Polygenic Scores Predict Alcohol Problems in an Independent Sample and Show Moderation by the Environment
Abstract
:1. Introduction
2. Experimental Section
2.1. Avon Longitudinal Study of Parents and Children
2.1.1. Alcohol Problems Factor Score
2.1.2. Genotyping
2.2. FinnTwin12
2.2.1. Alcohol Problems, Parental Knowledge, and Peer Deviance
2.2.2. Genotyping
2.3. Analytic Plan
2.3.1. Genome-Wide Association Analysis in the ALSPAC Sample
2.3.2. Calculation of Polygenic Scores in FinnTwin12
| Polygenic threshold | Number of autosomal SNPs meeting threshold in ALSPAC | Number (percent) of SNPs available in FinnTwin12 |
|---|---|---|
| p ≤ 0.05 | 125,969 | 113,992 (90.5%) |
| p ≤ 0.10 | 250, 244 | 226,789 (90.6%) |
| p ≤ 0.20 | 495,760 | 449,273 (90.6%) |
| p ≤ 0.30 | 739,758 | 670,293 (90.6%) |
| p ≤ 0.40 | 984,167 | 891,782 (90.6%) |
| p ≤ 0.50 | 1,231,165 | 1,115,557 (90.6%) |
2.3.3. Polygenic Association and Moderation Analyses in FinnTwin12
3. Results and Discussion
3.1. Descriptive Statistics and Zero-Order Correlations
| Variable | M | SD | Min | Max |
|---|---|---|---|---|
| Alcohol problems (age 14), range 0–30 | 0.29 | 0.96 | 0 | 8 |
| Parental knowledge (age 14), range 4–16 | 6.62 | 2.08 | 4 | 15 |
| Peer deviance (age 14), range 4–16 | 7.91 | 3.14 | 4 | 16 |
| Polygenic score (p≤ 0.05 threshold) | −0.07 | 0.02 | −0.13 | 0.00 |
3.2. Polygenic Associations with Alcohol Problems
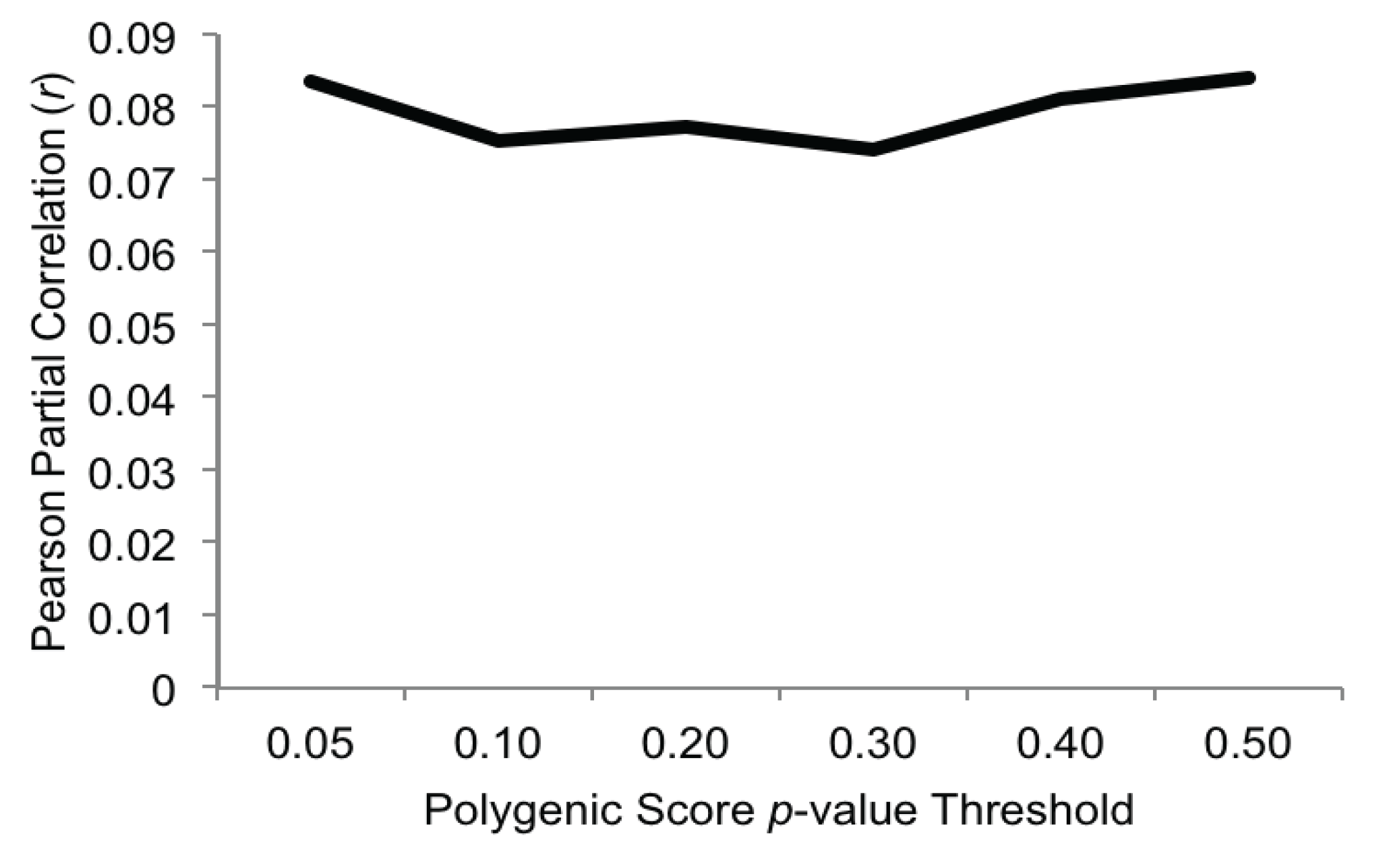
3.3. Gene-by-Environment Interactions
| Parental Knowledge | |||||
| b | SE | t | P | ΔR2 | |
| Intercept | 0.16 | 0.04 | 3.97 | <0.01 | -- |
| Sex | 0.23 | 0.06 | 4.17 | <0.01 | 0.006 |
| Polygenic score | 3.10 | 1.40 | 2.21 | 0.03 | 0.006 |
| Parental knowledge | 0.14 | 0.01 | 10.31 | <0.01 | 0.088 |
| Polygenic score × Parental knowledge | 1.54 | 0.68 | 2.27 | 0.02 | 0.003 |
| Peer Deviance | |||||
| b | SE | t | P | ΔR2 | |
| Intercept | 0.19 | 0.04 | 4.88 | <0.01 | -- |
| Sex | 0.17 | 0.05 | 3.07 | <0.01 | 0.006 |
| Polygenic score | 2.75 | 1.38 | 1.99 | 0.05 | 0.006 |
| Peer deviance | 0.11 | 0.01 | 12.43 | <0.01 | 0.120 |
| Polygenic score × Peer deviance | 0.94 | 0.44 | 2.11 | 0.04 | 0.003 |
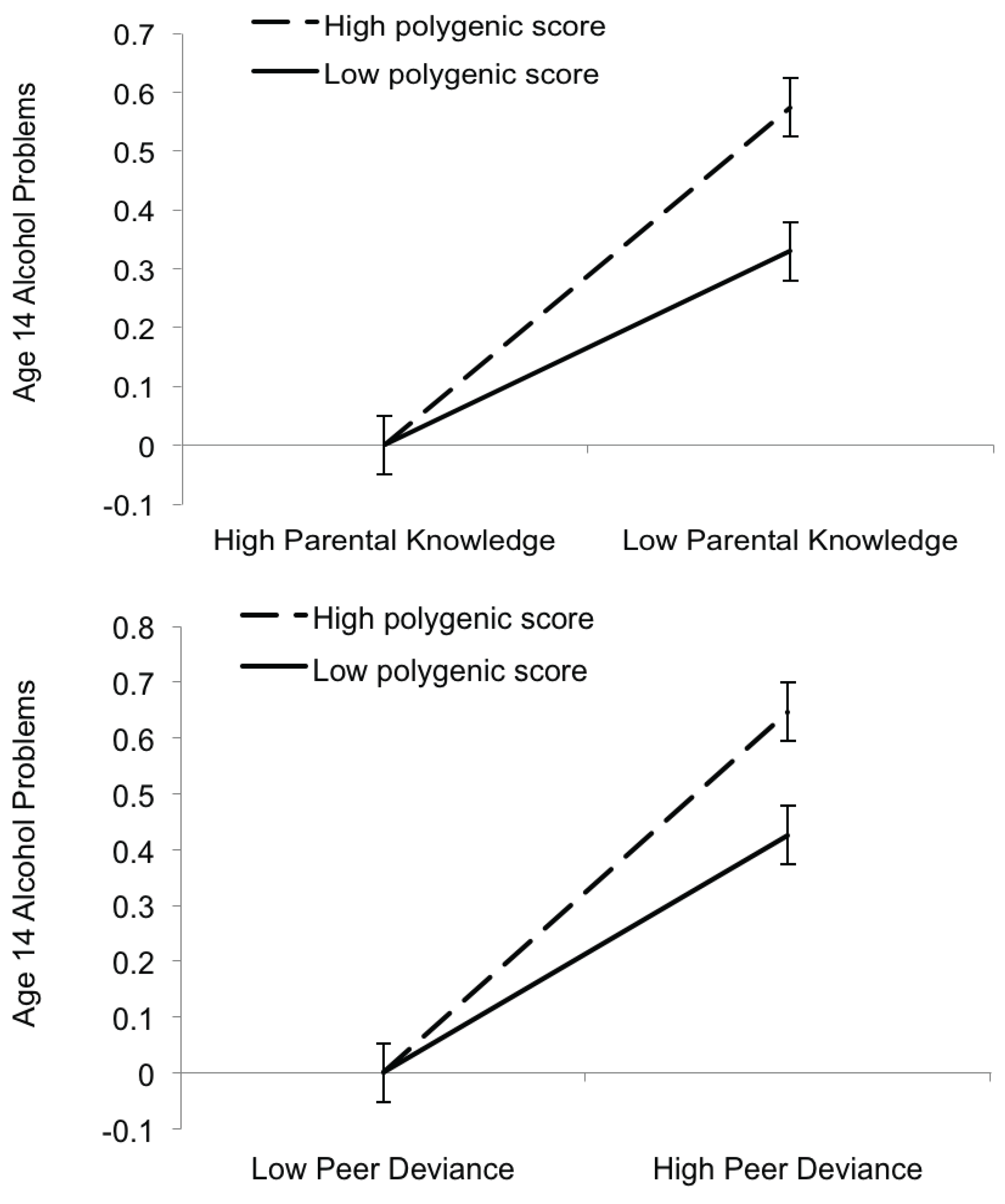
3.4. Set-Based Analyses Examining Enrichment for Gene-Environment Interaction among Top SNPs
3.5. Limitations
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Manolio, T.A.; Collins, F.S.; Cox, N.J.; Goldstein, D.B.; Hindorff, L.A.; Hunter, D.J.; McCarthy, M.I.; Ramos, E.M.; Cardon, L.R.; Chakravarti, A.; et al. Finding the missing heritability of complex diseases. Nature 2009, 461, 747–753. [Google Scholar] [CrossRef]
- Heath, A.C.; Bucholz, K.K.; Madden, P.A.F.; Dinwiddie, S.H.; Slutske, W.S.; Bierut, L.J.; Statham, D.J.; Dunne, M.P.; Whitfield, J.B.; Martin, N.G. Genetic and environmental contributions to alcohol dependence risk in a national twin sample: Consistency of findings in women and men. Psychol. Med. 1997, 27, 1381–1396. [Google Scholar] [CrossRef]
- Kendler, K.S.; Heath, A.C.; Neale, M.C.; Kessler, R.C.; Eaves, L.J. A population-based twin study of alcoholism in women. JAMA 1992, 268, 1877–1882. [Google Scholar] [CrossRef]
- Rose, R.J.; Dick, D.M.; Viken, R.J.; Kaprio, J. Gene-environment interaction in patterns of adolescent drinking: Regional residency moderates longitudinal influences in alcohol use. Alcohol. Clin. Exp. Res. 2001, 25, 637–643. [Google Scholar] [CrossRef]
- Prescott, C.A.; Kendler, K.S. Genetic and environmental contributions to alcohol abuse and dependence in a population-based sample of male twins. Am. J. Psychiatry 1999, 156, 34–40. [Google Scholar]
- Dick, D.M.; Viken, R.; Purcell, S.; Kaprio, J.; Pulkkinen, L.; Rose, R.J. Parental monitoring moderates the importance of genetic and environmental influences on adolescent smoking. J. Abnorm. Psychol. 2007, 116, 213–218. [Google Scholar] [CrossRef]
- Harden, K.P.; Hill, J.E.; Turkheimer, E.; Emery, R.E. Gene-environment correlation and interaction in peer effects on adolescent alcohol and tobacco use. Behav. Genet. 2008, 38, 339–347. [Google Scholar] [CrossRef]
- Button, T.M.M.; Corley, R.P.; Rhee, S.H.; Hewitt, J.K.; Young, S.E.; Stallings, M.C. Delinquent peer affiliation and conduct problems: A twin study. J. Abnorm. Psychol. 2007, 116, 554–564. [Google Scholar] [CrossRef]
- Hicks, B.M.; South, S.C.; DiRago, A.C.; Iacono, W.G.; McGue, M. Environmental adversity and increasing genetic risk for externalizing disorders. Arch. Gen. Psychiatry 2009, 66, 640–648. [Google Scholar] [CrossRef]
- Shanahan, M.J.; Hofer, S.M. Social context in gene-environment interactions: Retrospect and prospect. J. Gerontol. Ser. B Psychol. Sci. Soc. Sci. 2005, 60B, 65–76. [Google Scholar] [CrossRef]
- Hartz, S.M.; Short, S.E.; Saccone, N.L.; Culverhouse, R.; Chen, L.; Schwantes-An, T.H.; Coon, H.; Han, Y.; Stephens, S.H.; Sun, J.; et al. Increased genetic vulnerability to smoking at CHRNA5 in early-onset smokers. Arch. Gen. Psychiatry 2012, 69, 854–860. [Google Scholar] [CrossRef]
- Thomasson, H.R.; Edenberg, H.J.; Crabb, D.W.; Mai, X.L.; Jerome, R.E.; Li, T.K.; Wang, S.P.; Lin, Y.T.; Lu, R.B.; Yin, S.J. Alcohol and aldehyde dehydrogenase genotypes and alcoholism in Chinese men. Am. J. Hum. Genet. 1991, 48, 667–681. [Google Scholar]
- Luczak, S.E.; Glatt, S.J.; Wall, T.L. Meta-analyses of ALDH2 and ADH1B with alcohol dependence in Asians. Psychol. Bull. 2006, 132, 607–621. [Google Scholar]
- Whitfield, J.B. Alcohol dehydrogenase and alcohol dependence: Variation in genotype-associated risk between populations. Am. J. Hum. Genet. 2002, 71, 1247–1250. [Google Scholar] [CrossRef]
- Gelernter, J.; Kranzler, H.R.; Sherva, R.; Almasy, L.; Koesterer, R.; Smith, A.H.; Anton, R.; Preuss, U.W.; Ridinger, M.; Rujescu, D.; et al. Genome-wide association study of alcohol dependence:significant findings in African- and European-Americans including novel risk loci. Mol. Psychiatry 2014, 19, 41–49. [Google Scholar] [CrossRef]
- Edenberg, H.J.; Dick, D.M.; Xuei, X.; Tian, H.; Almasy, L.; Bauer, L.O.; Crowe, R.; Goate, A.; Hesselbrock, V.; Jones, K.A.; et al. Variations in GABRA2, encoding the α2 subunit of the GABA-A receptor are associated with alcohol dependence and with brain oscillations. Am. J. Hum. Genet. 2004, 74, 705–714. [Google Scholar] [CrossRef]
- Covault, J.; Gelernter, J.; Hesselbrock, V.; Nellissery, M.; Kranzler, H.R. Allelic and haplotypic association of GABRA2 with alcohol dependence. Am. J. Med. Genet. Part. B Neuropsychiatr. Genet. 2004, 129B, 104–109. [Google Scholar] [CrossRef]
- Duncan, L.; Keller, M.C. A critical review of the first ten years of measured gene-by-environment interaction research in psychiatry. Am. J. Psychiatry 2011, 168, 1041–1049. [Google Scholar]
- The International Schizophrenia Consortium Common polygenic variation contributes to risk of schizophrenia and bipolar disorder. Nature 2009, 460, 748–752.
- National Institute on Drug Abuse Monitoring the Future Study: Trends in Prevalence of Various Drugs (2009–2012). Available online: http://www.drugabuse.gov/related-topics/trends-statistics/monitoring-future/trends-in-prevalence-various-drugs/ (accessed on 10 December 2013).
- Substance Abuse and Mental Health Services Administration. Results from the 2012 National Survey on Drug Use and Health: Summary of National Findings; SAMHSA: Rockville, MD, USA, 2013.
- Kendler, K.S.; Schmitt, J.E.; Aggen, S.H.; Prescott, C.A. Genetic and environmental influences on alcohol, caffeine, cannabis, and nicotine use from adolescence to middle adulthood. Arch. Gen. Psychiatry 2008, 65, 674–682. [Google Scholar] [CrossRef]
- Golding, J.; Pembrey, M.; Jones, R.; ALSPAC Study Team. ALSPAC-The Avon Longitudinal Study of Parents and Children—I. Study methodology. Paediatr. Perinat. Epidemiol. 2001, 15, 74–87. [Google Scholar]
- Kaprio, J.; Pulkkinen, L.; Rose, R.J. Genetic and environmental factors in health-related behaviors: Studies on Finnish twins and twin families. Twin Res. 2002, 5, 366–371. [Google Scholar] [CrossRef]
- Boyd, A.; Golding, J.; Macleod, J.; Lawlor, D.A.; Fraser, A.; Henderson, J.; Molloy, L.; Ness, A.; Ring, S.; Smith, G.D. Cohort profile: The Children of the 90s’—The index offspring of the Avon Longitudinal Study of Parents and Children. Int. J. Epidemiol. 2012, 42, 1–17. [Google Scholar]
- Babor, T.F.; Higgins-Biddle, J.C.; Saunders, J.B.; Monteiro, M.G. AUDIT: The Alcohol Use Disorders Identification Test, 2nd ed.; World Health Organization: Geneva, Switzerland, 2001. [Google Scholar]
- American Psychiatric Association. Diagnostic and Statistical Manual of Mental Disorders (DSM-IV); American Psychiatric Association: Washington, DC, USA, 1994. [Google Scholar]
- Raghunathan, T.E.; Solenberger, P.W.; van Hoewyk, J. IVEware: Imputation and Variance Estimation Software; Institute for Social Research: Ann Arbor, MI, USA, 2002. [Google Scholar]
- Muthén, L.K.; Muthén, B.O. Mplus User’s Guide, 6th ed.; Muthén & Muthén: Los Angeles, CA, USA, 1998–2011. [Google Scholar]
- Li, Y.; Willer, C.J.; Ding, J.; Scheet, P.; Abecasis, G.R. MaCH: Using sequence and genotype data to estimate haplotypes and unobserved genotypes. Genet. Epidemiol. 2010, 34, 816–834. [Google Scholar] [CrossRef]
- Fatemifar, G.; Hoggart, C.J.; Paternoster, L.; Kemp, J.P.; Prokopenko, I.; Horikoshi, M.; Wright, V.J.; Tobias, J.H.; Richmond, S.; Zhurov, A.I.; et al. Genome-wide association study of primary tooth eruption identifies pleiotropic loci associated with height and craniofacial distances. Hum. Mol. Genet. 2013, 22, 3807–3817. [Google Scholar] [CrossRef]
- Bucholz, K.K.; Cadoret, R.; Cloninger, C.R.; Dinwiddie, S.H.; Hesselbrock, V.M.; Nurnberger, J.I., Jr.; Reich, T.; Schmidt, I.; Schuckit, M.A. A new, semi-structured psychiatric interview for use in genetic linkage studies: A report on the reliability of the SSAGA. J. Stud. Alcohol 1994, 55, 149–158. [Google Scholar]
- Chassin, L.; Pillow, D.R.; Curran, P.J.; Molina, B.S.G.; Barrera, M. Relation of parental alcoholism to early adolescent substance use: A test of 3 mediating mechanisms. J. Abnorm. Psychol. 1993, 102, 3–19. [Google Scholar]
- Broms, U.; Wedenoja, J.; Largeau, M.R.; Korhonen, T.; Pitkaniemi, J.; Keskitalo-Vuokko, K.; Happola, A.; Heikkila, K.H.; Heikkila, K.; Ripatti, S.; et al. Analysis of detailed phenotype profiles reveals CHRNA5-CHRNA3-CHRNB4 gene cluster association with several nicotine dependence traits. Nicotine Tob. Res. 2012, 14, 720–733. [Google Scholar] [CrossRef]
- Delaneau, O.; Marchini, J.; Zagury, J.F. A linear complexity phasing method for thousands of genomes. Nat. Methods 2012, 9, 179–181. [Google Scholar] [CrossRef]
- Howie, B.N.; Donnelly, P.; Marchini, J. A flexible and accurate genotype imputation method for the next generation of genome-wide association studies. PLoS Genet. 2009, 5, e1000529. [Google Scholar] [CrossRef]
- Chen, W.M.; Abecasis, G.R. Family-based association tests for genomewide association scans. Am. J. Hum. Genet. 2007, 81, 913–926. [Google Scholar] [CrossRef]
- Purcell, S.; Neale, B.; Todd-Brown, K.; Thomas, L.; Ferreira, M.; Bender, D.; Maller, J.; Sklar, P.; de Bakker, P.; Daly, M.; et al. PLINK: A tool set for whole-genome association and population-based linkage analyses. Am. J. Hum. Genet. 2007, 81, 559–575. [Google Scholar] [CrossRef]
- Evans, D.; Visscher, P.M.; Wray, N. Harnessing the information contained within genome-wide association studies to improve individual prediction of complex disease risk. Hum. Mol. Genet. 2009, 18, 3525–3531. [Google Scholar] [CrossRef]
- Vrieze, S.I.; McGue, M.; Miller, M.B.; Hicks, B.M.; Iacono, W.G. Three mutually informative ways to understand the genetic relationships among behavioral disinhibition, alcohol use, drug use, nicotine use/dependence, and their co-occurrence: Twin biometry, GCTA, and genome-wide scoring. Behav. Genet. 2013, 43, 97–107. [Google Scholar] [CrossRef]
- Dishion, T.J.; Owen, L.D. A longitudinal analysis of friendships and substance use: Bidirectional influence from adolescence to adulthood. Dev. Psychol. 2002, 38, 480–491. [Google Scholar] [CrossRef]
- Duncan, S.C.; Duncan, T.E.; Biglan, A.; Ary, D. Contributions of the social context to the development of adolescent substance use: A multivariate latent growth modeling approach. Drug Alcohol Depend. 1998, 50, 57–71. [Google Scholar] [CrossRef]
- Fanous, A.H.; Zhou, B.Y.; Aggen, S.H.; Bergen, S.E.; Amdur, R.L.; Duan, J.B.; Sanders, A.R.; Shi, J.X.; Mowry, B.J.; Olincy, A.; et al. Genome-wide association study of clinical dimensions of schizophrenia: Polygenic effect on disorganized symptoms. Am. J. Psychiatry 2012, 169, 1309–1317. [Google Scholar] [CrossRef]
- Yang, J.; Lee, S.H.; Goddard, M.E.; Visscher, P.M. GCTA: A tool for genome-wide complex trait analysis. Am. J. Hum. Genet. 2011, 88, 76–82. [Google Scholar] [CrossRef]
- Gibson, G. Rare and common variants: Twenty arguments. Nat. Rev. Genet. 2012, 13, 135–145. [Google Scholar] [CrossRef]
- Kendler, K.S.; Baker, J.H. Genetic influences on measures of the environment: A systematic review. Psychol. Med. 2007, 37, 615–626. [Google Scholar] [CrossRef]
- Thomas, D. Gene-environment-wide association studies: Emerging approaches. Nat. Rev. Genet. 2010, 11, 259–272. [Google Scholar] [CrossRef]
- Chen, L.S.; Johnson, E.O.; Breslau, N.; Hatsukami, D.; Saccone, N.L.; Grucza, R.A.; Wang, J.C.; Hinrichs, A.L.; Fox, L.; Goate, A.M.; et al. Interplay of genetic risk factors and parent monitoring in risk for nicotine dependence. Addiction 2009, 104, 1731–1740. [Google Scholar] [CrossRef]
- Brody, G.H.; Beach, S.R.H.; Philibert, R.A.; Chen, Y.F.; Murry, V.M. Prevention effects moderate the association of 5-HTTLPR and youth risk behavior initiation: Gene x environment hypotheses tested via a randomized prevention design. Child. Dev. 2009, 80, 645–661. [Google Scholar] [CrossRef]
- Hamburg, M.A.; Collins, F.S. The path to personalized medicine. N. Engl. J. Med. 2010, 363, 301–304. [Google Scholar] [CrossRef]
- Yan, J.; Aliev, F.; Webb, B.T.; Kendler, K.S.; Edenberg, H.J.; Agrawal, A.; Kos, M.Z.; Almasy, L.; Nurnberger, J.I., Jr.; Schuckit, M.A.; et al. Using genetic information from candidate gene and genome wide association studies in risk prediction for alcohol dependence. Addict. Biol. 2013. [Google Scholar] [CrossRef] [Green Version]
- Wang, L.; Jia, P.; Wolfinger, R.D.; Chen, X.; Zhao, Z. Gene set analysis of genome-wide association studies: Methodological issues and perspectives. Genomics 2011, 98, 1–8. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Salvatore, J.E.; Aliev, F.; Edwards, A.C.; Evans, D.M.; Macleod, J.; Hickman, M.; Lewis, G.; Kendler, K.S.; Loukola, A.; Korhonen, T.; et al. Polygenic Scores Predict Alcohol Problems in an Independent Sample and Show Moderation by the Environment. Genes 2014, 5, 330-346. https://doi.org/10.3390/genes5020330
Salvatore JE, Aliev F, Edwards AC, Evans DM, Macleod J, Hickman M, Lewis G, Kendler KS, Loukola A, Korhonen T, et al. Polygenic Scores Predict Alcohol Problems in an Independent Sample and Show Moderation by the Environment. Genes. 2014; 5(2):330-346. https://doi.org/10.3390/genes5020330
Chicago/Turabian StyleSalvatore, Jessica E., Fazil Aliev, Alexis C. Edwards, David M. Evans, John Macleod, Matthew Hickman, Glyn Lewis, Kenneth S. Kendler, Anu Loukola, Tellervo Korhonen, and et al. 2014. "Polygenic Scores Predict Alcohol Problems in an Independent Sample and Show Moderation by the Environment" Genes 5, no. 2: 330-346. https://doi.org/10.3390/genes5020330





