Next-Generation Bio-Products Sowing the Seeds of Success for Sustainable Agriculture
Abstract
:1. Introduction
1.1. The Plant as Meta-Organism
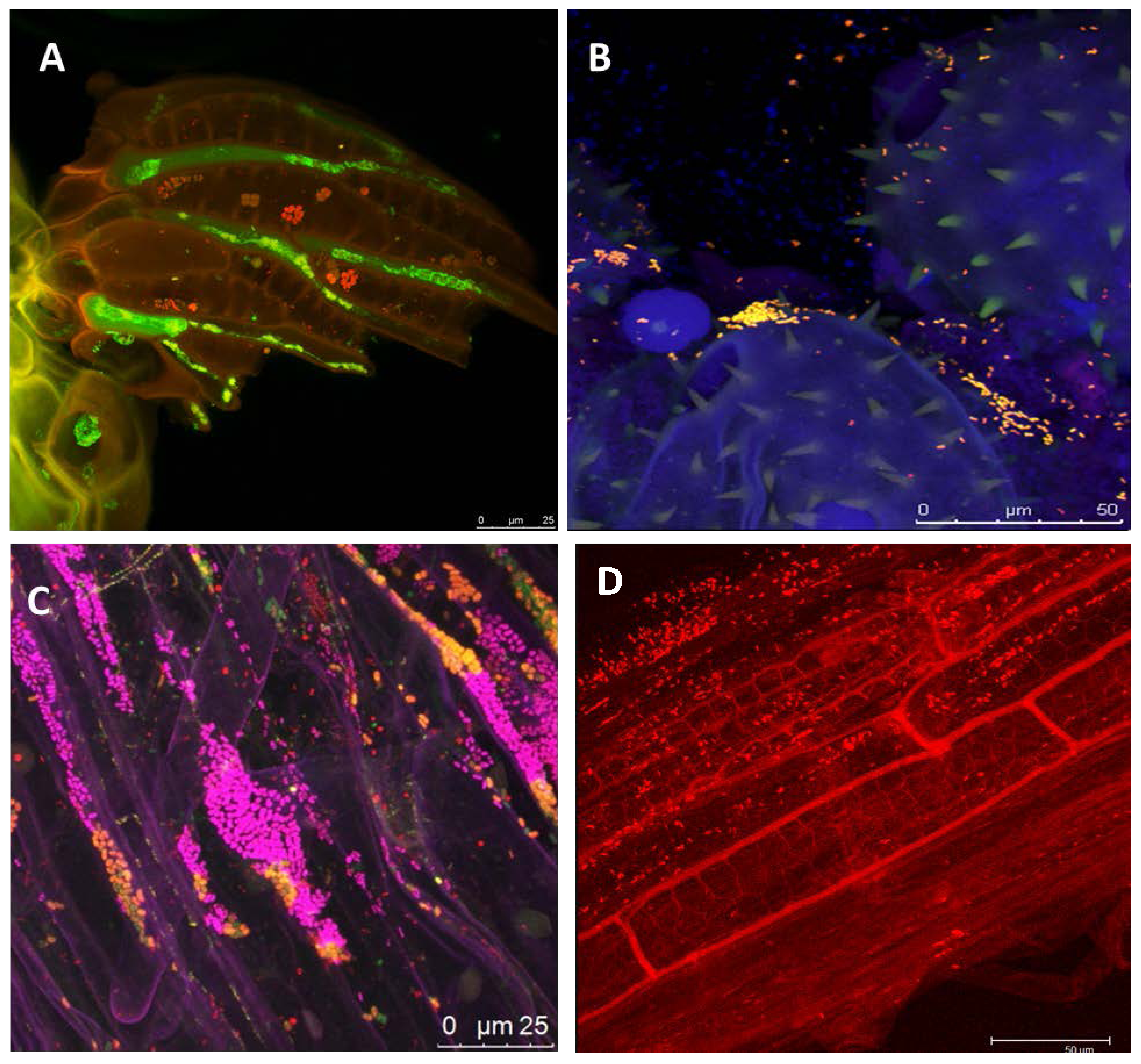
1.2. The Potential of Next-Generation Bio-Products

2. Conclusions
- New tools based on NGS techniques have a high impact on next-generation bio-products. They can improve the detection of new bio-resources for biocontrol and plant growth promoting agents, and also increase their performance and registration efficiency.
- Plants as meta-organisms create numerous new perspectives for sustainable agriculture, e.g., breeding, seed production, and seed treatment processes.
- Plants are a hitherto little explored source for new microbes, genes, and bioactive compounds due to the extent of specificity within the plant microbiome.
- New functions and interactions of plant-associated bacteria can be identified using NGS techniques such as quality enhancement or stress protection.
- Although, the application of single microorganisms is already successful, information on diversity and specificity of the plant microbiome suggest that microbial consortia or natural microbial transplants, e.g., from suppressive soil, can be more successful as single inoculants.
- Together, these points open new windows for sustainable agriculture by means of next-generation bio-products. This trend is currently visible in industrial and university research topics.
- A public dialogue is required in addition to research in order to promote the application of next-generation bio-products in the future for both agriculture and food.
Acknowledgments
Conflicts of Interest
References
- Hartmann, A.; Rothballer, M.; Schmid, M. Lorenz Hiltner, a pioneer in rhizosphere microbial ecology and soil bacteriology research. Plant Soil 2008, 312, 7–14. [Google Scholar] [CrossRef]
- Leveau, J.H.J. The magic and menace of metagenomics: Prospects for the study of plant growth-promoting rhizobacteria. Eur. J. Plant Pathol. 2007, 119, 279–300. [Google Scholar] [CrossRef]
- Jansson, J.K.; Neufeld, J.D.; Moran, M.A.; Gilbert, J.A. Omics for understanding microbial functional dynamics. Environ. Microbiol. 2012, 14, 1–3. [Google Scholar] [CrossRef]
- Sorensen, J.; Hauberg Nicolaisen, M.; Ron, E.; Simonet, P. Molecular tools in rhizosphere microbiology—From single-cell to whole-community analysis. Plant Soil 2009, 321, 483–512. [Google Scholar] [CrossRef]
- Berendsen, R.L.; Pieterse, C.M.; Bakker, P.A. The rhizosphere microbiome and plant health. Trends Plant Sci. 2012, 17, 478–486. [Google Scholar] [CrossRef]
- Berg, G.; Smalla, K. Plant species and soil type cooperatively shape the structure and function of microbial communities in the rhizosphere. FEMS Microbiol. Ecol. 2009, 68, 1–13. [Google Scholar] [CrossRef]
- Smalla, K.; Wieland, G.; Buchner, A.; Zock, A.; Parzy, J.; Kaiser, S.; Roskot, N.; Heuer, H.; Berg, G. Bulk and rhizosphere soil bacterial communities studied by denaturing gradient gel electrophoresis: Plant-dependent enrichment and seasonal shifts revealed. Appl. Environ. Microbiol. 2001, 67, 4742–4751. [Google Scholar]
- Berg, G.; Zachow, C.; Lottmann, J.; Götz, M.; Costa, R.; Smalla, K. Impact of plant species and site on rhizosphere-associated fungi antagonistic to Verticillium dahliae Kleb. Appl. Environ. Microbiol. 2005c, 71, 4203–4213. [Google Scholar] [CrossRef]
- Berg, G.; Opelt, K.; Zachow, C.; Lottmann, J.; Götz, M.; Costa, R.; Smalla, K. The rhizosphere effect on bacteria antagonistic towards the pathogenic fungus Verticillium differs depending on plant species and site. FEMS Microbiol. Ecol. 2006, 56, 250–261. [Google Scholar] [CrossRef]
- Bais, H.P.; Weir, T.L.; Perry, L.G.; Gilroy, S.; Vivanco, J.M. The role of root exudates in rhizosphere interactions with plants and other organisms. Annu. Rev. Plant Biol. 2006, 57, 233–266. [Google Scholar] [CrossRef]
- Doornbos, R.F.; van Loon, L.C.; Bakker, P.A.H.M. Impact of root exudates and plant defense signaling on bacterial communities in the rhizosphere. Agron. Sustain. 2012, 32, 227–243. [Google Scholar] [CrossRef]
- Hartmann, A.; Schmid, M.; van Tuinen, D.; Berg, G. Plant-driven selection of microbes. Plant Soil 2009, 321, 235–257. [Google Scholar] [CrossRef]
- Haichar, F.Z.; Marol, C.; Berge, O.; Rangel-Castro, J.I.; Prosser, J.I.; Balesdent, J.; Heulin, T.; Achouak, W. Plant host habitat and root exudates shape soil bacterial community structure. ISME J. 2008, 2, 1221–1230. [Google Scholar] [CrossRef]
- Lundberg, D.S.; Lebeis, S.L.; Paredes, S.H.; Yourstone, S.; Gehring, J.; Malfatti, S.; Tremblay, J.; Engelbrektson, A.; Kunin, V.; del Rio, T.G.; et al. Defining the core Arabidopsis thaliana root microbiome. Nature 2012, 488, 86–90. [Google Scholar] [CrossRef]
- Bulgarelli, D.; Rott, M.; Schlaeppi, K.; Ver Loren van Themaat , E.; Ahmadinejad, N.; Assenza, F.; Rauf, P.; Huettel, B.; Reinhardt, R.; Schmelzer, E.; et al. Revealing structure and assembly cues for Arabidopsis root-inhabiting bacterial microbiota. Nature 2012, 488, 91–95. [Google Scholar] [CrossRef]
- Vorholt, J.A. Microbial life in the phyllosphere. Nat. Rev. Microbiol. 2012, 10, 828–840. [Google Scholar] [CrossRef]
- Ryan, R.P.; Germaine, K.; Franks, A.; Ryan, D.J.; Dowling, D.N. Bacterial endophytes: Recent developments and applications. FEMS Microbiol. Lett. 2008, 278, 1–9. [Google Scholar] [CrossRef]
- Berg, G.; Krechel, A.; Ditz, M.; Sikora, R.A.; Ulrich, A.; Hallmann, J. Endophytic and ectophytic potato-associated bacterial communities differ in structure and antagonistic function against plant pathogenic fungi. FEMS Microbiol. Ecol. 2005, 51, 215–229. [Google Scholar] [CrossRef]
- Fürnkranz, M.; Lukesch, B.; Müller, H.; Huss, H.; Grube, M.; Berg, G. Microbial diversity inside pumpkins: Microhabitat-specific communities display a high antagonistic potential against phytopathogens. Microb. Ecol. 2012, 63, 418–428. [Google Scholar] [CrossRef]
- Bragina, A.; Berg, C.; Cardinale, M.; Shcherbakov, A.; Chebotar, V.; Berg, G. Sphagnum mosses harbour highly specific bacterial diversity during their whole lifecycle. ISME J. 2012, 6, 802–813. [Google Scholar] [CrossRef]
- Berg, G. Plant-microbe interactions promoting plant growth and health: Perspectives for controlled use of microorganisms in agriculture. J. Appl. Microbiol. Biotechnol. 2009, 84, 11–18. [Google Scholar] [CrossRef]
- Hirsch, P.R.; Mauchline, T.H. Who’s who in the plant root microbiome? Nat. Biotechnol. 2012, 30, 961–962. [Google Scholar] [CrossRef]
- Köberl, M.; Ramadan, E.M.; Roßmann, B.; Staver, C.; Fürnkranz, M.; Lukesch, B.; Grube, M.; Berg, G. Using ecological knowledge and molecular tools to develop effective and safe biocontrol strategies. In Pesticides in the Modern World/Book 5; E-book: Rijeka, Croatia, 2012. [Google Scholar]
- Van der Heijden, M.G.A.; Klironomos, J.N.; Ursic, M.; Moutoglis, P.; Streitwolf-Engel, R.; Boller, T.; Wiemken, A.; Sanders, I.R. Mycorrhizal fungal diversity determines plant biodiversity, ecosystem variability and productivity. Nature 1998, 396, 69–72. [Google Scholar] [CrossRef]
- Latz, E.; Eisenhauer, N.; Rall, B.C.; Allan, E.; Roscher, C.; Scheu, S.; Jousset, A. Plant diversity improves protection against soil-borne pathogens by fostering antagonistic bacterial communities. J. Ecol. 2012, 100, 597–604. [Google Scholar] [CrossRef]
- Opelt, K.; Berg, C.; Berg, G. The bryophyte genus Sphagnum is a reservoir for powerful and extraordinary antagonists and potentially facultative human pathogens. FEMS Microbiol. Ecol. 2007, 61, 38–53. [Google Scholar] [CrossRef]
- Berg, G.; Hartenberger, K.; Liebminger, S.; Zachow, C. Antagonistic endophytes from mistletoes as bio-resource to control plant as well as clean room pathogens. IOBC/wprs Bull. 2012, 79, 29–32. [Google Scholar]
- Zachow, C.; Berg, C.; Müller, H.; Meincke, R.; Komon-Zelazowska, M.; Druzhinina, I.S.; Kubicek, C.P.; Berg, G. Fungal diversity in the rhizosphere of endemic plant species of Tenerife (Canary Islands): Relationship to vegetation zones and environmental factors. ISME J. 2009, 3, 79–92. [Google Scholar] [CrossRef]
- Mazzola, M. Mechanisms of natural soil suppressiveness to soilborne diseases. Antonie Van Leeuwenhoek 2002, 81, 557–564. [Google Scholar] [CrossRef]
- Weller, D.M.; Raaijmakers, J.M.; McSpadden Gardener, B.B.; Thomashow, L.S. Microbial populations responsible for specific soil suppressiveness to plant pathogens. Annu. Rev. Phytopathol. 2002, 40, 309–348. [Google Scholar] [CrossRef]
- Mendes, R.; Kruijt, M.; de Bruijn, I.; Dekkers, E.; van der Voort, M.; Schneider, J.H.M.; Piceno, Y.M.; DeSantis, T.Z.; Andersen, G.L.; Bakker, P.A.; et al. Deciphering the rhizosphere microbiome for disease-suppressive bacteria. Science 2012, 332, 1097–1100. [Google Scholar]
- Schmid, F.; Moser, G.; Müller, H.; Berg, G. Functional and structural microbial diversity in organic and conventional viticulture: Organic farming benefits natural biocontrol agents. Appl. Environ. Microbiol. 2011, 77, 2188–2191. [Google Scholar] [CrossRef]
- Köberl, M.; Müller, H.; Ramadan, E.M.; Berg, G. Desert farming benefits from microbial potential in arid soils and promotes diversity and plant health. PLoS One 2011, 6, e24452. [Google Scholar]
- Grube, M.; Cardinale, M.; Vieira de Castro Junior, J.; Müller, H.; Berg, G. Species-specific structural and functional diversity of bacterial communities in lichen symbiosis. ISME J. 2009, 3, 1105–1115. [Google Scholar] [CrossRef]
- Ehlers, R.U. Regulation of Biological Control Agents; Springer: Heidelberg, Germany, 2011. [Google Scholar]
- Berg, G.; Eberl, L.; Hartmann, A. The rhizosphere as a reservoir for opportunistic human pathogenic bacteria. Environ. Microbiol. 2005, 7, 1673–1685. [Google Scholar] [CrossRef]
- Egamberdieva, D.; Kucharova, Z.; Davranov, K.; Berg, G.; Makarova, N.; Azarova, T.; Chebotar, V.; Tikhonovich, I.; Kamilova, F.; Validov, S.Z.; et al. Bacteria able to control foot and root rot and to promote growth of cucumber in salinated soils. Biol. Fertil. Soils 2011, 47, 197–205. [Google Scholar] [CrossRef]
- Roder, A.; Hoffmann, E.; Hagemann, M.; Berg, G. Synthesis of the compatible solutes glucosylglycerol and trehalose by salt-stressed cells of Stenotrophomonas strains. FEMS Microbiol. Lett. 2005, 7, 1853–1858. [Google Scholar]
- Ryan, R.P.; Monchy, S.; Cardinale, M.; Taghavi, S.; Crossman, L.; Avison, M.B.; Berg, G.; van der Lelie, D.; Dow, J.M. The versatility and adaptation of bacteria from the genus Stenotrophomonas. Nat. Rev. Microbiol. 2009, 7, 514–525. [Google Scholar] [CrossRef]
- Alavi, P.; Starcher, M.R.; Zachow, C.; Müller, H.; Berg, G. Root-microbe systems: The effect and mode of interaction of Stress Protecting Agent (SPA) Stenotrophomonas rhizophila DSM14405T. Front. Plant Sci. 2013, 4, 141. [Google Scholar]
- Alavi, P.; Starcher, M.R.; Thallinger, G.; Zachow, C.; Müller, H.; Berg, G. Stenotrophomonas comparative genomics reveal a fine line between beneficial and pathogenic bacteria. 2013; submitted. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Berg, G.; Zachow, C.; Müller, H.; Philipps, J.; Tilcher, R. Next-Generation Bio-Products Sowing the Seeds of Success for Sustainable Agriculture. Agronomy 2013, 3, 648-656. https://doi.org/10.3390/agronomy3040648
Berg G, Zachow C, Müller H, Philipps J, Tilcher R. Next-Generation Bio-Products Sowing the Seeds of Success for Sustainable Agriculture. Agronomy. 2013; 3(4):648-656. https://doi.org/10.3390/agronomy3040648
Chicago/Turabian StyleBerg, Gabriele, Christin Zachow, Henry Müller, Jörg Philipps, and Ralf Tilcher. 2013. "Next-Generation Bio-Products Sowing the Seeds of Success for Sustainable Agriculture" Agronomy 3, no. 4: 648-656. https://doi.org/10.3390/agronomy3040648
APA StyleBerg, G., Zachow, C., Müller, H., Philipps, J., & Tilcher, R. (2013). Next-Generation Bio-Products Sowing the Seeds of Success for Sustainable Agriculture. Agronomy, 3(4), 648-656. https://doi.org/10.3390/agronomy3040648



