Papillary Thyroid Carcinoma: A thorough Bioinformatic Analysis of Gene Expression and Clinical Data
Abstract
1. Introduction
2. Data and Methods
2.1. Data
2.2. Methods
- Network type: full STRING network (the edges indicate both functional and physical protein associations).
- Meaning of network edges: evidence (line color indicates the type of interaction evidence).
- Active interaction sources: Text mining, Experiments, Databases, Co-expression, Neighborhood, Gene Fusion, Co-occurrence.
- Minimum required interaction score: 0.4.
- Max number of interactors to show:
- -
- 1st shell: no more than 5.
- -
- 2nd shell: no more than 5.
3. Results
- Age at initial pathologic diagnosis
- Gender
- Race
- Primary thyroid gland neoplasm location
- Tumor stage
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
Abbreviations
| ATC | Anaplastic Thyroid Carcinomas |
| cDNA | complementary DNA |
| DEGs | Differentially Expressed Genes |
| GEO | Gene Expression Omnibus |
| GO | Gene Ontology |
| GTEx | Genotype-Tissue Expression |
| KEGG | Kyoto Encyclopedia of Genes and Genomes |
| KM | Kaplan–Meier |
| OS | Overall Survival |
| PCR | Polymerase Chain Reaction |
| PPI | Protein–Protein Interaction |
| PTC | Papillary Thyroid Carcinoma |
| RFE | Recursive Feature Elimination |
| RFS | Relapse Free Survival |
| RNA-seq | RNA sequencing |
| TCGA | the Cancer Genome Atlas |
| TCGA-THCA | The Cancer Genome Atlas Thyroid Cancer |
References
- Xue, J.M.; Liu, Y.; Wan, L.H.; Zhu, Y.X. Comprehensive analysis of differential gene expression to identify common gene signatures in multiple cancers. Med. Sci. Monit. Int. Med. J. Exp. Clin. Res. 2020, 26, e919953-1. [Google Scholar] [CrossRef] [PubMed]
- Mirza, B.; Wang, W.; Wang, J.; Choi, H.; Chung, N.C.; Ping, P. Machine learning and integrative analysis of biomedical big data. Genes 2019, 10, 87. [Google Scholar] [CrossRef]
- Martin, S.A.; Dehler, C.E.; Król, E. Transcriptomic responses in the fish intestine. Dev. Comp. Immunol. 2016, 64, 103–117. [Google Scholar] [CrossRef] [PubMed]
- Rao, M.S.; Van Vleet, T.R.; Ciurlionis, R.; Buck, W.R.; Mittelstadt, S.W.; Blomme, E.A.G.; Liguori, M.J. Comparison of RNA-Seq and Microarray Gene Expression Platforms for the Toxicogenomic Evaluation of Liver From Short-Term Rat Toxicity Studies. Front. Genet. 2019, 9, 636. [Google Scholar] [CrossRef]
- Li, L.; Ching, W.K.; Liu, Z.P. Robust biomarker screening from gene expression data by stable machine learning-recursive feature elimination methods. Comput. Biol. Chem. 2022, 100, 107747. [Google Scholar] [CrossRef]
- Bommert, A.; Welchowski, T.; Schmid, M.; Rahnenführer, J. Benchmark of filter methods for feature selection in high-dimensional gene expression survival data. Briefings Bioinform. 2022, 23, bbab354. [Google Scholar] [CrossRef] [PubMed]
- Petrini, I.; Cecchini, R.L.; Mascaró, M.; Ponzoni, I.; Carballido, J.A. Statistical Learning Analysis of Thyroid Cancer Microarray Data. In Proceedings of the Bioinformatics and Biomedical Engineering: 9th International Work-Conference, IWBBIO 2022, Maspalomas, Gran Canaria, Spain, 27–30 June 2022; Springer: Berlin/Heidelberg, Germany, 2022; pp. 90–102. [Google Scholar]
- Edgar, R.; Domrachev, M.; Lash, A.E. Gene Expression Omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res. 2002, 30, 207–210. [Google Scholar] [CrossRef]
- Barrett, T.; Wilhite, S.E.; Ledoux, P.; Evangelista, C.; Kim, I.F.; Tomashevsky, M.; Marshall, K.A.; Phillippy, K.H.; Sherman, P.M.; Holko, M.; et al. NCBI GEO: Archive for functional genomics data sets–update. Nucleic Acids Res. 2012, 41, D991–D995. [Google Scholar] [CrossRef]
- Cancer Genome Atlas Research Network. Integrated genomic characterization of papillary thyroid carcinoma. Cell 2014, 159, 676–690. [Google Scholar] [CrossRef]
- Tomas, G. Sporadic vs. Post-Chernobyl Papillary vs. Anaplastic Thyroid Cancers, geo, V1. 2012. Available online: https://www.omicsdi.org/dataset/geo/GSE29265 (accessed on 30 May 2023).
- Tomas, G.; Tarabichi, M.; Gacquer, D.; Dom, G.; Dumont, J.; Keutgen, X.; Fahey, T.; Maenhaut, C.; Detours, V. A general method to derive robust organ-specific gene expression-based differentiation indices: Application to thyroid cancer diagnostic. Oncogene 2012, 31, 4490–4498. [Google Scholar] [CrossRef] [PubMed]
- Dom, G.; Tarabichi, M.; Unger, K.; Thomas, G.; Oczko-Wojciechowska, M.; Bogdanova, T.; Jarząb, B.; Dumont, J.; Detours, V.; Maenhaut, C. A gene expression signature distinguishes normal tissues of sporadic and radiation-induced papillary thyroid carcinomas. Br. J. Cancer 2012, 107, 994–1000. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Robinson, M.D.; McCarthy, D.J.; Smyth, G.K. edgeR: A Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 2009, 26, 139–140. [Google Scholar] [CrossRef]
- Robinson, M.; Smyth, G. Small-sample estimation of negative binomial dispersion, with applications to SAGE data. Biostatistics 2008, 9, 321–332. [Google Scholar] [CrossRef]
- Benjamini, Y.; Hochberg, Y. Controlling the False Discovery Rate: A Practical and Powerful Approach to Multiple Testing. J. R. Stat. Soc. Ser. B (Methodol.) 1995, 57, 289–300. [Google Scholar] [CrossRef]
- Liu, C.; Wang, W.; Zhao, Q.; Shen, X.; Konan, M. A new feature selection method based on a validity index of feature subset. Pattern Recognit. Lett. 2017, 92, 1–8. [Google Scholar] [CrossRef]
- Szklarczyk, D.; Gable, A.L.; Lyon, D.; Junge, A.; Wyder, S.; Huerta-Cepas, J.; Simonovic, M.; Doncheva, N.T.; Morris, J.H.; Bork, P.; et al. STRING v11: Protein–protein association networks with increased coverage, supporting functional discovery in genome-wide experimental datasets. Nucleic Acids Res. 2019, 47, D607–D613. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Gable, A.L.; Nastou, K.C.; Lyon, D.; Kirsch, R.; Pyysalo, S.; Doncheva, N.T.; Legeay, M.; Fang, T.; Bork, P.; et al. The STRING database in 2021: Customizable protein–protein networks, and functional characterization of user-uploaded gene/measurement sets. Nucleic Acids Res. 2020, 49, D605–D612. [Google Scholar] [CrossRef] [PubMed]
- Lánczky, A.; Györffy, B. Web-Based Survival Analysis Tool Tailored for Medical Research (KMplot): Development and Implementation. J. Med. Internet Res. 2021, 23, e27633. [Google Scholar] [CrossRef]
- Nagy, Á.; Munkácsy, G.; Györffy, B. Pancancer survival analysis of cancer hallmark genes. Sci. Rep. 2021, 11, 6047. [Google Scholar] [CrossRef]
- Barros-Filho, M.C.; Marchi, F.A.; Pinto, C.A.; Rogatto, S.R.; Kowalski, L.P. High Diagnostic Accuracy Based on CLDN10, HMGA2, and LAMB3 Transcripts in Papillary Thyroid Carcinoma. J. Clin. Endocrinol. Metab. 2015, 100, E890–E899. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.X.; Chen, E.D.; Cai, Y.F.; Li, Q.; Jin, Y.X.; Jin, W.X.; Wang, Y.H.; Zheng, Z.C.; Xue, L.; Ouchen, W.; et al. A panel of four genes accurately differentiates benign from malignant thyroid nodules. J. Exp. Clin. Cancer Res. 2016, 35, 169. [Google Scholar] [CrossRef]
- Jin, Y.; Jin, W.; Zheng, Z.; Chen, E.; Wang, Q.; Wang, Y.; Wang, O.; Zhang, X. GABRB2 plays an important role in the lymph node metastasis of papillary thyroid cancer. Biochem. Biophys. Res. Commun. 2017, 492, 323–330. [Google Scholar] [CrossRef] [PubMed]
- Beltrami, C.; Reis, M.; Filho, M.; Marchi, F.; Kuasne, H.; Pinto, C.; Ambatipudi, S.; Herceg, Z.; Kowalski, L.; Rogatto, S. Integrated data analysis reveals potential drivers and pathways disrupted by DNA methylation in papillary thyroid carcinomas. Clin. Epigenetics 2017, 9, 45. [Google Scholar] [CrossRef] [PubMed]
- Yates, C.M.; Patel, A.; Oakley, K.; Helms, A.; Tuttle, R.M.; Francis, G.L. Erythropoietin in thyroid cancer. J. Endocrinol. Investig. 2006, 29, 320–329. [Google Scholar] [CrossRef] [PubMed]


| Sample Distribution | Source | Details | |
|---|---|---|---|
| GSE33630 | 11 anaplastic thyroid carcinomas (ATC), 49 papillary thyroid carcinomas (PTC) and 45 normal thyroids (N) | GEO | https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=gse33630 (accessed on 30 May 2023) |
| GSE29265 | 9 anaplastic thyroid carcinomas (ATC), 20 papillary thyroid carcinomas (PTC) and 20 normal thyroids (N) | GEO | https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=gse29265 (accessed on 30 May 2023) |
| TCGA-THCA | 1 Carcinoma, 1 Follicular adenocarcinoma, 1 Follicular carcinoma, minimally invasive, 4 Non encapsulated sclerosing carcinomas, 1 Oxyphilic adenocarcinoma, 444 Papillary adenocarcinomas, 43 Papillary carcinomas, columnar cell, 120 Papillary carcinomas, follicular variant | TCGA | https://www.cancer.gov/types/thyroid (accessed on 30 May 2023) https://xenabrowser.net/datapages/?cohort=GDC%20TCGA%20Thyroid%20Cancer%20(THCA)&removeHub=https%3A%2F%2Fxena.treehouse.gi.ucsc.edu%3A443 (accessed on 30 May 2023) |
| Assesed Genes | DEGs | Upregulated in PTC | Downregulated in PTC | |||
|---|---|---|---|---|---|---|
| GSE33630 | 23,521 | 11,196 | 6339 | 57% | 4857 | 43% |
| GSE29265 | 23,521 | 2497 | 1309 | 53% | 1188 | 47% |
| TCGA-THCA | 60,488 | 7941 | 2251 | 28% | 5690 | 72% |
| Overlap | 19,516 genes in common between the 3 assays | |||||
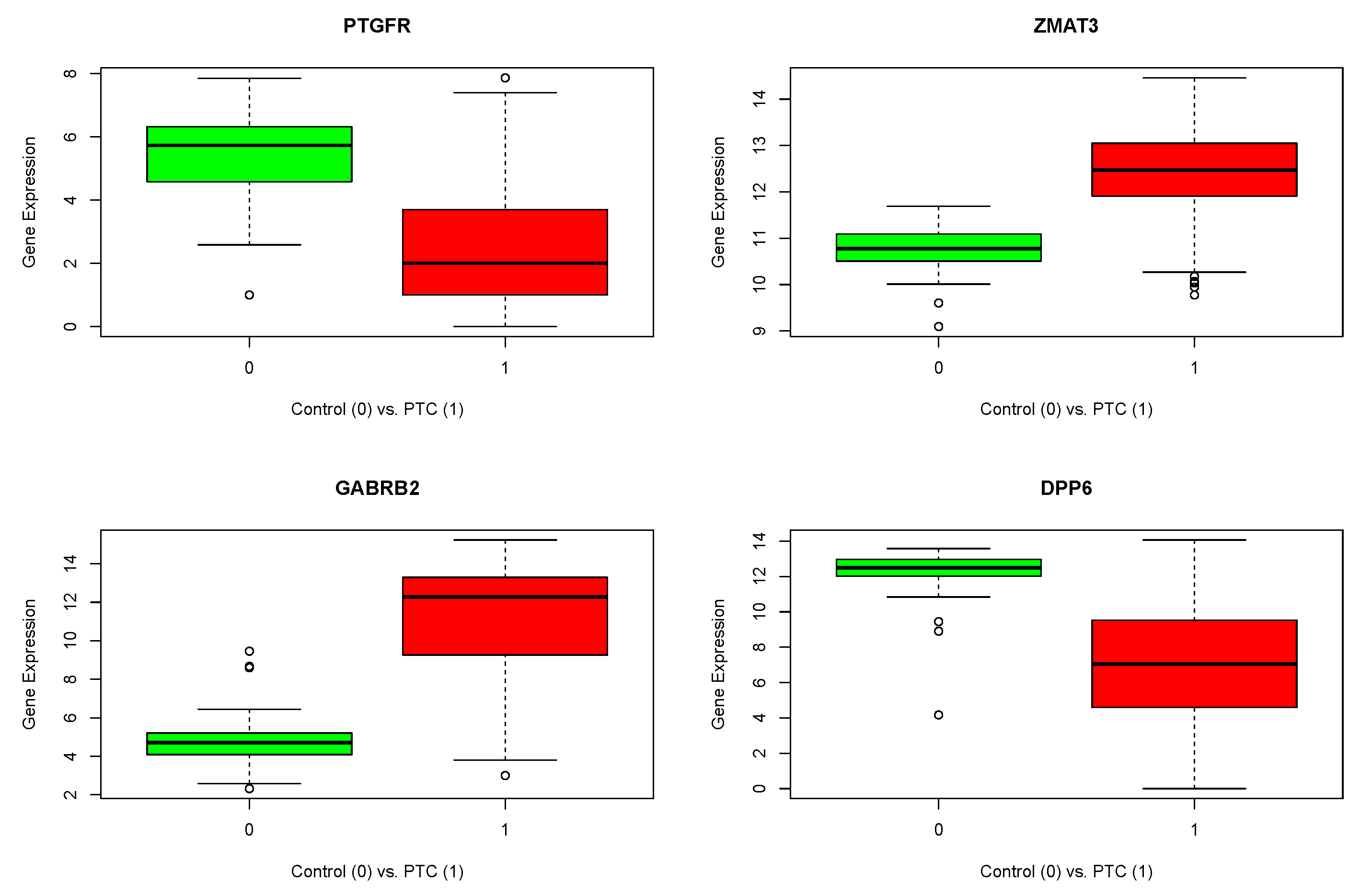
| Expression in PTC | DEGs (Gene Symbol) |
|---|---|
| Up | WNT4, EDARADD, MYT1L, ZNF385B, B3GNT7, ARPP21, RP11-757F18.5, WDR49, ZMAT3, LPP-AS2, FAM43A, PCDH10, C4orf45, GALNTL6, ARHGEF28, SPINK6, C5orf47, LY86-AS1, PDE1C, KCP, ASB10, ERICH1-AS1, TDH, PSKH2, C8orf88, C9orf66, TRPM3, ATOH7, SYCE1, SYT9, GLYATL2, TYR, FLJ12825, NAV3, FAM124A, ABCC4, RPS6KA5, LINC00521, ASPG, CYP19A1, PRR35, ASPHD1, CNTNAP4, LINC00304, TRPV3, CD300LG, RP11-700H6.1, TK1, NETO1, ZNF99, GABRB2, SIGLEC11, KLK4, ZSCAN4, CBS |
| Down | NEXN, PTGFR, RP11-498C9.17, AMER3, FN1, ADAMTS9, SLC7A14, GPR111, TAAR1, DPP6, RSPO2, SLC30A8, IFNE, TMEM252, AKR1E2, C10orf107, C10orf55, TECTB, PLEKHS1, SLC18A2, THRSP, PIANP, KRT73, C12orf74, FAM71C, RFX4, MYO16, RPPH1, GSC, NOX5, GAS2L2, B4GALNT2, STARD6, ZNF560, KLK8, NLRP11, PAX1, WFDC11, LKAAEAR1, FMR1NB |
| ENSID | Gen Symbol | logFC | logCPM | p Value | FDR | |
|---|---|---|---|---|---|---|
| 1 | ENSG00000122420.8 | PTGFR | −0.89475 | 13.48313 | 2.37 | 2.25 |
| 2 | ENSG00000130226.15 | DPP6 | −0.56845 | 14.50570 | 9.07 | 7.18 |
| 3 | ENSG00000145864.11 | GABRB2 | 1.42467 | 14.88948 | 1.98 | 1.88 |
| 4 | ENSG00000172667.9 | ZMAT3 | 0.40854 | 15.06364 | 7.30 | 2.77 |
| Tumor type | Thyroid |
| TCGA code | THCA |
| Samples with RNA-seq data | 502 |
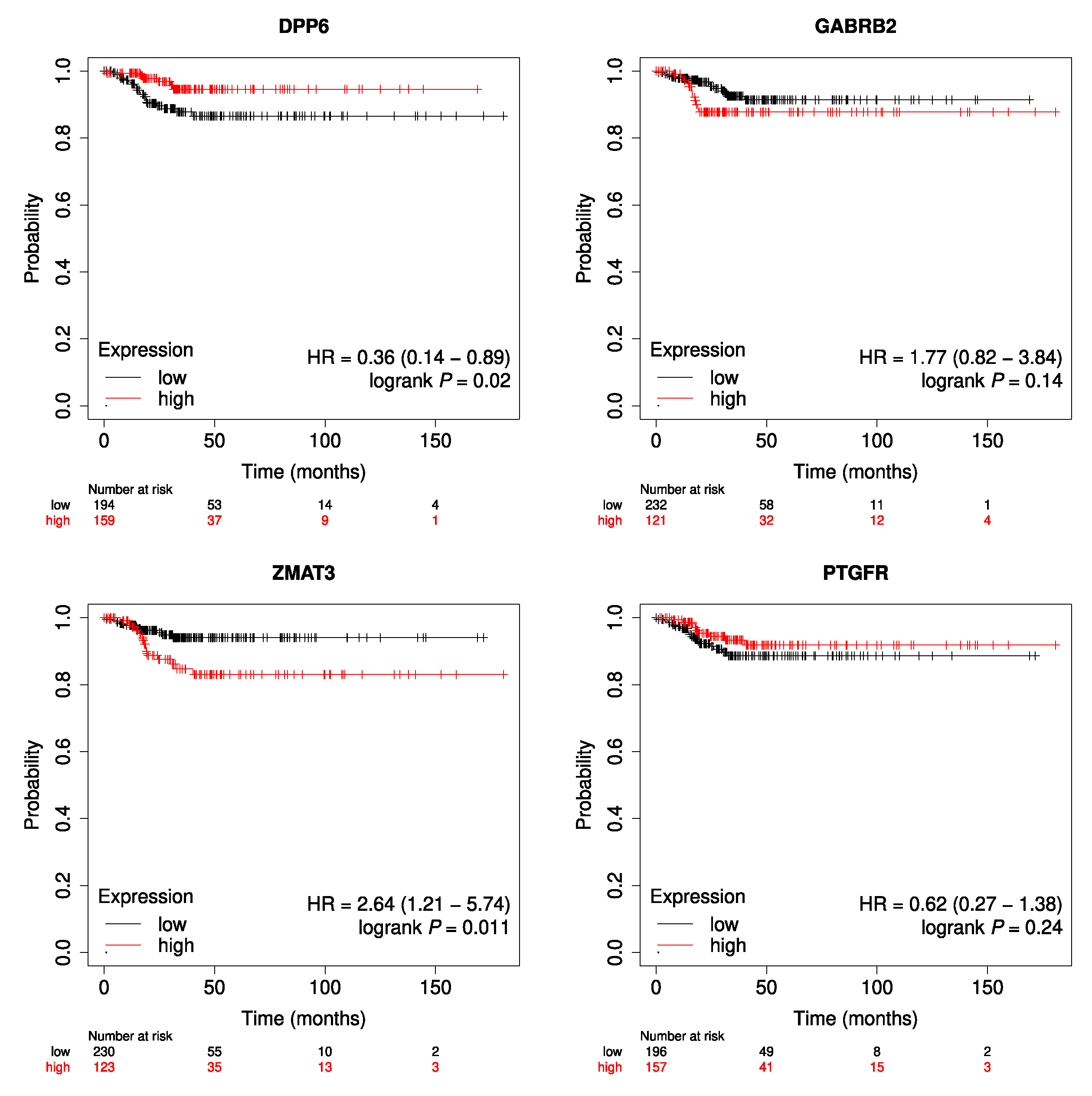
| Median survival–OS (months) | 31.47 |
| Events (n) | 16 |
| Median survival time in patients with an OS event | 34.03 |
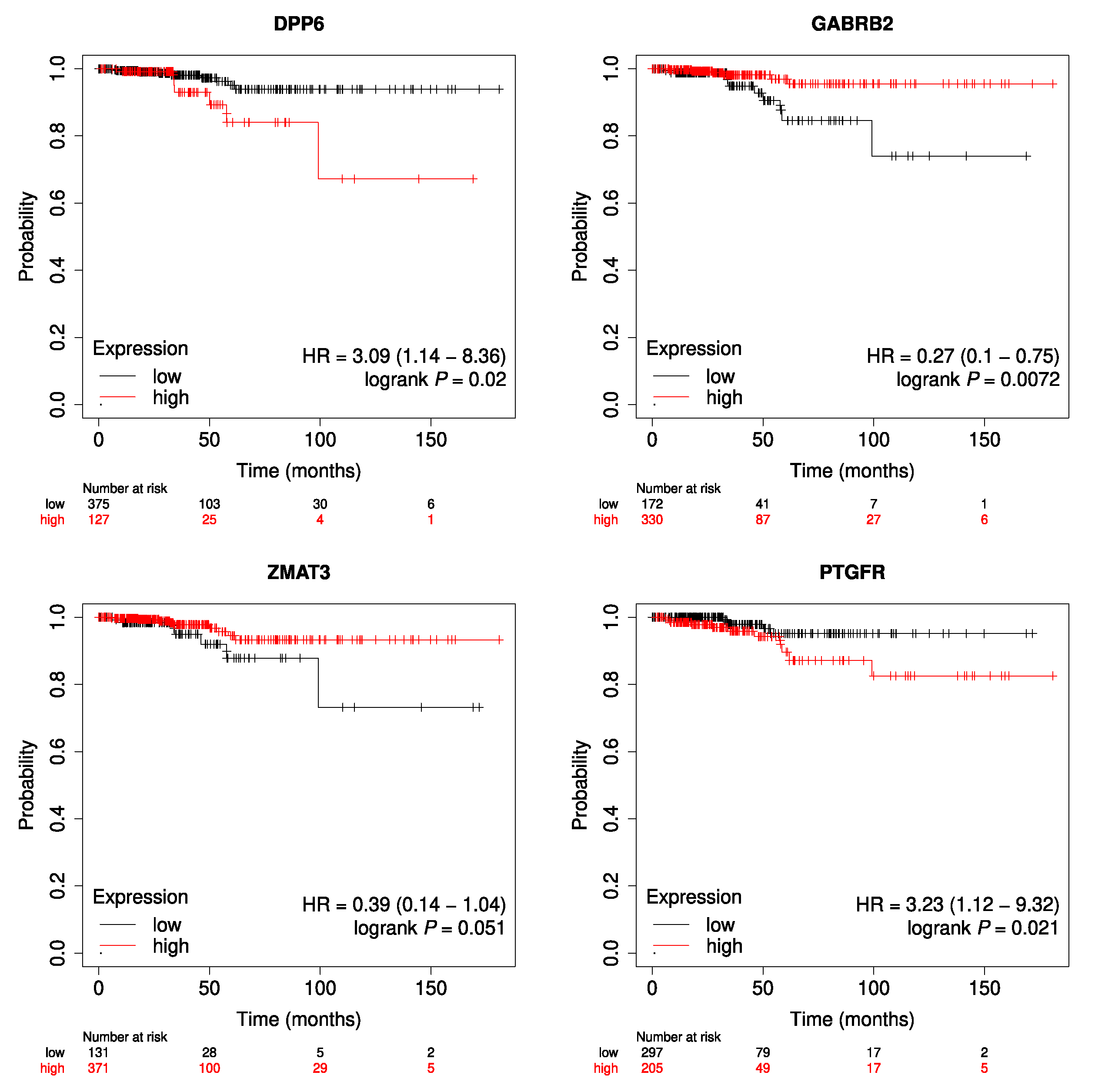
| Median survival–RFS (months) | 18.72 |
| Median survival in patients with a relapse (months) | 16.43 |
| Sex (F/M) | 367/135 |
| Stage (S0/S1/S2/S3/S4) | 0/281/52/112/55 |
| Grade (low/high) | – |
| Race (White/Asian/Black-African) | 332/51/27 |
| ZMAT3 | GABRB2 | DPP6 | PTGFR | |
|---|---|---|---|---|
| AGE at initial diagnosis | 0.001439/−0.100624 | 0.000313/−0.113823 | 0.378/0.027890 | 0.7956/−0.008471 |
| Gender | 0.688/0.162 | 0.848/0.037 | 0.635/0.225 | 0.159/1.986 |
| Race | 0.592/0.7 | 0.00033/5.338 | 0.0845/2.064 | 0.00632/3.631 |
| Primary thyroid gland neoplasm location | 0.191/1.535 | 0.0619/2.26 | 0.00776/3.511 | 0.148/1.705 |
| Tumor stage | 0.221/1.436 | 0.000372/5.269 | 0.00103/4.687 | 0.00168/4.401 |
| ZMAT3 | Df | Sum Sq | Mean Sq | F value | Pr(>F) | |
| Tumor stage | 4 | 4.14 | 1034 | 1463 | 0.21251 | |
| Gender | 1 | 0.12 | 0.123 | 0.174 | 0.67681 | |
| Primary Location | 4 | 4.25 | 1063 | 1503 | 0.20026 | |
| Race | 4 | 1.88 | 0.470 | 0.665 | 0.61672 | |
| AGE at initial d. | 1 | 6.88 | 6878 | 9729 | 0.00193 | ** |
| Residuals | 442 | 312.50 | 0.707 | |||
| GABRB2 | Df | Sum Sq | Mean Sq | F value | Pr(>F) | |
| Tumor stage | 4 | 138.1 | 34.54 | 5660 | 0.000189 | *** |
| Gender | 1 | 2.0 | 2.02 | 0.331 | 0.565269 | |
| Primary Location | 4 | 57.9 | 14.46 | 2370 | 0.051792 | . |
| Race | 4 | 125.5 | 31.38 | 5142 | 0.000466 | *** |
| AGE at initial d. | 1 | 80.4 | 80.41 | 13,178 | 0.000316 | *** |
| Residuals | 442 | 2697.0 | 6.10 | |||
| DPP6 | Df | Sum Sq | Mean Sq | F value | Pr(>F) | |
| Tumor stage | 4 | 172 | 43.01 | 4965 | 0.000636 | *** |
| Gender | 1 | 0 | 0.19 | 0.022 | 0.882296 | |
| Primary Location | 4 | 163 | 40.71 | 4700 | 0.001007 | ** |
| Race | 4 | 89 | 22.36 | 2582 | 0.036718 | * |
| AGE at initial d. | 1 | 66 | 65.83 | 7599 | 0.006083 | ** |
| Residuals | 442 | 3829 | 8.66 | |||
| PTGFR | Df | Sum Sq | Mean Sq | F value | Pr(>F) | |
| Tumor stage | 4 | 61.8 | 15,454 | 4509 | 0.0014 * | * |
| Gender | 1 | 8.6 | 8613 | 2513 | 0.1136 | |
| Primary Location | 4 | 22.9 | 5736 | 1673 | 0.1551 | ** |
| Race | 4 | 39.1 | 9774 | 2852 | 0.0235 * | * |
| AGE at initial d. | 1 | 1.6 | 1604 | 0.468 | 0.4942 | ** |
| Residuals | 442 | 1515.0 | 3428 | |||
| Signif. codes: | 0 ‘***’ | 0.001 ‘**’ | 0.01 ‘*’ | 0.05 ‘.’ | 0.1 ‘ ’ | 1 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Petrini, I.; Cecchini, R.L.; Mascaró, M.; Ponzoni, I.; Carballido, J.A. Papillary Thyroid Carcinoma: A thorough Bioinformatic Analysis of Gene Expression and Clinical Data. Genes 2023, 14, 1250. https://doi.org/10.3390/genes14061250
Petrini I, Cecchini RL, Mascaró M, Ponzoni I, Carballido JA. Papillary Thyroid Carcinoma: A thorough Bioinformatic Analysis of Gene Expression and Clinical Data. Genes. 2023; 14(6):1250. https://doi.org/10.3390/genes14061250
Chicago/Turabian StylePetrini, Iván, Rocío L. Cecchini, Marilina Mascaró, Ignacio Ponzoni, and Jessica A. Carballido. 2023. "Papillary Thyroid Carcinoma: A thorough Bioinformatic Analysis of Gene Expression and Clinical Data" Genes 14, no. 6: 1250. https://doi.org/10.3390/genes14061250
APA StylePetrini, I., Cecchini, R. L., Mascaró, M., Ponzoni, I., & Carballido, J. A. (2023). Papillary Thyroid Carcinoma: A thorough Bioinformatic Analysis of Gene Expression and Clinical Data. Genes, 14(6), 1250. https://doi.org/10.3390/genes14061250









