínez-Checa, F.; Quesada, E. Halomonas anticariensis sp. nov., from Fuente de Piedra, a saline-wetland, wildfowl reserve in Málaga, Southern Spain. Int. J. Syst. Evol. Microbiol. 2004, 54, 1329–1332. [Google Scholar] [CrossRef]
- Martínez-Cánovas, M.J.; Quesada, E.; Llamas, I.; Béjar, V. Halomonas ventosae sp. nov., a moderately halophilic, denitrifying, exopolysaccharide-producing bacterium. Int. J. Syst. Evol. Microbiol. 2004, 54, 733–737. [Google Scholar] [CrossRef]
- Quesada, E.; Valderrama, M.J.; Bejar, V.; Ventosa, A.; Gutierrez, M.C.; Ruiz-Berraquero, F.; Ramos-Cormenzana, A. Volcaniella eurihalina gen. nov., sp. nov., a moderately halophilic nonmotile Gram-negative rod. Int. J. Syst. Bacteriol. 1990, 40, 261–267. [Google Scholar] [CrossRef]
- Valderrama, M.J.; Quesada, E.; Béjar, V.; Ventosa, A.; Gutierrez, M.C.; Ruiz-Berraquero, F.; Ramos-Cormenzana, A. Deleya salina sp. nov., a moderately halophilic Gram-negative bacterium. Int. J. Syst. Bacteriol. 1991, 41, 377–384. [Google Scholar] [CrossRef]
- Martínez-Checa, F.; Béjar, V.; Martínez-Cánovas, M.J.; Llamas, I.; Quesada, E. Halomonas almeriensis sp. nov., a moderately halophilic, exopolysaccharide-producing bacterium from Cabo de Gata, Almería, south-east Spain. Int. J. Syst. Evol. Microbiol. 2005, 55, 2007–2011. [Google Scholar] [CrossRef]
- Mellado, E.; Moore, E.R.B.; Nieto, J.J.; Ventosa, A. Phylogenetic inferences and taxonomic consequences of 16S ribosomal DNA sequence comparison of Chromohalobacter marismortui, Volcaniella eurihalina, and Deleya salina and reclassification of V. eurihalina as Halomonas eurihalina comb. nov. Int. J. Syst.Bacteriol. 1995, 45, 712–716. [Google Scholar] [CrossRef]
- Steindler, L.; Venturi, V. Detection of quorum-sensing N-acyl homoserine lactone signal molecules by bacterial biosensors. FEMS Microbiol. Lett. 2007, 266, 1–9. [Google Scholar] [CrossRef]
- McClean, K.H.; Winson, M.K.; Fish, L.; Taylor, A.; Chhabra, S.R.; Cámara, M.; Daykin, M.; Lamb, J.H.; Swift, S.; Bycroft, B.W.; et al. Quorum sensing and Chromobacterium violaceum: Exploitation of violacein production and inhibition for the detection of N-acyl homoserine lactones. Microbiology 1997, 143, 3703–3711. [Google Scholar] [CrossRef]
- Shaw, P.D.; Ping, G.; Daly, S.L.; Cha, C.; Cronan, J.E., Jr.; Rinehart, K.L.; Farrand, S.K. Detecting and characterizing N-acyl-homoserine lactone signal molecules by thin-layer chromatography. Proc. Natl. Acad. Sci. USA 1997, 94, 6036–6041. [Google Scholar]
- García-Aljaro, C.; Eberl, L.; Riedel, K.; Blanch, A. Detection of quorum-sensing-related molecules in Vibrio scophthalmi. BMC Microbiol. 2008, 8, 138. [Google Scholar] [CrossRef]
- Yang, Q.; Han, Y.; Zhang, X.H. Detection of quorum sensing signal molecules in the family Vibrionaceae. J. Appl. Microbiol. 2011, 110, 1438–1448. [Google Scholar] [CrossRef]
- Parsek, M.R.; Greenberg, E.P. Acyl-homoserine lactone quorum sensing in Gram-negative bacteria: A signaling mechanism involved in associations with higher organisms. Proc. Natl. Acad. Sci. USA 2000, 97, 8789–8793. [Google Scholar] [CrossRef]
- Parsek, M.R.; Schaefer, A.L.; Greenberg, E.P. Analysis of random and site-directed mutations in rhlI, a Pseudomonas aeruginosa gene encoding an acylhomoserine lactone synthase. Mol. Microbiol. 1997, 26, 301–310. [Google Scholar] [CrossRef]
- Hanzelka, B.L.; Parsek, M.R.; Val, D.L.; Dunlap, P.V.; Cronan, J.E.; Greenberg, E.P. Acylhomoserine lactone synthase activity of the Vibrio fischeri AinS protein. J. Bacteriol. 1999, 181, 5766–5770. [Google Scholar]
- Milton, D.L.; Chalker, V.J.; Kirke, D.; Hardman, A.; Cámara, M.; Williams, P. The LuxM Homologue VanM from Vibrio anguillarum directs the synthesis of N-(3-hydroxyhexanoyl)-homoserine lactone and N-hexanoyl-homoserine lactone. J. Bacteriol. 2001, 183, 3537–3547. [Google Scholar] [CrossRef]
- Laue, B.E.; Jiang, Y.; Chhabra, S.R.; Jacob, S.; Stewart, G.S.A.B.; Hardman, A.; Downie, J.A.; O'Gara, F.; Williams, P. The biocontrol strain Pseudomonas fluorescens F113 produces the Rhizobium small bacteriocin, N-(3-hydroxy-7-cis-tetradecenoyl) homoserine lactone, via HdtS, a putative novel N-acylhomoserine lactone synthase. Microbiology 2000, 146, 2469–2480. [Google Scholar]
- Haynes, W.C.; Wickerham, L.J.; Hesseltine, C.W. Maintenance of cultures of industrially important microorganisms. Appl. Microbiol. 1955, 3, 361–368. [Google Scholar]
- Moraine, R.A.; Rogovin, P. Kinetics of polysaccharide B-1459 fermentation. Biotechnol. Bioeng. 1966, 8, 511–524. [Google Scholar] [CrossRef]
- Rodríguez-Valera, F.; Ruíz-Berraquero, F.; Ramos-Cormenzana, A. Characteristics of the heterotrophic bacterial populations in hypersaline environments of different salt concentrations. Microb. Ecol. 1981, 7, 235–243. [Google Scholar]
- Cha, C.; Gao, P.; Chen, Y.-C.; Shaw, P.D.; Farrand, S.K. Production of acyl-Homoserine lactone quorum-sensing signals by Gram-negative plant-associated bacteria. Mol. Plant. Microbe In. 1998, 11, 1119–1129. [Google Scholar] [CrossRef]
- Marketon, M.M.; Gronquist, M.R.; Eberhard, A.; González, J.E. Characterization of the Sinorhizobium meliloti sinR/sinI locus and the production of novel N-acyl homoserine lactones. J. Bacteriol. 2002, 184, 5686–5695. [Google Scholar] [CrossRef]
- Llamas, I.; Keshavan, N.; González, J.E. Use of Sinorhizobium meliloti as an indicator for specific detection of longchain N-acyl homoserine lactones. Appl. Environ. Microbiol. 2004, 70, 3715–3723. [Google Scholar] [CrossRef]
- Marmur, J. A procedure for the isolation of deoxyribonucleic acid from micro-organisms. J. Mol. Biol. 1961, 3, 208–218. [Google Scholar] [CrossRef]
- Martín-Platero, A.M.; Valdivia, E.; Maqueda, M.; Martínez-Bueno, M. Fast, convenient, and economical method for isolating genomic DNA from lactic acid bacteria using a modification of the protein “salting-out” procedure. Anal. Biochem. 2007, 366, 102–104. [Google Scholar] [CrossRef]
- Sambrook, J.; Russel, D.W. Molecular Cloning: A Laboratory Manual, 3rd Ed. ed; Cold Spring Harbor Laboratory Press: New York, USA, 2001. [Google Scholar]
- National Center for Biotechnology Information, N. Available online: http://www.ncbi.nlm.nih.gov/ (accessed on 15 November 2012).
- Tamura, K.; Dudley, J.; Nei, M.; Kumar, S. MEGA4: Molecular Evolutionary Genetics Analysis (MEGA) Software Version 4.0. Mol.Biol. Evol. 2007, 24, 1596–1599. [Google Scholar] [CrossRef]
- Thompson, J.D.; Gibson, T.J.; Plewniak, F.; Jeanmougin, F.; Higgins, D.G. The CLUSTAL_X windows interface: Flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res. 1997, 25, 4876–4882. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Tahrioui, A.; Schwab, M.; Quesada, E.; Llamas, I. Quorum Sensing in Some Representative Species of Halomonadaceae. Life 2013, 3, 260-275. https://doi.org/10.3390/life3010260
Tahrioui A, Schwab M, Quesada E, Llamas I. Quorum Sensing in Some Representative Species of Halomonadaceae. Life. 2013; 3(1):260-275. https://doi.org/10.3390/life3010260
Chicago/Turabian StyleTahrioui, Ali, Melanie Schwab, Emilia Quesada, and Inmaculada Llamas. 2013. "Quorum Sensing in Some Representative Species of Halomonadaceae" Life 3, no. 1: 260-275. https://doi.org/10.3390/life3010260
APA StyleTahrioui, A., Schwab, M., Quesada, E., & Llamas, I. (2013). Quorum Sensing in Some Representative Species of Halomonadaceae. Life, 3(1), 260-275. https://doi.org/10.3390/life3010260





