The Little Fly that Could: Wizardry and Artistry of Drosophila Genomics
Abstract
:1. Introduction
2. Drosophila as a Model
2.1. In Development
2.2. In Signaling
2.3. In Disease
3. Meet the Drosophila Genome
4. Genomes by the Dozen
5. Genomes by Population
6. Decoding the Genome’s Secrets
6.1. The modENCODE Project
6.2. The Transcriptome
6.3. Chromatin Landscape
6.4. Transcriptional Regulation
7. The Fruit Fly Toolkit
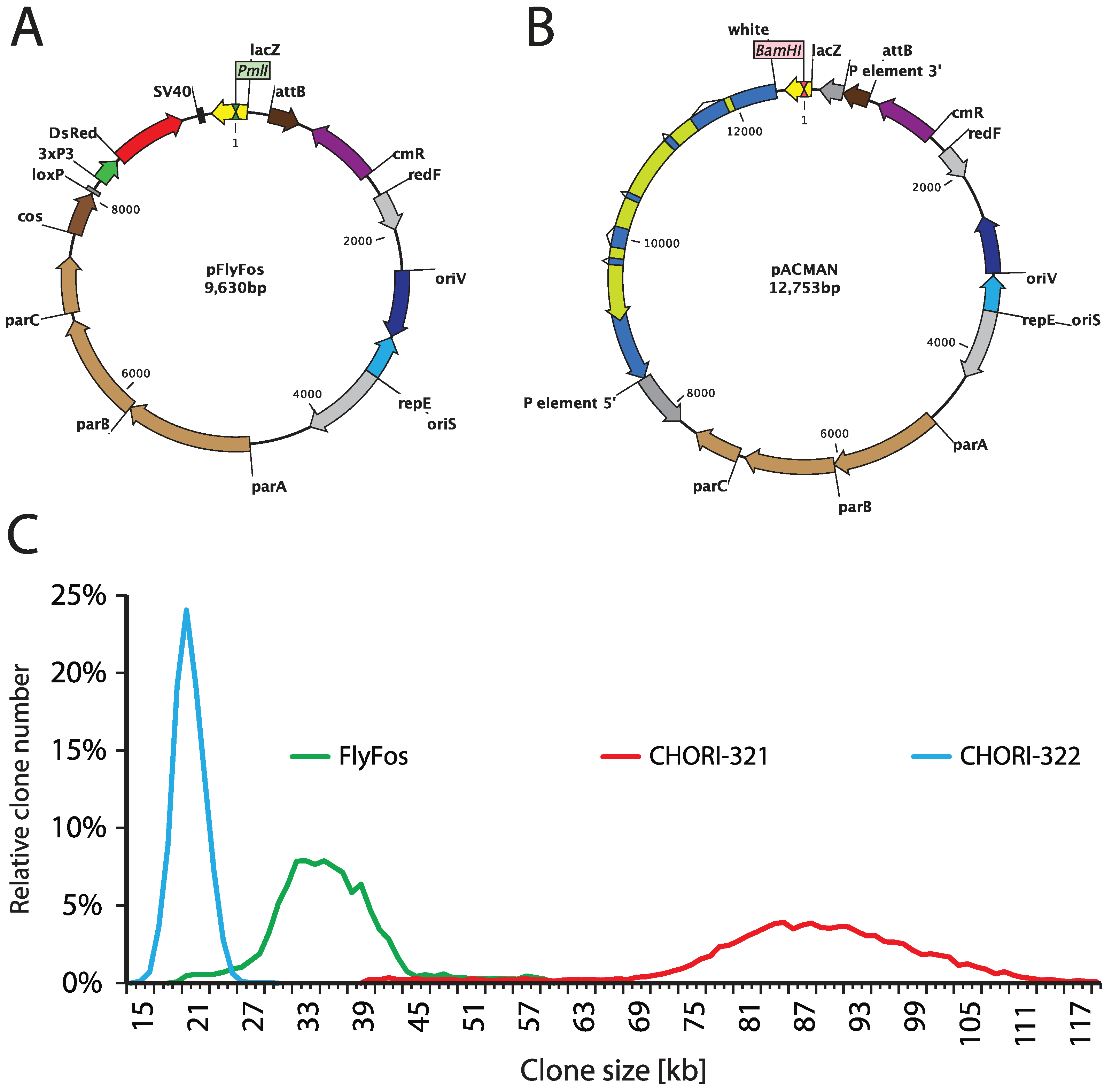
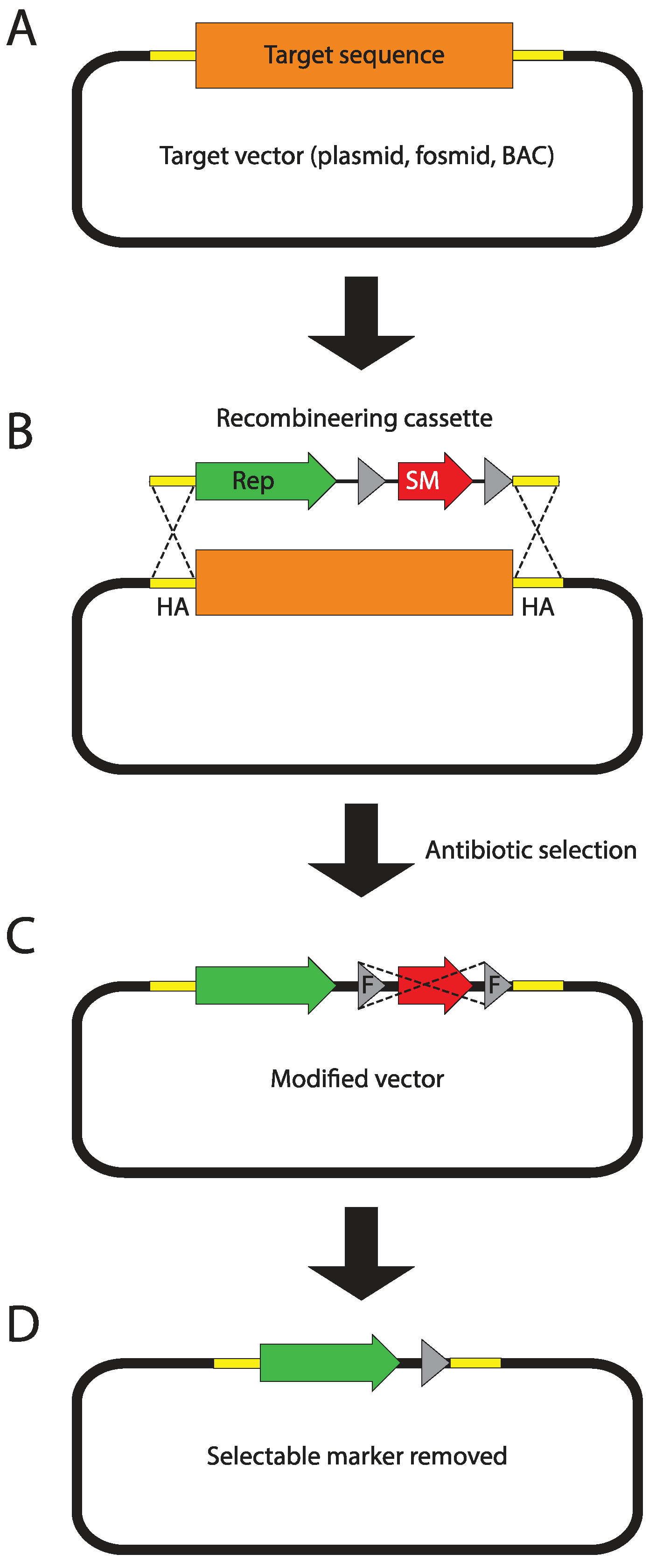
7.1. Getting Constructs in
7.2. Express What You Want, Where You Want, and When You Want
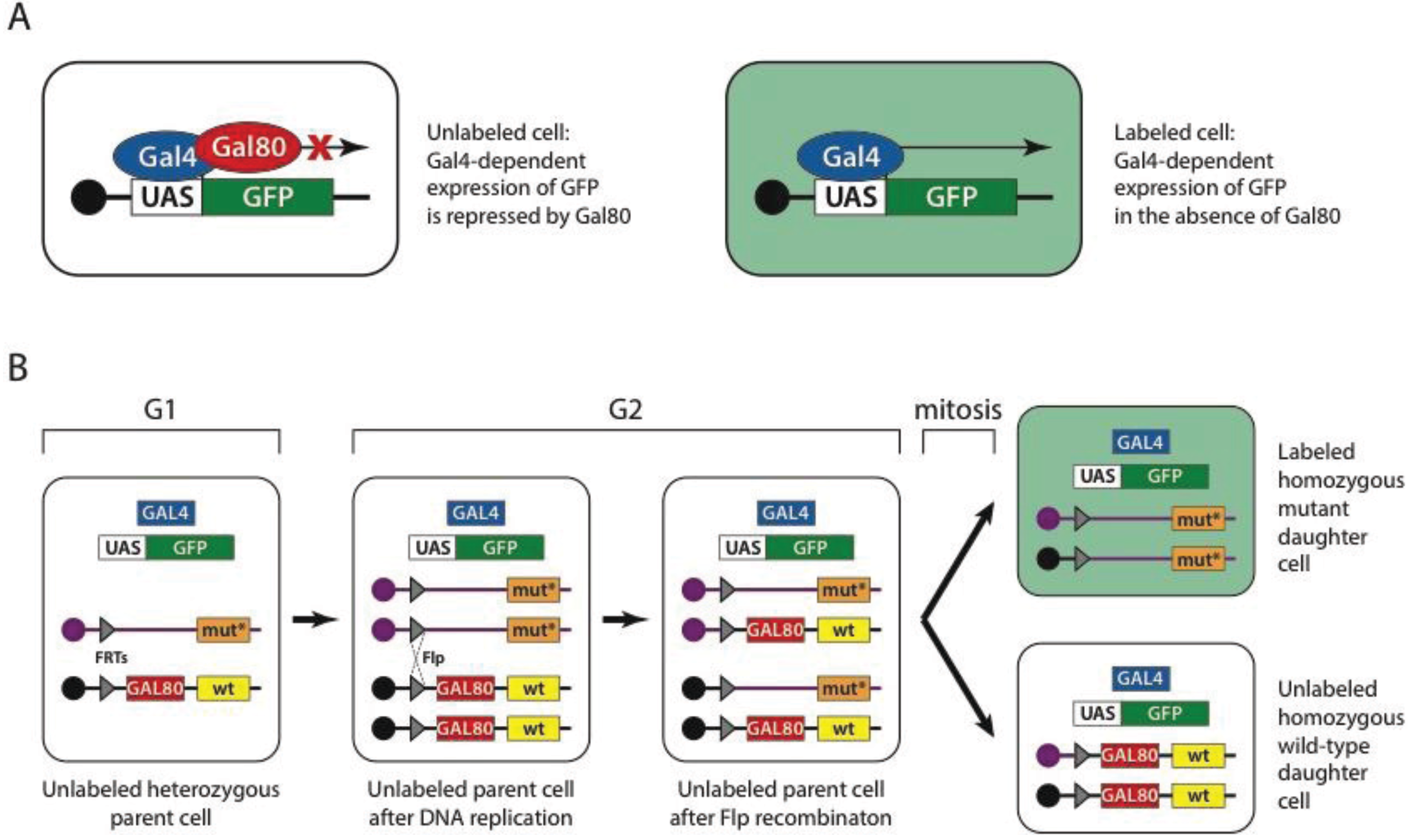
7.3. Mutant Tissue on Demand
7.4. Bright Rescue

8. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Fleischmann, R.D.; Adams, M.D.; White, O.; Clayton, R.A.; Kirkness, E.F.; Kerlavage, A.R.; Bult, C.J.; Tomb, J.F.; Dougherty, B.A.; Merrick, J.M.; et al. Whole-genome random sequencing and assembly of haemophilus influenzae Rd. Science 1995, 269, 496–512. [Google Scholar]
- Clayton, R.A.; White, O.; Ketchum, K.A.; Venter, J.C. The first genome from the third domain of life. Nature 1997, 387, 459–462. [Google Scholar] [CrossRef]
- C. elegans Sequencing Consortium. Genome sequence of the nematode C. elegans: A platform for investigating biology. Science 1998, 282, 2012–2018. [CrossRef]
- Adams, M.D.; Celniker, S.E.; Holt, R.A.; Evans, C.A.; Gocayne, J.D.; Amanatides, P.G.; Scherer, S.E.; Li, P.W.; Hoskins, R.A.; Galle, R.F.; et al. The genome sequence of drosophila melanogaster. Science 2000, 287, 2185–2195. [Google Scholar] [CrossRef]
- Lewis, E.B. A gene complex controlling segmentation in drosophila. Nature 1978, 276, 565–570. [Google Scholar] [CrossRef]
- Nusslein-Volhard, C.; Wieschaus, E. Mutations affecting segment number and polarity in drosophila. Nature 1980, 287, 795–801. [Google Scholar] [CrossRef]
- Gehring, W.J.; Hiromi, Y. Homeotic genes and the homeobox. Annu. Rev. Genet. 1986, 20, 147–173. [Google Scholar] [CrossRef]
- Lawrence, P.A.; Morata, G. Homeobox genes: Their function in drosophila segmentation and pattern formation. Cell 1994, 78, 181–189. [Google Scholar] [CrossRef]
- Krumlauf, R. Hox genes in vertebrate development. Cell 1994, 78, 191–201. [Google Scholar] [CrossRef]
- Quiring, R.; Walldorf, U.; Kloter, U.; Gehring, W.J. Homology of the eyeless gene of drosophila to the small eye gene in mice and aniridia in humans. Science 1994, 265, 785–789. [Google Scholar]
- Ton, C.C.; Hirvonen, H.; Miwa, H.; Weil, M.M.; Monaghan, P.; Jordan, T.; van Heyningen, V.; Hastie, N.D.; Meijers-Heijboer, H.; Drechsler, M.; et al. Positional cloning and characterization of a paired box- and homeobox-containing gene from the aniridia region. Cell 1991, 67, 1059–1074. [Google Scholar] [CrossRef]
- Hill, R.E.; Favor, J.; Hogan, B.L.; Ton, C.C.; Saunders, G.F.; Hanson, I.M.; Prosser, J.; Jordan, T.; Hastie, N.D.; van Heyningen, V. Mouse small eye results from mutations in a paired-like homeobox-containing gene. Nature 1991, 354, 522–525. [Google Scholar] [CrossRef]
- Halder, G.; Callaerts, P.; Flister, S.; Walldorf, U.; Kloter, U.; Gehring, W.J. Eyeless initiates the expression of both sine oculis and eyes absent during drosophila compound eye development. Development 1998, 125, 2181–2191. [Google Scholar]
- Kumar, J.P. The sine oculis homeobox (six) family of transcription factors as regulators of development and disease. Cell. Mol. Life Sci. 2009, 66, 565–583. [Google Scholar] [CrossRef]
- Jarman, A.P.; Grell, E.H.; Ackerman, L.; Jan, L.Y.; Jan, Y.N. Atonal is the proneural gene for drosophila photoreceptors. Nature 1994, 369, 398–400. [Google Scholar]
- Jarman, A.P.; Grau, Y.; Jan, L.Y.; Jan, Y.N. Atonal is a proneural gene that directs chordotonal organ formation in the drosophila peripheral nervous system. Cell 1993, 73, 1307–1321. [Google Scholar] [CrossRef]
- Brown, N.L.; Patel, S.; Brzezinski, J.; Glaser, T. Math5 is required for retinal ganglion cell and optic nerve formation. Development 2001, 128, 2497–2508. [Google Scholar]
- Bermingham, N.A.; Hassan, B.A.; Price, S.D.; Vollrath, M.A.; Ben-Arie, N.; Eatock, R.A.; Bellen, H.J.; Lysakowski, A.; Zoghbi, H.Y. Math1: An essential gene for the generation of inner ear hair cells. Science 1999, 284, 1837–1841. [Google Scholar] [CrossRef]
- Tabata, T.; Eaton, S.; Kornberg, T.B. The drosophila hedgehog gene is expressed specifically in posterior compartment cells and is a target of engrailed regulation. Genes Dev. 1992, 6, 2635–2645. [Google Scholar] [CrossRef]
- Lee, J.J.; von Kessler, D.P.; Parks, S.; Beachy, P.A. Secretion and localized transcription suggest a role in positional signaling for products of the segmentation gene hedgehog. Cell 1992, 71, 33–50. [Google Scholar] [CrossRef]
- Hooper, J.E.; Scott, M.P. The drosophila patched gene encodes a putative membrane protein required for segmental patterning. Cell 1989, 59, 751–765. [Google Scholar] [CrossRef]
- Nakano, Y.; Guerrero, I.; Hidalgo, A.; Taylor, A.; Whittle, J.R.; Ingham, P.W. A protein with several possible membrane-spanning domains encoded by the drosophila segment polarity gene patched. Nature 1989, 341, 508–513. [Google Scholar]
- Stone, D.M.; Hynes, M.; Armanini, M.; Swanson, T.A.; Gu, Q.; Johnson, R.L.; Scott, M.P.; Pennica, D.; Goddard, A.; Phillips, H.; et al. The tumour-suppressor gene patched encodes a candidate receptor for sonic hedgehog. Nature 1996, 384, 129–134. [Google Scholar] [CrossRef]
- Marigo, V.; Davey, R.A.; Zuo, Y.; Cunningham, J.M.; Tabin, C.J. Biochemical evidence that patched is the hedgehog receptor. Nature 1996, 384, 176–179. [Google Scholar] [CrossRef]
- Bhanot, P.; Brink, M.; Samos, C.H.; Hsieh, J.C.; Wang, Y.; Macke, J.P.; Andrew, D.; Nathans, J.; Nusse, R. A new member of the frizzled family from drosophila functions as a wingless receptor. Nature 1996, 382, 225–230. [Google Scholar] [CrossRef]
- Siegfried, E.; Chou, T.B.; Perrimon, N. Wingless signaling acts through zeste-white 3, the drosophila homolog of glycogen synthase kinase-3, to regulate engrailed and establish cell fate. Cell 1992, 71, 1167–1179. [Google Scholar] [CrossRef]
- Wehrli, M.; Dougan, S.T.; Caldwell, K.; O'Keefe, L.; Schwartz, S.; Vaizel-Ohayon, D.; Schejter, E.; Tomlinson, A.; DiNardo, S. Arrow encodes an ldl-receptor-related protein essential for wingless signalling. Nature 2000, 407, 527–530. [Google Scholar] [CrossRef]
- Brunner, E.; Peter, O.; Schweizer, L.; Basler, K. Pangolin encodes a lef-1 homologue that acts downstream of armadillo to transduce the wingless signal in drosophila. Nature 1997, 385, 829–833. [Google Scholar] [CrossRef]
- Adler, P.N. The genetic control of tissue polarity in drosophila. BioEssays 1992, 14, 735–741. [Google Scholar]
- Eaton, S.; Julicher, F. Cell flow and tissue polarity patterns. Curr. Opin. Genet. Dev. 2011, 21, 747–752. [Google Scholar] [CrossRef]
- Wharton, K.A.; Johansen, K.M.; Xu, T.; Artavanis-Tsakonas, S. Nucleotide sequence from the neurogenic locus notch implies a gene product that shares homology with proteins containing egf-like repeats. Cell 1985, 43, 567–581. [Google Scholar]
- Artavanis-Tsakonas, S.; Matsuno, K.; Fortini, M.E. Notch signaling. Science 1995, 268, 225–232. [Google Scholar]
- Artavanis-Tsakonas, S.; Rand, M.D.; Lake, R.J. Notch signaling: Cell fate control and signal integration in development. Science 1999, 284, 770–776. [Google Scholar] [CrossRef]
- Wu, S.; Huang, J.; Dong, J.; Pan, D. Hippo encodes a STE-20 family protein kinase that restricts cell proliferation and promotes apoptosis in conjunction with salvador and warts. Cell 2003, 114, 445–456. [Google Scholar]
- Harvey, K.F.; Pfleger, C.M.; Hariharan, I.K. The drosophila mst ortholog, hippo, restricts growth and cell proliferation and promotes apoptosis. Cell 2003, 114, 457–467. [Google Scholar]
- Udan, R.S.; Kango-Singh, M.; Nolo, R.; Tao, C.; Halder, G. Hippo promotes proliferation arrest and apoptosis in the salvador/warts pathway. Nat. Cell Biol. 2003, 5, 914–920. [Google Scholar] [CrossRef]
- Fortini, M.E.; Bonini, N.M. Modeling human neurodegenerative diseases in drosophila: On a wing and a prayer. Trends Genet. 2000, 16, 161–167. [Google Scholar] [CrossRef]
- Lloyd, T.E.; Taylor, J.P. Flightless flies: Drosophila models of neuromuscular disease. Ann. N. Y. Acad. Sci. 2010, 1184, e1–e20. [Google Scholar]
- Potter, C.J.; Turenchalk, G.S.; Xu, T. Drosophila in cancer research. An expanding role. Trends Genet. 2000, 16, 33–39. [Google Scholar] [CrossRef]
- Jackson, G.R.; Salecker, I.; Dong, X.; Yao, X.; Arnheim, N.; Faber, P.W.; MacDonald, M.E.; Zipursky, S.L. Polyglutamine-expanded human huntingtin transgenes induce degeneration of drosophila photoreceptor neurons. Neuron 1998, 21, 633–642. [Google Scholar] [CrossRef]
- Warrick, J.M.; Paulson, H.L.; Gray-Board, G.L.; Bui, Q.T.; Fischbeck, K.H.; Pittman, R.N.; Bonini, N.M. Expanded polyglutamine protein forms nuclear inclusions and causes neural degeneration in drosophila. Cell 1998, 93, 939–949. [Google Scholar]
- Morales, J.; Hiesinger, P.R.; Schroeder, A.J.; Kume, K.; Verstreken, P.; Jackson, F.R.; Nelson, D.L.; Hassan, B.A. Drosophila fragile x protein, dfxr, regulates neuronal morphology and function in the brain. Neuron 2002, 34, 961–972. [Google Scholar] [CrossRef]
- Luo, L.Q.; Martin-Morris, L.E.; White, K. Identification, secretion, and neural expression of appl, a drosophila protein similar to human amyloid protein precursor. J. Neurosci. 1990, 10, 3849–3861. [Google Scholar]
- Luo, L.; Tully, T.; White, K. Human amyloid precursor protein ameliorates behavioral deficit of flies deleted for appl gene. Neuron 1992, 9, 595–605. [Google Scholar] [CrossRef]
- Ye, Y.; Fortini, M.E. Apoptotic activities of wild-type and alzheimer’s disease-related mutant presenilins in drosophila melanogaster. J. Cell Biol. 1999, 146, 1351–1364. [Google Scholar] [CrossRef]
- Wittmann, C.W.; Wszolek, M.F.; Shulman, J.M.; Salvaterra, P.M.; Lewis, J.; Hutton, M.; Feany, M.B. Tauopathy in drosophila: Neurodegeneration without neurofibrillary tangles. Science 2001, 293, 711–714. [Google Scholar] [CrossRef]
- Feany, M.B.; Bender, W.W. A drosophila model of parkinson’s disease. Nature 2000, 404, 394–398. [Google Scholar] [CrossRef]
- Whitworth, A.J.; Theodore, D.A.; Greene, J.C.; Benes, H.; Wes, P.D.; Pallanck, L.J. Increased glutathione s-transferase activity rescues dopaminergic neuron loss in a drosophila model of parkinson’s disease. Proc. Natl. Acad. Sci. USA 2005, 102, 8024–8029. [Google Scholar]
- Parkes, T.L.; Elia, A.J.; Dickinson, D.; Hilliker, A.J.; Phillips, J.P.; Boulianne, G.L. Extension of drosophila lifespan by overexpression of human sod1 in motorneurons. Nat. Genet. 1998, 19, 171–174. [Google Scholar]
- Shahidullah, M.; Le Marchand, S.J.; Fei, H.; Zhang, J.; Pandey, U.B.; Dalva, M.B.; Pasinelli, P.; Levitan, I.B. Defects in synapse structure and function precede motor neuron degeneration in drosophila models of fus-related als. J. Neurosci. 2013, 33, 19590–19598. [Google Scholar] [CrossRef]
- Van der Plas, M.C.; Pilgram, G.S.; Plomp, J.J.; de Jong, A.; Fradkin, L.G.; Noordermeer, J.N. Dystrophin is required for appropriate retrograde control of neurotransmitter release at the drosophila neuromuscular junction. J. Neurosci. 2006, 26, 333–344. [Google Scholar]
- Moore, M.S.; DeZazzo, J.; Luk, A.Y.; Tully, T.; Singh, C.M.; Heberlein, U. Ethanol intoxication in drosophila: Genetic and pharmacological evidence for regulation by the camp signaling pathway. Cell 1998, 93, 997–1007. [Google Scholar] [CrossRef]
- Kaun, K.R.; Azanchi, R.; Maung, Z.; Hirsh, J.; Heberlein, U. A drosophila model for alcohol reward. Nat. Neurosci. 2011, 14, 612–619. [Google Scholar]
- McClung, C.; Hirsh, J. Stereotypic behavioral responses to free-base cocaine and the development of behavioral sensitization in drosophila. Curr. Biol. 1998, 8, 109–112. [Google Scholar] [CrossRef]
- Al-Anzi, B.; Sapin, V.; Waters, C.; Zinn, K.; Wyman, R.J.; Benzer, S. Obesity-blocking neurons in drosophila. Neuron 2009, 63, 329–341. [Google Scholar] [CrossRef]
- Baker, K.D.; Thummel, C.S. Diabetic larvae and obese flies-emerging studies of metabolism in drosophila. Cell Metab. 2007, 6, 257–266. [Google Scholar] [CrossRef]
- Wolf, M.J.; Amrein, H.; Izatt, J.A.; Choma, M.A.; Reedy, M.C.; Rockman, H.A. Drosophila as a model for the identification of genes causing adult human heart disease. Proc. Natl. Acad. Sci. USA 2006, 103, 1394–1399. [Google Scholar] [CrossRef]
- Roeder, T.; Isermann, K.; Kallsen, K.; Uliczka, K.; Wagner, C. A drosophila asthma model—What the fly tells us about inflammatory diseases of the lung. Adv. Exp. Med. Biol. 2012, 710, 37–47. [Google Scholar] [CrossRef]
- Xu, T.; Wang, W.; Zhang, S.; Stewart, R.A.; Yu, W. Identifying tumor suppressors in genetic mosaics: The drosophila lats gene encodes a putative protein kinase. Development 1995, 121, 1053–1063. [Google Scholar]
- Pagliarini, R.A.; Xu, T. A genetic screen in drosophila for metastatic behavior. Science 2003, 302, 1227–1231. [Google Scholar] [CrossRef]
- Januschke, J.; Gonzalez, C. Drosophila asymmetric division, polarity and cancer. Oncogene 2008, 27, 6994–7002. [Google Scholar] [CrossRef]
- Brumby, A.M.; Richardson, H.E. Using drosophila melanogaster to map human cancer pathways. Nat. Rev. Cancer 2005, 5, 626–639. [Google Scholar] [CrossRef]
- Vidal, M.; Cagan, R.L. Drosophila models for cancer research. Curr. Opin. Genet. Dev. 2006, 16, 10–16. [Google Scholar] [CrossRef]
- Miles, W.O.; Dyson, N.J.; Walker, J.A. Modeling tumor invasion and metastasis in drosophila. Dis. Models Mech. 2011, 4, 753–761. [Google Scholar] [CrossRef]
- Gonzalez, C. Drosophila melanogaster: A model and a tool to investigate malignancy and identify new therapeutics. Nat. Rev. Cancer 2013, 13, 172–183. [Google Scholar] [CrossRef]
- Bosco, G.; Campbell, P.; Leiva-Neto, J.T.; Markow, T.A. Analysis of drosophila species genome size and satellite DNA content reveals significant differences among strains as well as between species. Genetics 2007, 177, 1277–1290. [Google Scholar] [CrossRef]
- Hoskins, R.A.; Smith, C.D.; Carlson, J.W.; Carvalho, A.B.; Halpern, A.; Kaminker, J.S.; Kennedy, C.; Mungall, C.J.; Sullivan, B.A.; Sutton, G.G.; et al. Heterochromatic sequences in a drosophila whole-genome shotgun assembly. Genome Biol. 2002. [Google Scholar] [CrossRef] [Green Version]
- Misra, S.; Crosby, M.A.; Mungall, C.J.; Matthews, B.B.; Campbell, K.S.; Hradecky, P.; Huang, Y.; Kaminker, J.S.; Millburn, G.H.; Prochnik, S.E.; et al. Annotation of the drosophila melanogaster euchromatic genome: A systematic review. Genome Biol. 2002. [Google Scholar] [CrossRef] [Green Version]
- Stapleton, M.; Carlson, J.; Brokstein, P.; Yu, C.; Champe, M.; George, R.; Guarin, H.; Kronmiller, B.; Pacleb, J.; Park, S.; et al. A drosophila full-length cdna resource. Genome Biol. 2002. [Google Scholar] [CrossRef]
- Tomancak, P.; Beaton, A.; Weiszmann, R.; Kwan, E.; Shu, S.; Lewis, S.E.; Richards, S.; Ashburner, M.; Hartenstein, V.; Celniker, S.E.; et al. Systematic determination of patterns of gene expression during drosophila embryogenesis. Genome Biol. 2002. [Google Scholar] [CrossRef] [Green Version]
- Kaminker, J.S.; Bergman, C.M.; Kronmiller, B.; Carlson, J.; Svirskas, R.; Patel, S.; Frise, E.; Wheeler, D.A.; Lewis, S.E.; Rubin, G.M.; et al. The transposable elements of the drosophila melanogaster euchromatin: A genomics perspective. Genome Biol. 2002. [Google Scholar] [CrossRef] [Green Version]
- Ohler, U.; Liao, G.C.; Niemann, H.; Rubin, G.M. Computational analysis of core promoters in the drosophila genome. Genome Biol. 2002. [Google Scholar] [CrossRef]
- Marygold, S.J.; Leyland, P.C.; Seal, R.L.; Goodman, J.L.; Thurmond, J.; Strelets, V.B.; Wilson, R.J.; the FlyBase consortium. Flybase: Improvements to the bibliography. Nucleic Acids Res. 2013, 41, D751–D757. [Google Scholar] [CrossRef]
- Daines, B.; Wang, H.; Wang, L.; Li, Y.; Han, Y.; Emmert, D.; Gelbart, W.; Wang, X.; Li, W.; Gibbs, R.; et al. The drosophila melanogaster transcriptome by paired-end RNA sequencing. Genome Res. 2011, 21, 315–324. [Google Scholar] [CrossRef]
- Mungall, C.J.; Misra, S.; Berman, B.P.; Carlson, J.; Frise, E.; Harris, N.; Marshall, B.; Shu, S.; Kaminker, J.S.; Prochnik, S.E.; et al. An integrated computational pipeline and database to support whole-genome sequence annotation. Genome Biol. 2002. [Google Scholar] [CrossRef]
- Lewis, S.E.; Searle, S.M.; Harris, N.; Gibson, M.; Lyer, V.; Richter, J.; Wiel, C.; Bayraktaroglir, L.; Birney, E.; Crosby, M.A.; et al. Apollo: A sequence annotation editor. Genome Biol. 2002. [Google Scholar] [CrossRef]
- Reese, M.G.; Kulp, D.; Tammana, H.; Haussler, D. Genie—Gene finding in drosophila melanogaster. Genome Res. 2000, 10, 529–538. [Google Scholar] [CrossRef]
- Wood, V.; Gwilliam, R.; Rajandream, M.A.; Lyne, M.; Lyne, R.; Stewart, A.; Sgouros, J.; Peat, N.; Hayles, J.; Baker, S.; et al. The genome sequence of schizosaccharomyces pombe. Nature 2002, 415, 871–880. [Google Scholar] [CrossRef] [Green Version]
- Stein, L.D.; Bao, Z.; Blasiar, D.; Blumenthal, T.; Brent, M.R.; Chen, N.; Chinwalla, A.; Clarke, L.; Clee, C.; Coghlan, A.; et al. The genome sequence of caenorhabditis briggsae: A platform for comparative genomics. PLoS Biol. 2003, 1, e45. [Google Scholar]
- Richards, S.; Liu, Y.; Bettencourt, B.R.; Hradecky, P.; Letovsky, S.; Nielsen, R.; Thornton, K.; Hubisz, M.J.; Chen, R.; Meisel, R.P.; et al. Comparative genome sequencing of drosophila pseudoobscura: Chromosomal, gene, and cis-element evolution. Genome Res. 2005, 15, 1–18. [Google Scholar] [CrossRef]
- Drosophila 12 Genomes Consortium; Clark, A.G.; Eisen, M.B.; Smith, D.R.; Bergman, C.M.; Oliver, B.; Markow, T.A.; Kaufman, T.C.; Kellis, M.; Gelbart, W.; et al. Evolution of genes and genomes on the drosophila phylogeny. Nature 2007, 450, 203–218. [Google Scholar] [CrossRef] [Green Version]
- Lin, M.F.; Carlson, J.W.; Crosby, M.A.; Matthews, B.B.; Yu, C.; Park, S.; Wan, K.H.; Schroeder, A.J.; Gramates, L.S.; St Pierre, S.E.; et al. Revisiting the protein-coding gene catalog of drosophila melanogaster using 12 fly genomes. Genome Res. 2007, 17, 1823–1836. [Google Scholar] [CrossRef]
- Stark, A.; Lin, M.F.; Kheradpour, P.; Pedersen, J.S.; Parts, L.; Carlson, J.W.; Crosby, M.A.; Rasmussen, M.D.; Roy, S.; Deoras, A.N.; et al. Discovery of functional elements in 12 drosophila genomes using evolutionary signatures. Nature 2007, 450, 219–232. [Google Scholar] [CrossRef]
- Majoros, W.H.; Ohler, U. Modeling the evolution of regulatory elements by simultaneous detection and alignment with phylogenetic pair hmms. PLoS Comput. Biol. 2010, 6, e1001037. [Google Scholar] [CrossRef]
- Arunachalam, M.; Jayasurya, K.; Tomancak, P.; Ohler, U. An alignment-free method to identify candidate orthologous enhancers in multiple drosophila genomes. Bioinformatics 2010, 26, 2109–2115. [Google Scholar] [CrossRef]
- Aerts, S.; Quan, X.J.; Claeys, A.; Naval Sanchez, M.; Tate, P.; Yan, J.; Hassan, B.A. Robust target gene discovery through transcriptome perturbations and genome-wide enhancer predictions in drosophila uncovers a regulatory basis for sensory specification. PLoS Biol. 2010, 8, e1000435. [Google Scholar] [CrossRef]
- Mackay, T.F. Quantitative trait loci in drosophila. Nat. Rev. Genet. 2001, 2, 11–20. [Google Scholar] [CrossRef]
- Hindorff, L.A.; Sethupathy, P.; Junkins, H.A.; Ramos, E.M.; Mehta, J.P.; Collins, F.S.; Manolio, T.A. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl. Acad. Sci. USA 2009, 106, 9362–9367. [Google Scholar] [CrossRef]
- Mackay, T.F.; Richards, S.; Stone, E.A.; Barbadilla, A.; Ayroles, J.F.; Zhu, D.; Casillas, S.; Han, Y.; Magwire, M.M.; Cridland, J.M.; et al. The drosophila melanogaster genetic reference panel. Nature 2012, 482, 173–178. [Google Scholar]
- King, E.G.; Merkes, C.M.; McNeil, C.L.; Hoofer, S.R.; Sen, S.; Broman, K.W.; Long, A.D.; Macdonald, S.J. Genetic dissection of a model complex trait using the drosophila synthetic population resource. Genome Res. 2012, 22, 1558–1566. [Google Scholar] [CrossRef]
- Ayroles, J.F.; Carbone, M.A.; Stone, E.A.; Jordan, K.W.; Lyman, R.F.; Magwire, M.M.; Rollmann, S.M.; Duncan, L.H.; Lawrence, F.; Anholt, R.R.; et al. Systems genetics of complex traits in drosophila melanogaster. Nat. Genet. 2009, 41, 299–307. [Google Scholar] [CrossRef]
- Massouras, A.; Waszak, S.M.; Albarca-Aguilera, M.; Hens, K.; Holcombe, W.; Ayroles, J.F.; Dermitzakis, E.T.; Stone, E.A.; Jensen, J.D.; Mackay, T.F.; et al. Genomic variation and its impact on gene expression in drosophila melanogaster. PLoS Genet. 2012, 8, e1003055. [Google Scholar] [CrossRef]
- Jordan, K.W.; Craver, K.L.; Magwire, M.M.; Cubilla, C.E.; Mackay, T.F.; Anholt, R.R. Genome-wide association for sensitivity to chronic oxidative stress in drosophila melanogaster. PLoS One 2012, 7, e38722. [Google Scholar]
- Jumbo-Lucioni, P.; Bu, S.; Harbison, S.T.; Slaughter, J.C.; Mackay, T.F.; Moellering, D.R.; de Luca, M. Nuclear genomic control of naturally occurring variation in mitochondrial function in drosophila melanogaster. BMC Genomics 2002. [Google Scholar] [CrossRef]
- Magwire, M.M.; Fabian, D.K.; Schweyen, H.; Cao, C.; Longdon, B.; Bayer, F.; Jiggins, F.M. Genome-wide association studies reveal a simple genetic basis of resistance to naturally coevolving viruses in drosophila melanogaster. PLoS Genet. 2012, 8, e1003057. [Google Scholar] [CrossRef]
- Harbison, S.T.; McCoy, L.J.; Mackay, T.F. Genome-wide association study of sleep in drosophila melanogaster. BMC Genomics 2013. [Google Scholar] [CrossRef]
- King, E.G.; Macdonald, S.J.; Long, A.D. Properties and power of the drosophila synthetic population resource for the routine dissection of complex traits. Genetics 2012, 191, 935–949. [Google Scholar] [CrossRef]
- ENCODE Project Consortium. The encode (encyclopedia of DNA elements) project. Science 2004, 306, 636–640. [Google Scholar] [CrossRef]
- Gerstein, M.B.; Lu, Z.J.; van Nostrand, E.L.; Cheng, C.; Arshinoff, B.I.; Liu, T.; Yip, K.Y.; Robilotto, R.; Rechtsteiner, A.; Ikegami, K.; et al. Integrative analysis of the caenorhabditis elegans genome by the modencode project. Science 2010, 330, 1775–1787. [Google Scholar] [CrossRef]
- modENCODE Consortium; Roy, S.; Ernst, J.; Kharchenko, P.V.; Kheradpour, P.; Negre, N.; Eaton, M.L.; Landolin, J.M.; Bristow, C.A.; Ma, L.; et al. Identification of functional elements and regulatory circuits by drosophila modencode. Science 2010, 330, 1787–1797. [Google Scholar] [CrossRef]
- Riddle, N.C.; Minoda, A.; Kharchenko, P.V.; Alekseyenko, A.A.; Schwartz, Y.B.; Tolstorukov, M.Y.; Gorchakov, A.A.; Jaffe, J.D.; Kennedy, C.; Linder-Basso, D.; et al. Plasticity in patterns of histone modifications and chromosomal proteins in drosophila heterochromatin. Genome Res. 2011, 21, 147–163. [Google Scholar] [CrossRef]
- Kharchenko, P.V.; Alekseyenko, A.A.; Schwartz, Y.B.; Minoda, A.; Riddle, N.C.; Ernst, J.; Sabo, P.J.; Larschan, E.; Gorchakov, A.A.; Gu, T.; et al. Comprehensive analysis of the chromatin landscape in drosophila melanogaster. Nature 2011, 471, 480–485. [Google Scholar] [CrossRef]
- Nègre, N.; Brown, C.D.; Ma, L.; Bristow, C.A.; Miller, S.W.; Wagner, U.; Kheradpour, P.; Eaton, M.L.; Loriaux, P.; Sealfon, R.; et al. A cis-regulatory map of the drosophila genome. Nature 2011, 471, 527–531. [Google Scholar] [CrossRef] [Green Version]
- Spradling, A.C.; Rubin, G.M. Transposition of cloned p elements into drosophila germ line chromosomes. Science 1982, 218, 341–347. [Google Scholar]
- Searles, L.L.; Jokerst, R.S.; Bingham, P.M.; Voelker, R.A.; Greenleaf, A.L. Molecular cloning of sequences from a drosophila RNA polymerase ii locus by p element transposon tagging. Cell 1982, 31, 585–592. [Google Scholar] [CrossRef]
- Rubin, G.M.; Spradling, A.C. Genetic transformation of drosophila with transposable element vectors. Science 1982, 218, 348–353. [Google Scholar]
- Cooley, L.; Kelley, R.; Spradling, A. Insertional mutagenesis of the drosophila genome with single P elements. Science 1988, 239, 1121–1128. [Google Scholar]
- Cooley, L.; Thompson, D.; Spradling, A.C. Constructing deletions with defined endpoints in drosophila. Proc. Natl. Acad. Sci. USA 1990, 87, 3170–3173. [Google Scholar] [CrossRef]
- O'Kane, C.J.; Gehring, W.J. Detection in situ of genomic regulatory elements in drosophila. Proc. Natl. Acad. Sci. USA 1987, 84, 9123–9127. [Google Scholar] [CrossRef]
- Bellen, H.J.; Levis, R.W.; Liao, G.; He, Y.; Carlson, J.W.; Tsang, G.; Evans-Holm, M.; Hiesinger, P.R.; Schulze, K.L.; Rubin, G.M.; et al. The bdgp gene disruption project: Single transposon insertions associated with 40% of drosophila genes. Genetics 2004, 167, 761–781. [Google Scholar] [CrossRef]
- Bellen, H.J.; Levis, R.W.; He, Y.; Carlson, J.W.; Evans-Holm, M.; Bae, E.; Kim, J.; Metaxakis, A.; Savakis, C.; Schulze, K.L.; et al. The drosophila gene disruption project: Progress using transposons with distinctive site specificities. Genetics 2011, 188, 731–743. [Google Scholar] [CrossRef]
- Jacobson, J.W.; Medhora, M.M.; Hartl, D.L. Molecular structure of a somatically unstable transposable element in drosophila. Proc. Natl. Acad. Sci. USA 1986, 83, 8684–8688. [Google Scholar] [CrossRef]
- Garza, D.; Medhora, M.; Koga, A.; Hartl, D.L. Introduction of the transposable element mariner into the germline of drosophila melanogaster. Genetics 1991, 128, 303–310. [Google Scholar]
- Franz, G.; Savakis, C. Minos, a new transposable element from drosophila hydei, is a member of the tc1-like family of transposons. Nucleic Acids Res. 1991, 19, 6646. [Google Scholar] [CrossRef]
- Loukeris, T.G.; Arca, B.; Livadaras, I.; Dialektaki, G.; Savakis, C. Introduction of the transposable element minos into the germ line of drosophila melanogaster. Proc. Natl. Acad. Sci. USA 1995, 92, 9485–9489. [Google Scholar] [CrossRef]
- Handler, A.M.; McCombs, S.D.; Fraser, M.J.; Saul, S.H. The lepidopteran transposon vector, piggybac, mediates germ-line transformation in the mediterranean fruit fly. Proc. Natl. Acad. Sci. USA 1998, 95, 7520–7525. [Google Scholar] [CrossRef]
- Handler, A.M.; Harrell, R.A., 2nd. Germline transformation of drosophila melanogaster with the piggybac transposon vector. Insect Mol. Biol. 1999, 8, 449–457. [Google Scholar] [CrossRef]
- Spradling, A.C.; Rubin, G.M. The effect of chromosomal position on the expression of the drosophila xanthine dehydrogenase gene. Cell 1983, 34, 47–57. [Google Scholar] [CrossRef]
- Levis, R.; Hazelrigg, T.; Rubin, G.M. Effects of genomic position on the expression of transduced copies of the white gene of drosophila. Science 1985, 229, 558–561. [Google Scholar]
- Taillebourg, E.; Dura, J.M. A novel mechanism for p element homing in drosophila. Proc. Natl. Acad. Sci. USA 1999, 96, 6856–6861. [Google Scholar] [CrossRef]
- Cheng, Y.; Kwon, D.Y.; Arai, A.L.; Mucci, D.; Kassis, J.A. P-element homing is facilitated by engrailed polycomb-group response elements in drosophila melanogaster. PLoS One 2012, 7, e30437. [Google Scholar]
- Thorpe, H.M.; Smith, M.C. In vitro site-specific integration of bacteriophage DNA catalyzed by a recombinase of the resolvase/invertase family. Proc. Natl. Acad. Sci. USA 1998, 95, 5505–5510. [Google Scholar] [CrossRef]
- Groth, A.C.; Fish, M.; Nusse, R.; Calos, M.P. Construction of transgenic drosophila by using the site-specific integrase from phage phic31. Genetics 2004, 166, 1775–1782. [Google Scholar]
- Venken, K.J.; He, Y.; Hoskins, R.A.; Bellen, H.J. P[acman]: A bac transgenic platform for targeted insertion of large DNA fragments in d. Melanogaster. Science 2006, 314, 1747–1751. [Google Scholar] [CrossRef]
- Bischof, J.; Maeda, R.K.; Hediger, M.; Karch, F.; Basler, K. An optimized transgenesis system for drosophila using germ-line-specific phic31 integrases. Proc. Natl. Acad. Sci. USA 2007, 104, 3312–3317. [Google Scholar] [CrossRef]
- Markstein, M.; Pitsouli, C.; Villalta, C.; Celniker, S.E.; Perrimon, N. Exploiting position effects and the gypsy retrovirus insulator to engineer precisely expressed transgenes. Nat. Genet. 2008, 40, 476–483. [Google Scholar] [CrossRef]
- Brand, A.H.; Perrimon, N. Targeted gene expression as a means of altering cell fates and generating dominant phenotypes. Development 1993, 118, 401–415. [Google Scholar]
- Manseau, L.; Baradaran, A.; Brower, D.; Budhu, A.; Elefant, F.; Phan, H.; Philp, A.V.; Yang, M.; Glover, D.; Kaiser, K.; et al. Gal4 enhancer traps expressed in the embryo, larval brain, imaginal discs, and ovary of drosophila. Dev. Dyn. 1997, 209, 310–322. [Google Scholar] [CrossRef]
- Hayashi, S.; Ito, K.; Sado, Y.; Taniguchi, M.; Akimoto, A.; Takeuchi, H.; Aigaki, T.; Matsuzaki, F.; Nakagoshi, H.; Tanimura, T.; et al. Getdb, a database compiling expression patterns and molecular locations of a collection of gal4 enhancer traps. Genesis 2002, 34, 58–61. [Google Scholar] [CrossRef]
- Horn, C.; Offen, N.; Nystedt, S.; Hacker, U.; Wimmer, E.A. Piggybac-based insertional mutagenesis and enhancer detection as a tool for functional insect genomics. Genetics 2003, 163, 647–661. [Google Scholar]
- Jenett, A.; Rubin, G.M.; Ngo, T.T.; Shepherd, D.; Murphy, C.; Dionne, H.; Pfeiffer, B.D.; Cavallaro, A.; Hall, D.; Jeter, J.; et al. A gal4-driver line resource for drosophila neurobiology. Cell Rep. 2012, 2, 991–1001. [Google Scholar] [CrossRef]
- Stark, A.; Dickson, B.J. Vt (vienna tile) gal4 Driver Lines. Available online: http://stockcenter.vdrc.at/control/vtlibrary/ (accessed on 10 January 2014).
- Suster, M.L.; Seugnet, L.; Bate, M.; Sokolowski, M.B. Refining gal4-driven transgene expression in drosophila with a gal80 enhancer-trap. Genesis 2004, 39, 240–245. [Google Scholar] [CrossRef]
- McGuire, S.E.; Le, P.T.; Osborn, A.J.; Matsumoto, K.; Davis, R.L. Spatiotemporal rescue of memory dysfunction in drosophila. Science 2003, 302, 1765–1768. [Google Scholar] [CrossRef]
- Lee, T.; Luo, L. Mosaic analysis with a repressible cell marker for studies of gene function in neuronal morphogenesis. Neuron 1999, 22, 451–461. [Google Scholar] [CrossRef]
- Han, D.D.; Stein, D.; Stevens, L.M. Investigating the function of follicular subpopulations during drosophila oogenesis through hormone-dependent enhancer-targeted cell ablation. Development 2000, 127, 573–583. [Google Scholar]
- Osterwalder, T.; Yoon, K.S.; White, B.H.; Keshishian, H. A conditional tissue-specific transgene expression system using inducible gal4. Proc. Natl. Acad. Sci. USA 2001, 98, 12596–12601. [Google Scholar] [CrossRef]
- Roman, G.; Endo, K.; Zong, L.; Davis, R.L. P[switch], a system for spatial and temporal control of gene expression in drosophila melanogaster. Proc. Natl. Acad. Sci. USA 2001, 98, 12602–12607. [Google Scholar] [CrossRef]
- Stebbins, M.J.; Urlinger, S.; Byrne, G.; Bello, B.; Hillen, W.; Yin, J.C. Tetracycline-inducible systems for drosophila. Proc. Natl. Acad. Sci. USA 2001, 98, 10775–10780. [Google Scholar] [CrossRef]
- Lai, S.L.; Lee, T. Genetic mosaic with dual binary transcriptional systems in drosophila. Nat. Neurosci. 2006, 9, 703–709. [Google Scholar] [CrossRef]
- Potter, C.J.; Tasic, B.; Russler, E.V.; Liang, L.; Luo, L. The q system: A repressible binary system for transgene expression, lineage tracing, and mosaic analysis. Cell 2010, 141, 536–548. [Google Scholar] [CrossRef]
- Yagi, R.; Mayer, F.; Basler, K. Refined lexa transactivators and their use in combination with the drosophila gal4 system. Proc. Natl. Acad. Sci. USA 2010, 107, 16166–16171. [Google Scholar] [CrossRef]
- Dang, D.T.; Perrimon, N. Use of a yeast site-specific recombinase to generate embryonic mosaics in drosophila. Dev. Genet. 1992, 13, 367–375. [Google Scholar] [CrossRef]
- Harrison, D.A.; Perrimon, N. Simple and efficient generation of marked clones in drosophila. Curr. Biol. 1993, 3, 424–433. [Google Scholar] [CrossRef]
- Golic, K.G. Site-specific recombination between homologous chromosomes in drosophila. Science 1991, 252, 958–961. [Google Scholar]
- Yu, H.H.; Chen, C.H.; Shi, L.; Huang, Y.; Lee, T. Twin-spot marcm to reveal the developmental origin and identity of neurons. Nat. Neurosci. 2009, 12, 947–953. [Google Scholar] [CrossRef]
- Venken, K.J.; Schulze, K.L.; Haelterman, N.A.; Pan, H.; He, Y.; Evans-Holm, M.; Carlson, J.W.; Levis, R.W.; Spradling, A.C.; Hoskins, R.A.; et al. Mimic: A highly versatile transposon insertion resource for engineering drosophila melanogaster genes. Nat. Methods 2011, 8, 737–743. [Google Scholar] [CrossRef]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef]
- Hannon, G.J. RNA interference. Nature 2002, 418, 244–251. [Google Scholar] [CrossRef]
- Kennerdell, J.R.; Carthew, R.W. Use of dsrna-mediated genetic interference to demonstrate that frizzled and frizzled 2 act in the wingless pathway. Cell 1998, 95, 1017–1026. [Google Scholar] [CrossRef]
- Misquitta, L.; Paterson, B.M. Targeted disruption of gene function in drosophila by RNA interference (RNA-i): A role for nautilus in embryonic somatic muscle formation. Proc. Natl. Acad. Sci. USA 1999, 96, 1451–1456. [Google Scholar] [CrossRef]
- Hammond, S.M.; Bernstein, E.; Beach, D.; Hannon, G.J. An RNA-directed nuclease mediates post-transcriptional gene silencing in drosophila cells. Nature 2000, 404, 293–296. [Google Scholar] [CrossRef]
- Kennerdell, J.R.; Carthew, R.W. Heritable gene silencing in drosophila using double-stranded RNA. Nat. Biotechnol. 2000, 18, 896–898. [Google Scholar] [CrossRef]
- Clemens, J.C.; Worby, C.A.; Simonson-Leff, N.; Muda, M.; Maehama, T.; Hemmings, B.A.; Dixon, J.E. Use of double-stranded RNA interference in drosophila cell lines to dissect signal transduction pathways. Proc. Natl. Acad. Sci. USA 2000, 97, 6499–6503. [Google Scholar]
- Kiger, A.A.; Baum, B.; Jones, S.; Jones, M.R.; Coulson, A.; Echeverri, C.; Perrimon, N. A functional genomic analysis of cell morphology using RNA interference. J. Biol. 2003. [Google Scholar] [CrossRef] [Green Version]
- Lum, L.; Yao, S.; Mozer, B.; Rovescalli, A.; von Kessler, D.; Nirenberg, M.; Beachy, P.A. Identification of hedgehog pathway components by RNAi in drosophila cultured cells. Science 2003, 299, 2039–2045. [Google Scholar] [CrossRef]
- Boutros, M.; Kiger, A.A.; Armknecht, S.; Kerr, K.; Hild, M.; Koch, B.; Haas, S.A.; Paro, R.; Perrimon, N.; Heidelberg Fly Array, C. Genome-wide RNAi analysis of growth and viability in drosophila cells. Science 2004, 303, 832–835. [Google Scholar] [CrossRef]
- Foley, E.; O’Farrell, P.H. Functional dissection of an innate immune response by a genome-wide RNAi screen. PLoS Biol. 2004, 2, E203. [Google Scholar] [CrossRef] [Green Version]
- Dietzl, G.; Chen, D.; Schnorrer, F.; Su, K.-C.; Barinova, Y.; Fellner, M.; Gasser, B.; Kinsey, K.; Oppel, S.; Scheiblauer, S.; et al. A genome-wide transgenic RNAi library for conditional gene inactivation in drosophila. Nature 2007, 448, 151–156. [Google Scholar] [CrossRef]
- Ueda, R. RNAi Fly—A Comprehensive RNAi-Mutant Fly Bank. Available online: http://www.shigen.nig.ac.jp/fly/nigfly/about/aboutrnai.jsp/ (accessed on 13 January 2014).
- Ni, J.-Q.; Liu, L.-P.; Binari, R.; Hardy, R.; Shim, H.-S.; Cavallaro, A.; Booker, M.; Pfeiffer, B.D.; Markstein, M.; Wang, H.; et al. A drosophila resource of transgenic RNAi lines for neurogenetics. Genetics 2009, 182, 1089–1100. [Google Scholar] [CrossRef]
- Schnorrer, F.; Schonbauer, C.; Langer, C.C.; Dietzl, G.; Novatchkova, M.; Schernhuber, K.; Fellner, M.; Azaryan, A.; Radolf, M.; Stark, A.; et al. Systematic genetic analysis of muscle morphogenesis and function in drosophila. Nature 2010, 464, 287–291. [Google Scholar] [CrossRef]
- Neely, G.G.; Kuba, K.; Cammarato, A.; Isobe, K.; Amann, S.; Zhang, L.; Murata, M.; Elmen, L.; Gupta, V.; Arora, S.; et al. A global in vivo drosophila RNAi screen identifies not3 as a conserved regulator of heart function. Cell 2010, 141, 142–153. [Google Scholar] [CrossRef]
- Pospisilik, J.A.; Schramek, D.; Schnidar, H.; Cronin, S.J.; Nehme, N.T.; Zhang, X.; Knauf, C.; Cani, P.D.; Aumayr, K.; Todoric, J.; et al. Drosophila genome-wide obesity screen reveals hedgehog as a determinant of brown versus white adipose cell fate. Cell 2010, 140, 148–160. [Google Scholar] [CrossRef]
- Neely, G.G.; Hess, A.; Costigan, M.; Keene, A.C.; Goulas, S.; Langeslag, M.; Griffin, R.S.; Belfer, I.; Dai, F.; Smith, S.B.; et al. A genome-wide drosophila screen for heat nociception identifies alpha2delta3 as an evolutionarily conserved pain gene. Cell 2010, 143, 628–638. [Google Scholar]
- Ghosh, A.; Kling, T.; Snaidero, N.; Sampaio, J.L.; Shevchenko, A.; Gras, H.; Geurten, B.; Gopfert, M.C.; Schulz, J.B.; Voigt, A.; et al. A global in vivo drosophila RNAi screen identifies a key role of ceramide phosphoethanolamine for glial ensheathment of axons. PLoS Genet. 2013, 9, e1003980. [Google Scholar]
- Czech, B.; Preall, J.B.; McGinn, J.; Hannon, G.J. A transcriptome-wide RNAi screen in the drosophila ovary reveals factors of the germline piRNA pathway. Mol. Cell 2013, 50, 749–761. [Google Scholar] [CrossRef]
- Kondo, S.; Booker, M.; Perrimon, N. Cross-species RNAi rescue platform in drosophila melanogaster. Genetics 2009, 183, 1165–1173. [Google Scholar]
- Langer, C.C.H.; Ejsmont, R.K.; Schönbauer, C.; Schnorrer, F.; Tomancak, P. In vivo RNAi rescue in drosophila melanogaster with genomic transgenes from drosophila pseudoobscura. PLoS One 2010, 5, e8928. [Google Scholar]
- Schulz, J.G.; David, G.; Hassan, B.A. A novel method for tissue-specific RNAi rescue in drosophila. Nucleic Acids Res. 2009, 37, e93. [Google Scholar]
- Lawrence, P.A.; Bodmer, R.; Vincent, J.P. Segmental patterning of heart precursors in drosophila. Development 1995, 121, 4303–4308. [Google Scholar]
- Murray, M.J.; Merritt, D.J.; Brand, A.H.; Whitington, P.M. In vivo dynamics of axon pathfinding in the drosophilia CNS: A time-lapse study of an identified motorneuron. J. Neurobiol. 1998, 37, 607–621. [Google Scholar] [CrossRef]
- Murphy, K.C. Use of bacteriophage lambda recombination functions to promote gene replacement in escherichia coli. J. Bacteriol. 1998, 180, 2063–2071. [Google Scholar]
- Zhang, Y.; Buchholz, F.; Muyrers, J.P.; Stewart, A.F. A new logic for DNA engineering using recombination in escherichia coli. Nat. Genet. 1998, 20, 123–128. [Google Scholar]
- Muyrers, J.P.; Zhang, Y.; Testa, G.; Stewart, A.F. Rapid modification of bacterial artificial chromosomes by et-recombination. Nucleic Acids Res. 1999, 27, 1555–1557. [Google Scholar] [CrossRef]
- Yu, D.; Ellis, H.M.; Lee, E.C.; Jenkins, N.A.; Copeland, N.G.; Court, D.L. An efficient recombination system for chromosome engineering in Escherichia coli. Proc. Natl. Acad. Sci. USA 2000, 97, 5978–5983. [Google Scholar]
- Venken, K.J.T.; Carlson, J.W.; Schulze, K.L.; Pan, H.; He, Y.; Spokony, R.; Wan, K.H.; Koriabine, M.; de Jong, P.J.; White, K.P.; et al. Versatile P[acman] bac libraries for transgenesis studies in drosophila melanogaster. Nat. Methods 2009, 6, 431–434. [Google Scholar]
- Ejsmont, R.K.; Sarov, M.; Winkler, S.; Lipinski, K.A.; Tomancak, P. A toolkit for high-throughput, cross-species gene engineering in drosophila. Nat. Methods 2009, 6, 435–437. [Google Scholar]
- Horn, C.; Jaunich, B.; Wimmer, E.A. Highly sensitive, fluorescent transformation marker for drosophila transgenesis. Dev. Genes Evol. 2000, 210, 623–629. [Google Scholar] [CrossRef]
- Poser, I.; Sarov, M.; Hutchins, J.R.; Heriche, J.K.; Toyoda, Y.; Pozniakovsky, A.; Weigl, D.; Nitzsche, A.; Hegemann, B.; Bird, A.W.; et al. Bac transgeneomics: A high-throughput method for exploration of protein function in mammals. Nat. Methods 2008, 5, 409–415. [Google Scholar]
- Wu, J.S.; Luo, L. A protocol for mosaic analysis with a repressible cell marker (Marcm) in drosophila. Nat. Protoc. 2006, 1, 2583–2589. [Google Scholar]
- Rong, Y.S.; Golic, K.G. Gene targeting by homologous recombination in drosophila. Science 2000, 288, 2013–2018. [Google Scholar] [CrossRef]
- Gong, W.J.; Golic, K.G. Ends-out, or replacement, gene targeting in drosophila. Proc. Natl. Acad. Sci. USA 2003, 100, 2556–2561. [Google Scholar] [CrossRef]
- Choi, C.M.; Vilain, S.; Langen, M.; van Kelst, S.; de Geest, N.; Yan, J.; Verstreken, P.; Hassan, B.A. Conditional mutagenesis in drosophila. Science 2009. [Google Scholar] [CrossRef]
- Huang, J.; Zhou, W.; Dong, W.; Watson, A.M.; Hong, Y. From the cover: Directed, efficient, and versatile modifications of the drosophila genome by genomic engineering. Proc. Natl. Acad. Sci. USA 2009, 106, 8284–8289. [Google Scholar]
- Gloor, G.B.; Nassif, N.A.; Johnson-Schlitz, D.M.; Preston, C.R.; Engels, W.R. Targeted gene replacement in drosophila via p element-induced gap repair. Science 1991, 253, 1110–1117. [Google Scholar] [CrossRef]
- Kim, Y.G.; Cha, J.; Chandrasegaran, S. Hybrid restriction enzymes: Zinc finger fusions to Fok I cleavage domain. Proc. Natl. Acad. Sci. USA 1996, 93, 1156–1160. [Google Scholar] [CrossRef]
- Bibikova, M.; Golic, M.; Golic, K.G.; Carroll, D. Targeted chromosomal cleavage and mutagenesis in drosophila using zinc-finger nucleases. Genetics 2002, 161, 1169–1175. [Google Scholar]
- Auer, T.O.; Duroure, K.; de Cian, A.; Concordet, J.P.; Del Bene, F. Highly efficient crispr/cas9-mediated knock-in in zebrafish by homology-independent DNA repair. Genome Res. 2014, 24, 142–153. [Google Scholar] [CrossRef]
- Beumer, K.; Bhattacharyya, G.; Bibikova, M.; Trautman, J.K.; Carroll, D. Efficient gene targeting in drosophila with zinc-finger nucleases. Genetics 2006, 172, 2391–2403. [Google Scholar]
- Maeder, M.L.; Thibodeau-Beganny, S.; Osiak, A.; Wright, D.A.; Anthony, R.M.; Eichtinger, M.; Jiang, T.; Foley, J.E.; Winfrey, R.J.; Townsend, J.A.; et al. Rapid “open-source” engineering of customized zinc-finger nucleases for highly efficient gene modification. Mol. Cell 2008, 31, 294–301. [Google Scholar] [CrossRef]
- Sander, J.D.; Dahlborg, E.J.; Goodwin, M.J.; Cade, L.; Zhang, F.; Cifuentes, D.; Curtin, S.J.; Blackburn, J.S.; Thibodeau-Beganny, S.; Qi, Y.; et al. Selection-free zinc-finger-nuclease engineering by context-dependent assembly (coda). Nat. Methods 2011, 8, 67–69. [Google Scholar] [CrossRef]
- Kim, S.; Lee, M.J.; Kim, H.; Kang, M.; Kim, J.S. Preassembled zinc-finger arrays for rapid construction of zfns. Nat. Methods 2011. [Google Scholar] [CrossRef]
- Doyon, Y.; Vo, T.D.; Mendel, M.C.; Greenberg, S.G.; Wang, J.; Xia, D.F.; Miller, J.C.; Urnov, F.D.; Gregory, P.D.; Holmes, M.C. Enhancing zinc-finger-nuclease activity with improved obligate heterodimeric architectures. Nat. Methods 2011, 8, 74–79. [Google Scholar]
- Mahfouz, M.M.; Li, L.; Shamimuzzaman, M.; Wibowo, A.; Fang, X.; Zhu, J.K. De novo-engineered transcription activator-like effector (tale) hybrid nuclease with novel DNA binding specificity creates double-strand breaks. Proc. Natl. Acad. Sci. USA 2011, 108, 2623–2628. [Google Scholar]
- Miller, J.C.; Tan, S.; Qiao, G.; Barlow, K.A.; Wang, J.; Xia, D.F.; Meng, X.; Paschon, D.E.; Leung, E.; Hinkley, S.J.; et al. A tale nuclease architecture for efficient genome editing. Nat. Biotechnol. 2011, 29, 143–148. [Google Scholar] [CrossRef]
- Morbitzer, R.; Elsaesser, J.; Hausner, J.; Lahaye, T. Assembly of custom tale-type DNA binding domains by modular cloning. Nucleic Acids Res. 2011, 39, 5790–5799. [Google Scholar] [CrossRef]
- Moscou, M.J.; Bogdanove, A.J. A simple cipher governs DNA recognition by tal effectors. Science 2009. [Google Scholar] [CrossRef]
- Brouns, S.J.; Jore, M.M.; Lundgren, M.; Westra, E.R.; Slijkhuis, R.J.; Snijders, A.P.; Dickman, M.J.; Makarova, K.S.; Koonin, E.V.; van der Oost, J. Small crispr rnas guide antiviral defense in prokaryotes. Science 2008, 321, 960–964. [Google Scholar] [CrossRef]
- Gasiunas, G.; Barrangou, R.; Horvath, P.; Siksnys, V. Cas9-crrna ribonucleoprotein complex mediates specific DNA cleavage for adaptive immunity in bacteria. Proc. Natl. Acad. Sci. USA 2012, 109, E2579–E2586. [Google Scholar]
- Jinek, M.; Chylinski, K.; Fonfara, I.; Hauer, M.; Doudna, J.A.; Charpentier, E. A programmable dual-RNA-guided DNA endonuclease in adaptive bacterial immunity. Science 2012, 337, 816–821. [Google Scholar] [CrossRef]
- Bassett, A.R.; Tibbit, C.; Ponting, C.P.; Liu, J.L. Highly efficient targeted mutagenesis of drosophila with the crispr/cas9 system. Cell Rep. 2013, 4, 220–228. [Google Scholar] [CrossRef]
- Gratz, S.J.; Cummings, A.M.; Nguyen, J.N.; Hamm, D.C.; Donohue, L.K.; Harrison, M.M.; Wildonger, J.; O’Connor-Giles, K.M. Genome engineering of drosophila with the crispr RNA-guided cas9 nuclease. Genetics 2013, 194, 1029–1035. [Google Scholar] [CrossRef]
- Yu, Z.; Ren, M.; Wang, Z.; Zhang, B.; Rong, Y.S.; Jiao, R.; Gao, G. Highly efficient genome modifications mediated by crispr/cas9 in drosophila. Genetics 2013, 195, 289–291. [Google Scholar] [CrossRef]
- Sebo, Z.L.; Lee, H.B.; Peng, Y.; Guo, Y. A simplified and efficient germline-specific crispr/cas9 system for drosophila genomic engineering. Fly 2014, 8, 52–57. [Google Scholar]
- Ren, X.; Sun, J.; Housden, B.E.; Hu, Y.; Roesel, C.; Lin, S.; Liu, L.-P.; Yang, Z.; Mao, D.; Sun, L.; et al. Optimized gene editing technology for drosophila melanogaster using germ line-specific cas9. Proc. Natl. Acad. Sci. USA 2013, 110, 19012–19017. [Google Scholar] [CrossRef]
- Gratz, S.J.; Ukken, F.P.; Rubinstein, C.D.; Thiede, G.; Donohue, L.K.; Cummings, A.M.; O’Connor-Giles, K.M. Highly specific and efficient crispr/cas9-catalyzed homology-directed repair in drosophila. Genetics 2014. [Google Scholar] [CrossRef]
- Kondo, S.; Ueda, R. Highly improved gene targeting by germline-specific cas9 expression in drosophila. Genetics 2013, 195, 715–721. [Google Scholar] [CrossRef]
- Baena-Lopez, L.A.; Alexandre, C.; Mitchell, A.; Pasakarnis, L.; Vincent, J.P. Accelerated homologous recombination and subsequent genome modification in drosophila. Development 2013, 140, 4818–4825. [Google Scholar] [CrossRef]
- Del Valle Rodriguez, A.; Didiano, D.; Desplan, C. Power tools for gene expression and clonal analysis in drosophila. Nat. Methods 2012, 9, 47–55. [Google Scholar]
- Venken, K.J.; Bellen, H.J. Genome-wide manipulations of drosophila melanogaster with transposons, flp recombinase, and phic31 integrase. Methods Mol. Biol. 2012, 859, 203–228. [Google Scholar] [CrossRef]
- Gratz, S.J.; Wildonger, J.; Harrison, M.M.; O’Connor-Giles, K.M. Crispr/cas9-mediated genome engineering and the promise of designer flies on demand. Fly 2013. [Google Scholar] [CrossRef]
- Reiter, L.T.; Potocki, L.; Chien, S.; Gribskov, M.; Bier, E. A systematic analysis of human disease-associated gene sequences in drosophila melanogaster. Genome Res. 2001, 11, 1114–1125. [Google Scholar] [CrossRef]
- Chien, S.; Reiter, L.T.; Bier, E.; Gribskov, M. Homophila: Human disease gene cognates in drosophila. Nucleic Acids Res. 2002, 30, 149–151. [Google Scholar] [CrossRef]
- Richter, A.; Boch, J. Designer tales team up for highly efficient gene induction. Nat. Methods 2013, 10, 207–208. [Google Scholar] [CrossRef]
- Crocker, J.; Stern, D.L. Tale-mediated modulation of transcriptional enhancers in vivo. Nat. Methods 2013, 10, 762–767. [Google Scholar] [CrossRef]
- Mali, P.; Esvelt, K.M.; Church, G.M. Cas9 as a versatile tool for engineering biology. Nat. Methods 2013, 10, 957–963. [Google Scholar] [CrossRef]
- modENCODE Consortium Modencode Comparative Genomics Whitepaper. Available online: http://www.genome.gov/Pages/Research/Sequencing/SeqProposals/modENCODE_ComparativeGenomics_WhitePaper.pdf (accessed on 16 January 2014).
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Ejsmont, R.K.; Hassan, B.A. The Little Fly that Could: Wizardry and Artistry of Drosophila Genomics. Genes 2014, 5, 385-414. https://doi.org/10.3390/genes5020385
Ejsmont RK, Hassan BA. The Little Fly that Could: Wizardry and Artistry of Drosophila Genomics. Genes. 2014; 5(2):385-414. https://doi.org/10.3390/genes5020385
Chicago/Turabian StyleEjsmont, Radoslaw K., and Bassem A. Hassan. 2014. "The Little Fly that Could: Wizardry and Artistry of Drosophila Genomics" Genes 5, no. 2: 385-414. https://doi.org/10.3390/genes5020385



