Potential for Natural Killer Cell-Mediated Antibody-Dependent Cellular Cytotoxicity for Control of Human Cytomegalovirus
Abstract
:1. Introduction
2. NK Cell Control of HCMV
| HCMV Gene/locus | Comment |
|---|---|
| UL18 | MHC class I homologue. Binds to inhibitory receptor LIR-1, as well as an unknown NK activating receptor [27,28] |
| SP-UL40 | Upregulates HLA-E, the ligand for the inhibitory receptor, CD94-NKG2A [29,30,31] |
| UL83 | Directly binds to the activating receptor, NKp30 [32] |
| UL16 | Retains the NKG2D ligands, MICB, ULBP1, 2, 4, 5 (immature form) and 6, intracellularly [33,34,35,36,37,38,39] |
| miR-UL112 | Suppresses gene expression of the NKG2D ligand, MICB [40,41] |
| UL142 | Retains the NKG2D ligands, MICA and ULBP3, intracellularly [42,43,44,45] |
| UL141 | Downregulates surface expression of PVR (CD155) and Nectin-2 (CD112), ligands for DNAM-1 (CD226), TACTILE (CD96) and TIGIT [46,47] |
| Retains TRAILR2, the ligand for TRAIL, intracellularly [48,49] |
3. Mechanisms of Action of NK Cells
4. HCMV UL40 and HLA-E
5. NKG2C+ NK Cell Expansions in HCMV
6. Functional Responses of NKG2C+ NK Cells
7. Role of CD16 in NK Cell Responses
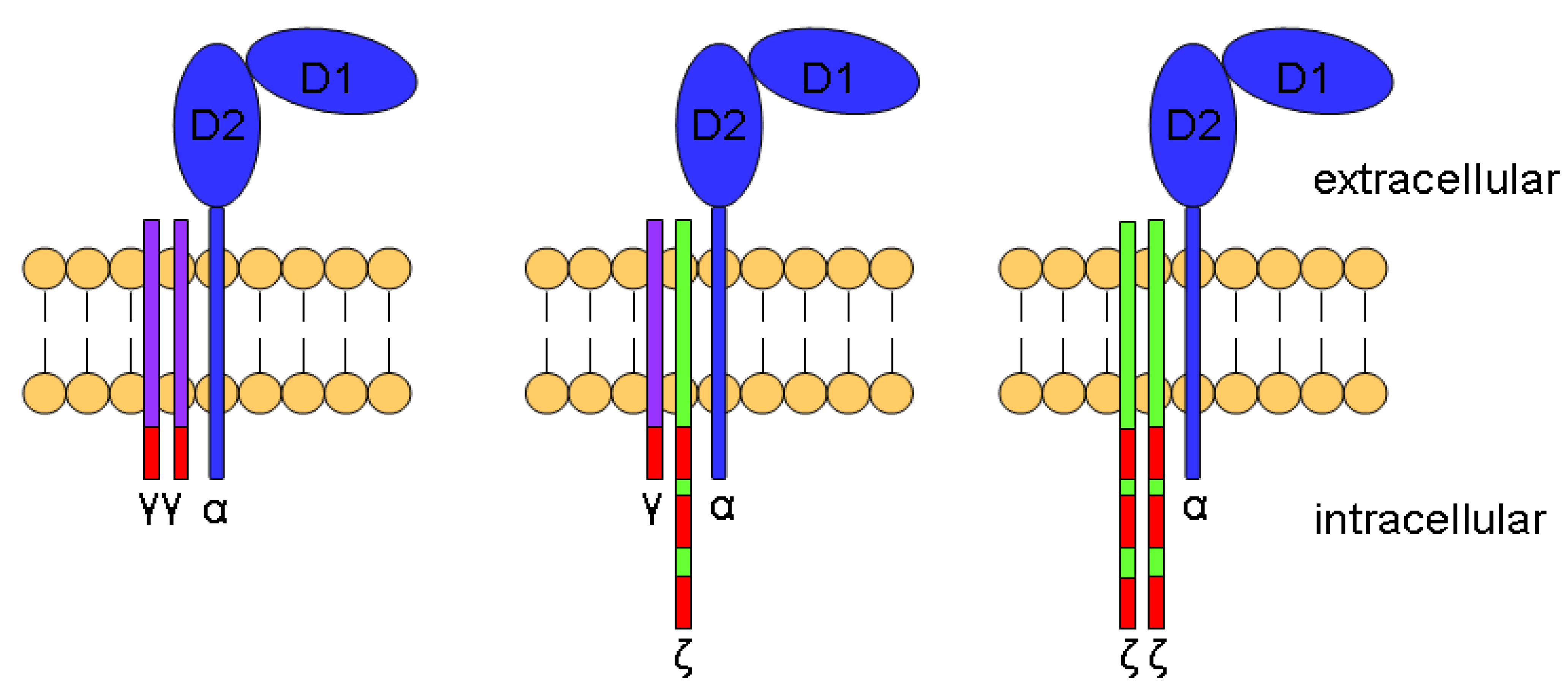
8. FcεRI-γ Chain-Deficient NK Cells
9. Clinical Importance of NK Cell-Mediated ADCC in Vaccine Design and Therapeutics
10. Conclusions
Conflicts of Interest
References
- Patel, R.; Snydman, D.R.; Rubin, R.H.; Ho, M.; Pescovitz, M.; Martin, M.; Paya, C.V. Cytomegalovirus prophylaxis in solid organ transplant recipients. Transplantation 1996, 61, 1279–1289. [Google Scholar] [CrossRef]
- Mocarski, E.S.; Thomas, S.T.; Griffiths, P.; Pass, R.F. Cytomegaloviruses in Fields Virology; Knipe, D.M., Howley, P., Eds.; Lippincott Williams and Wilkins: Philadelphia, PA, USA, 2013; Volume 2. [Google Scholar]
- Baldanti, F.; Lurain, N.; Gerna, G. Clinical and biologic aspects of human cytomegalovirus resistance to antiviral drugs. Hum. Immunol. 2004, 65, 403–409. [Google Scholar] [CrossRef]
- Dollard, S.C.; Grosse, S.D.; Ross, D.S. New estimates of the prevalence of neurological and sensory sequelae and mortality associated with congenital cytomegalovirus infection. Rev. Med. Virol. 2007, 17, 355–363. [Google Scholar] [CrossRef]
- Stratton, K.R.; Durch, J.S.; Lawrence, R.S. Vaccines for the 21st Century; National Academies Press: Washington, DC, USA, 2000. [Google Scholar]
- Avetisyan, G.; Aschan, J.; Hagglund, H.; Ringden, O.; Ljungman, P. Evaluation of intervention strategy based on CMV-specific immune responses after allogeneic SCT. Bone Marrow Transplant. 2007, 40, 865–869. [Google Scholar] [CrossRef]
- Barron, M.A.; Gao, D.; Springer, K.L.; Patterson, J.A.; Brunvand, M.W.; McSweeney, P.A.; Zeng, C.; Baron, A.E.; Weinberg, A. Relationship of reconstituted adaptive and innate cytomegalovirus (CMV)-specific immune responses with CMV viremia in hematopoietic stem cell transplant recipients. Clin. Infect. Dis. 2009, 49, 1777–1783. [Google Scholar] [CrossRef]
- Cwynarski, K.; Ainsworth, J.; Cobbold, M.; Wagner, S.; Mahendra, P.; Apperley, J.; Goldman, J.; Craddock, C.; Moss, P.A. Direct visualization of cytomegalovirus-specific T-cell reconstitution after allogeneic stem cell transplantation. Blood 2001, 97, 1232–1240. [Google Scholar] [CrossRef]
- Tormo, N.; Solano, C.; Benet, I.; Clari, M.A.; Nieto, J.; de la Camara, R.; Lopez, J.; Lopez-Aldeguer, N.; Hernandez-Boluda, J.C.; Remigia, M.J.; et al. Lack of prompt expansion of cytomegalovirus pp65 and IE-1-specific IFNgamma CD8+ and CD4+ T cells is associated with rising levels of pp65 antigenemia and DNAemia during pre-emptive therapy in allogeneic hematopoietic stem cell transplant recipients. Bone Marrow Transplant. 2010, 45, 543–549. [Google Scholar] [CrossRef]
- Sylwester, A.W.; Mitchell, B.L.; Edgar, J.B.; Taormina, C.; Pelte, C.; Ruchti, F.; Sleath, P.R.; Grabstein, K.H.; Hosken, N.A.; Kern, F.; et al. Broadly targeted human cytomegalovirus-specific CD4+ and CD8+ T cells dominate the memory compartments of exposed subjects. J. Exp. Med. 2005, 202, 673–685. [Google Scholar] [CrossRef]
- Einsele, H.; Roosnek, E.; Rufer, N.; Sinzger, C.; Riegler, S.; Loffler, J.; Grigoleit, U.; Moris, A.; Rammensee, H.G.; Kanz, L.; et al. Infusion of cytomegalovirus (CMV)-specific T cells for the treatment of CMV infection not responding to antiviral chemotherapy. Blood 2002, 99, 3916–3922. [Google Scholar] [CrossRef]
- Peggs, K.S.; Verfuerth, S.; Pizzey, A.; Khan, N.; Guiver, M.; Moss, P.A.; Mackinnon, S. Adoptive cellular therapy for early cytomegalovirus infection after allogeneic stem-cell transplantation with virus-specific T-cell lines. Lancet 2003, 362, 1375–1377. [Google Scholar] [CrossRef]
- Riddell, S.R.; Watanabe, K.S.; Goodrich, J.M.; Li, C.R.; Agha, M.E.; Greenberg, P.D. Restoration of viral immunity in immunodeficient humans by the adoptive transfer of T cell clones. Science 1992, 257, 238–241. [Google Scholar]
- Walter, E.A.; Greenberg, P.D.; Gilbert, M.J.; Finch, R.J.; Watanabe, K.S.; Thomas, E.D.; Riddell, S.R. Reconstitution of cellular immunity against cytomegalovirus in recipients of allogeneic bone marrow by transfer of T-cell clones from the donor. N. Engl. J. Med. 1995, 333, 1038–1044. [Google Scholar] [CrossRef]
- Biron, C.A.; Byron, K.S.; Sullivan, J.L. Severe Herpesvirus Infections in an Adolescent without Natural-Killer Cells. N. Engl. J. Med. 1989, 320, 1731–1735. [Google Scholar] [CrossRef]
- Orange, J.S. Natural killer cell deficiency. J. Allergy Clin. Immunol. 2013, 132, 515–525. [Google Scholar] [CrossRef]
- Orange, J.S.; Brodeur, S.R.; Jain, A.; Bonilla, F.A.; Schneider, L.C.; Kretschmer, R.; Nurko, S.; Koehler, J.R.; Rasmussen, W.L.; Fergusson, B.M.; et al. Deficiency of natural cytotoxicity in patients with IKK gamma/NEMO mutations. Faseb J. 2002, 16, A1242–A1242. [Google Scholar] [CrossRef]
- Cunningham-Rundles, C. Human B cell defects in perspective. Immunol. Res. 2012, 54, 227–232. [Google Scholar] [CrossRef]
- Aspalter, R.M.; Sewell, W.A.C.; Dolman, K.; Farrant, J.; Webster, A.D.B. Deficiency in circulating natural killer (NK) cell subsets in common variable immunodeficiency and X-linked agammaglobulinaemia. Clin. Exp. Immunol. 2000, 121, 506–514. [Google Scholar]
- Witte, T.; Werwitzke, S.; Schmidt, R.E. CMV complications in common variable immunodeficiency. Immunobiology 2000, 202, 194–198. [Google Scholar] [CrossRef]
- Rai, N.; Thakur, N. Congenital CMV With LAD Type 1 and NK Cell Deficiency. J. Pediatr. Hematol. Oncol. 2013, 35, 468–469. [Google Scholar] [CrossRef]
- Long, E.O.; Kim, H.S.; Liu, D.; Peterson, M.E.; Rajagopalan, S. Controlling natural killer cell responses: Integration of signals for activation and inhibition. Annu. Rev. Immunol. 2013, 31, 227–258. [Google Scholar] [CrossRef]
- Guma, M.; Angulo, A.; Vilches, C.; Gomez-Lozano, N.; Malats, N.; Lopez-Botet, M. Imprint of human cytomegalovirus infection on the NK cell receptor repertoire. Blood 2004, 104, 3664–3671. [Google Scholar] [CrossRef]
- Orange, J.S.; Brodeur, S.R.; Jain, A.; Bonilla, F.A.; Schneider, L.C.; Kretschmer, R.; Nurko, S.; Rasmussen, W.L.; Kohler, J.R.; Gellis, S.E.; et al. Deficient natural killer cell cytotoxicity in patients with IKK-gamma/NEMO mutations. J. Clin. Invest. 2002, 109, 1501–1509. [Google Scholar]
- Gazit, R.; Garty, B.Z.; Monselise, Y.; Hoffer, V.; Finkelstein, Y.; Markel, G.; Katz, G.; Hanna, J.; Achdout, H.; Gruda, R.; et al. Expression of KIR2DL1 on the entire NK cell population: a possible novel immunodeficiency syndrome. Blood 2004, 103, 1965–1966. [Google Scholar] [CrossRef]
- Kuijpers, T.W.; Baars, P.A.; Dantin, C.; van den Burg, M.; van Lier, R.A.; Roosnek, E. Human NK cells can control CMV infection in the absence of T cells. Blood 2008, 112, 914–915. [Google Scholar] [CrossRef]
- Prod’homme, V.; Griffin, C.; Aicheler, R.J.; Wang, E.C.; McSharry, B.P.; Rickards, C.R.; Stanton, R.J.; Borysiewicz, L.K.; Lopez-Botet, M.; Wilkinson, G.W.; et al. The human cytomegalovirus MHC class I homolog UL18 inhibits LIR-1+ but activates LIR-1- NK cells. J. Immunol. 2007, 178, 4473–4481. [Google Scholar]
- Cosman, D.; Fanger, N.; Borges, L.; Kubin, M.; Chin, W.; Peterson, L.; Hsu, M.L. A novel immunoglobulin superfamily receptor for cellular and viral MHC class I molecules. Immunobiology 1997, 7, 273–282. [Google Scholar]
- Tomasec, P.; Braud, V.M.; Rickards, C.; Powell, M.B.; McSharry, B.P.; Gadola, S.; Cerundolo, V.; Borysiewicz, L.K.; McMichael, A.J.; Wilkinson, G.W. Surface expression of HLA-E, an inhibitor of natural killer cells, enhanced by human cytomegalovirus gpUL40. Science 2000, 287, 1031. [Google Scholar] [CrossRef]
- Wang, E.C.; McSharry, B.; Retiere, C.; Tomasec, P.; Williams, S.; Borysiewicz, L.K.; Braud, V.M.; Wilkinson, G.W. UL40-mediated NK evasion during productive infection with human cytomegalovirus. Proc. Natl. Acad. Sci. USA 2002, 99, 7570–7575. [Google Scholar] [CrossRef]
- Ulbrecht, M.; Martinozzi, S.; Grzeschik, M.; Hengel, H.; Ellwart, J.W.; Pla, M.; Weiss, E.H. Cutting edge: The human cytomegalovirus UL40 gene product contains a ligand for HLA-E and prevents NK cell-mediated lysis. J. Immunol. 2000, 164, 5019–5022. [Google Scholar]
- Arnon, T.I.; Achdout, H.; Levi, O.; Markel, G.; Saleh, N.; Katz, G.; Gazit, R.; Gonen-Gross, T.; Hanna, J.; Nahari, E.; et al. Inhibition of the NKp30 activating receptor by pp65 of human cytomegalovirus. Nat. Immunol. 2005, 6, 515–523. [Google Scholar]
- Welte, S.A.; Sinzger, C.; Lutz, S.Z.; Singh-Jasuja, H.; Sampaio, K.L.; Eknigk, U.; Rammensee, H.G.; Steinle, A. Selective intracellular retention of virally induced NKG2D ligands by the human cytomegalovirus UL16 glycoprotein. Eur. J. Immunol. 2003, 33, 194–203. [Google Scholar] [CrossRef]
- Wu, J.; Chalupny, N.J.; Manley, T.J.; Riddell, S.R.; Cosman, D.; Spies, T. Intracellular retention of the MHC class I-related chain B ligand of NKG2D by the human cytomegalovirus UL16 glycoprotein. J. Immunol. 2003, 170, 4196–4200. [Google Scholar]
- Dunn, C.; Chalupny, N.J.; Sutherland, C.L.; Dosch, S.; Sivakumar, P.V.; Johnson, D.C.; Cosman, D. Human cytomegalovirus glycoprotein UL16 causes intracellular sequestration of NKG2D ligands, protecting against natural killer cell cytotoxicity. J. Exp. Med. 2003, 197, 1427–1439. [Google Scholar] [CrossRef]
- Rolle, A.; Mousavi-Jazi, M.; Eriksson, M.; Odeberg, J.; Soderberg-Naucler, C.; Cosman, D.; Karre, K.; Cerboni, C. Effects of human cytomegalovirus infection on ligands for the activating NKG2D receptor of NK cells: up-regulation of UL16-binding protein (ULBP)1 and ULBP2 is counteracted by the viral UL16 protein. J. Immunol. 2003, 171, 902–908. [Google Scholar]
- Vales-Gomez, M.; Browne, H.; Reyburn, H.T. Expression of the UL16 glycoprotein of Human Cytomegalovirus protects the virus-infected cell from attack by natural killer cells. BMC Immunol. 2003, 4, 4. [Google Scholar] [CrossRef]
- Kubin, M.; Cassiano, L.; Chalupny, J.; Chin, W.; Cosman, D.; Fanslow, W.; Mullberg, J.; Rousseau, A.M.; Ulrich, D.; Armitage, R. ULBP1, 2, 3: Novel MHC class I-related molecules that bind to human cytomegalovirus glycoprotein UL16, activate NK cells. Eur. J. Immunol. 2001, 31, 1428–1437. [Google Scholar] [CrossRef]
- Spreu, J.; Stehle, T.; Steinle, A. Human cytomegalovirus-encoded UL16 discriminates MIC molecules by their alpha2 domains. J. Immunol. 2006, 177, 3143–3149. [Google Scholar]
- Stern-Ginossar, N.; Elefant, N.; Zimmermann, A.; Wolf, D.G.; Saleh, N.; Biton, M.; Horwitz, E.; Prokocimer, Z.; Prichard, M.; Hahn, G.; et al. Host immune system gene targeting by a viral miRNA. Science 2007, 317, 376–381. [Google Scholar] [CrossRef]
- Nachmani, D.; Lankry, D.; Wolf, D.G.; Mandelboim, O. The human cytomegalovirus microRNA miR-UL112 acts synergistically with a cellular microRNA to escape immune elimination. Nat. Immunol. 2010, 11, 806–813. [Google Scholar]
- Wills, M.R.; Ashiru, O.; Reeves, M.B.; Okecha, G.; Trowsdale, J.; Tomasec, P.; Wilkinson, J.; Sinclair, G.W.; Sissons, J.G. Human cytomegalovirus encodes an MHC class I-like molecule (UL142) that functions to inhibit NK cell lysis. J. Immunol. 2005, 175, 7457–7465. [Google Scholar]
- Chalupny, N.J.; Rein-Weston, A.; Dosch, S.; Cosman, D. Down-regulation of the NKG2D ligand MICA by the human cytomegalovirus glycoprotein UL142. Biophys. Res. Commun. 2006, 346, 175–181. [Google Scholar] [CrossRef]
- Ashiru, O.; Bennett, N.J.; Boyle, L.H.; Thomas, M.; Trowsdale, J.; Wills, M.R. NKG2D ligand MICA is retained in the cis-Golgi apparatus by human cytomegalovirus protein UL142. J. Virol. 2009, 83, 12345–12354. [Google Scholar] [CrossRef]
- Bennett, N.J.; Ashiru, O.; Morgan, F.J.; Pang, Y.; Okecha, G.; Eagle, R.A.; Trowsdale, J.; Sissons, J.G.; Wills, M.R. Intracellular sequestration of the NKG2D ligand ULBP3 by human cytomegalovirus. J. Immunol. 2010, 185, 1093–1102. [Google Scholar]
- Tomasec, P.; Wang, E.C.; Davison, A.J.; Vojtesek, B.; Armstrong, M.; Griffin, C.; McSharry, B.P.; Morris, R.J.; Llewellyn-Lacey, S.; Rickards, C.; et al. Downregulation of natural killer cell-activating ligand CD155 by human cytomegalovirus UL141. Nat. Immunol. 2005, 6, 181–188. [Google Scholar]
- Prod’homme, V.; Sugrue, D.M.; Stanton, R.J.; Nomoto, A.; Davies, J.; Rickards, C.R.; Cochrane, D.; Moore, M.; Wilkinson, G.W.; Tomasec, P. Human cytomegalovirus UL141 promotes efficient downregulation of the natural killer cell activating ligand CD112. J. Gen. Virol. 2010, 91, 2034–2039. [Google Scholar] [CrossRef]
- Smith, W.; Tomasec, P.; Aicheler, R.; Loewendorf, A.; Nemcovicova, I.; Wang, E.C.; Stanton, R.J.; Macauley, M.; Norris, P.; Willen, L.; et al. Human cytomegalovirus glycoprotein UL141 targets the TRAIL death receptors to thwart host innate antiviral defenses. Cell Host Microbe 2013, 13, 324–335. [Google Scholar] [CrossRef]
- Nemcovicova, I.; Benedict, C.A.; Zajonc, D.M. Structure of human cytomegalovirus UL141 binding to TRAIL-R2 reveals novel, non-canonical death receptor interactions. PLoS Pathog. 2013, 9, e1003224. [Google Scholar] [CrossRef]
- Vivier, E.; Tomasello, E.; Baratin, M.; Walzer, T.; Ugolini, S. Functions of natural killer cells. Nat. Immunol. 2008, 9, 503–510. [Google Scholar]
- Jones, T.R.; Wiertz, E.J.; Sun, L.; Fish, K.N.; Nelson, J.A.; Ploegh, H.L. Human cytomegalovirus US3 impairs transport and maturation of major histocompatibility complex class I heavy chains. Proc. Natl. Acad. Sci. USA 1996, 93, 11327–11333. [Google Scholar] [CrossRef]
- Wiertz, E.J.; Jones, T.R.; Sun, L.; Bogyo, M.; Geuze, H.J.; Ploegh, H.L. The human cytomegalovirus US11 gene product dislocates MHC class I heavy chains from the endoplasmic reticulum to the cytosol. Cell 1996, 84, 769–779. [Google Scholar] [CrossRef]
- Jones, T.R.; Hanson, L.K.; Sun, L.; Slater, J.S.; Stenberg, R.M.; Campbell, A.E. Multiple independent loci within the human cytomegalovirus unique short region down-regulate expression of major histocompatibility complex class I heavy chains. J. Virol. 1995, 69, 4830–4841. [Google Scholar]
- Machold, R.P.; Wiertz, E.J.; Jones, T.R.; Ploegh, H.L. The HCMV gene products US11 and US2 differ in their ability to attack allelic forms of murine major histocompatibility complex (MHC) class I heavy chains. J. Exp. Med. 1997, 185, 363–366. [Google Scholar] [CrossRef]
- Ahn, K.; Gruhler, A.; Galocha, B.; Jones, T.R.; Wiertz, E.J.; Ploegh, H.L.; Peterson, P.A.; Yang, Y.; Fruh, K. The ER-luminal domain of the HCMV glycoprotein US6 inhibits peptide translocation by TAP. Immunity 1997, 6, 613–621. [Google Scholar] [CrossRef]
- Braud, V.; Jones, E.Y.; McMichael, A. The human major histocompatibility complex class Ib molecule HLA-E binds signal sequence-derived peptides with primary anchor residues at positions 2 and 9. Eur. J. Immunol. 1997, 27, 1164–1169. [Google Scholar] [CrossRef]
- Millo, E.; Pietra, G.; Armirotti, A.; Vacca, P.; Mingari, M.C.; Moretta, L.; Damonte, G. Purification and HPLC-MS analysis of a naturally processed HCMV-derived peptide isolated from the HEK-293T/HLA-E+/Ul40+ cell transfectants and presented at the cell surface in the context of HLA-E. J. Immunol. Meth. 2007, 322, 128–136. [Google Scholar] [CrossRef]
- Borrego, F.; Ulbrecht, M.; Weiss, E.H.; Coligan, J.E.; Brooks, A.G. Recognition of human histocompatibility leukocyte antigen (HLA)-E complexed with HLA class I signal sequence-derived peptides by CD94/NKG2 confers protection from natural killer cell-mediated lysis. J. Exp. Med. 1998, 187, 813–818. [Google Scholar] [CrossRef] [Green Version]
- Braud, V.M.; Allan, D.S.; O’Callaghan, C.A.; Soderstrom, K.; D’Andrea, A.; Ogg, G.S.; Lazetic, S.; Young, N.T.; Bell, J.I.; Phillips, J.H.; et al. HLA-E binds to natural killer cell receptors CD94/NKG2A, B and C. Nature 1998, 391, 795–799. [Google Scholar] [CrossRef]
- Lee, N.; Llano, M.; Carretero, M.; Ishitani, A.; Navarro, F.; Lopez-Botet, M.; Geraghty, D.E. HLA-E is a major ligand for the natural killer inhibitory receptor CD94/NKG2A. Proc. Natl. Acad. Sci. USA 1998, 95, 5199–5204. [Google Scholar] [CrossRef]
- Kaiser, B.K.; Barahmand-Pour, F.; Paulsene, W.; Medley, S.; Geraghty, D.E.; Strong, R.K. Interactions between NKG2x immunoreceptors and HLA-E ligands display overlapping affinities and thermodynamics. J. Immunol. 2005, 174, 2878–2884. [Google Scholar]
- Miller, J.D.; Weber, D.A.; Ibegbu, C.; Pohl, J.; Altman, J.D.; Jensen, P.E. Analysis of HLA-E peptide-binding specificity and contact residues in bound peptide required for recognition by CD94/NKG2. J. Immunol. 2003, 171, 1369–1375. [Google Scholar]
- Sullivan, L.C.; Clements, C.S.; Beddoe, T.; Johnson, D.; Hoare, H.L.; Lin, J.; Huyton, T.; Hopkins, E.J.; Reid, H.H.; Wilce, M.C.; et al. The heterodimeric assembly of the CD94-NKG2 receptor family and implications for human leukocyte antigen-E recognition. Immunity 2007, 27, 900–911. [Google Scholar] [CrossRef]
- Vales-Gomez, M.; Reyburn, H.T.; Erskine, R.A.; Lopez-Botet, M.; Strominger, J.L. Kinetics and peptide dependency of the binding of the inhibitory NK receptor CD94/NKG2-A and the activating receptor CD94/NKG2-C to HLA-E. EMBO J. 1999, 18, 4250–4260. [Google Scholar] [CrossRef]
- Heatley, S.L.; Pietra, G.; Lin, J.; Widjaja, J.M.; Harpur, C.M.; Lester, S.; Rossjohn, J.; Szer, J.; Schwarer, A.; Bradstock, K.; et al. Polymorphism in human cytomegalovirus UL40 impacts on recognition of human leukocyte antigen-E (HLA-E) by natural killer cells. J. Biol. Chem. 2013, 288, 8679–8690. [Google Scholar] [CrossRef]
- Llano, M.; Lee, N.; Navarro, F.; Garcia, P.; Albar, J.P.; Geraghty, D.E.; Lopez-Botet, M. HLA-E-bound peptides influence recognition by inhibitory and triggering CD94/NKG2 receptors: preferential response to an HLA-G-derived nonamer. Eur. J. Immunol. 1998, 28, 2854–2863. [Google Scholar] [CrossRef]
- Bjorkstrom, N.K.; Lindgren, T.; Stoltz, M.; Fauriat, C.; Braun, M.; Evander, M.; Michaelsson, J.; Malmberg, K.J.; Klingstrom, J.; Ahlm, C.; et al. Rapid expansion and long-term persistence of elevated NK cell numbers in humans infected with hantavirus. J. Exp. Med. 2011, 208, 13–21. [Google Scholar] [CrossRef]
- Petitdemange, C.; Becquart, P.; Wauquier, N.; Beziat, V.; Debre, P.; Leroy, E.M.; Vieillard, V. Unconventional repertoire profile is imprinted during acute chikungunya infection for natural killer cells polarization toward cytotoxicity. PLoS Pathog. 2011, 7, e1002268. [Google Scholar] [CrossRef]
- Brunetta, E.; Fogli, M.; Varchetta, S.; Bozzo, L.; Hudspeth, K.L.; Marcenaro, E.; Moretta, A.; Mavilio, D. Chronic HIV-1 viremia reverses NKG2A/NKG2C ratio on natural killer cells in patients with human cytomegalovirus co-infection. AIDS 2010, 24, 27–34. [Google Scholar] [CrossRef]
- Beziat, V.; Dalgard, O.; Asselah, T.; Halfon, P.; Bedossa, P.; Boudifa, A.; Hervier, B.; Theodorou, I.; Martinot, M.; Debre, P.; et al. CMV drives clonal expansion of NKG2C+ NK cells expressing self-specific KIRs in chronic hepatitis patients. Eur. J. Immunol. 2012, 42, 447–457. [Google Scholar] [CrossRef]
- Monsivais-Urenda, A.; Noyola-Cherpitel, D.; Hernandez-Salinas, A.; Garcia-Sepulveda, C.; Romo, N.; Baranda, L.; Lopez-Botet, M.; Gonzalez-Amaro, R. Influence of human cytomegalovirus infection on the NK cell receptor repertoire in children. Eur. J. Immunol. 2010, 40, 1418–1427. [Google Scholar] [CrossRef]
- Petersen, L.; Roug, A.S.; Skovbo, A.; Thysen, A.H.; Eskelund, C.W.; Hokland, M.E. The CD94/NKG2C-expressing NK cell subset is augmented in chronic lymphocytic leukemia patients with positive human cytomegalovirus serostatus. Viral. Immunol. 2009, 22, 333–337. [Google Scholar] [CrossRef]
- Guma, M.; Cabrera, C.; Erkizia, I.; Bofill, M.; Clotet, B.; Ruiz, L.; Lopez-Botet, M. Human cytomegalovirus infection is associated with increased proportions of NK cells that express the CD94/NKG2C receptor in aviremic HIV-1-positive patients. J. Infect. Dis. 2006, 194, 38–41. [Google Scholar] [CrossRef]
- Noyola, D.E.; Fortuny, C.; Muntasell, A.; Noguera-Julian, A.; Munoz-Almagro, C.; Alarcon, A.; Juncosa, T.; Moraru, M.; Vilches, C.; Lopez-Botet, M. Influence of congenital human cytomegalovirus infection and the NKG2C genotype on NK-cell subset distribution in children. Eur. J. Immunol. 2012, 42, 3256–3266. [Google Scholar] [CrossRef]
- Foley, B.; Cooley, S.; Verneris, M.R.; Pitt, M.; Curtsinger, J.; Luo, X.; Lopez-Verges, S.; Lanier, L.L.; Weisdorf, D.; Miller, J.S. Cytomegalovirus reactivation after allogeneic transplantation promotes a lasting increase in educated NKG2C+ natural killer cells with potent function. Blood 2012, 119, 2665–2674. [Google Scholar] [CrossRef]
- Chiesa, D.M.; Falco, M.; Podesta, M.; Locatelli, F.; Moretta, L.; Frassoni, F.; Moretta, A. Phenotypic and functional heterogeneity of human NK cells developing after umbilical cord blood transplantation: A role for human cytomegalovirus? Blood 2012, 119, 399–410. [Google Scholar]
- Lopez-Verges, S.; Milush, J.M.; Schwartz, B.S.; Pando, M.J.; Jarjoura, J.; York, V.A.; Houchins, J.P.; Miller, S.; Kang, S.M.; Norris, P.J.; et al. Expansion of a unique CD57(+)NKG2Chi natural killer cell subset during acute human cytomegalovirus infection. Proc. Natl. Acad. Sci. USA 2011, 108, 14725–14732. [Google Scholar]
- Charoudeh, H.N.; Terszowski, G.; Czaja, K.; Gonzalez, A.; Schmitter, K.; Stern, M. Modulation of the natural killer cell KIR repertoire by cytomegalovirus infection. Eur. J. Immunol. 2013, 43, 480–487. [Google Scholar] [CrossRef]
- Beziat, V.; Liu, L.L.; Malmberg, J.A.; Ivarsson, M.A.; Sohlberg, E.; Bjorklund, A.T.; Retiere, C.; Sverremark-Ekstrom, E.; Traherne, J.; Ljungman, P.; et al. NK cell responses to cytomegalovirus infection lead to stable imprints in the human KIR repertoire and involve activating KIRs. Blood 2013, 121, 2678–2688. [Google Scholar] [CrossRef]
- Djaoud, Z.; David, G.; Bressollette, C.; Willem, C.; Rettman, P.; Gagne, K.; Legrand, N.; Mehlal, S.; Cesbron, A.; Imbert-Marcille, B.M.; et al. Amplified NKG2C+ NK cells in cytomegalovirus (CMV) infection preferentially express killer cell Ig-like receptor 2DL: functional impact in controlling CMV-infected dendritic cells. J. Immunol. 2013, 191, 2708–2716. [Google Scholar] [CrossRef]
- Moraru, M.; Cisneros, E.; Gomez-Lozano, N.; de Pablo, R.; Portero, F.; Canizares, M.; Vaquero, M.; Roustan, G.; Millan, I.; Lopez-Botet, M.; et al. Host genetic factors in susceptibility to herpes simplex type 1 virus infection: contribution of polymorphic genes at the interface of innate and adaptive immunity. J. Immunol. 2012, 188, 4412–4420. [Google Scholar] [CrossRef]
- Muntasell, A.; Vilches, C.; Angulo, A.; Lopez-Botet, M. Adaptive reconfiguration of the human NK-cell compartment in response to cytomegalovirus: a different perspective of the host-pathogen interaction. Eur. J. Immunol. 2013, 43, 1133–1141. [Google Scholar] [CrossRef]
- Hikami, K.; Tsuchiya, N.; Yabe, T.; Tokunaga, K. Variations of human killer cell lectin-like receptors: Common occurrence of NKG2-C deletion in the general population. Genes Immun. 2003, 4, 160–167. [Google Scholar] [CrossRef]
- Miyashita, R.; Tsuchiya, N.; Hikami, K.; Kuroki, K.; Fukazawa, T.; Bijl, M.; Kallenberg, C.G.; Hashimoto, H.; Yabe, T.; Tokunaga, K. Molecular genetic analyses of human NKG2C (KLRC2) gene deletion. Int. Immunol. 2004, 16, 163–168. [Google Scholar] [CrossRef]
- Moraru, M.; Canizares, M.; Muntasell, A.; de Pablo, R.; Lopez-Botet, M.; Vilches, C. Assessment of copy-number variation in the NKG2C receptor gene in a single-tube and characterization of a reference cell panel, using standard polymerase chain reaction. Tissue Antigens 2012, 80, 184–187. [Google Scholar] [CrossRef]
- Munoz-Cobo, B.; Solano, C.; Benet, I.; Costa, E.; Remigia, M.J.; de la Camara, R.; Nieto, J.; Lopez, J.; Amat, P.; Garcia-Noblejas, A.; et al. Functional profile of cytomegalovirus (CMV)-specific CD8+ T cells and kinetics of NKG2C+ NK cells associated with the resolution of CMV DNAemia in allogeneic stem cell transplant recipients. J. Med. Virol. 2012, 84, 259–267. [Google Scholar] [CrossRef]
- Wu, Z.; Sinzger, C.; Frascaroli, G.; Reichel, J.; Bayer, C.; Wang, L.; Schirmbeck, R.; Mertens, T. Human cytomegalovirus-induced NKG2C(hi) CD57(hi) natural killer cells are effectors dependent on humoral antiviral immunity. J. Virol. 2013, 87, 7717–7725. [Google Scholar] [CrossRef]
- Guma, M.; Budt, M.; Saez, A.; Brckalo, T.; Hengel, H.; Angulo, A.; Lopez-Botet, M. Expansion of CD94/NKG2C+ NK cells in response to human cytomegalovirus-infected fibroblasts. Blood 2006, 107, 3624–3631. [Google Scholar] [CrossRef]
- Beziat, V.; Hervier, B.; Achour, A.; Boutolleau, D.; Marfain-Koka, A.; Vieillard, V. Human NKG2A overrides NKG2C effector functions to prevent autoreactivity of NK cells. Blood 2011, 117, 4394–4396. [Google Scholar] [CrossRef]
- Foley, B.; Cooley, S.; Verneris, M.R.; Curtsinger, J.; Luo, X.; Waller, E.K.; Anasetti, C.; Weisdorf, D.; Miller, J.S. Human cytomegalovirus (CMV)-induced memory-like NKG2C(+) NK cells are transplantable and expand in vivo in response to recipient CMV antigen. J. Immunol. 2012, 189, 5082–5088. [Google Scholar] [CrossRef]
- Daeron, M. Fc receptor biology. Annu. Rev. Immunol. 1997, 15, 203–234. [Google Scholar] [CrossRef]
- Ravetch, J.V.; Perussia, B. Alternative membrane forms of Fc gamma RIII(CD16) on human natural killer cells and neutrophils. Cell type-specific expression of two genes that differ in single nucleotide substitutions. J. Exp. Med. 1989, 170, 481–497. [Google Scholar] [CrossRef]
- Wirthmueller, U.; Kurosaki, T.; Murakami, M.S.; Ravetch, J.V. Signal transduction by Fc gamma RIII (CD16) is mediated through the gamma chain. J. Exp. Med. 1992, 175, 1381–1390. [Google Scholar] [CrossRef]
- Takai, T.; Li, M.; Sylvestre, D.; Clynes, R.; Ravetch, J.V. FcR gamma chain deletion results in pleiotrophic effector cell defects. Cell 1994, 76, 519–529. [Google Scholar] [CrossRef]
- Hazenbos, W.L.; Gessner, J.E.; Hofhuis, F.M.; Kuipers, H.; Meyer, D.; Heijnen, I.A.; Schmidt, R.E.; Sandor, M.; Capel, P.J.; Daeron, M.; et al. Impaired IgG-dependent anaphylaxis and Arthus reaction in Fc gamma RIII (CD16) deficient mice. Immunity 1996, 5, 181–188. [Google Scholar] [CrossRef]
- Hwang, I.; Zhang, T.; Scott, J.M.; Kim, A.R.; Lee, T.; Kakarla, T.; Kim, A.; Sunwoo, J.B.; Kim, S. Identification of human NK cells that are deficient for signaling adaptor FcRgamma and specialized for antibody-dependent immune functions. Int. Immunol. 2012, 24, 793–802. [Google Scholar] [CrossRef]
- Zhang, T.; Scott, J.M.; Hwang, I.; Kim, S. Cutting edge: antibody-dependent memory-like NK cells distinguished by FcRgamma deficiency. J. Immunol. 2013, 190, 1402–1406. [Google Scholar] [CrossRef]
- Mandelboim, O.; Lieberman, N.; Lev, M.; Paul, L.; Arnon, T.I.; Bushkin, Y.; Davis, D.M.; Strominger, J.L.; Yewdell, J.W.; Porgador, A. Recognition of haemagglutinins on virus-infected cells by NKp46 activates lysis by human NK cells. Nature 2001, 409, 1055–1060. [Google Scholar] [CrossRef]
- Hecht, M.L.; Rosental, B.; Horlacher, T.; Hershkovitz, O.; de Paz, J.L.; Noti, C.; Schauer, S.; Porgador, A.; Seeberger, P.H. Natural cytotoxicity receptors NKp30, NKp44 and NKp46 bind to different heparan sulfate/heparin sequences. J. Proteome Res. 2009, 8, 712–720. [Google Scholar] [CrossRef]
- Romo, N.; Magri, G.; Muntasell, A.; Heredia, G.; Baia, D.; Angulo, A.; Guma, M.; Lopez-Botet, M. Natural killer cell-mediated response to human cytomegalovirus-infected macrophages is modulated by their functional polarization. J. Leukoc. Biol. 2011, 90, 717–726. [Google Scholar] [CrossRef]
- Magri, G.; Muntasell, A.; Romo, N.; Saez-Borderias, A.; Pende, D.; Geraghty, D.E.; Hengel, H.; Angulo, A.; Moretta, A.; Lopez-Botet, M. NKp46 and DNAM-1 NK-cell receptors drive the response to human cytomegalovirus-infected myeloid dendritic cells overcoming viral immune evasion strategies. Blood 2011, 117, 848–856. [Google Scholar] [CrossRef]
- Pessino, A.; Sivori, S.; Bottino, C.; Malaspina, A.; Morelli, L.; Moretta, L.; Biassoni, R.; Moretta, A. Molecular cloning of NKp46: A novel member of the immunoglobulin superfamily involved in triggering of natural cytotoxicity. J. Exp. Med. 1998, 188, 953–960. [Google Scholar]
- Moretta, A.; Biassoni, R.; Bottino, C.; Mingari, M.C.; Moretta, L. Natural cytotoxicity receptors that trigger human NK-cell-mediated cytolysis. Immunol. Today 2000, 21, 228–234. [Google Scholar] [CrossRef]
- Nigro, G.; Adler, S.P.; Torre, L.R.; Best, A.M. Passive immunization during pregnancy for congenital cytomegalovirus infection. N. Engl. J. Med. 2005, 353, 1350–1362. [Google Scholar] [CrossRef]
- Snydman, D.R.; Werner, B.G.; Dougherty, N.N.; Griffith, J.; Rubin, R.H.; Dienstag, J.L.; Rohrer, R.H.; Freeman, R.; Jenkins, R.; Lewis, W.D.; et al. Cytomegalovirus immune globulin prophylaxis in liver transplantation. A randomized, double-blind, placebo-controlled trial. Ann. Intern. Med. 1993, 119, 984–991. [Google Scholar] [CrossRef]
- Snydman, D.R.; Werner, B.G.; Heinze-Lacey, B.; Berardi, V.P.; Tilney, N.L.; Kirkman, R.L.; Milford, E.L.; Cho, S.I.; Bush, H.L.J.; Levey, A.S.; et al. Use of cytomegalovirus immune globulin to prevent cytomegalovirus disease in renal-transplant recipients. N. Engl. J. Med. 1987, 317, 1049–1054. [Google Scholar] [CrossRef]
- Metselaar, H.J.; Rothbarth, P.H.; Brouwer, R.M.; Wenting, G.J.; Jeekel, J.; Weimar, W. Prevention of cytomegalovirus-related death by passive immunization. A double-blind placebo-controlled study in kidney transplant recipients treated for rejection. Transplantation 1989, 48, 264–266. [Google Scholar] [CrossRef]
- Alexander, B.T.; Hladnik, L.M.; Augustin, K.M.; Casabar, E.; McKinnon, P.S.; Reichley, R.M.; Ritchie, D.J.; Westervelt, P.; Dubberke, E.R. Use of cytomegalovirus intravenous immune globulin for the adjunctive treatment of cytomegalovirus in hematopoietic stem cell transplant recipients. Pharmacotherapy 2010, 30, 554–561. [Google Scholar] [CrossRef]
- Tsevat, J.; Snydman, D.R.; Pauker, S.G.; Durand-Zaleski, I.; Werner, B.G.; Levey, A.S. Which renal transplant patients should receive cytomegalovirus immune globulin? A cost-effectiveness analysis. Transplantation 1991, 52, 259–265. [Google Scholar]
- Raanani, P.; Gafter-Gvili, A.; Paul, M.; Ben-Bassat, I.; Leibovici, L.; Shpilberg, O. Immunoglobulin prophylaxis in hematological malignancies and hematopoietic stem cell transplantation. Cochrane Database Syst. Rev. 2008, 4, CD006501. [Google Scholar]
- Britt, W.J.; Vugler, L.; Butfiloski, E.J.; Stephens, E.B. Cell surface expression of human cytomegalovirus (HCMV) gp55–116 (gB): Use of HCMV-recombinant vaccinia virus-infected cells in analysis of the human neutralizing antibody response. J. Virol. 1990, 64, 1079–1085. [Google Scholar]
- Adler, B.; Scrivano, L.; Ruzcics, Z.; Rupp, B.; Sinzger, C.; Koszinowski, U. Role of human cytomegalovirus UL131A in cell type-specific virus entry and release. J. Gen. Virol. 2006, 87, 2451–2460. [Google Scholar] [CrossRef]
- Revello, M.G.; Gerna, G. Human cytomegalovirus tropism for endothelial/epithelial cells: Scientific background and clinical implications. Rev. Med. Virol. 2010, 20, 136–155. [Google Scholar] [CrossRef]
- Lilleri, D.; Kabanova, A.; Revello, M.G.; Percivalle, E.; Sarasini, A.; Genini, E.; Sallusto, F.; Lanzavecchia, A.; Corti, D.; Gerna, G. Fetal human cytomegalovirus transmission correlates with delayed maternal antibodies to gH/gL/pUL128–130–131 complex during primary infection. PLoS One 2013, 8, e59863. [Google Scholar]
- Macagno, A.; Bernasconi, N.L.; Vanzetta, F.; Dander, E.; Sarasini, A.; Revello, M.G.; Gerna, G.; Sallusto, F.; Lanzavecchia, A. Isolation of human monoclonal antibodies that potently neutralize human cytomegalovirus infection by targeting different epitopes on the gH/gL/UL128–131A complex. J. Virol. 2010, 84, 1005–1013. [Google Scholar] [CrossRef]
- Hahn, G.; Revello, M.G.; Patrone, M.; Percivalle, E.; Campanini, G.; Sarasini, A.; Wagner, M.; Gallina, A.; Milanesi, G.; Koszinowski, U.; et al. Human cytomegalovirus UL131–128 genes are indispensable for virus growth in endothelial cells and virus transfer to leukocytes. J. Virol. 2004, 78, 10023–10033. [Google Scholar] [CrossRef]
- Ryckman, B.J.; Rainish, B.L.; Chase, M.C.; Borton, J.A.; Nelson, J.A.; Jarvis, M.A.; Johnson, D.C. Characterization of the human cytomegalovirus gH/gL/UL128–131 complex that mediates entry into epithelial and endothelial cells. J. Virol. 2008, 82, 60–70. [Google Scholar] [CrossRef]
- Wang, D.; Shenk, T. Human cytomegalovirus virion protein complex required for epithelial and endothelial cell tropism. Proc. Natl. Acad. Sci. USA 2005, 102, 18153–18158. [Google Scholar] [CrossRef]
- Wang, D.; Shenk, T. Human cytomegalovirus UL131 open reading frame is required for epithelial cell tropism. J. Virol. 2005, 79, 10330–10338. [Google Scholar] [CrossRef]
- Genini, E.; Percivalle, E.; Sarasini, A.; Revello, M.G.; Baldanti, F.; Gerna, G. Serum antibody response to the gH/gL/pUL128–131 five-protein complex of human cytomegalovirus (HCMV) in primary and reactivated HCMV infections. J. Clin. Virol. 2011, 52, 113–118. [Google Scholar] [CrossRef]
- Fouts, A.E.; Chan, P.; Stephan, J.P.; Vandlen, R.; Feierbach, B. Antibodies against the gH/gL/UL128/UL130/UL131 complex comprise the majority of the anti-cytomegalovirus (anti-CMV) neutralizing antibody response in CMV hyperimmune globulin. J. Virol. 2012, 86, 7444–7447. [Google Scholar] [CrossRef]
- Plotkin, S.A.; Starr, S.E.; Friedman, H.M.; Brayman, K.; Harris, S.; Jackson, S.; Tustin, N.B.; Grossman, R.; Dafoe, D.; Barker, C. Effect of Towne live virus vaccine on cytomegalovirus disease after renal transplant. A controlled trial. Ann. Intern. Med. 1991, 114, 525–531. [Google Scholar] [CrossRef]
- Plotkin, S.A.; Higgins, R.; Kurtz, J.B.; Morris, P.J.; Campbell, D.A.J.; Shope, T.C.; Spector, S.A.; Dankner, W.M. Multicenter trial of Towne strain attenuated virus vaccine in seronegative renal transplant recipients. Transplantation 1994, 58, 1176–1178. [Google Scholar]
- Adler, S.P.; Starr, S.E.; Plotkin, S.A.; Hempfling, S.H.; Buis, J.; Manning, M.L.; Best, A.M. Immunity induced by primary human cytomegalovirus infection protects against secondary infection among women of childbearing age. J. Infect. Dis. 1995, 171, 26–32. [Google Scholar] [CrossRef]
- Sabbaj, S.; Pass, R.F.; Goepfert, P.A.; Pichon, S. Glycoprotein B vaccine is capable of boosting both antibody and CD4 T-cell responses to cytomegalovirus in chronically infected women. J. Infect. Dis. 2011, 203, 1534–1541. [Google Scholar] [CrossRef]
- Pass, R.F.; Zhang, C.; Evans, A.; Simpson, T.; Andrews, W.; Huang, M.L.; Corey, L.; Hill, J.; Davis, E.; Flanigan, C.; et al. Vaccine prevention of maternal cytomegalovirus infection. N. Engl. J. Med. 2009, 360, 1191–1199. [Google Scholar] [CrossRef]
- Pass, R.F.; Duliege, A.M.; Boppana, S.; Sekulovich, R.; Percell, S.; Britt, W.; Burke, R.L. A subunit cytomegalovirus vaccine based on recombinant envelope glycoprotein B and a new adjuvant. J. Infect. Dis. 1999, 180, 970–975. [Google Scholar] [CrossRef]
- Griffiths, P.D.; Stanton, A.; McCarrell, E.; Smith, C.; Osman, M.; Harber, M.; Davenport, A.; Jones, G.; Wheeler, D.C.; O’Beirne, J.; et al. Cytomegalovirus glycoprotein-B vaccine with MF59 adjuvant in transplant recipients: a phase 2 randomised placebo-controlled trial. Lancet 2011, 377, 1256–1263. [Google Scholar] [CrossRef]
- Nokta, M.; Tolpin, M.D.; Nadler, P.I.; Pollard, R.B. Human monoclonal anti-cytomegalovirus (CMV) antibody (MSL 109): Enhancement of in vitro foscarnet- and ganciclovir-induced inhibition of CMV replication. Antivir. Res. 1994, 24, 17–26. [Google Scholar] [CrossRef]
- Manley, K.; Anderson, J.; Yang, F.; Szustakowski, J.; Oakeley, E.J.; Compton, T.; Feire, A.L. Human cytomegalovirus escapes a naturally occurring neutralizing antibody by incorporating it into assembling virions. Cell Host Microbe 2011, 10, 197–209. [Google Scholar] [CrossRef]
- Jabs, D.A.; Gilpin, A.M.; Min, Y.I.; Erice, A.; Kempen, J.H.; Quinn, T.C. HIV and cytomegalovirus viral load and clinical outcomes in AIDS and cytomegalovirus retinitis patients: Monoclonal Antibody Cytomegalovirus Retinitis Trial. AIDS 2002, 16, 877–887. [Google Scholar] [CrossRef]
- Boeckh, M.; Bowden, R.A.; Storer, B.; Chao, N.J.; Spielberger, R.; Tierney, D.K.; Gallez-Hawkins, G.; Cunningham, T.; Blume, K.G.; Levitt, D.; et al. Randomized, placebo-controlled, double-blind study of a cytomegalovirus-specific monoclonal antibody (MSL-109) for prevention of cytomegalovirus infection after allogeneic hematopoietic stem cell transplantation. Biol. Blood Marrow Transplant. 2001, 7, 343–351. [Google Scholar] [CrossRef]
- Jegaskanda, S.; Job, E.R.; Kramski, M.; Laurie, K.; Isitman, G.; de Rose, R.; Winnall, W.R.; Stratov, I.; Brooks, A.G.; Reading, P.C.; et al. Cross-reactive influenza-specific antibody-dependent cellular cytotoxicity antibodies in the absence of neutralizing antibodies. J. Immunol. 2013, 190, 1837–1848. [Google Scholar] [CrossRef]
- Jia, M.; Li, D.; He, X.; Zhao, Y.; Peng, H.; Ma, P.; Hong, K.; Liang, H.; Shao, Y. Impaired natural killer cell-induced antibody-dependent cell-mediated cytotoxicity is associated with human immunodeficiency virus-1 disease progression. Clin. Exp. Immunol. 2013, 171, 107–116. [Google Scholar]
- Hansen, S.G.; Powers, C.J.; Richards, R.; Ventura, A.B.; Ford, J.C.; Siess, D.; Axthelm, M.K.; Nelson, J.A.; Jarvis, M.A.; Picker, L.J.; et al. Evasion of CD8+ T cells is critical for superinfection by cytomegalovirus. Science 2010, 328, 102–106. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Aicheler, R.J.; Wang, E.C.Y.; Tomasec, P.; Wilkinson, G.W.G.; Stanton, R.J. Potential for Natural Killer Cell-Mediated Antibody-Dependent Cellular Cytotoxicity for Control of Human Cytomegalovirus. Antibodies 2013, 2, 617-635. https://doi.org/10.3390/antib2040617
Aicheler RJ, Wang ECY, Tomasec P, Wilkinson GWG, Stanton RJ. Potential for Natural Killer Cell-Mediated Antibody-Dependent Cellular Cytotoxicity for Control of Human Cytomegalovirus. Antibodies. 2013; 2(4):617-635. https://doi.org/10.3390/antib2040617
Chicago/Turabian StyleAicheler, Rebecca J., Eddie C. Y. Wang, Peter Tomasec, Gavin W. G. Wilkinson, and Richard J. Stanton. 2013. "Potential for Natural Killer Cell-Mediated Antibody-Dependent Cellular Cytotoxicity for Control of Human Cytomegalovirus" Antibodies 2, no. 4: 617-635. https://doi.org/10.3390/antib2040617




