2.2. Fly Stocks
fatfd/CyO (Kieran Harvey, Peter MacCallum Cancer Centre, Melbourne, Australia); GMR-GAL4 (Bloomington Stock Centre, Bloomington, IN, USA); UAS-aPKCCAAXWT (Sonsoles Campuzano, Centro de Biología Molecular Severo Ochoa, Madrid, Spain); hpo5.1/CyO, dRASSFX36/TM6B (Nicholas Tapon, Cancer Research, London, UK); FRT82B, kibra4/TM6B, FRT82B, kibra1/TM6B (Hugo Stocker, Institute of Molecular Systems Biology, Zurich, Switzerland); dachsGC13, FRT40/CyO, dachs1/CyO (Kenneth Irvine, The State University of New Jersey, New Brusnwick NJ, USA); diap1-GFP1.8 (Duojia Pan, Johns Hopkins University School of Medicine, Baltimore, MD, USA); diap1-lacZ reporter, thic58 (Bruce Hay, California Institute of Technology, Pasadena, CA, USA); actin < CD2 < GAL4, UAS-GFP (Bruce Edgar, Fred Hutchinson Cancer Research Center, Seattle, WA, USA); eyFLP; UAS-RFP, tubgal4, FRT82B, tubgal80/TM6B (Virender Sahota, Peter MacCallum Cancer Centre, Melbourne, Australia) this study; merlin (mer)-RNAi14228R−1 (National Institute of Genetics (NIG), Shizuoka , Japan); mob4-RNAi110742, cka-RNAi106971, CTTNBP2-RNAi31377, CCM3-RNAi106841, lgl-RNAi51247 (Vienna Drosophila RNAi Centre (VDRC), Vienna, Austria).
2.3. Immunohistochemistry, Imaging and Antibodies
Larval and pupal discs were dissected and fixed as previously described [
7]. For Hpo, dRASSF, Cka and Mob4 staining, tissues were fixed in paraformaldehyde lysine periodate (PLP). Labeled samples were cleared through 80% glycerol and mounted. For
Figure 1,
Figure 2E,
Figure 3A,C, Appendix
Figure A1B,C, and
Figure 4A,B images were collected on Biorad MRC1000 (Bio-Rad Laboratories, Hercules, CA, USA).
Figure 2B,
Figure 3B and Appendix
Figure A1A were collected on an Olympus FV1000 (Olympus, Center Valley, PA, USA). Images in
Figure 2C,D and
Figure 4G–I were taken on a Nikon Eclipse 90i (Nikon, New York, NY, USA). Images were processed using Fiji, and assembled with Adobe Photoshop CS6 and Adobe Illustrator CS6. Adult eyes were imaged with a Scitec Infinity1 camera (Lumenera, Ottawa, Canada).
Antibodies used were mouse anti-β-galactosidase (Sigma, St. Louis, MO, USA, 1:500), rat anti-Hpo (1:100), rabbit anti-dRASSF (1:200) both from Nicholas Tapon, Cancer Research, UK, rabbit anti-Cka (Wei Du, University of Chicago, Chicago, IL, USA 1:1000) rabbit anti-Lgl (Dennis Strand, Johannes Gutenberg University, Germany. 1:1000), guinea-pig anti-Mob4 (Joost Schulte, University of British Columbia, Vancouver, Canada, 1:1000), rat anti-Shg (E-cadherin, Developmental Studies Hybridoma Bank, DSHB, University of Iowa, IA, USA. 1:50), rabbit anti-aPKC (Santa Cruz Biotechnology, Dallas, TX, USA, human PKCζ 1:500).
2.6. Statistical Analysis of Signal Intensity
To determine the relative diap1-GFP1.8 expression in eye discs, images of each disc were taken with identical confocal settings. Using Image J, a ratio of average pixel intensity was determined for diap1-GFP1.8 immunofluorescence in an area posterior and anterior to the morphogenetic furrow that colocalized with DAPI staining. Data was analyzed using unpaired t-test (Welch corrected).
Figure 1.
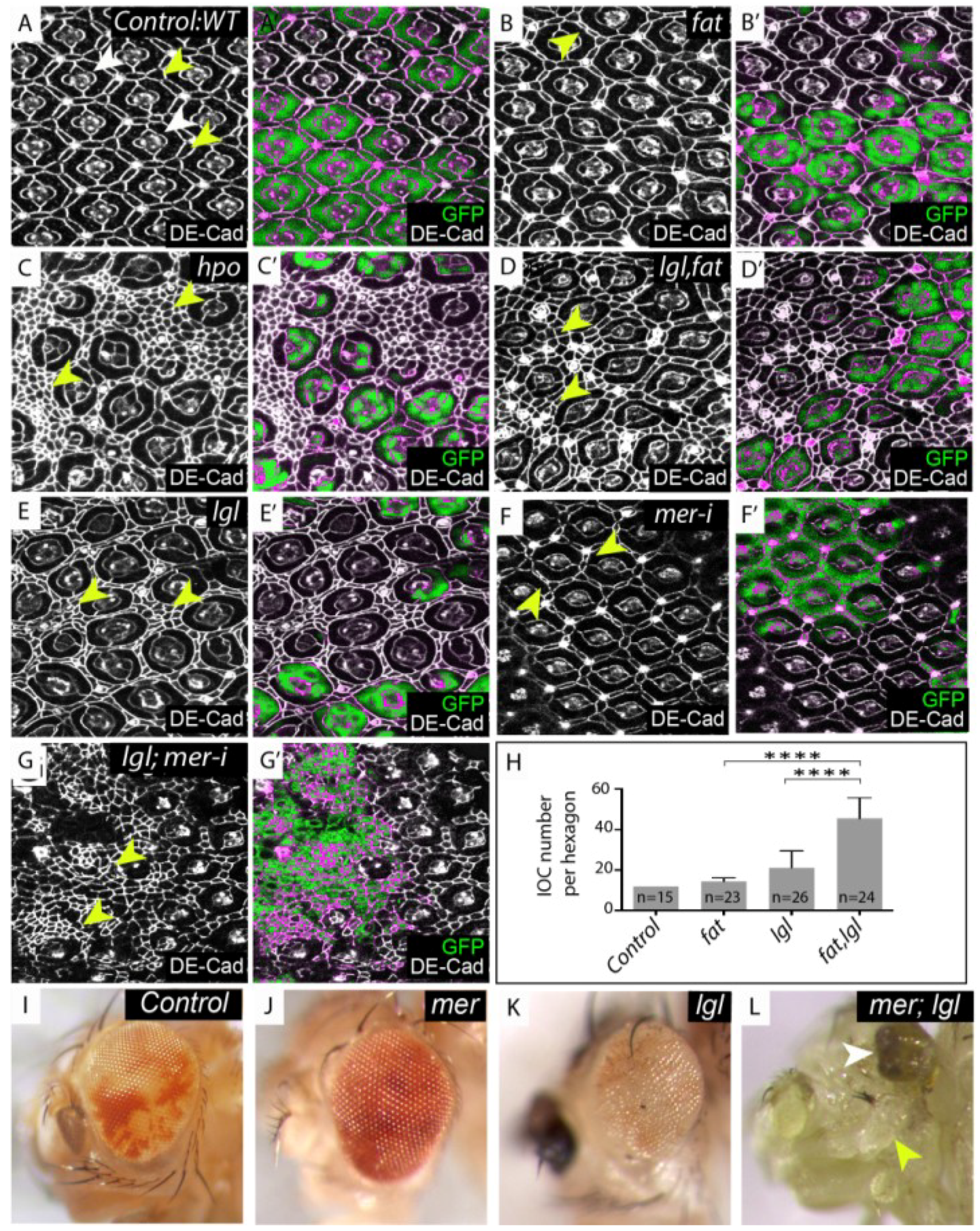
lgl regulates the Hippo (Hpo) pathway in parallel with fat and merlin. (A–G) Mosaic mid-pupal retinae stained with Drosophila E-cadherin (DE-Cad) antibodies that localize to the adherens junction and mark cell outlines. (A–E) mutant tissue lacks the expression of green fluorescent protein (GFP) and is merged with DE-Cad expression in (A'–E'). (A) Control, wildtype retina containing mosaic tissue expressing GFP and GFP negative tissue, white arrowheads denote photoreceptor and cone cells, yellow arrowheads denote IOCs; (B) Retina with fat−/− tissue or (E) lgl−/− tissue. The mutant area (GFP negative) shows a few extra IOC’s (yellow arrowheads); (C) hpo−/− clone (GFP negative); (D) lgl, fat−/− double mutant tissue (GFP negative) displays a substantial increase in IOC number (yellow arrowheads); (F) mer-RNAi depleted tissue (marked by the presence of GFP) shows a few additional IOC’s (yellow arrowheads); (G) lgl−/−; mer-RNAi tissue (marked by the presence of GFP) display excess IOC’s (yellow arrowheads); (H) Mean number of IOCs per ommatidia. **** indicates p < 0.0001. Error bars represent Standard Deviation. n equals the number of ommatidia counted; (I–L) Adult male eye images; (I) Control, mosaic eye; (J) mer−/−, mosaic eye (mer−/− tissue is pale red); (K) lgl−/− mosaic eye (lgl−/− tissue is white); (L) mer; lgl double mutant mosaic eye (note the decrease in adult retinal tissue (white arrowhead) and increase in head capsule tissue (yellow arrowhead).
Figure 1.
lgl regulates the Hippo (Hpo) pathway in parallel with fat and merlin. (A–G) Mosaic mid-pupal retinae stained with Drosophila E-cadherin (DE-Cad) antibodies that localize to the adherens junction and mark cell outlines. (A–E) mutant tissue lacks the expression of green fluorescent protein (GFP) and is merged with DE-Cad expression in (A'–E'). (A) Control, wildtype retina containing mosaic tissue expressing GFP and GFP negative tissue, white arrowheads denote photoreceptor and cone cells, yellow arrowheads denote IOCs; (B) Retina with fat−/− tissue or (E) lgl−/− tissue. The mutant area (GFP negative) shows a few extra IOC’s (yellow arrowheads); (C) hpo−/− clone (GFP negative); (D) lgl, fat−/− double mutant tissue (GFP negative) displays a substantial increase in IOC number (yellow arrowheads); (F) mer-RNAi depleted tissue (marked by the presence of GFP) shows a few additional IOC’s (yellow arrowheads); (G) lgl−/−; mer-RNAi tissue (marked by the presence of GFP) display excess IOC’s (yellow arrowheads); (H) Mean number of IOCs per ommatidia. **** indicates p < 0.0001. Error bars represent Standard Deviation. n equals the number of ommatidia counted; (I–L) Adult male eye images; (I) Control, mosaic eye; (J) mer−/−, mosaic eye (mer−/− tissue is pale red); (K) lgl−/− mosaic eye (lgl−/− tissue is white); (L) mer; lgl double mutant mosaic eye (note the decrease in adult retinal tissue (white arrowhead) and increase in head capsule tissue (yellow arrowhead).
![Cancers 06 00879 g001]()
(A) Control: eyFLP; FRT40/UbiGFP, FRT40; (B) fat: eyFLP; fatfd, FRT40/UbiGFP, FRT40; (C) hpo: eyFLP, FRT42D, hpo5.1/FRT42D, UbiGFP; (D) lgl, fat: eyFLP, lgl27S3, fatfd, FRT40/UbiGFP, FRT40; (E) lgl: eyFLP, lgl27S3, FRT40/UbiGFP, FRT40; (F) mer-i: eyFLP, UAS-GFP; FRT40/tubgal80, FRT40; UAS mer-RNAi14228R/tubgal4; (G) lgl;mer-i: eyFLP, UAS-GFP; lgl27S3, FRT40/gal80, FRT40; UAS-mer-RNAi14228R/tubgal4; (I) Control: eyFLP; FRT40/UbiGFP FRT40; (J) mer: y w Mer4 FRT19A/Y; FRT40A/P[w+ Mer+], Ubi-GFP, FRT40A; eyGAL4, UAS-flp/+; (K) lgl: eyFLP; lgl27S3, FRT40/UbiGFP, FRT40; (L) mer; lgl: y w Mer4 FRT19A/Y; P[w+ Mer+] Ubi-GFP FRT40A/lgl27S3 FRT40A; eyGAL4, UAS-flp/+.
Figure 2.
lgl regulates the Hpo pathway independently of Fat-Dachs and Kibra/Expanded/Merlin branches. (A) Western Blot showing Wts protein levels; (B, C, E) Planar sections of larval eye discs; (D) Sideview of larval eye disc. White scale bar represents 50 µM. Yellow bar denotes the morphogenetic furrow (MF), posterior is to the left in planar sections, in this and all other figures; (F–K) Adult female eye images; (L) Graphical representation of adult eye size presented in (F–K); (A) Western Blot analysis of protein extracts derived from control and Lgl depleted larval eye-antennal discs probed for expression of Warts (Wts). There is no change in Wts protein levels in lgl-depleted tissue; (B) dachs1: lgl−/−, dachsGC13/GC13, diap1-lacZ mosaic discs stained for β-gal (red) show increased diap1-lacZ staining in lgl−/−, dachsGC13/GC13 clones (arrowheads). lgl−/−, dachsGC13/GC13 tissue identified by lack of Lgl antibody staining (green); (C,C',D,D') Hpo (white) and (E,E') dRASSF (white) staining in kibra, mer-RNAi mosaic eye discs. In kibra, mer-RNAi tissue (RFP positive, green, mutant tissue indicated by arrowheads) Hpo and dRASSF concentration and localization are normal compared to wildtype tissue (RFP negative); (C"–E") Overlay of RFP and antibody staining, pink highlights mutant RFP positive tissue; (C'''–E''') Green dots outline RFP clone boundaries; (F–K) Reducing the level of kibra increases the size of GMR > aPKCCA or lgl−/− mosaic adult eyes; (F) Control adult eye; (G) GMR > aPKCCA adult eye; (H) kibra−/− tissue is white; (I) kibra−/− tissue that expresses aPKCCA shows synergistic increase in adult eye size; (J) lgl−/− mosaic adult eye (lgl−/− tissue is white); (K) lgl−/−; kibra−/− mosaic adult eye (double mutant tissue is white, arrowhead denotes head cuticle tissue); (L) Quantification of adult eye size (F–K).
Figure 2.
lgl regulates the Hpo pathway independently of Fat-Dachs and Kibra/Expanded/Merlin branches. (A) Western Blot showing Wts protein levels; (B, C, E) Planar sections of larval eye discs; (D) Sideview of larval eye disc. White scale bar represents 50 µM. Yellow bar denotes the morphogenetic furrow (MF), posterior is to the left in planar sections, in this and all other figures; (F–K) Adult female eye images; (L) Graphical representation of adult eye size presented in (F–K); (A) Western Blot analysis of protein extracts derived from control and Lgl depleted larval eye-antennal discs probed for expression of Warts (Wts). There is no change in Wts protein levels in lgl-depleted tissue; (B) dachs1: lgl−/−, dachsGC13/GC13, diap1-lacZ mosaic discs stained for β-gal (red) show increased diap1-lacZ staining in lgl−/−, dachsGC13/GC13 clones (arrowheads). lgl−/−, dachsGC13/GC13 tissue identified by lack of Lgl antibody staining (green); (C,C',D,D') Hpo (white) and (E,E') dRASSF (white) staining in kibra, mer-RNAi mosaic eye discs. In kibra, mer-RNAi tissue (RFP positive, green, mutant tissue indicated by arrowheads) Hpo and dRASSF concentration and localization are normal compared to wildtype tissue (RFP negative); (C"–E") Overlay of RFP and antibody staining, pink highlights mutant RFP positive tissue; (C'''–E''') Green dots outline RFP clone boundaries; (F–K) Reducing the level of kibra increases the size of GMR > aPKCCA or lgl−/− mosaic adult eyes; (F) Control adult eye; (G) GMR > aPKCCA adult eye; (H) kibra−/− tissue is white; (I) kibra−/− tissue that expresses aPKCCA shows synergistic increase in adult eye size; (J) lgl−/− mosaic adult eye (lgl−/− tissue is white); (K) lgl−/−; kibra−/− mosaic adult eye (double mutant tissue is white, arrowhead denotes head cuticle tissue); (L) Quantification of adult eye size (F–K).
![Cancers 06 00879 g002]()
(A) Control: eyFLP; actin < CD2 < GAL4; UAS-GFP; lgli: eyFLP; actin < CD2 < GAL4; UAS-GFP/UAS-lgl-RNAi51247; (B) dachs1: lgl, dachsGC13: eyFLP; lgl27S3, dachsGC13, FRT40/dachs1, FRT40; diap1-lacZ reporter (thic58)/+; (C–E) kibra, mer-i: eyFLP; FRT82B, kibra4, UAS-merlin RNAi14228R−1/UAS-mRFP, tubgal4, FRT82B, tubgal80; (F) Control: GMR-GAL4/+; (G) GMR > aPKCCA: GMR-GAL4 > UAS-aPKCCAAXWT/+; (H) kibra: eyFLP; FRT82B, kibra1/FRT82B, P{w+} cell lethal; (I) GMR > aPKCCA; kibra: eyFLP; GMR-GAL4 > UAS-aPKCCAAXWT/+; FRT82B, kibra1/FRT82B, P{w+} cell lethal; (J) lgl: eyFLP; lgl27S3, FRT40/Ubi-GFP, FRT40; (K) lgl, kibra: eyFLP; lgl27S3, FRT40/Ubi-GFP, FRT40; FRT82B, kibra1/FRT82B, P{w+} cell lethal; (L) Mean eye size represented by the area * indicates p < 0.05; **** indicates p < 0.0001. Error bars represent Standard Deviation. n = 12 eyes per genotype.
Figure 3.
Members of the dSTRIPAK complex are mislocalized in lgl−/− tissue and genetically interact with GMR>aPKCCA. (A, C, H, I) Planar sections of larval eye discs; (B) Cross section of larval eye disc. Yellow line indicates morphogenetic furrow. White scale bar represents 50 µM; (A, A', A''') lgl−/− mosaic eye disc stained for Cka (white) and Hpo (white) respectively. Mutant clones display apical accumulation of Hpo and Cka (arrowheads); (B, B') lgl−/− mosaic eye disc, stained for Cka (white) displays apical accumulation and basolateral mislocalization of Cka (mutant tissue GFP negative, white arrowheads indicate Cka mislocalization); (B''') Green dots outline GFP clone boundaries; (C, C', C''') lgl−/− mosaic eye disc, stained for Mob4 (white) and Hpo (white) respectively. Only moderate accumulation of Mob4 was observed compared to Hpo (mutant tissue GFP negative, arrowheads indicate Mob4 mislocalization); (A"–C") Overlay of GFP and antibody staining, pink highlights wildtype GFP positive tissue; (D–G) Adult female eye images of GMR > aPKCCA and depletion of dSTRIPAK components; (D) CTTNBP2; (E) CCM3i; (F) ckai; (G) mob4. Reducing the levels of mob4, but not other dSTRIPAK components, in conjunction with GMR > aPKCCA alters tissue growth.
Figure 3.
Members of the dSTRIPAK complex are mislocalized in lgl−/− tissue and genetically interact with GMR>aPKCCA. (A, C, H, I) Planar sections of larval eye discs; (B) Cross section of larval eye disc. Yellow line indicates morphogenetic furrow. White scale bar represents 50 µM; (A, A', A''') lgl−/− mosaic eye disc stained for Cka (white) and Hpo (white) respectively. Mutant clones display apical accumulation of Hpo and Cka (arrowheads); (B, B') lgl−/− mosaic eye disc, stained for Cka (white) displays apical accumulation and basolateral mislocalization of Cka (mutant tissue GFP negative, white arrowheads indicate Cka mislocalization); (B''') Green dots outline GFP clone boundaries; (C, C', C''') lgl−/− mosaic eye disc, stained for Mob4 (white) and Hpo (white) respectively. Only moderate accumulation of Mob4 was observed compared to Hpo (mutant tissue GFP negative, arrowheads indicate Mob4 mislocalization); (A"–C") Overlay of GFP and antibody staining, pink highlights wildtype GFP positive tissue; (D–G) Adult female eye images of GMR > aPKCCA and depletion of dSTRIPAK components; (D) CTTNBP2; (E) CCM3i; (F) ckai; (G) mob4. Reducing the levels of mob4, but not other dSTRIPAK components, in conjunction with GMR > aPKCCA alters tissue growth.
![Cancers 06 00879 g003]()
(A–C) lgl: eyFLP; lgl27S3, FRT40/UbiGFP, FRT40; (D) GMR > aPKCCA/CTTNBP2: GMR-GAL4 > UAS-aPKCCAAXWT/UAS-CTTNBP2-RNAi31377; (E) GMR > aPKCCA/CCM3i: GMR-GAL4 > UAS-aPKCCAAXWT/UAS-CCM3-RNAi106841; (F) GMR > aPKCCA/ckai: GMR-GAL4 > UAS-aPKCCAAXWT/UAS-cka-RNAi106971; (G) GMR > aPKCCA/mob4i: GMR-GAL4>UAS-aPKCCAAXWT/UAS-mob4-RNAi11074.
Figure 4.
Hpo mislocalization in lgl−/− mosiac tissue is independent of dRASSF and dRASSF acts as a tumor suppressor. (A, G–I) Planar sections of larval eye discs; (B) Cross section of larval eye disc. Yellow line indicates morphogenetic furrow. White scale bar represents 50 µM. (C–E) Adult female eye images. (A) lgl; dRASSF double mutant tissue (GFP negative) displays apical accumulation of Hpo (white, arrowhead indicates double mutant, GFP negative tissue) compared to dRASSF mutant tissue (GFP positive) and is mislocalized basolaterally in lgl; dRASSF double mutant tissue (B arrowhead). aPKC staining (white) is altered where the tissue shows obvious folds (A, white asterik) but is still localized apically in lgl; dRASSF double mutant tissue (B); (C, C', D, D', E, E') Adult female eye images, side and ventral views respectively; (C,C') dRASSF; (D, D') lgl; dRASSF; (E, E') GMR > aPKCCA, dRASSF. The eye size of lgl mosaic and GMR > aPKCCA overexpressing adult eyes is increased when dRASSF is removed; (F) Western blot of protein extracts from eye discs detecting pERK, total ERK and tubulin levels; (F') Graphical representation of signal intensity from Western blots. There is a two fold increase in pERK/ERK levels in GMR > aPKCCA mutant tissue. However, removal of dRASSF does not alter the relative ratio of pERK/ERK in either a wildtype or GMR > aPKCCA tissue; (G–H) dRASSF, GMR > aPKCCA, GMR > aPKCCA; dRASSF crossed with diap1-GFP1.8. GFP expression (green) reports diap1 expression. Compared with the control; (G) dRASSF; (H) GMR > aPKCCA; and (I) GMR > aPKCCA; dRASSF discs show elevated levels of diap1-GFP1.8 expression but there is no increase in diap1-GFP1.8 expression between GMR > aPKCCA and GMR > aPKCCA; dRASSF tissue; (J) Graph of the ratio of intensity from posterior versus anterior regions of diap1-GFP1.8 expressing samples, taken from apical section. The upregulation of diap1-GFP1.8 in GMR > aPKCCA and GMR > aPKCCA; dRASSF tissue is significant compared to dRASSF alone but there is no significant difference between GMR > aPKCCA and GMR > aPKCCA; dRASSF tissues. n = 5 discs per sample. Error bars represent Standard Error of the Mean. ** indicates p < 0.001.
Figure 4.
Hpo mislocalization in lgl−/− mosiac tissue is independent of dRASSF and dRASSF acts as a tumor suppressor. (A, G–I) Planar sections of larval eye discs; (B) Cross section of larval eye disc. Yellow line indicates morphogenetic furrow. White scale bar represents 50 µM. (C–E) Adult female eye images. (A) lgl; dRASSF double mutant tissue (GFP negative) displays apical accumulation of Hpo (white, arrowhead indicates double mutant, GFP negative tissue) compared to dRASSF mutant tissue (GFP positive) and is mislocalized basolaterally in lgl; dRASSF double mutant tissue (B arrowhead). aPKC staining (white) is altered where the tissue shows obvious folds (A, white asterik) but is still localized apically in lgl; dRASSF double mutant tissue (B); (C, C', D, D', E, E') Adult female eye images, side and ventral views respectively; (C,C') dRASSF; (D, D') lgl; dRASSF; (E, E') GMR > aPKCCA, dRASSF. The eye size of lgl mosaic and GMR > aPKCCA overexpressing adult eyes is increased when dRASSF is removed; (F) Western blot of protein extracts from eye discs detecting pERK, total ERK and tubulin levels; (F') Graphical representation of signal intensity from Western blots. There is a two fold increase in pERK/ERK levels in GMR > aPKCCA mutant tissue. However, removal of dRASSF does not alter the relative ratio of pERK/ERK in either a wildtype or GMR > aPKCCA tissue; (G–H) dRASSF, GMR > aPKCCA, GMR > aPKCCA; dRASSF crossed with diap1-GFP1.8. GFP expression (green) reports diap1 expression. Compared with the control; (G) dRASSF; (H) GMR > aPKCCA; and (I) GMR > aPKCCA; dRASSF discs show elevated levels of diap1-GFP1.8 expression but there is no increase in diap1-GFP1.8 expression between GMR > aPKCCA and GMR > aPKCCA; dRASSF tissue; (J) Graph of the ratio of intensity from posterior versus anterior regions of diap1-GFP1.8 expressing samples, taken from apical section. The upregulation of diap1-GFP1.8 in GMR > aPKCCA and GMR > aPKCCA; dRASSF tissue is significant compared to dRASSF alone but there is no significant difference between GMR > aPKCCA and GMR > aPKCCA; dRASSF tissues. n = 5 discs per sample. Error bars represent Standard Error of the Mean. ** indicates p < 0.001.
![Cancers 06 00879 g004]()
(A,B) lgl; dRASSF: eyFLP; lgl27S3, FRT40/UbiGFP, FRT40; dRASSFX36/X36; (C) Control dRASSF: dRASSFX36/X36; (D) lgl; dRASSF: eyFLP; lgl27S3, FRT40/UbiGFP, FRT40; dRASSFX36/X36; (E) GMR>aPKCCA; dRASSF: GMR-GAL4 > UAS-aPKCCAAXWT; dRASSFX36/X36; (F) Control: w1118, dRASSF: dRASSFX36/X36, GMR>aPKCCA: GMR-GAL4 > UAS-aPKCCAAXWT GMR>aPKCCA; dRASSF, GMR-GAL4 > UAS-aPKCCAAXWT; dRASSFX36/X36; (G) Control dRASSF: GMR-GAL4 > diap1-GFP1.8/+; dRASSF X36/X36; (H) GMR > aPKCCA: GMR-GAL4 > diap1-GFP1.8/UAS-aPKCCAAXWT; (I) GMR>aPKCCA; dRASSF: GMR-GAL4 > diap1-GFP1.8/UAS-aPKCCAAXWT; dRASSF X36/X36.
2.7. Sample Preparation and Western Blot Analysis
Eye-antennal discs were dissected from
w1118,
dRASSFX36/X36,
GMR > aPKCCAAXWT,
GMR > aPKCCAAXWT;
dRASSFX36/X36 or
ey-FLP and
eyFLP;
lgli actin-FLP-out larvae, homogenized in 0.1 M Tris-HCl pH 6.8, 2% SDS. 5 mM EDTA, 5 mM DTT buffer containing Complete protease inhibitor cocktail (Roche, Basel, Switzerland), 1 mM Na
3(VO)
4, 5 mM NaF
2. In
Figure 2A, protein concentration was determined by Lowry assays. Samples containing 20 mg of protein were electrophoresed on a 10% SDS-PAGE gel and transferred to Immobilon-FL polyvinylidene difluoride membranes. In
Figure 4F, 11 eye disc pairs were dissected per sample and electrophoresed as above. Antibodies used in
Figure 2A were rabbit anti-Lgl (Dennis Strand, Johannes Gutenberg University, Germany. 1:1000), rabbit anti-Wts (Kenneth Irvine, The State University of New Jersey, NJ, USA. 1:500) and mouse anti-actin (Developmental Studies Hybridoma Bank, DSHB, University of Iowa, IA, USA. 1:100) and in
Figure 4F were mouse anti-pERK (Sigma, St. Louis, MO, USA., 1:10,000), anti rabbit-ERK 137 F5 (Cell Signalling, Danvers, MA, USA 1:1000) and mouse anti-α-tubulin (Calbiochem, MERCK, Darmstadt, Germany 1:10,000). The signal intensity of bands in
Figure 2A and
Figure 4F was measured in Fiji and relative pERK/ERK levels were normalized to α-tubulin.