Host Plant Specific Control of 2,4-Diacetylphloroglucinol Production in the Rhizosphere
Abstract
:1. Introduction
2. Results and Discussion
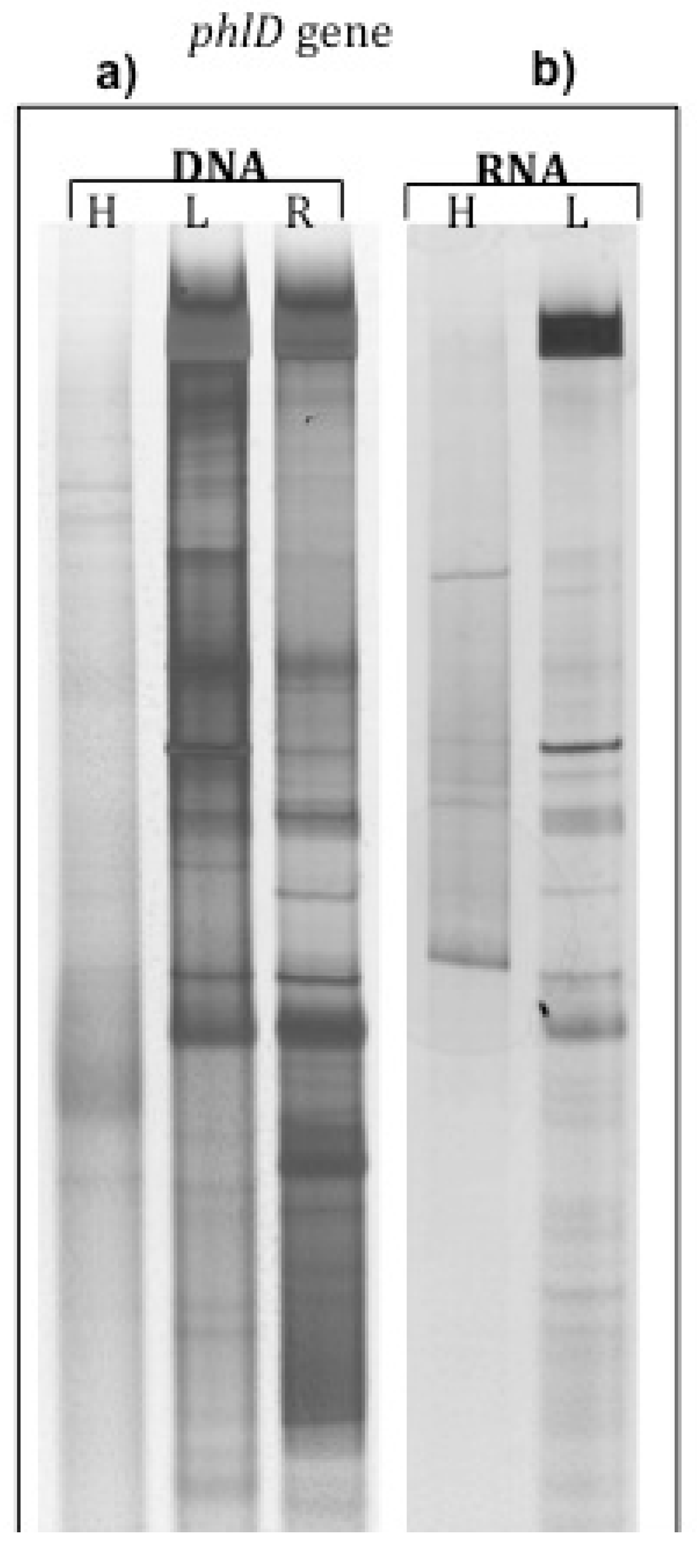
2.1. phlD Gene Expression in the Rhizosphere of A. Thaliana Grown Under Natural Conditions
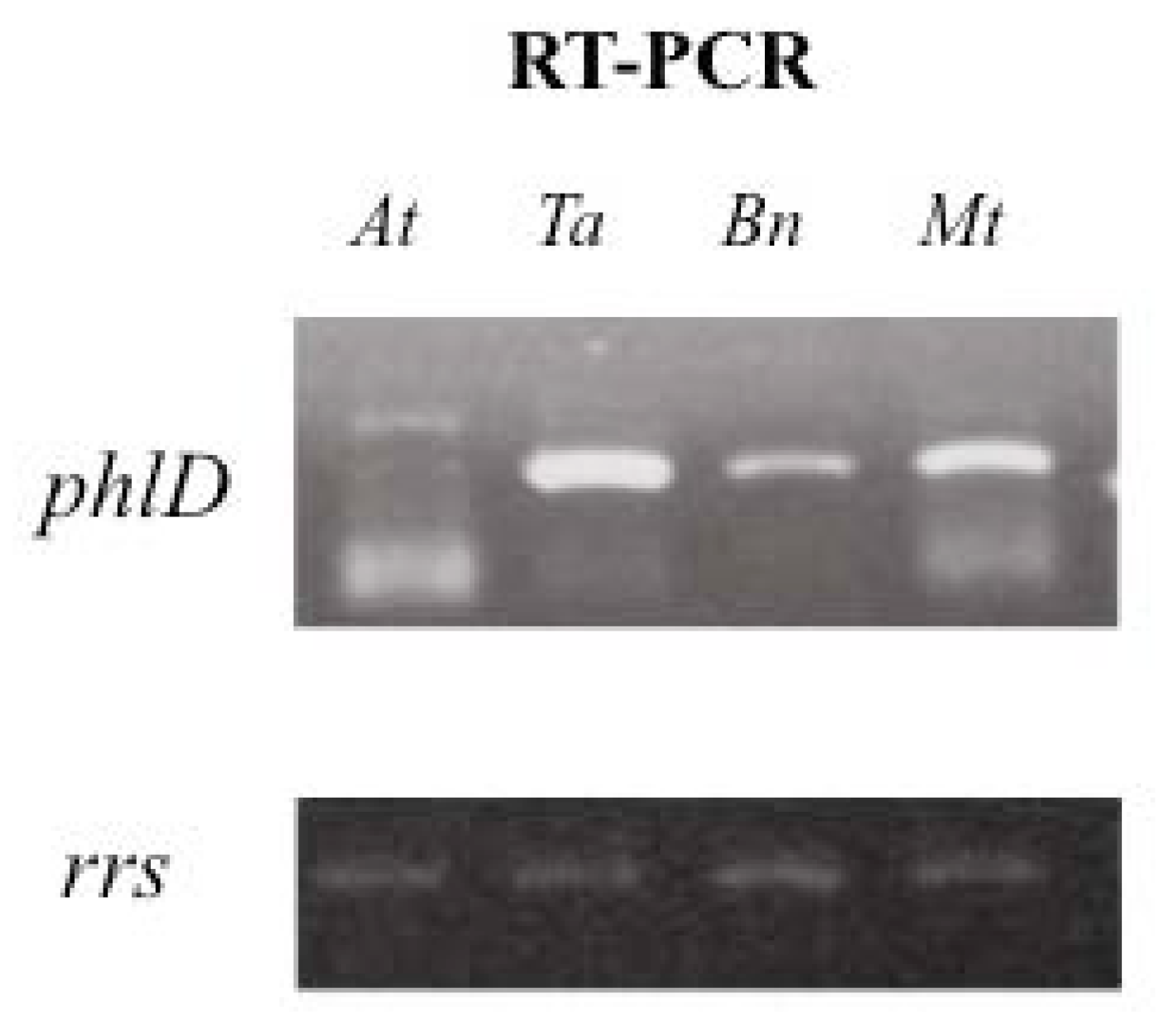
2.2. Impact of Root Exudates of Different Plant Species on phlD Gene Expression with Pseudomonas brassicacearum
| Rhizosphere soil | Root DNA | Rhizosphere soil | ||
|---|---|---|---|---|
| Light-DNA | Heavy-DNA | 12C-RNA | 13C-RNA | |
|
|
|
|
|
3. Experimental Section
3.1. Plant Growth & 13C Labeling
3.2. DNA Extraction and Gradient Fractionation
3.3. RNA Extraction and CsTFA Centrifugation
3.4. PCR and RT-PCR Amplification of phlD Gene
3.5. Denaturing Gradient Gel Electrophoresis (DGGE) Fingerprinting and the Recovery of DNA Template from DGGE Bands
3.6. Sequencing and Phylogenetic Analysis
3.7. Seeds Sterilization and Plant Growth in vitro
3.8. RT-PCR Amplification of phlD Gene
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Bais, H.P.; Weir, T.L.; Perry, L.G.; Gilroy, S.; Vivanco, J.M. The role of root exudates in rhizosphere interactions with plants and other organisms. Annu. Rev. Plant Biol. 2006, 57, 233–266. [Google Scholar] [CrossRef]
- Haichar, F.Z.; Roncato, M.A.; Achouak, W. Stable isotope probing of bacterial community structure and gene expression in the rhizosphere of Arabidopsis thaliana. FEMS Microb. Ecol. 2012, 81, 291–302. [Google Scholar] [CrossRef]
- Haichar, F.Z.; Marol, C.; Berge, O.; Rangel-Castro, J.; Prosser, J.I.; Balesdent, J.; Heulin, T.; Achouak, W. Plant host habitat and root exudates shape soil bacterial community structure. ISME J. 2008, 2, 1221–1230. [Google Scholar] [CrossRef]
- Jousset, A.; Rochat, L.; Lanoue, A.; Bonkowski, M.; Keel, C.; Scheu, S. Plants respond to pathogen infection by enhancing the antifungal gene expression of root-associated bacteria. MPMI. 2011, 24, 352–358. [Google Scholar] [CrossRef]
- Haas, D.; Défago, G. Biological control of soil-borne pathogens by Fluorescent pseudomonads. Nat. Rev. Microbiol. 2005, 3, 307–319. [Google Scholar] [CrossRef]
- Almario, J.; Moenne-Loccoz, Y.; Muller, D. Monitoring of the relation between 2,4-diacetylphloroglucinol-producing Pseudomonas and Thielaviopsis basicola populations by real-time PCR in tobacco black root-rot suppressive and conducive soils. Soil Biol. Biochem. 2012, 57, 44–155. [Google Scholar]
- Achouak, W.; Sutra, L.; Heulin, T.; Meyer, J.-M.; Fromin, N.; Degraeve, S.; Christen, R; Gardan, L. Description of Pseudomonas brassicacearum sp. nov. and Pseudomonas thivervalensis sp. nov., two root-associated bacteria isolated from Arabidopsis thaliana and Brassica napus. Int. J. Syst. Evol. Microbiol. 2000, 50, 9–18. [Google Scholar] [CrossRef]
- Persello-Cartieaux, F.; David, P.; Sarrobert, C.; Thibaud, MC.; Achouak, W.; Robaglia, C.; Nussaume, L. Utilization of mutants to analyze the interaction between Arabidopsis thaliana and its naturally root-associated Pseudomonas. Planta 2001, 212, 190–198. [Google Scholar] [CrossRef]
- Lalaouna, D.; Fochesato, S.; Sanchez, L.; Schmitt-Kopplin, P.; Haas, D.; Heulin, T.; Achouak, W. Phenotypic switching involves GacS/GacAdependent Rsm small RNAs in Pseudomonas brassicacearum. Appl. Environ. Microbiol. 2012, 78, 1658–1665. [Google Scholar] [CrossRef]
- Haas, D.; Keel, C. Regulation of antibiotic production in root-colonizing Pseudomonas spp. and relevance for biological control of plant disease. Annu. Rev. Phytopathol. 2003, 41, 117–153. [Google Scholar] [CrossRef]
- Laville, J.; Voisard, C.; Keel, C.; Mauhofer, M.; Défago, G.; Haas, D. Global control in Pseudomonas fluorescens mediating antibiotic synthesis and suppression of black root rot of tobacco. Proc. Natl. Acad. Sci. USA. 1992, 89, 1562–1566. [Google Scholar]
- de Werra, P.; Huser, A.; Tabacchi, R.; Keel, C.; Maurhofer, M. Plant- and microbe-derived compounds affect the expression of genes encoding antifungal compounds in a pseudomonad with biocontrol activity. Appl. Environ. Microbiol. 2011, 77, 2807–2812. [Google Scholar] [CrossRef]
- Picard, C.; Di Cello, F.; Ventura, M.; Fani, R.; Guckert, A. Frequency and biodiversity of 2,4-diacetylphloroglucinol producing bacteria isolated from the maize rhizosphere at different stages of plant growth. Appl. Environ. Microbiol. 2000, 66, 948–955. [Google Scholar] [CrossRef]
- Bergsma-Vlami, M.; Prins, M.E.; Raaijmakers, J.M. Influence of plant species on population dynamics, genotypic diversity and antibiotic production in the rhizosphere by indigenous Pseudomonas spp. FEMS Microbiol. Ecol. 2005, 52, 59–69. [Google Scholar] [CrossRef]
- Achouak, W.; Conrod, S.; Cohen, V.; Heulin, T. Phenotypic variation of Pseudomonas brassicacearum as a plant root-colonisation strategy. MPMI. 2004, 17, 872–879. [Google Scholar] [CrossRef]
- Keel, C.; Schnider, U.; Maurhofer, M.; Voisard, C.; Laville, J.; Burger, P.; Wirthner, P.; Haas, D.; Défago, G. Suppression of root diseases by Pseudomonas fluorescens CHA0, importance of the bacterial secondary metabolite 2,4-diacetylphloroglucinol. MPMI. 1992, 5, 4–13. [Google Scholar] [CrossRef]
- Brazelton, J.N.; Pfeufer, E.E.; Sweat, T.A.; Gardener, B.B.; Coenen, C. 2,4-Diacetylphloroglucinol alters plant root development. Mol. Plant-Microb. Interact. 2008, 21, 1349–1358. [Google Scholar] [CrossRef]
- Duffy, B.K.; Défago, G. Zinc improves biocontrol of Fusarium crown and root rot of tomato by Pseudomonas fluorescens and represses the production of pathogen metabolites inhibitory to bacterial antibiotic biosynthesis. Phytopathology 1997, 87, 1250–1257. [Google Scholar] [CrossRef]
- Schnider-Keel, U.; Seematter, A.; Maurhofer, M.; Blumer, C.; Duffy, B.; Gigot-Bonnefoy, C.; Reimmann, C.; Notz, R.; Défago, G.; Haas, D.; Keel, C. Autoinduction of 2,4-diacetylphloroglucinol biosynthesis in the biocontrol agent Pseudomonas fluorescens CHA0 and repression by the bacterial metabolites salicylate and pyoluteorin. J. Bact. 2000, 182, 1215–1225. [Google Scholar] [CrossRef]
- Chabeaud, P.; de Groot, A.; Bitter, W.; Tommassen, J.; Heulin, T.; Achouak, W. Phase-variable expression of an operon encoding extracellular alkaline protease, serine protease homologue and lipase in Pseudomonas brassicacearum. J. Bacteriol. 2001, 183, 2117–2120. [Google Scholar] [CrossRef]
- Fromin, N.; Achouak, W.; Thiery, J.M.; Heulin, T. The genotypic diversity of Pseudomonas brassicacearum populations isolated from roots of Arabidopsis thaliana, influence of plant genotype. FEMS Microbiol. Ecol. 2001, 37, 21–29. [Google Scholar] [CrossRef]
- Ross, I.; Alami, Y.; Harvey, P.R.; Achouak, W.; Ryder, M. Genetic diversity and biological control activity of novel species related to pseudomonads isolated from wheat field soils in South Australia. Appl. Environ. Microbiol. 2000, 66, 1609–1616. [Google Scholar] [CrossRef]
- Weller, D.M.; Mavrodi, D.V.; van Pelt, J.A.; Pieterse, C.M.J.; van Loon, L.C.; Bakker, P.A.H.M. Induced systemic resistance in Arabidopsis thaliana against Pseudomonas syringae pv. tomato by 2,4-diacetylphloroglucinolproducing Pseudomonas fluorescens. Phytopathology. 2012, 102, 403–412. [Google Scholar]
- Phillips, D.A.; Fox, T.C.; King, M.D.; Bhuvaneswari, T.V.; Teuber, L.R. Microbial products trigger amino acid exudation from plant roots. Plant Physiol. 2004, 136, 2887–2894. [Google Scholar] [CrossRef]
- Raaijmakers, J.M.; Weller, D.M. Natural plant protection by 2,4-diacetylphloroglucinol-producing Pseudomonas spp. in take-all decline soils. MPMI. 1998, 11, 144–152. [Google Scholar] [CrossRef]
- De Souza, J.T.; Raaijmakers, J.M. Polymorphisms within the prnD and pltC genes from pyrrolnitrin and pyoluteorin-producing Pseudomonas and Burkholderia spp. FEMS Microbiol. Ecol. 2003, 43, 21–34. [Google Scholar]
- Derrien, D.; Marol, C.; Balesdent, J. The dynamics of neutral sugars in the rhizosphere of wheat, an approach by 13C pulse-labelling and GC/C/IRMS. Plant Soil 2004, 267, 243–253. [Google Scholar] [CrossRef]
- Lueders, T.; Manefield, M.; Friedrich, M.W. Enhanced sensitivity of DNA-and rRNA-based stable isotope probing by fractionation and quantitative analysis of isopycnic centrifugation gradients. Environ. Microbiol. 2004, 6, 73–78. [Google Scholar]
- Haichar, F.Z.; Achouak, W.; Christen, R.; Heulin, T.; Marol, C.; Marais., M.F.; Mougel, C.; Ranjard, L.; Balesdent, J.; Berge, O. Identification of cellulolytic bacteria in soil by stable isotope probing. Environ. Microbiol. 2007, 9, 625–634. [Google Scholar] [CrossRef]
- Rangel-Castro, J.I.; Killham, K.; Ostle, N.; Nicol, G.W.; Anderson, I.C.; Scrimgeour, C.M.; Killham, K.; Meharg, A.A. Stable isotope probing analysis of the influence of liming on root exudates utilization by soil microorganisms. Environ. Microbiol. 2005, 7, 828–838. [Google Scholar] [CrossRef]
- McCaig, A.E.; Glover, L.A.; Prosser, J.I. Numerical analysis of grassland bacterial community structure under different land management regimes by using 16S ribosomal DNA sequence data and denaturing gradient gel electrophoresis banding patterns. Appl. Environ. Microbiol. 2001, 67, 4554–4559. [Google Scholar] [CrossRef]
- Altschul, S.F.; Gish, W.; Miller, W.; Myers, W.; Lipman, D.J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Haichar, F.E.Z.; Fochesato, S.; Achouak, W. Host Plant Specific Control of 2,4-Diacetylphloroglucinol Production in the Rhizosphere. Agronomy 2013, 3, 621-631. https://doi.org/10.3390/agronomy3040621
Haichar FEZ, Fochesato S, Achouak W. Host Plant Specific Control of 2,4-Diacetylphloroglucinol Production in the Rhizosphere. Agronomy. 2013; 3(4):621-631. https://doi.org/10.3390/agronomy3040621
Chicago/Turabian StyleHaichar, Feth El Zahar, Sylvain Fochesato, and Wafa Achouak. 2013. "Host Plant Specific Control of 2,4-Diacetylphloroglucinol Production in the Rhizosphere" Agronomy 3, no. 4: 621-631. https://doi.org/10.3390/agronomy3040621




