Mouse Lymphoblastic Leukemias Induced by Aberrant Prdm14 Expression Demonstrate Widespread Copy Number Alterations Also Found in Human ALL
Abstract
:1. Introduction
2. Results and Discussion
| Sample # | Tumorlocation | IgHrearrangement | TCRrearrangement | Histological abnormalities | BCR/TCR-based tumor type |
|---|---|---|---|---|---|
| 91 (control) | Spleen | N/A | N/A | None | N/A |
| 92 (control) | Spleen | N/A | N/A | None | N/A |
| 93 (control) | Spleen | N/A | N/A | None | N/A |
| MPr14-111 | Lymph node | No | No | LL | Common Lymphoid Progenitor (CLP)-like |
| MPr14-143 | Spleen | Yes | No | LL with erythroblastosis | Precursor-B |
| MPr14-147 | Spleen | Yes | Yes | LL | Mixed |
| MPr14-148 | Lymph node | No | Yes | LL with erythroblastosis | Precursor-T |
| MPr14-185 | Lymph node | Yes | No | LL | Precursor-B |
| MPr14-189 | Lymph node | Yes | No | LL | Precursor-B |
| MPr14-196 | Spleen | Unknown | Unknown | LL with erythroblastosis | Unclassified |
| MPr14-197 | Spleen | Yes | Yes | LL | Mixed |
| MPr14-217 | Lymph node | Yes | No | LL | Precursor-B |
| MPr14-218 | Lymph node | Yes | No | LL-pleiomorphic | Precursor-B |
| MPr14-228 | Spleen | Yes | Yes | LL with increased megakaryocytes | Mixed |
| MPr14-258 | Spleen | No | Yes | LL | Precursor-T |
2.1. PRDM14-Induced Tumors Have Numerous, Recurrent Copy Number Alterations with Occasional Aneuploidy

2.2. Deleted Genes Confirm Role of B-Cell Developmental Disruption in PRDM14-Induced Lymphomagenesis
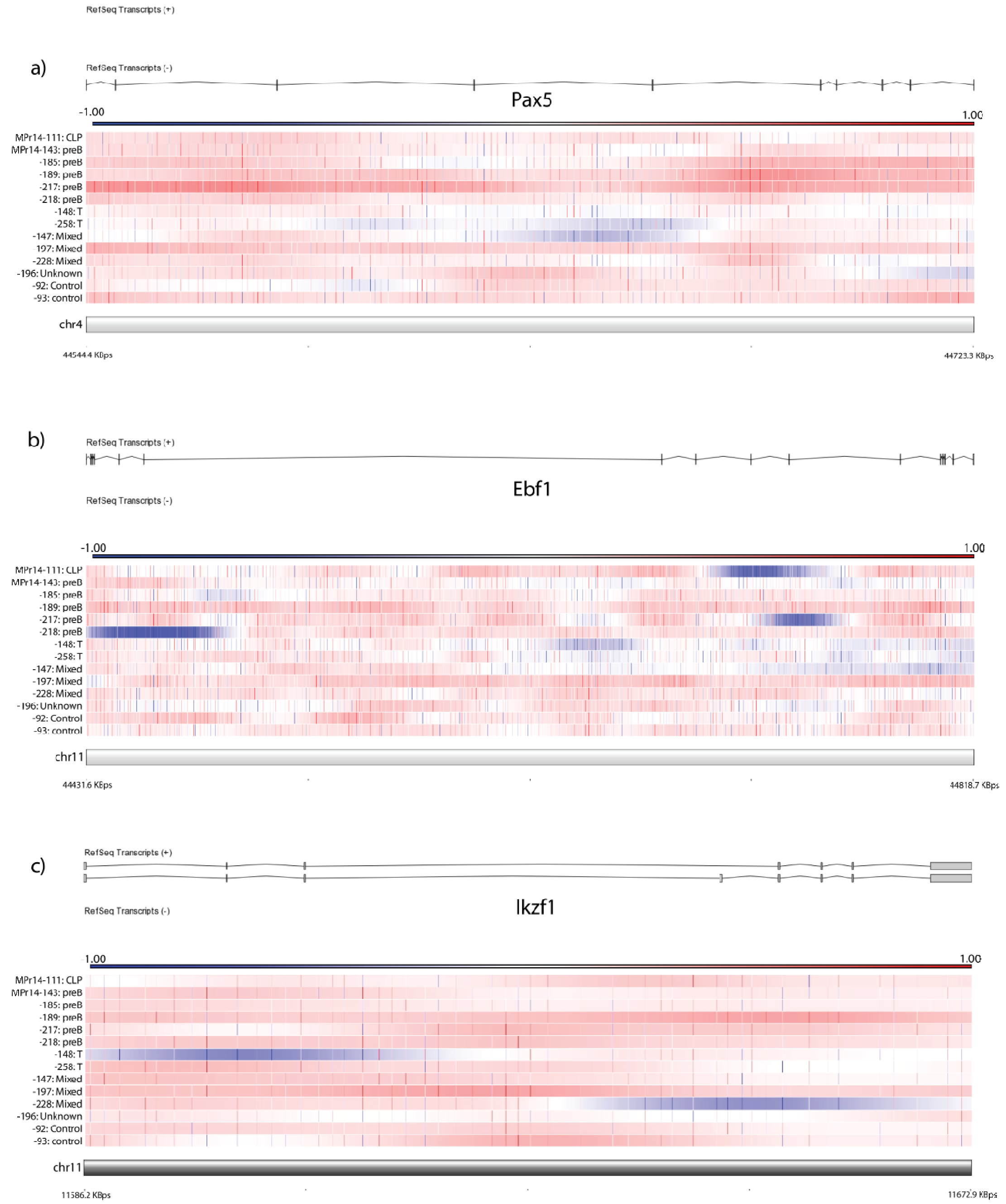
2.3. Deletions and Amplifications Occur in Characterized Tumor Suppressors and Oncogenes
| Tumor # | Cytogenetics | BCR/TCR-based tumor type | Oncogene amplifications | Tumor suppressor deletions | Fragile site gene CNA | MLL rearrangement partners amplified |
|---|---|---|---|---|---|---|
| MPr14-111 | 40,XX | CLP-like | Ebf1, Cdkn2a | Lpp | ||
| MPr14-143 | 40,XX | Precursor-B | Eg5 | |||
| MPr14-185 | 43,XY,+14,+15,+17 | Precursor-B | Eg5, Tbx2 | Cdkn2a | Mllt10 | |
| MPr14-189 | 40,XY,del(18qD1→qter), del(19qter→qB) | Precursor-B | Tbx2 | Mllt10, Ell, Mllt3 | ||
| MPr14-217 | 40,XY | Precursor-B | Notch1, Tcf3, Tbx2 | Ebf1 | Grid2 | |
| MPr14-218 | 41,XY,+1 | Precursor-B | Ebf1 | Wwox | Ell | |
| MPr14-148 | 40,XX | Precursor-T | Eg5 | |||
| MPr14-258 | 40,XX | Precursor-T | Eg5 | Ell, Eps15, Lpp | ||
| MPr14-147 | 42,XX,+14,+15 | Mixed | Eg5 | Pax5 | Grid2 | Mllt10 |
| MPr14-197 | 40,XY | Mixed | Mycl1, Eg5, Tbx2 | Cdkn2a, Fhit | Fhit | Mllt10, Lpp |
| MPr14-228 | 41,XY,+2 | Mixed | Lmo1 | Ikzf1 | Wwox | Mllt10, Ell, Lpp |
| MPr14-196 | 40,XY | Unclassified | Lmo1, Eg5 | Cdkn2a |

2.4. T-lineage Lymphoblastic Leukemia/Lymphoma Demonstrate Fewer Candidate Driver Mutations than Precursor B or Mixed Lineage Leukemias
2.5. Gene Amplifications and Deletions Occur at PRDM14 Binding Loci
2.6. Extensive Amplification of Known Cancer Rearrangement Partners
2.7. Genes Involved in Maintaining Chromosomal Stability Have Decreased Expression Prior to Leukemia Formation, and DNA Damage Response Fails to Activate
2.8. Discussion
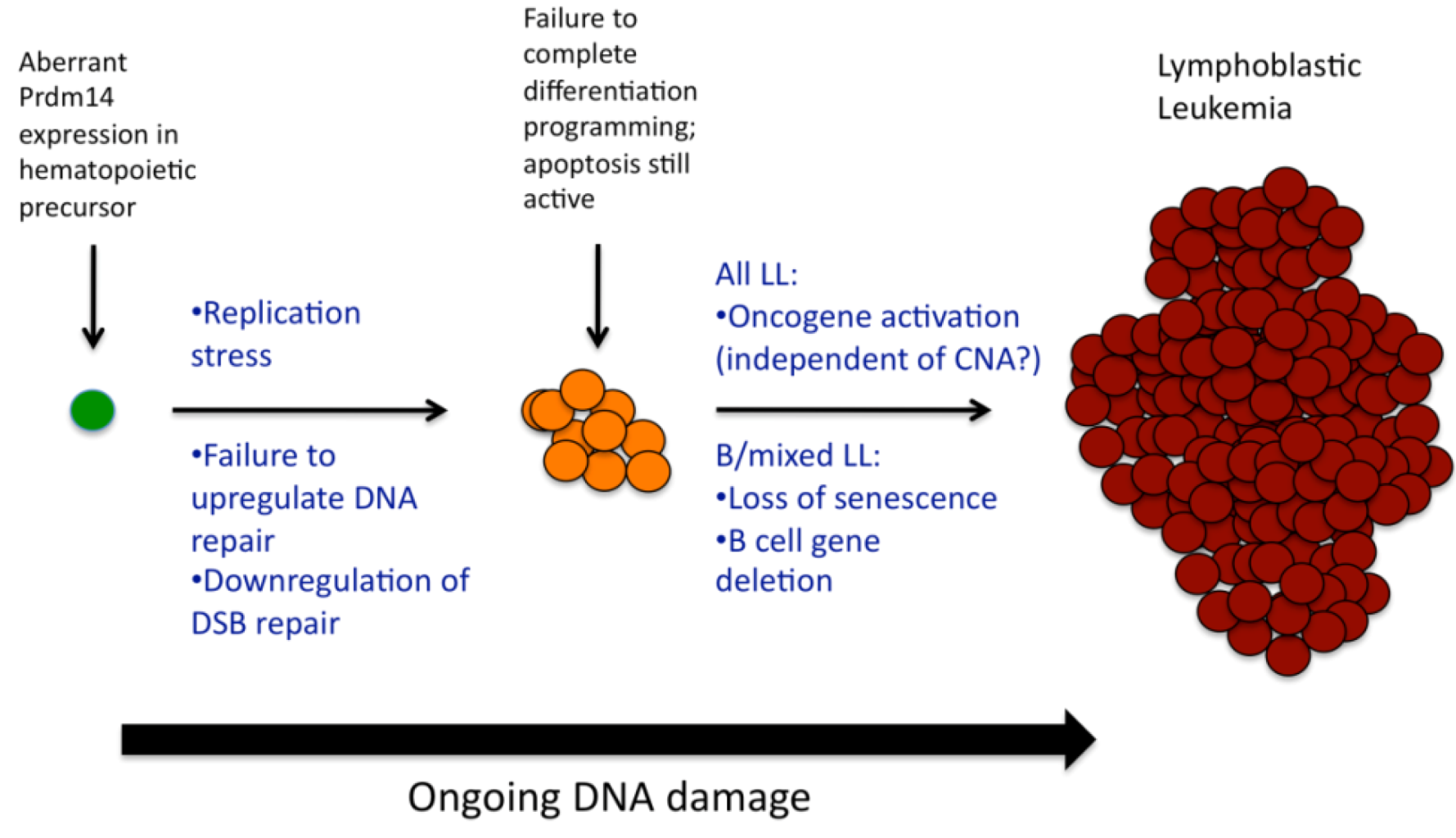
3. Experimental Section
3.1. Animal Care
3.2. Transduction of Prdm14 into Stem-Cell Enriched Bone Marrow Cells
3.3. Array CGH
3.4. Gene Expression Array
3.5. Gene Expression Studies
4. Conclusions
Acknowledgments
Supplementary Materials
References
- Mullighan, C.G.; Goorha, S.; Radtke, I.; Miller, C.B.; Coustan-Smith, E.; Dalton, J.D.; Girtman, K.; Mathew, S.; Ma, J.; Pounds, S.B.; et al. Genome-wide analysis of genetic alterations in acute lymphoblastic leukaemia. Nature 2007, 446, 758–764. [Google Scholar] [CrossRef]
- Mullighan, C.G.; Phillips, L.A.; Su, X.; Ma, J.; Miller, C.B.; Shurtleff, S.A.; Downing, J.R. Genomic analysis of the clonal origins of relapsed acute lymphoblastic leukemia. Science 2008, 322, 1377–1380. [Google Scholar]
- Mullighan, C.G.; Su, X.; Zhang, J.; Radtke, I.; Phillips, L.A.; Miller, C.B.; Ma, J.; Liu, W.; Cheng, C.; Schulman, B.A.; et al. Deletion of IKZF1 and prognosis in acute lymphoblastic leukemia. N. Engl. J. Med. 2009, 360, 470–480. [Google Scholar] [CrossRef]
- Schatz, D.G.; Ji, Y. Recombination centres and the orchestration of V(D)J recombination. Nat. Rev. Immunol. 2011, 11, 251–263. [Google Scholar] [CrossRef]
- Novara, F.; Beri, S.; Bernardo, M.E.; Bellazzi, R.; Malovini, A.; Ciccone, R.; Cometa, A.M.; Locatelli, F.; Giorda, R.; Zuffardi, O. Different molecular mechanisms causing 9p21 deletions in acute lymphoblastic leukemia of childhood. Hum. Genet. 2009, 126, 511–520. [Google Scholar] [CrossRef]
- Hauer, J.; Mullighan, C.; Morillon, E.; Wang, G.; Bruneau, J.; Brousse, N.; Lelorc’h, M.; Romana, S.; Boudil, A.; Tiedau, D.; et al. Loss of p19Arf in a Rag1−/− B-cell precursor population initiates acute B-lymphoblastic leukemia. Blood 2011, 118, 544–553. [Google Scholar] [CrossRef]
- Heltemes-Harris, L.M.; Willette, M.J.; Ramsey, L.B.; Qiu, Y.H.; Neeley, E.S.; Zhang, N.; Thomas, D.A.; Koeuth, T.; Baechler, E.C.; Kornblau, S.M.; et al. Ebf1 or Pax5 haploinsufficiency synergizes with STAT5 activation to initiate acute lymphoblastic leukemia. J. Exp. Med. 2011, 208, 1135–1149. [Google Scholar] [CrossRef]
- Maser, R.S.; Choudhury, B.; Campbell, P.J.; Feng, B.; Wong, K.K.; Protopopov, A.; O’Neil, J.; Gutierrez, A.; Ivanova, E.; Perna, I.; et al. Chromosomally unstable mouse tumours have genomic alterations similar to diverse human cancers. Nature 2007, 447, 966–971. [Google Scholar] [CrossRef]
- Sander, S.; Bullinger, L.; Karlsson, A.; Giuriato, S.; Hernandez-Boussard, T.; Felsher, D.W.; Pollack, J.R. Comparative genomic hybridization on mouse cDNA microarrays and its application to a murine lymphoma model. Oncogene 2005, 24, 6101–6107. [Google Scholar] [CrossRef]
- Weiser, K.C.; Liu, B.; Hansen, G.M.; Skapura, D.; Hentges, K.E.; Yarlagadda, S.; Morse Iii, H.C.; Justice, M.J. Retroviral insertions in the VISION database identify molecular pathways in mouse lymphoid leukemia and lymphoma. Mamm. Genome 2007, 18, 709–722. [Google Scholar] [CrossRef]
- Dettman, E.J.; Justice, M.J. The zinc finger SET domain gene Prdm14 is overexpressed in lymphoblastic lymphomas with retroviral insertions at Evi32. PLoS One 2008, 3, e3823. [Google Scholar]
- Dettman, E.J.; Simko, S.J.; Ayanga, B.; Carofino, B.L.; Margolin, J.F.; Morse, H.C., 3rd; Justice, M.J. Prdm14 initiates lymphoblastic leukemia after expanding a population of cells resembling common lymphoid progenitors. Oncogene 2011, 30, 2859–2873. [Google Scholar] [CrossRef]
- Nishikawa, N.; Toyota, M.; Suzuki, H.; Honma, T.; Fujikane, T.; Ohmura, T.; Nishidate, T.; Ohe-Toyota, M.; Maruyama, R.; Sonoda, T.; et al. Gene amplification and overexpression of PRDM14 in breast cancers. Cancer Res. 2007, 67, 9649–9657. [Google Scholar]
- Yamaji, M.; Seki, Y.; Kurimoto, K.; Yabuta, Y.; Yuasa, M.; Shigeta, M.; Yamanaka, K.; Ohinata, Y.; Saitou, M. Critical function of Prdm14 for the establishment of the germ cell lineage in mice. Nat. Genet. 2008, 40, 1016–1022. [Google Scholar]
- Gillich, A.; Bao, S.; Grabole, N.; Hayashi, K.; Trotter, M.W.; Pasque, V.; Magnusdottir, E.; Surani, M.A. Epiblast stem cell-based system reveals reprogramming synergy of germline factors. Cell Stem Cell 2012, 10, 425–439. [Google Scholar] [CrossRef]
- Ma, Z.; Swigut, T.; Valouev, A.; Rada-Iglesias, A.; Wysocka, J. Sequence-specific regulator Prdm14 safeguards mouse ESCs from entering extraembryonic endoderm fates. Nat. Struct. Mol. Biol. 2010, 18, 120–127. [Google Scholar]
- Chia, N.Y.; Chan, Y.S.; Feng, B.; Lu, X.; Orlov, Y.L.; Moreau, D.; Kumar, P.; Yang, L.; Jiang, J.; Lau, M.S.; et al. A genome-wide RNAi screen reveals determinants of human embryonic stem cell identity. Nature 2010, 468, 316–320. [Google Scholar]
- Boyer, L.A.; Lee, T.I.; Cole, M.F.; Johnstone, S.E.; Levine, S.S.; Zucker, J.P.; Guenther, M.G.; Kumar, R.M.; Murray, H.L.; Jenner, R.G.; et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 2005, 122, 947–956. [Google Scholar] [CrossRef]
- Greber, B.; Lehrach, H.; Adjaye, J. Silencing of core transcription factors in human EC cells highlights the importance of autocrine FGF signaling for self-renewal. BMC Dev. Biol. 2007, 7, 46. [Google Scholar] [CrossRef]
- Mayshar, Y.; Ben-David, U.; Lavon, N.; Biancotti, J.C.; Yakir, B.; Clark, A.T.; Plath, K.; Lowry, W.E.; Benvenisty, N. Identification and classification of chromosomal aberrations in human induced pluripotent stem cells. Cell Stem Cell 2010, 7, 521–531. [Google Scholar] [CrossRef]
- Pasi, C.E.; Dereli-Oz, A.; Negrini, S.; Friedli, M.; Fragola, G.; Lombardo, A.; Van Houwe, G.; Naldini, L.; Casola, S.; Testa, G.; et al. Genomic instability in induced stem cells. Cell Death Differ. 2011, 18, 745–753. [Google Scholar] [CrossRef]
- Momcilovic, O.; Knobloch, L.; Fornsaglio, J.; Varum, S.; Easley, C.; Schatten, G. DNA damage responses in human induced pluripotent stem cells and embryonic stem cells. PLoS One 2010, 5, e13410. [Google Scholar]
- Castillo, A.; Morse, H.C., 3rd; Godfrey, V.L.; Naeem, R.; Justice, M.J. Overexpression of Eg5 causes genomic instability and tumor formation in mice. Cancer Res. 2007, 67, 10138–10147. [Google Scholar]
- Ozeri-Galai, E.; Bester, A.C.; Kerem, B. The complex basis underlying common fragile site instability in cancer. Trends Genet. 2012, 28, 295–302. [Google Scholar] [CrossRef]
- Berquam-Vrieze, K.E.; Nannapaneni, K.; Brett, B.T.; Holmfeldt, L.; Ma, J.; Zagorodna, O.; Jenkins, N.A.; Copeland, N.G.; Meyerholz, D.K.; Knudson, C.M.; et al. Cell of origin strongly influences genetic selection in a mouse model of T-ALL. Blood 2011, 118, 4646–4656. [Google Scholar] [CrossRef]
- Tremblay, M.; Tremblay, C.S.; Herblot, S.; Aplan, P.D.; Hebert, J.; Perreault, C.; Hoang, T. Modeling T-cell acute lymphoblastic leukemia induced by the SCL and LMO1 oncogenes. Genes Dev. 2010, 24, 1093–1105. [Google Scholar] [CrossRef]
- Futreal, P.A.; Coin, L.; Marshall, M.; Down, T.; Hubbard, T.; Wooster, R.; Rahman, N.; Stratton, M.R. A census of human cancer genes. Nat. Rev. Cancer 2004, 4, 177–183. [Google Scholar] [CrossRef]
- Meyer, C.; Kowarz, E.; Hofmann, J.; Renneville, A.; Zuna, J.; Trka, J.; Ben Abdelali, R.; Macintyre, E.; de Braekeleer, E.; de Braekeleer, M.; et al. New insights to the MLL recombinome of acute leukemias. Leukemia 2009, 23, 1490–1499. [Google Scholar] [CrossRef]
- Gu, Y.; Jin, S.; Gao, Y.; Weaver, D.T.; Alt, F.W. Ku70-deficient embryonic stem cells have increased ionizing radiosensitivity, defective DNA end-binding activity, and inability to support V(D)J recombination. Proc. Natl. Acad. Sci. USA 1997, 94, 8076–8081. [Google Scholar] [CrossRef]
- Reliene, R.; Yamamoto, M.L.; Rao, P.N.; Schiestl, R.H. Genomic instability in mice is greater in Fanconi anemia caused by deficiency of Fancd2 than Fancg. Cancer Res. 2010, 70, 9703–9710. [Google Scholar]
- Suhasini, A.N.; Brosh, R.M., Jr. Fanconi anemia and Bloom’s syndrome crosstalk through FANCJ-BLM helicase interaction. Trends Genet. 2012, 28, 7–13. [Google Scholar] [CrossRef]
- Sivasubramaniam, S.; Sun, X.; Pan, Y.R.; Wang, S.; Lee, E.Y. Cep164 is a mediator protein required for the maintenance of genomic stability through modulation of MDC1, RPA, and CHK1. Genes Dev. 2008, 22, 587–600. [Google Scholar] [CrossRef]
- Huang, D.W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar]
- Kelly, K.O.; Dernburg, A.F.; Stanfield, G.M.; Villeneuve, A.M. Caenorhabditis elegans msh-5 is required for both normal and radiation-induced meiotic crossing over but not for completion of meiosis. Genetics 2000, 156, 617–630. [Google Scholar]
- Keeney, S.; Giroux, C.N.; Kleckner, N. Meiosis-specific DNA double-strand breaks are catalyzed by Spo11, a member of a widely conserved protein family. Cell 1997, 88, 375–384. [Google Scholar] [CrossRef]
- Gorgoulis, V.G.; Vassiliou, L.V.; Karakaidos, P.; Zacharatos, P.; Kotsinas, A.; Liloglou, T.; Venere, M.; Ditullio, R.A., Jr.; Kastrinakis, N.G.; Levy, B.; et al. Activation of the DNA damage checkpoint and genomic instability in human precancerous lesions. Nature 2005, 434, 907–913. [Google Scholar] [CrossRef]
- Ribes-Zamora, A.; Mihalek, I.; Lichtarge, O.; Bertuch, A.A. Distinct faces of the Ku heterodimer mediate DNA repair and telomeric functions. Nat. Struct. Mol. Biol. 2007, 14, 301–307. [Google Scholar]
- Jacobs, J.J.; Keblusek, P.; Robanus-Maandag, E.; Kristel, P.; Lingbeek, M.; Nederlof, P.M.; van Welsem, T.; van de Vijver, M.J.; Koh, E.Y.; Daley, G.Q.; et al. Senescence bypass screen identifies TBX2, which represses Cdkn2a (p19(ARF)) and is amplified in a subset of human breast cancers. Nat. Genet. 2000, 26, 291–299. [Google Scholar] [CrossRef]
- Prince, S.; Carreira, S.; Vance, K.W.; Abrahams, A.; Goding, C.R. Tbx2 directly represses the expression of the p21(WAF1) cyclin-dependent kinase inhibitor. Cancer Res. 2004, 64, 1669–1674. [Google Scholar] [CrossRef]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar]
- Livak, K.J.; Schmittgen, T.D. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) Method. Methods 2001, 25, 402–408. [Google Scholar] [CrossRef]
Supplementary Files
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Simko, S.J.; Voicu, H.; Carofino, B.L.; Justice, M.J. Mouse Lymphoblastic Leukemias Induced by Aberrant Prdm14 Expression Demonstrate Widespread Copy Number Alterations Also Found in Human ALL. Cancers 2012, 4, 1050-1066. https://doi.org/10.3390/cancers4041050
Simko SJ, Voicu H, Carofino BL, Justice MJ. Mouse Lymphoblastic Leukemias Induced by Aberrant Prdm14 Expression Demonstrate Widespread Copy Number Alterations Also Found in Human ALL. Cancers. 2012; 4(4):1050-1066. https://doi.org/10.3390/cancers4041050
Chicago/Turabian StyleSimko, Stephen J., Horatiu Voicu, Brandi L. Carofino, and Monica J. Justice. 2012. "Mouse Lymphoblastic Leukemias Induced by Aberrant Prdm14 Expression Demonstrate Widespread Copy Number Alterations Also Found in Human ALL" Cancers 4, no. 4: 1050-1066. https://doi.org/10.3390/cancers4041050




