2.1. Composition of Resident Microbial Communities in the Strawberry Phyllosphere
The resident phyllosphere microbiome as determined by culture-dependent and -independent techniques varied considerably between the two years of experiment, but also between the three phenological stages.
According to plate counts, the resident culturable leaf microbiota consisted of 7.76 × 10
4 to 2.09 × 10
5 CFU g DW
−1 bacteria and 1.15 × 10
3 to 1.29 × 10
5 CFU g DW
−1 fungi in total, as well as 3.24 × 10
3 to 3.24 × 10
4 CFU g DW
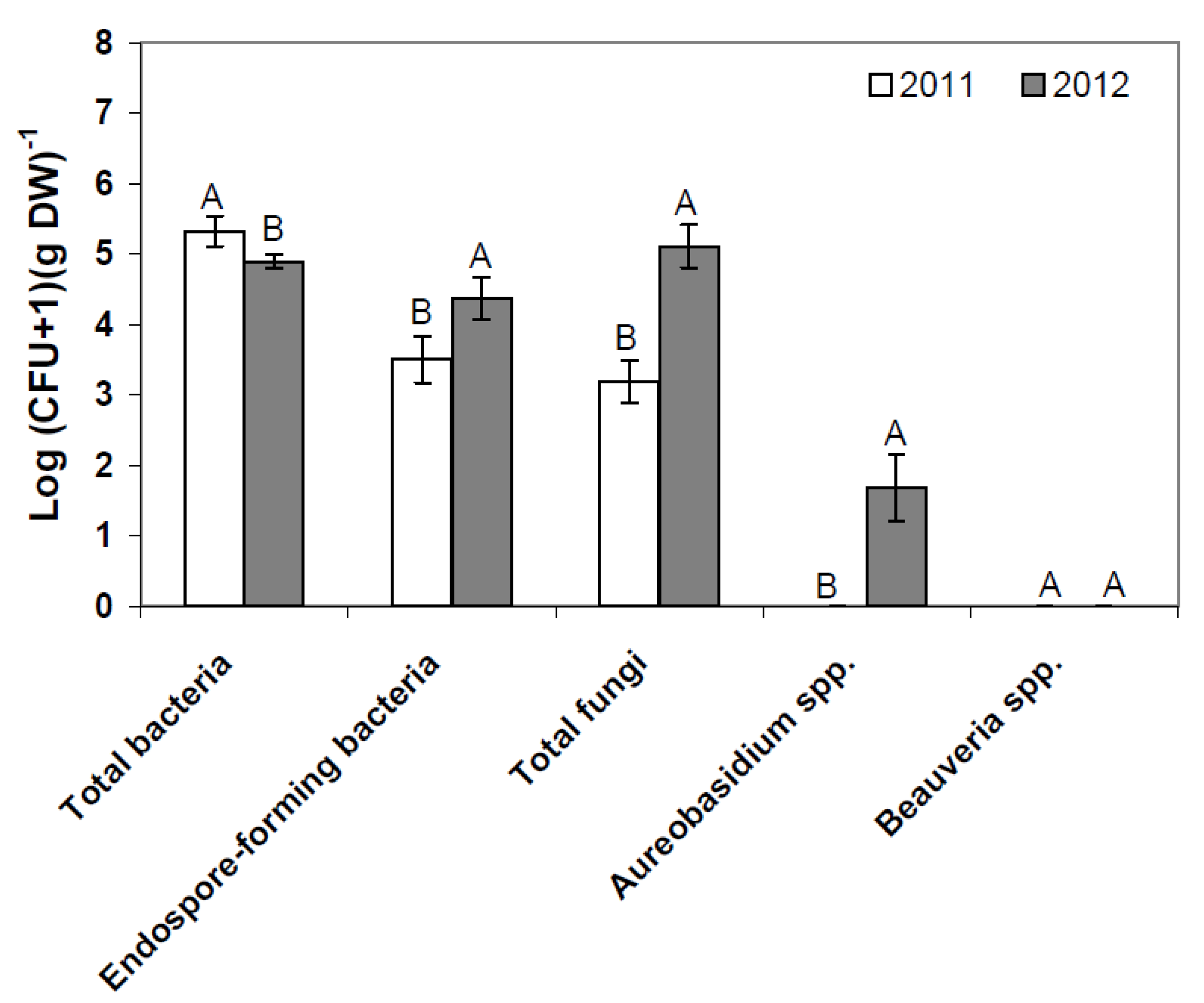
−1 endospore-forming bacteria in flowering strawberry plants (BBCH 60) in 2011 and 2012 (
Figure 1). At this phenological growth stage, the numbers of resident total bacteria, total fungi and endospore-forming bacteria significantly differed between 2011 and 2012, respectively. Neither
Aureobasidium spp., nor
Beauveria spp., were detected at this stage in 2011, but
Aureobasidium spp. were found at the same stage in 2012, albeit at a low level (
Figure 1).
Table 1.
Mean numbers of 454 pyrosequencing reads in differentially treated leaf samples obtained at different phenological stages in 2011 and 2012.
Table 1.
Mean numbers of 454 pyrosequencing reads in differentially treated leaf samples obtained at different phenological stages in 2011 and 2012.
| | Fungal ITS rRNA sequences | | Bacterial 16S rRNA sequences |
|---|
| | 2011 | | 2012 | | 2011 | | 2012 |
| Phenological stage a | Mean b | SEM c | | Mean | SEM | | Mean | SEM | | Mean | SEM |
| Untreated control (water) |
| BBCH 60 | 1217 | 443 | | 936 | 411 | | 357 d | 31 | | 276 | 80 |
| BBCH 73 | 1201 | 234 | | 996 | 98 | | 234 e | 66 | | 761 | 229 |
| BBCH 93 | 324 d | 165 | | 1609 | 397 | | 357 d | 113 | | 1703 | 1193 |
| Single strain treatment (
A. pullulans) |
| BBCH 60 | 1272 | 255 | | 1412 | 301 | | 636 e | 215 | | 501 d | 339 |
| BBCH 73 | 816 d | 512 | | 1275 | 52 | | 316 f | NA | | 464 | 108 |
| BBCH 93 | 380 | 97 | | 1496 | 247 | | 229 d | 9 | | 1496 | 235 |
| Multiple strain treatment (
A. pullulans/B. bassiana) |
| BBCH 60 | 1631 | 407 | | 1344 | 277 | | 103 d | 38 | | 952 | 306 |
| BBCH 73 | 482 d | 308 | | 1763 d | 150 | | NA g | NA | | 403 e | 155 |
| BBCH 93 | 340 | 172 | | 1059 | 90 | | 188 | 85 | | 1059 | 365 |
Figure 1.
Resident phyllosphere microbiome of flowering strawberries (BBCH 60) in 2011 and 2012 as determined by plate counts (pooled data, each N = 36). Error bars represent SEM values. Columns marked with the same letters are not significantly different within each microbial group (α = 0.05).
Figure 1.
Resident phyllosphere microbiome of flowering strawberries (BBCH 60) in 2011 and 2012 as determined by plate counts (pooled data, each N = 36). Error bars represent SEM values. Columns marked with the same letters are not significantly different within each microbial group (α = 0.05).
In untreated leaf samples, the amount of total resident bacteria significantly decreased (
p = 0.008) when comparing flowering and fruit-setting strawberry plants and subsequently increased (
p = 0.006) when old leaves died off (BBCH 93) in 2011 (see also
Figure 2). In 2012, counts of total bacteria were similar in the control samples during flowering and fruit-setting and significantly increased at BBCH 93 (
p = 0.002). An increase in the abundance of culturable bacteria in decaying leaves is expected [
18]. The counts of resident endospore-forming bacteria did not differ at the first two stages in 2011, but significantly decreased from the second to the third phenological stage (
p < 0.001). In 2012, counts of endospore-forming bacteria gradually decreased from flowering plants to plants with old, dying leaves (
p < 0.001). The amount of total resident fungi was similar at the first two phenological stages in 2011, but increased as expected on decaying leaves in 2011 (
p < 0.001), which is in line with the function of fungi during decomposition [
26]. In 2012, counts of total fungi were initially high during flowering and significantly decreased from flowering to the fruit-setting stage (
p < 0.001). The amount of total fungi, however, significantly increased on decaying leaves as well (
p < 0.001). Interestingly,
Aureobasidium spp. counts increased on leaves from untreated plots from the first through the third stage in 2011 (
p > 0.05).
A. pullulans is ubitiquous in the environment [
27,
28]. As no labeled strain was used in the treated plots, it is difficult to tell, if occurrence of
A. pullulans on untreated strawberry leaves is due to natural inoculation or transmission by wind from the treated plots. However,
Aureobasidium were detected as resident organisms on leaves on flowering plants in 2012, but not at later stages.
Figure 2.
Culturable phyllosphere microbiota in strawberries at two distinct phenological stages (BBCH 73 and BBCH 93) during two subsequent years (2011, 2012) as determined by plate counts (N = 12). Aureobasidium pullulans was introduced as a model organism either as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) through commercial formulations. The two phenological stages represent short- (BBCH 73, after four BCA applications) and long-term effects (BBCH 93, four weeks after adjournment of BCA applications). The control (W) was treated with water from a lotic reservoir. Error bars represent SEM values. Columns marked with the same letters are not significantly different within each microbial group (α = 0.05).
Figure 2.
Culturable phyllosphere microbiota in strawberries at two distinct phenological stages (BBCH 73 and BBCH 93) during two subsequent years (2011, 2012) as determined by plate counts (N = 12). Aureobasidium pullulans was introduced as a model organism either as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) through commercial formulations. The two phenological stages represent short- (BBCH 73, after four BCA applications) and long-term effects (BBCH 93, four weeks after adjournment of BCA applications). The control (W) was treated with water from a lotic reservoir. Error bars represent SEM values. Columns marked with the same letters are not significantly different within each microbial group (α = 0.05).
According to 454 pyrosequencing, the composition of the resident leaf microbiome can be described as follows. The resident fungal communities of leaf samples were mainly composed of the orders Filobasidiales, Cystofilobasidiales and Capnodiales in flowering strawberry plants in both years (BBCH 60;
Table 2 and
Table 3), whereas the spectrum of highly abundant fungal orders at this growing stage was extended by Helotiales and Pleosporales in 2012.
Cryptococcus (32.9% to 40.0%) represented the most abundant fungal genus (
Figure 3), followed by
Cystofilobasidium, in 2011. The genus
Cryptococcus was also most abundant across all untreated leaf samples at this growth stage in 2012. Compared to 2011, the genus
Cystofilobasidium was less abundant at BBCH 60 in 2012.
Table 2.
Relative abundance of ITS rRNA reads assigned to fungal orders in the strawberry phyllosphere at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) in 2011 as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) through commercial formulations. The control (W) was treated with water from a lotic reservoir.
Table 2.
Relative abundance of ITS rRNA reads assigned to fungal orders in the strawberry phyllosphere at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) in 2011 as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) through commercial formulations. The control (W) was treated with water from a lotic reservoir.
| | BBCH 60 (prior to BCA applications) | | BBCH 73 (after four BCA applications) | | BBCH 93 (four weeks after last BCA application) |
|---|
| | W a | | AP a | | AP/BB a | | W | | AP | | AP/BB | | W | | AP | | AP/BB |
|---|
| Fungal order | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM |
| Filobasidiales | 0.336 | 0.021 | | 0.329 | 0.025 | | 0.400 | 0.026 | | 0.088 | 0.013 | | 0.088 | 0.023 | | 0.050 | 0.009 | | 0.389 | 0.026 | | 0.270 | 0.036 | | 0.311 | 0.021 |
| Dothideales | 0.026 | 0.007 | | 0.020 | 0.003 | | 0.018 | 0.004 | | 0.093 | 0.008 | | 0.297 | 0.080 | | 0.519 | 0.024 | | 0.049 | 0.021 | | 0.234 | 0.029 | | 0.139 | 0.028 |
| Capnodiales | 0.106 | 0.011 | | 0.092 | 0.012 | | 0.069 | 0.008 | | 0.080 | 0.012 | | 0.035 | 0.007 | | 0.024 | 0.005 | | 0.224 | 0.026 | | 0.198 | 0.028 | | 0.138 | 0.044 |
| Cystofilobasidiales | 0.162 | 0.020 | | 0.176 | 0.027 | | 0.166 | 0.032 | | 0.016 | 0.002 | | 0.009 | 0.003 | | 0.007 | 0.004 | | 0.044 | 0.046 | | 0.011 | 0.003 | | 0.006 | 0.002 |
| Pleosporales | 0.033 | 0.006 | | 0.035 | 0.008 | | 0.027 | 0.001 | | 0.096 | 0.010 | | 0.103 | 0.035 | | 0.040 | 0.004 | | 0.047 | 0.020 | | 0.076 | 0.022 | | 0.114 | 0.034 |
| Sporidiobolales | 0.015 | 0.002 | | 0.011 | 0.001 | | 0.021 | 0.004 | | 0.017 | 0.003 | | 0.005 | 0.003 | | 0.011 | 0.000 | | 0.031 | 0.008 | | 0.033 | 0.013 | | 0.032 | 0.005 |
| Chaetothyriales | 0.003 | 0.001 | | 0.003 | 0.001 | | 0.003 | 0.001 | | 0.068 | 0.012 | | 0.036 | 0.003 | | 0.037 | 0.010 | | 0.003 | 0.003 | | 0.004 | 0.001 | | 0.006 | 0.002 |
| Helotiales | 0.011 | 0.004 | | 0.008 | 0.002 | | 0.006 | 0.001 | | 0.010 | 0.001 | | 0.004 | 0.002 | | 0.020 | 0.006 | | 0.053 | 0.021 | | 0.017 | 0.008 | | 0.041 | 0.007 |
| Hypocreales | 0.014 | 0.001 | | 0.032 | 0.013 | | 0.007 | 0.001 | | 0.011 | 0.004 | | 0.010 | 0.002 | | 0.008 | 0.003 | | 0.020 | 0.012 | | 0.013 | 0.006 | | 0.016 | 0.008 |
| Tremellales | 0.027 | 0.005 | | 0.019 | 0.002 | | 0.035 | 0.006 | | 0.003 | 0.001 | | 0.008 | 0.003 | | 0.001 | 0.001 | | 0.001 | 0.002 | | 0.001 | 0.001 | | 0.002 | 0.001 |
| Lecanorales | 0.002 | 0.001 | | 0.004 | 0.001 | | 0.003 | 0.001 | | 0.034 | 0.004 | | 0.013 | 0.005 | | 0.023 | 0.006 | | 0.000 | 0.000 | | 0.003 | 0.002 | | 0.002 | 0.002 |
| Agaricales | 0.003 | 0.001 | | 0.002 | 0.000 | | 0.002 | 0.000 | | 0.001 | 0.000 | | 0.001 | 0.001 | | 0.002 | 0.001 | | 0.043 | 0.047 | | 0.016 | 0.002 | | 0.041 | 0.009 |
| Taphrinales | 0.009 | 0.003 | | 0.013 | 0.004 | | 0.013 | 0.005 | | 0.007 | 0.002 | | 0.002 | 0.001 | | 0.002 | 0.001 | | 0.001 | 0.001 | | 0.012 | 0.003 | | 0.013 | 0.004 |
| Teloschistales | 0.004 | 0.001 | | 0.002 | 0.001 | | 0.001 | 0.000 | | 0.027 | 0.004 | | 0.016 | 0.001 | | 0.006 | 0.002 | | 0.000 | 0.001 | | 0.000 | 0.000 | | 0.003 | 0.001 |
| Erysiphales | 0.002 | 0.001 | | 0.003 | 0.001 | | 0.004 | 0.001 | | 0.017 | 0.001 | | 0.016 | 0.005 | | 0.002 | 0.001 | | 0.000 | 0.001 | | 0.001 | 0.001 | | 0.004 | 0.002 |
| Xylariales | 0.007 | 0.002 | | 0.002 | 0.000 | | 0.003 | 0.001 | | 0.008 | 0.002 | | 0.011 | 0.007 | | 0.009 | 0.002 | | 0.000 | 0.000 | | 0.002 | 0.001 | | 0.000 | 0.000 |
| Pucciniales | 0.000 | 0.000 | | 0.001 | 0.000 | | 0.000 | 0.000 | | 0.017 | 0.003 | | 0.010 | 0.005 | | 0.009 | 0.004 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Others b | 0.242 | 0.020 | | 0.247 | 0.028 | | 0.222 | 0.016 | | 0.408 | 0.023 | | 0.335 | 0.055 | | 0.228 | 0.031 | | 0.094 | 0.047 | | 0.108 | 0.020 | | 0.131 | 0.040 |
Table 3.
Relative abundance of ITS rRNA reads assigned to fungal orders in the strawberry phyllosphere at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) in 2012 as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) through commercial formulations. The control (W) was treated with water from a lotic reservoir.
Table 3.
Relative abundance of ITS rRNA reads assigned to fungal orders in the strawberry phyllosphere at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) in 2012 as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) through commercial formulations. The control (W) was treated with water from a lotic reservoir.
| | BBCH 60 (prior to BCA applications) | | BBCH 73 (after four BCA applications) | | BBCH 93 (four weeks after last BCA application) |
|---|
| | W a | | AP a | | AP/BB a | | W | | AP | | AP/BB | | W | | AP | | AP/BB |
|---|
| Order | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM |
| Filobasidiales | 0.214 | 0.009 | | 0.190 | 0.021 | | 0.348 | 0.018 | | 0.308 | 0.029 | | 0.258 | 0.021 | | 0.257 | 0.028 | | 0.234 | 0.010 | | 0.204 | 0.022 | | 0.209 | 0.018 |
| Capnodiales | 0.271 | 0.024 | | 0.333 | 0.029 | | 0.141 | 0.008 | | 0.178 | 0.036 | | 0.179 | 0.014 | | 0.112 | 0.010 | | 0.134 | 0.021 | | 0.107 | 0.015 | | 0.111 | 0.005 |
| Helotiales | 0.100 | 0.009 | | 0.119 | 0.022 | | 0.063 | 0.012 | | 0.059 | 0.008 | | 0.102 | 0.027 | | 0.047 | 0.014 | | 0.267 | 0.008 | | 0.360 | 0.052 | | 0.234 | 0.035 |
| Pleosporales | 0.087 | 0.013 | | 0.066 | 0.010 | | 0.066 | 0.010 | | 0.063 | 0.004 | | 0.083 | 0.006 | | 0.042 | 0.010 | | 0.066 | 0.014 | | 0.053 | 0.006 | | 0.064 | 0.007 |
| Cystofilobasidiales | 0.106 | 0.012 | | 0.107 | 0.017 | | 0.083 | 0.006 | | 0.074 | 0.018 | | 0.103 | 0.023 | | 0.056 | 0.016 | | 0.020 | 0.005 | | 0.011 | 0.003 | | 0.012 | 0.002 |
| Dothideales | 0.016 | 0.005 | | 0.017 | 0.003 | | 0.023 | 0.005 | | 0.025 | 0.001 | | 0.015 | 0.005 | | 0.338 | 0.020 | | 0.005 | 0.002 | | 0.043 | 0.003 | | 0.077 | 0.013 |
| Sporidiobolales | 0.038 | 0.005 | | 0.027 | 0.001 | | 0.016 | 0.003 | | 0.018 | 0.003 | | 0.034 | 0.008 | | 0.012 | 0.001 | | 0.035 | 0.005 | | 0.024 | 0.004 | | 0.029 | 0.001 |
| Agaricostilbales | 0.001 | 0.001 | | 0.003 | 0.002 | | 0.003 | 0.001 | | 0.003 | 0.000 | | 0.004 | 0.001 | | 0.001 | 0.000 | | 0.031 | 0.006 | | 0.032 | 0.005 | | 0.039 | 0.003 |
| Diaporthales | 0.014 | 0.007 | | 0.007 | 0.003 | | 0.021 | 0.007 | | 0.025 | 0.006 | | 0.023 | 0.009 | | 0.010 | 0.005 | | 0.006 | 0.003 | | 0.003 | 0.001 | | 0.003 | 0.000 |
| Xylariales | 0.006 | 0.003 | | 0.011 | 0.004 | | 0.014 | 0.003 | | 0.026 | 0.009 | | 0.015 | 0.004 | | 0.011 | 0.006 | | 0.002 | 0.001 | | 0.001 | 0.001 | | 0.001 | 0.000 |
| Hypocreales | 0.009 | 0.004 | | 0.013 | 0.003 | | 0.018 | 0.006 | | 0.013 | 0.005 | | 0.012 | 0.005 | | 0.006 | 0.002 | | 0.002 | 0.000 | | 0.002 | 0.000 | | 0.002 | 0.001 |
| Taphrinales | 0.004 | 0.003 | | 0.003 | 0.001 | | 0.009 | 0.001 | | 0.005 | 0.001 | | 0.004 | 0.001 | | 0.003 | 0.000 | | 0.014 | 0.002 | | 0.012 | 0.002 | | 0.018 | 0.003 |
| Agaricales | 0.000 | 0.000 | | 0.001 | 0.000 | | 0.013 | 0.003 | | 0.011 | 0.002 | | 0.004 | 0.003 | | 0.004 | 0.001 | | 0.006 | 0.001 | | 0.009 | 0.006 | | 0.008 | 0.002 |
| Others b | 0.133 | 0.002 | | 0.103 | 0.001 | | 0.184 | 0.001 | | 0.193 | 0.002 | | 0.164 | 0.001 | | 0.101 | 0.000 | | 0.178 | 0.001 | | 0.139 | 0.000 | | 0.193 | 0.001 |
Figure 3.
Most abundant fungal genera in the strawberry phyllosphere at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) during two subsequent years (2011, 2012) as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) and as determined by 454 pyrosequencing. The control (W) was treated with water from a lotic reservoir. Results from all treatments at BBCH 60 (prior to BCA applications) represent the resident fungal community. The color legend refers to the relative abundance (% of total sequences) of fungal OTUs.
Figure 3.
Most abundant fungal genera in the strawberry phyllosphere at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) during two subsequent years (2011, 2012) as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) and as determined by 454 pyrosequencing. The control (W) was treated with water from a lotic reservoir. Results from all treatments at BBCH 60 (prior to BCA applications) represent the resident fungal community. The color legend refers to the relative abundance (% of total sequences) of fungal OTUs.
The composition of fungal communities did not differ between the untreated leaf samples in flowering strawberry plants in 2011 (ANOSIM:
p = 0.255;
α = 0.05;
R = 0.07). In contrast, in 2012 the composition of fungal communities in leaf samples obtained from plots intended for multiple strain treatments significantly differed from leaf samples from plots designated for treatments with water (
p = 0.0316;
α = 0.05;
R = 1) and
A. pullulans (
p = 0.0304;
α = 0.05;
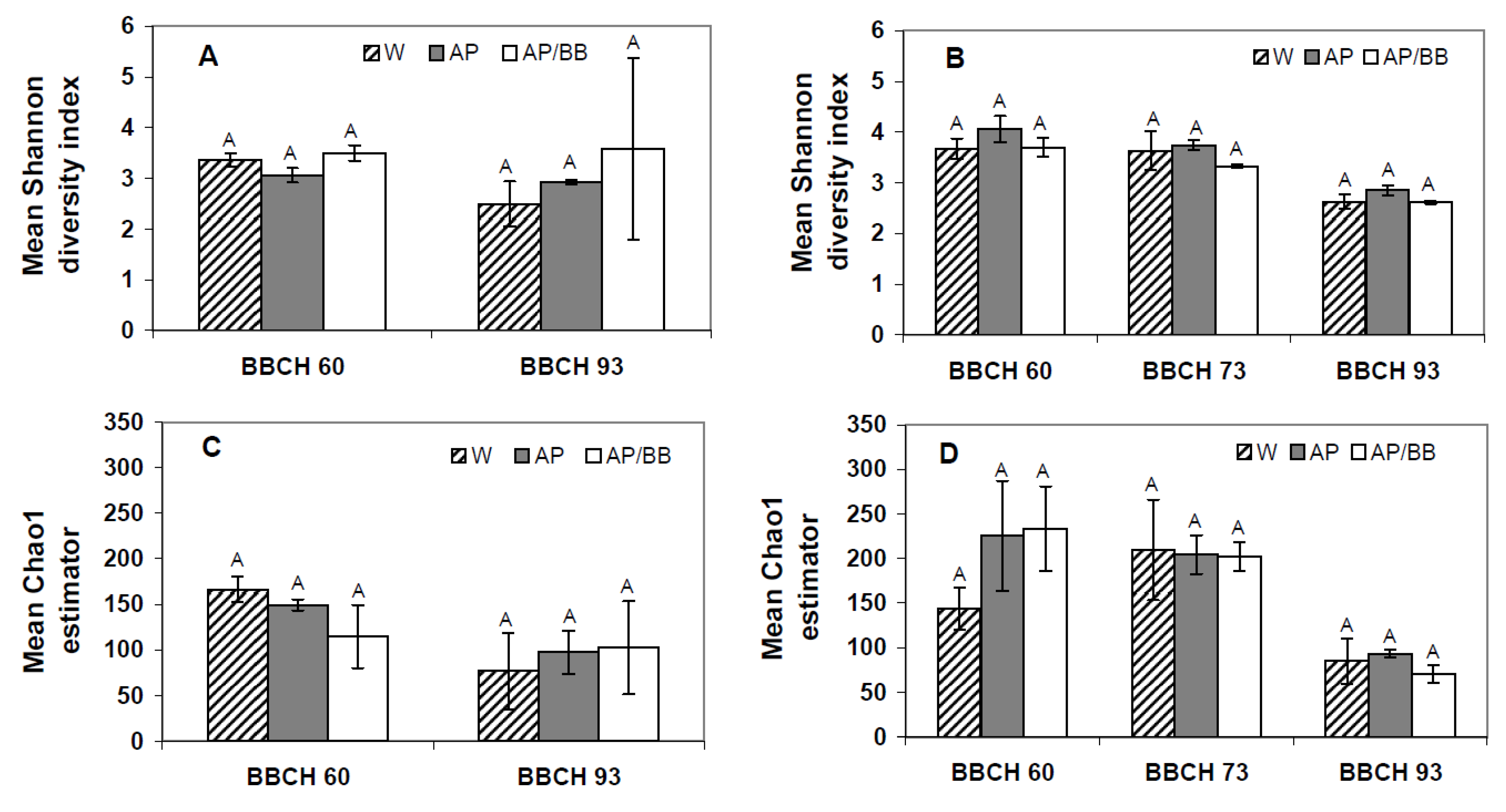
R = 1), respectively. These contrasting results from the two years at the early phenological stage is also pronounced by diversity index (Shannon diversity indices) and genus richness (Chao1), which were similar between all plots in 2011 (
Figure 4A,C) but displayed a significantly higher fungal richness in leaf samples obtained from plots intended for the co-inoculation treatment than in the other leaf samples collected from flowering strawberry plants in 2012 (
Figure 4D). Overall diversity of fungal communities, however, was similar among all leaf samples in 2012.
Figure 4.
Shannon diversity indices and Chao1 estimators describing the biodiversity and taxa richness of fungal phyllosphere communities at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) in 2011 (A,C) and 2012 (B,D). The control (W) was treated with water from a lotic reservoir. Error bars represent SEM values. Columns marked with the same letters are not significantly different within each phenological stage (α = 0.05).
Figure 4.
Shannon diversity indices and Chao1 estimators describing the biodiversity and taxa richness of fungal phyllosphere communities at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) in 2011 (A,C) and 2012 (B,D). The control (W) was treated with water from a lotic reservoir. Error bars represent SEM values. Columns marked with the same letters are not significantly different within each phenological stage (α = 0.05).
The resident leaf microbiome of fruit-setting plants was dominated by the fungal orders Filobasidiales, Dothideales, Capnodiales and Pleosporales in 2011 (
Table 2). In these samples, the abundance of
Cryptococcus decreased to 8.7%, while the genus
Aureobasidium became similarly abundant (
Figure 3). In 2012, the fungal orders Filobasidiales and Capnodiales together accounted for 48.6% of the total resident fungal community on leaves collected from fruit-setting strawberries in the control samples (
Table 3). At this stage in 2012, the genus
Cryptococcus was the major genus in the control samples. When old dying leaves were scrutinized in 2011, the resident fungal community was dominated by the orders Filobasidiales and Capnodiales in the control samples (
Table 2), while also a third order, the Helotiales, was highly represented at this stage in 2012 (
Table 3). The abundance of
Cryptococcus made up 38.3% (2011) and 23.3% (2012) in the non-treated leaf samples (
Figure 3). Thus, in both years, the genus
Cryptococcus was highly abundant in the phyllosphere of strawberries throughout the season. This yeast is well adapted to the phyllosphere and is commonly isolated from the phyllosphere of different crops [
20,
29,
30] as well as from ripe strawberry fruit [
31]. Likewise, the yeast-like fungus
A. pullulans is considered as an ubiquitous and well adapted resident of the phyllosphere [
28,
32]. The absence of
Aureobasidium spp. in plate counts from untreated leaf samples at later phenological stages in 2012 was, therefore, unexpected.
The resident bacterial phyllosphere communities of flowering plants were essentially composed of the orders Sphingobacteriales, Burkholderiales and Actinomycetales in 2011 (
Table 4), whereas the Rickettsiales and Actinomycetales represented the most abundant orders at this stage in 2012 (
Table 5). The bacterial genus
Hymenobacter was a dominant bacterial resident in the phyllosphere during flowering (BBCH 60) in 2011, but displayed low abundance in 2012 (
Figure 5). Although the occurrence of the genus
Hymenobacter has been reported from the plant’s phyllosphere [
33,
34], there is still little knowledge of its role in the phyllosphere. Furthermore, the relative abundance of the genus
Bacillus was high in leaf samples from plots designated for later multiple strain treatments (15.8%) in flowering plants in 2012, while it was below 4.0% for the other leaf samples. As a consequence, diversity and genus richness as well as the composition of bacterial communities (ANOSIM:
p = 0.065;
α = 0.05;
R = 0.325) were similar among all untreated leaf samples during flowering in 2011 (
Figure 6A,C), whereas the bacterial composition in leaf samples from plots designated for multiple strain treatments significantly differed from those designated for water (ANOSIM:
p = 0.030;
α = 0.05;
R = 0.771) and single treatments with
A. pullulans (ANOSIM:
p = 0.029;
α = 0.05;
R = 0.833) at this phenological stage in 2012. Diversity as well as genus richness, though, was similar between the untreated leaf samples collected during flowering in 2012 (
Figure 6B,D).
The resident leaf microbiome was mainly composed of the Enterobacteriales and Burkholderiales during fruit-setting (BBCH 73) in 2012 (
Table 5) with
Yersinia,
Microcoleus and
Pseudomonas as the most abundant bacterial genera (
Figure 5). The orders Enterobacteriales, Sphingomonadales and Pseudomonadales were highly abundant in leaf samples from control plots at BBCH 93 in 2011 (
Table 4). Accordingly, the genera
Sphingomonas and
Pseudomonas were predominant in these leaf samples, whereas the abundance of the genus
Hymenobacter decreased to 2.4% (
Figure 5). The bacterial order Enterobacteriales was predominant in control leaf samples at BBCH 93 in 2012, followed by the Sphingomonadales and Burkholderiales (
Table 5). The genera
Erwinia,
Sphingomonas and
Pseudomonas together accounted for about 44% of the total resident bacterial community in the control samples at this sampling date.
Table 4.
Relative abundance of 16S rRNA reads assigned to bacterial orders in the strawberry phyllosphere at two distinct phenological stages (BBCH 60 and BBCH 93) in 2011 as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) through commercial formulations. The control (W) was treated with water from a lotic reservoir.
Table 4.
Relative abundance of 16S rRNA reads assigned to bacterial orders in the strawberry phyllosphere at two distinct phenological stages (BBCH 60 and BBCH 93) in 2011 as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) through commercial formulations. The control (W) was treated with water from a lotic reservoir.
| | BBCH 60 (prior to BCA applications) | | BBCH 93 (four weeks after last BCA application) |
|---|
| | W a | | AP a | | AP/BB a | | W | | AP | | AP/BB |
|---|
| Bacterial order | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM |
| Burkholderiales | 0.183 | 0.052 | | 0.153 | 0.072 | | 0.092 | 0.015 | | 0.074 | 0.037 | | 0.144 | 0.040 | | 0.130 | 0.031 |
| Sphingobacteriales | 0.213 | 0.050 | | 0.200 | 0.046 | | 0.167 | 0.002 | | 0.017 | 0.017 | | 0.112 | 0.056 | | 0.053 | 0.017 |
| Actinomycetales | 0.130 | 0.010 | | 0.071 | 0.014 | | 0.160 | 0.010 | | 0.057 | 0.033 | | 0.044 | 0.018 | | 0.102 | 0.023 |
| Sphingomonadales | 0.058 | 0.029 | | 0.041 | 0.016 | | 0.097 | 0.026 | | 0.220 | 0.022 | | 0.106 | 0.028 | | 0.162 | 0.037 |
| Rickettsiales | 0.049 | 0.005 | | 0.089 | 0.046 | | 0.084 | 0.038 | | 0.007 | 0.007 | | 0.043 | 0.034 | | 0.028 | 0.012 |
| Pseudomonadales | 0.006 | 0.002 | | 0.000 | 0.000 | | 0.050 | 0.043 | | 0.189 | 0.129 | | 0.169 | 0.029 | | 0.156 | 0.044 |
| Enterobacteriales | 0.010 | 0.008 | | 0.002 | 0.002 | | 0.000 | 0.000 | | 0.229 | 0.092 | | 0.158 | 0.069 | | 0.089 | 0.019 |
| Rhizobiales | 0.033 | 0.008 | | 0.015 | 0.001 | | 0.029 | 0.014 | | 0.056 | 0.021 | | 0.090 | 0.008 | | 0.107 | 0.024 |
| Lactobacillales | 0.002 | 0.002 | | 0.114 | 0.114 | | 0.026 | 0.010 | | 0.015 | 0.008 | | 0.023 | 0.011 | | 0.033 | 0.014 |
| Rhodospirillales | 0.021 | 0.008 | | 0.024 | 0.005 | | 0.026 | 0.010 | | 0.020 | 0.018 | | 0.004 | 0.004 | | 0.027 | 0.008 |
| Bacillales | 0.033 | 0.011 | | 0.012 | 0.001 | | 0.033 | 0.017 | | 0.007 | 0.006 | | 0.019 | 0.019 | | 0.011 | 0.004 |
| Pasteurellales | 0.002 | 0.002 | | 0.071 | 0.056 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.002 | 0.002 | | 0.009 | 0.007 |
| Exiguobacterales | 0.090 | 0.090 | | 0.009 | 0.002 | | 0.011 | 0.004 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Rhodobacterales | 0.004 | 0.002 | | 0.005 | 0.005 | | 0.026 | 0.005 | | 0.008 | 0.006 | | 0.012 | 0.012 | | 0.003 | 0.003 |
| Myxococcales | 0.005 | 0.003 | | 0.002 | 0.001 | | 0.015 | 0.001 | | 0.001 | 0.001 | | 0.008 | 0.006 | | 0.005 | 0.003 |
| Chroococcales | 0.014 | 0.008 | | 0.024 | 0.021 | | 0.011 | 0.011 | | 0.001 | 0.001 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Bifidobacteriales | 0.001 | 0.001 | | 0.034 | 0.032 | | 0.004 | 0.004 | | 0.000 | 0.000 | | 0.002 | 0.002 | | 0.007 | 0.007 |
| Neisseriales | 0.003 | 0.002 | | 0.039 | 0.036 | | 0.004 | 0.004 | | 0.000 | 0.000 | | 0.002 | 0.002 | | 0.002 | 0.002 |
| Bacteroidales | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.059 | 0.059 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Flavobacteriales | 0.000 | 0.000 | | 0.001 | 0.001 | | 0.004 | 0.004 | | 0.007 | 0.006 | | 0.017 | 0.017 | | 0.002 | 0.002 |
| Xanthomonadales | 0.006 | 0.003 | | 0.002 | 0.002 | | 0.011 | 0.004 | | 0.001 | 0.001 | | 0.007 | 0.006 | | 0.001 | 0.001 |
| Solirubrobacterales | 0.006 | 0.004 | | 0.007 | 0.003 | | 0.008 | 0.008 | | 0.001 | 0.001 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| iii1-15 | 0.003 | 0.002 | | 0.002 | 0.001 | | 0.019 | 0.012 | | 0.001 | 0.001 | | 0.000 | 0.000 | | 0.003 | 0.003 |
| Acidimicrobiales | 0.003 | 0.002 | | 0.001 | 0.001 | | 0.008 | 0.008 | | 0.001 | 0.001 | | 0.002 | 0.002 | | 0.005 | 0.003 |
| 0319-7L14 | 0.003 | 0.002 | | 0.000 | 0.000 | | 0.011 | 0.004 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.001 | 0.001 |
| Others b | 0.123 | 0.040 | | 0.080 | 0.016 | | 0.107 | 0.061 | | 0.030 | 0.020 | | 0.037 | 0.030 | | 0.062 | 0.037 |
Table 5.
Relative abundance of 16S rRNA reads assigned to bacterial orders in the strawberry phyllosphere at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) in 2012 as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) through commercial formulations. The control (W) was treated with water from a lotic reservoir.
Table 5.
Relative abundance of 16S rRNA reads assigned to bacterial orders in the strawberry phyllosphere at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) in 2012 as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) through commercial formulations. The control (W) was treated with water from a lotic reservoir.
| | BBCH 60 (prior to BCA applications) | | BBCH 73 (after four BCA applications) | | BBCH 93 (four weeks after last BCA application) |
|---|
| | W a | | AP a | | AP/BB a | | W | | AP | | AP/BB | | W | | AP | | AP/BB |
|---|
| Bacterial order | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM | | Mean | SEM |
| Sphingomonadales | 0.039 | 0.004 | | 0.034 | 0.010 | | 0.087 | 0.029 | | 0.070 | 0.016 | | 0.037 | 0.009 | | 0.122 | 0.082 | | 0.182 | 0.028 | | 0.223 | 0.017 | | 0.314 | 0.052 |
| Enterobacteriales | 0.034 | 0.021 | | 0.005 | 0.001 | | 0.091 | 0.043 | | 0.151 | 0.099 | | 0.009 | 0.004 | | 0.012 | 0.008 | | 0.385 | 0.082 | | 0.169 | 0.034 | | 0.164 | 0.059 |
| Burkholderiales | 0.057 | 0.010 | | 0.070 | 0.011 | | 0.103 | 0.009 | | 0.111 | 0.014 | | 0.080 | 0.015 | | 0.172 | 0.023 | | 0.125 | 0.021 | | 0.110 | 0.013 | | 0.130 | 0.015 |
| Rickettsiales | 0.194 | 0.037 | | 0.173 | 0.017 | | 0.053 | 0.023 | | 0.084 | 0.041 | | 0.186 | 0.050 | | 0.106 | 0.019 | | 0.012 | 0.005 | | 0.012 | 0.006 | | 0.026 | 0.013 |
| Sphingobacteriales | 0.045 | 0.006 | | 0.077 | 0.018 | | 0.096 | 0.016 | | 0.068 | 0.016 | | 0.097 | 0.031 | | 0.188 | 0.002 | | 0.053 | 0.011 | | 0.093 | 0.018 | | 0.077 | 0.027 |
| Rhizobiales | 0.069 | 0.012 | | 0.067 | 0.007 | | 0.083 | 0.023 | | 0.045 | 0.009 | | 0.050 | 0.006 | | 0.031 | 0.001 | | 0.090 | 0.017 | | 0.132 | 0.008 | | 0.120 | 0.012 |
| Actinomycetales | 0.157 | 0.015 | | 0.123 | 0.014 | | 0.062 | 0.012 | | 0.079 | 0.021 | | 0.138 | 0.014 | | 0.048 | 0.024 | | 0.018 | 0.004 | | 0.018 | 0.004 | | 0.023 | 0.006 |
| Pseudomonadales | 0.020 | 0.008 | | 0.014 | 0.003 | | 0.031 | 0.009 | | 0.084 | 0.072 | | 0.037 | 0.015 | | 0.032 | 0.004 | | 0.084 | 0.040 | | 0.188 | 0.034 | | 0.109 | 0.040 |
| Chroococcales | 0.051 | 0.014 | | 0.024 | 0.013 | | 0.045 | 0.012 | | 0.073 | 0.025 | | 0.074 | 0.040 | | 0.116 | 0.094 | | 0.001 | 0.001 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Bacillales | 0.040 | 0.009 | | 0.029 | 0.009 | | 0.170 | 0.012 | | 0.029 | 0.007 | | 0.033 | 0.008 | | 0.025 | 0.016 | | 0.001 | 0.001 | | 0.000 | 0.000 | | 0.001 | 0.001 |
| Rhodospirillales | 0.042 | 0.010 | | 0.037 | 0.000 | | 0.025 | 0.005 | | 0.034 | 0.010 | | 0.031 | 0.007 | | 0.028 | 0.000 | | 0.002 | 0.001 | | 0.001 | 0.001 | | 0.001 | 0.001 |
| Xanthomonadales | 0.022 | 0.004 | | 0.019 | 0.010 | | 0.010 | 0.004 | | 0.010 | 0.004 | | 0.017 | 0.006 | | 0.012 | 0.008 | | 0.001 | 0.001 | | 0.003 | 0.001 | | 0.004 | 0.003 |
| iii1-15 | 0.010 | 0.002 | | 0.034 | 0.006 | | 0.006 | 0.001 | | 0.011 | 0.005 | | 0.014 | 0.006 | | 0.004 | 0.004 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Rhodobacterales | 0.005 | 0.003 | | 0.007 | 0.004 | | 0.006 | 0.003 | | 0.002 | 0.001 | | 0.006 | 0.003 | | 0.000 | 0.000 | | 0.009 | 0.004 | | 0.012 | 0.004 | | 0.009 | 0.002 |
| Myxococcales | 0.006 | 0.003 | | 0.014 | 0.005 | | 0.008 | 0.003 | | 0.008 | 0.003 | | 0.010 | 0.004 | | 0.007 | 0.001 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Solirubrobacterales | 0.019 | 0.005 | | 0.014 | 0.002 | | 0.005 | 0.002 | | 0.009 | 0.003 | | 0.004 | 0.002 | | 0.002 | 0.002 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Acidimicrobiales | 0.016 | 0.006 | | 0.011 | 0.003 | | 0.006 | 0.001 | | 0.007 | 0.003 | | 0.011 | 0.002 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Gaiellales | 0.013 | 0.006 | | 0.011 | 0.005 | | 0.007 | 0.001 | | 0.006 | 0.003 | | 0.010 | 0.005 | | 0.003 | 0.003 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Oceanospirillales | 0.020 | 0.020 | | 0.000 | 0.000 | | 0.001 | 0.000 | | 0.001 | 0.001 | | 0.018 | 0.018 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Flavobacteriales | 0.003 | 0.001 | | 0.013 | 0.008 | | 0.002 | 0.001 | | 0.002 | 0.001 | | 0.002 | 0.001 | | 0.002 | 0.002 | | 0.001 | 0.001 | | 0.003 | 0.001 | | 0.002 | 0.001 |
| Chthoniobacterales | 0.002 | 0.002 | | 0.011 | 0.003 | | 0.001 | 0.000 | | 0.001 | 0.001 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 | | 0.000 | 0.000 |
| Others b | 0.138 | 0.050 | | 0.210 | 0.072 | | 0.102 | 0.022 | | 0.120 | 0.044 | | 0.135 | 0.030 | | 0.090 | 0.023 | | 0.037 | 0.020 | | 0.036 | 0.010 | | 0.022 | 0.005 |
Figure 5.
Most abundant bacterial genera in the strawberry phyllosphere at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) during two subsequent years (2011, 2012) as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) and as determined by 454 pyrosequencing. The control (W) was treated with water from a lotic reservoir. Results from all treatments at BBCH 60 (prior to BCA applications) represent the resident bacterial community. Data for the second sampling (BBCH 73, after four BCA applications) in 2011 were insufficient and are not shown. The color legend refers to the relative abundance (% of total sequences) of bacterial OTUs.
Figure 5.
Most abundant bacterial genera in the strawberry phyllosphere at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) during two subsequent years (2011, 2012) as influenced by treatments with Aureobasidium pullulans as a single strain treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) and as determined by 454 pyrosequencing. The control (W) was treated with water from a lotic reservoir. Results from all treatments at BBCH 60 (prior to BCA applications) represent the resident bacterial community. Data for the second sampling (BBCH 73, after four BCA applications) in 2011 were insufficient and are not shown. The color legend refers to the relative abundance (% of total sequences) of bacterial OTUs.
Figure 6.
Shannon diversity indices and Chao1 estimators describing the biodiversity and taxa richness of bacterial phyllosphere communities at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) as influenced by treatments with Aureobasidium pullulans as a single treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) in 2011 (A,C) and 2012 (B,D). The control (W) was treated with water from a lotic reservoir. Error bars represent SEM values. Columns marked with the same letters are not significantly different within each phenological stage (α = 0.05).
Figure 6.
Shannon diversity indices and Chao1 estimators describing the biodiversity and taxa richness of bacterial phyllosphere communities at three distinct phenological stages (BBCH 60, BBCH 73, and BBCH 93) as influenced by treatments with Aureobasidium pullulans as a single treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) in 2011 (A,C) and 2012 (B,D). The control (W) was treated with water from a lotic reservoir. Error bars represent SEM values. Columns marked with the same letters are not significantly different within each phenological stage (α = 0.05).
The results from both plate counts and 454 pyrosequencing suggest a general change in the resident leaf microbiome during the course of the growing season of strawberries. Seasonal changes of the microbial communities in the phyllosphere of strawberries have previously been observed [
22]. It is suggested that the changes in microbial phyllosphere communities are driven by changing environmental conditions in general (e.g., permanent increase of day length, temperature, and irradiation) and by environmental events such as heavy rainfall [
9,
18]. In addition, the ageing of leaves also contributes to seasonal changes in microbial phyllosphere communities as already shown for the phyllosphere of sugar beet [
29], mango [
20], and cottonwood trees [
35]. Leaf age affects leaf properties, both with respect to the leaf morphology [
19] and with respect to physiological properties, such as photosynthetic activity [
36]. These changes affect the size of the organic compound pool, sites of exudation and the quality of organic compounds exudated over the cuticle and, in turn, affect the microbial community composition. In the present study, the number of resident fungi was considerably increased at the end of the growing season both in 2011 and 2012, which most probably resulted from a general increase in nutrient leakage in ageing or dying leaves [
18].
In addition to seasonal changes, considerable differences in culturable and non-culturable microbial communities were observed between 2011 and 2012. The difference between 2011 and 2012 was most pronounced for bacterial communities as determined by 454 pyrosequencing.
2.2. Effects of Introducing BCAs to the Strawberry Phyllosphere
According to plate counts,
Aureobasidium spp. were significantly increased in BCA-treated leaf samples at fruit-setting stage (BBCH 73),
i.e., after four BCA applications and when old leaves were dying (BBCH 93),
i.e., four weeks after adjournment of BCA applications in 2011 (
Figure 2). In contrast,
Aureobasidium counts were neither significantly elevated in BCA-treated leaf samples at BBCH 73 nor at BBCH 93 in 2012 as compared to the controls.
In addition, according to 454 pyrosequencing, the relative abundance of
Aureobasidium was significantly higher in
A. pullulans-treated leaf samples than in control samples (
p = 0.024) at BBCH 73 in 2011. At this stage, leaf samples from plots treated with multiple strains not only showed significantly increased abundances of
Aureobasidium as compared to control leaf samples (
p < 0.001) but also as compared to single strain treatments with
A. pullulans (
p = 0.022,
Figure 7). In 2012, the relative abundance of
Aureobasidium was only significantly increased in co-inoculated leaf samples during fruit-setting as compared to control (
p < 0.001) and
A. pullulans-treated leaf samples (
p < 0.001). Four weeks after the last BCA application (BBCH 93), pronounced effects, in terms of relative abundance of
Aureobasidium spp., were observed during the first experimental year, whereas its relative abundance was comparably low (<8%) in all leaf samples in the second year (
Figure 7).
Both, plate counts as well as 454 pyrosequencing results indicated that the introduced yeast-like fungus
A. pullulans established in 2011 during fruit development and endured in the strawberry’s phyllosphere for at least four weeks. As
A. pullulans is a wide-spread colonizer of the phyllosphere and well adapted to this habitat [
28], this result was anticipated. Likewise, the limited establishment of
A. pullulans in 2012 is puzzling.
As the co-inoculated BCA, namely
B. bassiana, did not establish epiphytically in the phyllosphere in both 2011 and 2012 (
Figure 2), which is in accordance to the results from a previous study [
22], we conclude that the co-application of the entomopathogenic fungus
B. bassiana did not impair the establishment of
A. pullulans. Indeed, the relative abundance of
A. pullulans sometimes was significantly higher in multiple strain treatments than in single treatments with
A. pullulans as determined by 454 pyrosequencing (
Figure 7). In future experiments, it needs to be validated if co-application of
B. bassiana might promote the establishment of
A. pullulans, for instance indirectly by the potential endophytic growth of the co-applied fungus
B. bassiana [
37,
38].
As for the resident microbial communities, the differences in the establishment of the model organism
A. pullulans might be explained by (i) environmental factors and (ii) plant physiological factors [
7,
9], as well as (iii) the fitness of the introduced organism. In addition, introduced biological control agents have to compete with the resident microorganisms for nutrients and space to successfully immigrate and colonize the phyllosphere [
10,
11]. This paper illustrates the dynamics of both microbial residents and transients in the strawberry phyllosphere during the course of the crop. It might therefore be tempting to explain the present results by changing conditions during leaf maturation and shifting microbial functions of the residents. In this context, the role of
Hymenobacter spp. and of representatives of the fungal order Helotiales would be interesting to further examine. Does
Hymenobacter support the establishment of
A. pullulans in the phyllosphere? Do representatives of Helotiales counteract the BCA’s adhesion to the strawberry phyllosphere?
Figure 7.
Relative abundance of ITS rRNA sequences assigned to the genus Aureobasidium in strawberry leaf samples at BBCH 73 (after four BCA applications) and BBCH 93 (four weeks after the last BCA application) as influenced by treatments with Aureobasidium pullulans as a single treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) in 2011 and 2012. The control (W) was treated with water from a lotic reservoir. Error bars represent SEM values. Columns marked with the same letters are not significantly different within each phenological stage (α = 0.05).
Figure 7.
Relative abundance of ITS rRNA sequences assigned to the genus Aureobasidium in strawberry leaf samples at BBCH 73 (after four BCA applications) and BBCH 93 (four weeks after the last BCA application) as influenced by treatments with Aureobasidium pullulans as a single treatment (AP) or co-inoculated with Beauveria bassiana (AP/BB) in 2011 and 2012. The control (W) was treated with water from a lotic reservoir. Error bars represent SEM values. Columns marked with the same letters are not significantly different within each phenological stage (α = 0.05).
The results of the present study may also be seen in the light of the postulates of Cook
et al. [
17] on competitive displacement of non-target residents. As no plant pathogen was involved into the present study, the potential of such displacement caused by
A. pullulans may be followed by the different treatments. Due to impaired establishment of the model organism during the second year of experiments (2012), the second year data set is not appropriate to conclude on the organism’s short- or long-term impact and potential competitive displacement of non-target organisms in the strawberry phyllosphere. In the first experimental year (2011), plate counts indicated an increase in numbers of total culturable fungi in BCA-treated leaf samples shortly after repeated BCA applications,
i.e., at BBCH 73 (
Figure 2) as compared to the untreated control. However, as the increased fungal counts probably resulted from the introduction of the yeast-like fungus, we suggest that foliar treatments with the model BCA as single strain treatment or co-inoculated with
B. bassiana have neither a clear short-term nor a long-term impact on the culturable bacterial and fungal phyllosphere communities. These results are in line with findings from Russel
et al. [
24] and Sylla
et al. [
22].
Analysis using culture-independent tools verified the successful introduction of the model organism as the relative abundance of the order Dothideales and the genus
Aureobasidium spp. was much more pronounced in BCA-treated leaf samples than in control samples after four applications of the BCAs (BBCH 73) in 2011 (
Table 2 and
Figure 2). Despite of comparable abundances in the remaining orders and genera in BCA-treated samples compared with the control plots, overall composition of fungal communities in the control samples differed from fungal communities in leaf samples previously treated with
A. pullulans (ANOSIM:
p = 0.027;
α = 0.05;
R = 0.574) and with
A. pullulans co-inoculated with
B. bassiana (ANOSIM:
p = 0.030;
α = 0.05;
R = 1), displaying a significant decrease in fungal diversity shortly after the treatments with the respective BCAs (
Figure 4A). Fungal richness was, however, not affected (
Figure 4C). From the present data we conclude, that short-term displacement of the resident microbiome may occur as a consequence to introduction of
A. pullulans at condition that the BCA establishes successfully. This needs to be verified in future studies, as this finding has not previously been reported for
A. pullulans.
Four weeks after the adjourned BCA applications (BBCH 93) in 2011, the abundance of Dothideales was still increased in BCA-treated leaf samples (
Table 2). Among all leaf samples,
Cryptococcus and
Aureobasidium made up the major fungal genera at this stage, whereas the abundance of
Aureobasidium was highest in
A. pullulans-treated leaf samples (
Figure 3). At this stage, the fungal community composition was still significantly different in leaf samples previously treated with
A. pullulans as single strain treatment as compared to control samples (ANOSIM:
p = 0.029;
α = 0.05;
R = 0.510). This finding indicates that also long-term impacts of introduced
A. pullulans strains on the phyllosphere microbiome may occur if the BCA establishes throughout the progression of the crop.
The yeast-like fungus
A. pullulans is regarded as a promising candidate for biological control of nectrotrophic fungal pathogens (e.g.,
B. cinerea), which require high amounts of exogenous nutrients for germination and mycelial growth [
32], as it strongly competes for nutrients [
15]. We therefore suggest that the displacement of resident fungi might be explained by the good competitiveness of the established BCA in this habitat. It was also reported that other modes of action than competition for nutrients (e.g., induced resistance, enzyme production) might be generally involved in the biological control of plant pathogens [
16,
39] and, consequently, could be also involved in the interactions with the resident microorganisms. There is, however, still not enough knowledge if these mechanisms work
in situ/in the field.
As 454 pyrosequencing data were insufficient for the leaf samples collected at BBCH 73,
i.e., after four BCA treatments, no conclusions on short-term impacts of introduced BCAs on the non-culturable bacterial communities could be made in 2011. Four weeks after the last BCA application (BBCH 93) in 2011, the composition (ANOSIM:
p = 0.797;
α = 0.05;
R = −0.111) as well as diversity and richness of bacterial communities was not significantly affected by previous treatments with
A. pullulans as single strain treatment or co-inoculated with
B. bassiana as compared to control leaf samples (
Figure 6A,C). Interestingly, no long-term effects of the introduced BCAs on the bacterial communities in the phyllosphere were detected by means of 454 pyrosequencing in this study. This finding indicates that bacterial communities might be more stable than fungal communities in the phyllosphere of strawberries under field conditions, although one needs to keep in mind that the entire bacterial microbiota was probably not reflected in 2011. Likewise, Kim
et al. [
23] and Sylla
et al. [
22] did not observe any significant changes in bacterial communities by culture-independent techniques as a consequence of BCA applications under field conditions. In contrast, treatments with
Bacillus thuringiensis significantly altered the composition of bacteria in the phyllosphere of greenhouse-grown pepper [
21].
The results of the present study suggest that the co-application of B. bassiana does not clearly affect the interactions between A. pullulans and the resident strawberry leaf microbiota.
2.3. Interactions between Environmental Conditions and Microbial Dynamics in the Phyllosphere
The phyllosphere is a highly open and dynamic environment, where microorganisms are strongly affected by environmental conditions such as rainfall, temperature and radiation [
7,
18].
In the present study, the mean air temperatures did not considerably differ during the whole season in 2011 and 2012 (
Table 6). In 2011, however, mean temperatures were higher three days prior to the third sampling than in 2012. Accumulated precipitation, however, was considerably higher during the entire growing season as well as the days prior to each leaf sampling in 2012 than in 2011.
Table 6.
Temperatures and precipitation during the experiments in 2011 and 2012.
Table 6.
Temperatures and precipitation during the experiments in 2011 and 2012.
| | Mean temperature [°C] | | Accumulated precipitation [mm] |
|---|
| 2011 | 2012 | | 2011 | 2012 |
|---|
| During the whole season | 17.0 a | 16.0 a | | 103 b | 153 b |
| 7 d prior to 1st sampling (BBCH 60) | NA c | 8.3 | | 3.0 | 7.3 |
| 3 d prior to 1st sampling (BBCH 60) | NA c | 5.6 | | 0.0 | 2.3 |
| 1 d prior to 1st sampling (BBCH 60) | NA c | 5.7 | | 0.0 | 2.0 |
| 7 d prior to 2nd sampling (BBCH 73) | 14.4 | 14.9 | | 8.2 | 12.6 |
| 3 d prior to 2nd sampling (BBCH 73) | 11.4 | 11.2 | | 0.0 | 0.6 |
| 1 d prior to 2nd sampling (BBCH 73) | 9.8 | 10.3 | | 0.0 | 0.6 |
| 7 d prior to 3rd sampling (BBCH 93) | 19.6 | 21.2 | | 7.4 | 14.1 |
| 3 d prior to 3rd sampling (BBCH 93) | 24.3 | 18.8 | | 0.0 | 5.1 |
| 1 d prior to 3rd sampling (BBCH 93) | 26.0 | 21.5 | | 0.0 | 0.0 |
High amounts of precipitation correlated negatively to the establishment of
A. pullulans. In a short- and long-term perspective, the variations in
Aureobasidium counts were explained to 27.4% by the total amount of precipitation and to 37.7% by the amount of precipitation seven days before leaf sampling (
p < 0.001). In addition, the phyllosphere microbiomes differed between the two years as assessed by culture-dependent (
Figure 1 and
Figure 2) and culture-independent methods (
Table 2,
Table 3,
Table 4 and
Table 5), either with or without phyllosphere applied BCAs. This was most apparent for the genus
Hymenobacter,which almost vanished during flowering in 2012,
i.e., prior to BCA applications, as compared to the same phenological stage in the preceding year. When
A. pullulans was introduced, either as single strain treatment or co-inoculated with
B. bassiana, the abundance of the genus
Aureobasidium was most conservative in 2011 as compared to the poor abundance of
A. pullulans in 2012. Accordingly, the displacement effects on resident fungi in the year 2011 as determined by 454 pyrosequencing can be explained by the established
A. pullulans strains. The observed differences in fungal communities between control leaf samples and leaf samples treated with
A. pullulans co-inoculated with
B. bassiana (ANOSIM:
p = 0.029;
α = 0.05;
R = 0.963) at BBCH 73,
i.e., after four applications of the BCAs, however, cannot be explained by the introduced model BCA.
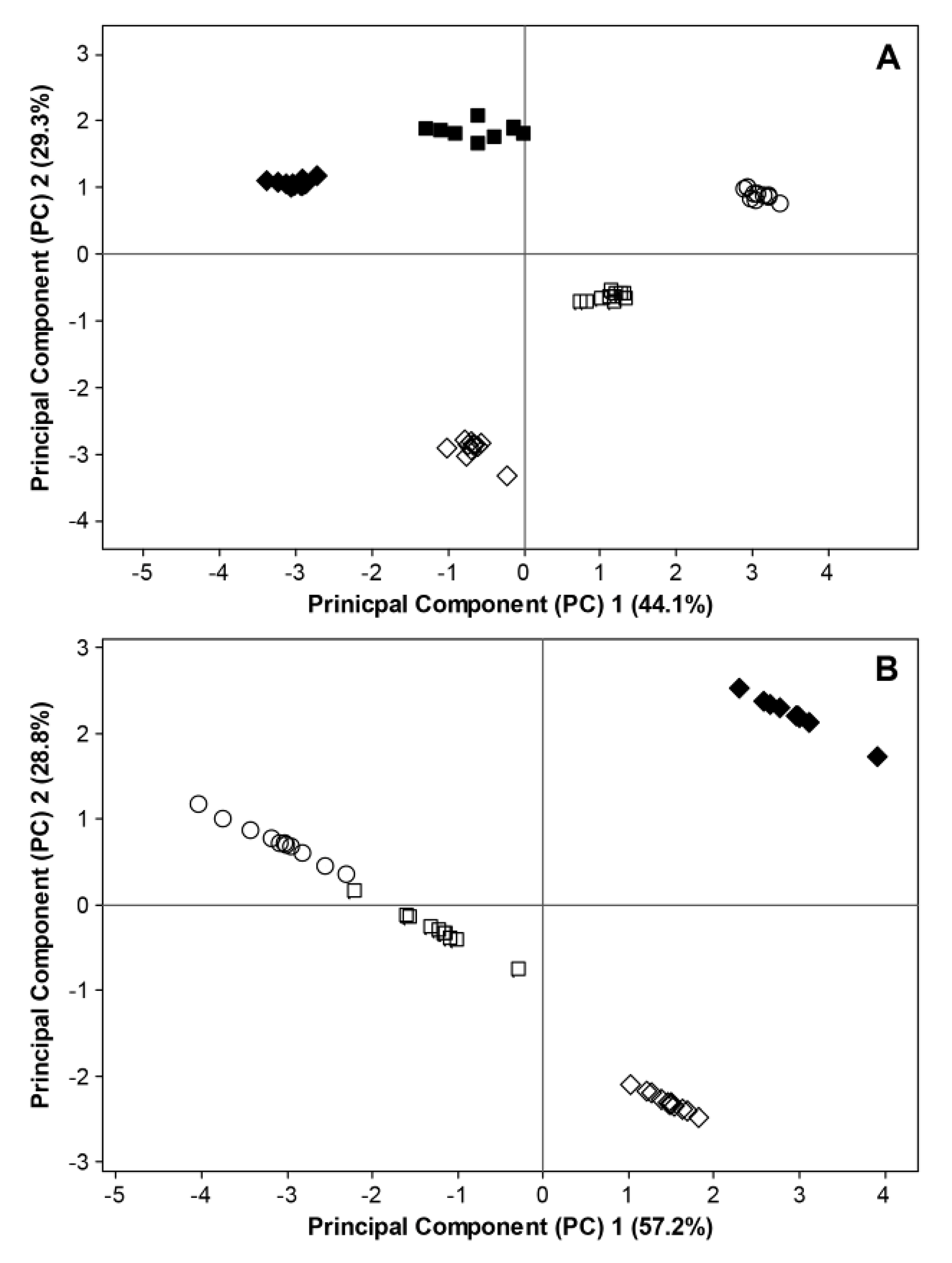
The interactions between phenological stages, weather conditions (precipitation and temperature) prior to sampling as well as microbial diversity and richness were also displayed by principal component analysis, where temperature and precipitation showed to have a strong influence on principal component (PC) 1 and PC 2, respectively (
Figure 8). The samples for fungal ITS rRNA sequences were grouped according to the phenological growth stages in both 2011 and 2012 (
Figure 8A), whereas the variation of the data could mainly be explained by temperature in 2011 and by both temperature and precipitation in 2012. The samples for 16S rRNA sequences were clearly separated at BBCH 93 between 2011 and 2012 (
Figure 8B).
Figure 8.
Principal component analysis (PCA) of interactivities in the phyllosphere microbiome of organically grown strawberries on the basis of Shannon diversity indices, Chao1 estimators, temperature and precipitation data for (
A) fungal ITS rRNA sequences and (
B) bacterial 16S rRNA sequences at different phenological stages (○ = BBCH 60, □ = BBCH 73,
![Agronomy 03 00704 i001]()
= BBCH 93). Filled and open symbols represent data generated in 2011 and 2012, respectively. For 2011, data considering fungal ITS sequences and bacterial 16S rRNA sequences are excluded at BBCH 60 due to missing temperature data. Data for bacterial 16S rRNA sequences are excluded at BBCH 73 due to insufficient bioinformatic data.
Figure 8.
Principal component analysis (PCA) of interactivities in the phyllosphere microbiome of organically grown strawberries on the basis of Shannon diversity indices, Chao1 estimators, temperature and precipitation data for (
A) fungal ITS rRNA sequences and (
B) bacterial 16S rRNA sequences at different phenological stages (○ = BBCH 60, □ = BBCH 73,
![Agronomy 03 00704 i001]()
= BBCH 93). Filled and open symbols represent data generated in 2011 and 2012, respectively. For 2011, data considering fungal ITS sequences and bacterial 16S rRNA sequences are excluded at BBCH 60 due to missing temperature data. Data for bacterial 16S rRNA sequences are excluded at BBCH 73 due to insufficient bioinformatic data.
The strong dependence of community structure on environmental factors has repeatedly been shown [
40,
41]. As no data on boundary layer, nor on plant physiological parameters, were recorded in the present study, it is difficult to determine decisive factors or mechanisms. It is tempting to assume that the model organism was washed off. However, precipitation also affects light intensity and spectrum. At the present stage, it therefore would be unwise to ascribe the observed variations exclusively to mechanical interactions. We, therefore, recommend for future studies, to expand data collection to measurements of boundary layer conditions, further on-line monitoring of environmental conditions, including light intensities, light spectrum, UV-light, wind, and plant physiological responses.
An overhasty conclusion would be to reduce the discussion on the poor establishment of the model organism A. pullulans in the second experimental year to detrimental environmental conditions. It is worth to note, that the present experiment was conducted at the same site during two subsequent years and that the plants, from which the leaf samples were collected in the second year, were exposed for a second year to BCA treatments. Samples from flowering plants in year two (2012) displayed differences in their microbial community composition and structure between the three groups of planned treatments. No leaf samples were taken at a very early stage of the season which could have shed light on the intriguing question if these differences might be perennial long-term effects due to introduction of A. pullulans as a single or multiple strain treatment. This is an interesting question for the use of BCAs in perennial crops and should be further investigated.
The present study reveals new insights on the establishment of A. pullulans as a single or multiple strain treatment in a strawberry crop and to the inoculants’ impact on the resident microbiome. Artificial inoculation of a plant pathogen and plant disease attack would by nature affect the resident microbial community composition and structure. Nevertheless, it points to two important new aspects that need to be included in future studies of short- and long-term effects of BCAs and to explain findings of impaired establishment of BCA, namely continuous monitoring of (i) meteorological and abiotic boundary layer parameters, as well as (ii) plant physiological parameters of importance for leaf colonization. For future studies, we furthermore recommend to consider more than three samplings to more effectively study the resident microbiota of the phyllosphere and the interactions with the introduced BCAs.





 = BBCH 93). Filled and open symbols represent data generated in 2011 and 2012, respectively. For 2011, data considering fungal ITS sequences and bacterial 16S rRNA sequences are excluded at BBCH 60 due to missing temperature data. Data for bacterial 16S rRNA sequences are excluded at BBCH 73 due to insufficient bioinformatic data.
= BBCH 93). Filled and open symbols represent data generated in 2011 and 2012, respectively. For 2011, data considering fungal ITS sequences and bacterial 16S rRNA sequences are excluded at BBCH 60 due to missing temperature data. Data for bacterial 16S rRNA sequences are excluded at BBCH 73 due to insufficient bioinformatic data.
 = BBCH 93). Filled and open symbols represent data generated in 2011 and 2012, respectively. For 2011, data considering fungal ITS sequences and bacterial 16S rRNA sequences are excluded at BBCH 60 due to missing temperature data. Data for bacterial 16S rRNA sequences are excluded at BBCH 73 due to insufficient bioinformatic data.
= BBCH 93). Filled and open symbols represent data generated in 2011 and 2012, respectively. For 2011, data considering fungal ITS sequences and bacterial 16S rRNA sequences are excluded at BBCH 60 due to missing temperature data. Data for bacterial 16S rRNA sequences are excluded at BBCH 73 due to insufficient bioinformatic data.