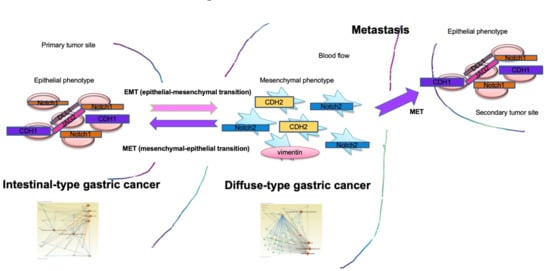
Molecular Network Profiling in Intestinal- and Diffuse-Type Gastric Cancer
Abstract
:Simple Summary
Abstract
1. Introduction
2. Results
2.1. Genes Altered in Intestinal- and Diffuse-Type GC
2.2. Networks Generated from Genes Altered in Intestinal- and Diffuse-Type GC
2.3. Regulator Effect Networks Related to Cancer in Intestinal- and Diffuse-Type GC
2.4. MicroRNA (miRNA)-Related Regulator Effect Networks in Intestinal- and Diffuse-Type GC
2.5. Upstream Regulators in Intestinal- and Diffuse-Type GC
2.6. Gene Ontology (GO) (Biological Process) and EMT-Related Processes of Genes Regulated in Intestinal- and Diffuse-Type GC
2.7. Prediction Model for Molecular Networks of Intestinal- and Diffuse-Type GC
2.8. EMT Molecular Pathway and Diffuse-Type GC Mapping
3. Discussion
4. Materials and Methods
4.1. Data Collection
4.2. Network Analysis
4.3. Gene Ontology (GO) Analysis
4.4. AI Prediction Modeling
4.5. Data Visualization
4.6. Statistical Analysis
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Perrot-Applanat, M.; Vacher, S.; Pimpie, C.; Chemlali, W.; Derieux, S.; Pocard, M.; Bieche, I. Differential gene expression in growth factors, epithelial mesenchymal transition and chemotaxis in the diffuse type compared with the intestinal type of gastric cancer. Oncol. Lett. 2019, 18, 674–686. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Tanabe, S.; Aoyagi, K.; Yokozaki, H.; Sasaki, H. Gene expression signatures for identifying diffuse-type gastric cancer associated with epithelial-mesenchymal transition. Int. J. Oncol. 2014, 44, 1955–1970. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Assumpção, P.P.; Barra, W.F.; Ishak, G.; Coelho, L.G.V.; Coimbra, F.J.F.; Freitas, H.C.; Dias-Neto, E.; Camargo, M.C.; Szklo, M. The diffuse-type gastric cancer epidemiology enigma. BMC Gastroenterol. 2020, 20, 223. [Google Scholar] [CrossRef] [PubMed]
- Hoang, T.; Ganesan, A.K.; Hiyama, D.; Dayyani, F. Gene mutations distinguishing gastric from colorectal and esophageal adenocarcinomas. J. Gastrointest Oncol. 2020, 11, 45–54. [Google Scholar] [CrossRef]
- Sohn, S.H.; Kim, B.; Sul, H.J.; Kim, Y.J.; Kim, H.S.; Kim, H.; Seo, J.B.; Koh, Y.; Zang, D.Y. INC280 inhibits Wnt/β-catenin and EMT signaling pathways and its induce apoptosis in diffuse gastric cancer positive for c-MET amplification. BMC Res. Notes 2019, 12, 125. [Google Scholar] [CrossRef] [Green Version]
- Tanabe, S.; Quader, S.; Cabral, H.; Ono, R. Interplay of EMT and CSC in Cancer and the Potential Therapeutic Strategies. Front. Pharmacol. 2020, 11, 904. [Google Scholar] [CrossRef]
- Zhang, Y.; Weinberg, R.A. Epithelial-to-mesenchymal transition in cancer: Complexity and opportunities. Front. Med. 2018, 12, 361–373. [Google Scholar] [CrossRef] [Green Version]
- Yang, J.; Antin, P.; Berx, G.; Blanpain, C.; Brabletz, T.; Bronner, M.; Campbell, K.; Cano, A.; Casanova, J.; Christofori, G.; et al. Guidelines and definitions for research on epithelial-mesenchymal transition. Nat. Rev. Mol. Cell Biol. 2020, 21, 341–352. [Google Scholar] [CrossRef] [Green Version]
- Simeone, P.; Trerotola, M.; Franck, J.; Cardon, T.; Marchisio, M.; Fournier, I.; Salzet, M.; Maffia, M.; Vergara, D. The multiverse nature of epithelial to mesenchymal transition. Semin. Cancer Biol. 2019, 58, 1–10. [Google Scholar] [CrossRef]
- Stefania, D.; Vergara, D. The Many-Faced Program of Epithelial-Mesenchymal Transition: A System Biology-Based View. Front. Oncol. 2017, 7, 274. [Google Scholar] [CrossRef]
- Tanabe, S.; Kawabata, T.; Aoyagi, K.; Yokozaki, H.; Sasaki, H. Gene expression and pathway analysis of CTNNB1 in cancer and stem cells. World J. Stem Cells 2016, 8, 384. [Google Scholar] [CrossRef] [PubMed]
- Cao, B.; Zhao, Y.; Zhang, Z.; Li, H.; Xing, J.; Guo, S.; Qiu, X.; Zhang, S.; Min, L.; Zhu, S. Gene regulatory network construction identified NFYA as a diffuse subtype-specific prognostic factor in gastric cancer. Int. J. Oncol. 2018, 53, 1857–1868. [Google Scholar] [CrossRef] [PubMed]
- Bass, A.J.; Thorsson, V.; Shmulevich, I.; Reynolds, S.M.; Miller, M.; Bernard, B.; Hinoue, T.; Laird, P.W.; Curtis, C.; Shen, H.; et al. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 2014, 513, 202–209. [Google Scholar] [CrossRef] [Green Version]
- Cerami, E.; Gao, J.; Dogrusoz, U.; Gross, B.E.; Sumer, S.O.; Aksoy, B.A.; Jacobsen, A.; Byrne, C.J.; Heuer, M.L.; Larsson, E.; et al. The cBio Cancer Genomics Portal: An Open Platform for Exploring Multidimensional Cancer Genomics Data. Cancer Discov. 2012, 2, 401–404. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gao, J.; Aksoy, B.A.; Dogrusoz, U.; Dresdner, G.; Gross, B.; Sumer, S.O.; Sun, Y.; Jacobsen, A.; Sinha, R.; Larsson, E.; et al. Integrative analysis of complex cancer genomics and clinical profiles using the cBioPortal. Sci. Signal. 2013, 6, pl1. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Hoadley, K.A.; Yau, C.; Hinoue, T.; Wolf, D.M.; Lazar, A.J.; Drill, E.; Shen, R.; Taylor, A.M.; Cherniack, A.D.; Thorsson, V.; et al. Cell-of-Origin Patterns Dominate the Molecular Classification of 10,000 Tumors from 33 Types of Cancer. Cell 2018, 173, 291–304. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Korpal, M.; Ell, B.J.; Buffa, F.M.; Ibrahim, T.; Blanco, M.A.; Celià-Terrassa, T.; Mercatali, L.; Khan, Z.; Goodarzi, H.; Hua, Y.; et al. Direct targeting of Sec23a by miR-200s influences cancer cell secretome and promotes metastatic colonization. Nat. Med. 2011, 17, 1101–1108. [Google Scholar] [CrossRef] [Green Version]
- Sun, Z.; Zhou, S.; Tang, J.; Ye, T.; Li, J.; Liu, D.; Zhou, J.; Wang, J.; Rosie Xing, H. Sec23a mediates miR-200c augmented oligometastatic to polymetastatic progression. EBioMedicine 2018, 37, 47–55. [Google Scholar] [CrossRef] [Green Version]
- Ma, L.; Young, J.; Prabhala, H.; Pan, E.; Mestdagh, P.; Muth, D.; Teruya-Feldstein, J.; Reinhardt, F.; Onder, T.T.; Valastyan, S.; et al. miR-9, a MYC/MYCN-activated microRNA, regulates E-cadherin and cancer metastasis. Nat. Cell Biol. 2010, 12, 247–256. [Google Scholar] [CrossRef] [Green Version]
- Pan, Y.; Li, J.; Zhang, Y.; Wang, N.; Liang, H.; Liu, Y.; Zhang, C.Y.; Zen, K.; Gu, H. Slug-upregulated miR-221 promotes breast cancer progression through suppressing E-cadherin expression. Sci Rep. 2016, 6, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Choi, P.W.; Ng, S.W. The Functions of MicroRNA-200 Family in Ovarian Cancer: Beyond Epithelial-Mesenchymal Transition. Int. J. Mol. Sci. 2017, 18, 1207. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Grossman, R.L.; Heath, A.P.; Ferretti, V.; Varmus, H.E.; Lowy, D.R.; Kibbe, W.A.; Staudt, L.M. Toward a Shared Vision for Cancer Genomic Data. New Engl. J. Med. 2016, 375, 1109–1112. [Google Scholar] [CrossRef] [PubMed]
- Krämer, A.; Green, J.; Pollard, J., Jr.; Tugendreich, S. Causal analysis approaches in Ingenuity Pathway Analysis. Bioinformatics 2014, 30, 523–530. [Google Scholar] [CrossRef] [PubMed]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protoc. 2009, 4, 44–57. [Google Scholar] [CrossRef] [PubMed]
- Huang da, W.; Sherman, B.T.; Lempicki, R.A. Bioinformatics enrichment tools: Paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 2009, 37, 1–13. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Breiman, L. Random Forests. Mach. Learn. 2001, 45, 5–32. [Google Scholar] [CrossRef] [Green Version]
- Friedman, J.H. Greedy function approximation: A gradient boosting machine. Ann. Statist. 2001, 29, 1189–1232. [Google Scholar] [CrossRef]







| Gene Symbol | Gene Name | GOTERM_BP_DIRECT |
|---|---|---|
| MSL3P1 | male-specific lethal 3 homolog (Drosophila) pseudogene 1 | GO:0006338~chromatin remodeling, GO:0006342~chromatin silencing, GO:0006351~transcription, DNA-templated, GO:0016575~histone deacetylation, GO:0043967~histone H4 acetylation, GO:0043968~histone H2A acetylation, |
| CKS1B | CDC28 protein kinase regulatory subunit 1B | GO:0007049~cell cycle, GO:0007346~regulation of mitotic cell cycle, GO:0008283~cell proliferation, GO:0044772~mitotic cell cycle phase transition, GO:0045737~positive regulation of cyclin-dependent protein serine/threonine kinase activity, GO:0045893~positive regulation of transcription, DNA-templated, GO:0051301~cell division, |
| DDX27 | DEAD-box helicase 27 | GO:0006364~rRNA processing, GO:0010501~RNA secondary structure unwinding, |
| GET4 | golgi to ER traffic protein 4 | GO:0006810~transport, GO:0051220~cytoplasmic sequestering of protein, GO:0071816~tail-anchored membrane protein insertion into ER membrane, GO:1904378~maintenance of unfolded protein involved in ERAD pathway, |
| CSE1L | chromosome segregation 1 like | GO:0006606~protein import into nucleus, GO:0006611~protein export from nucleus, GO:0006915~apoptotic process, GO:0008283~cell proliferation, |
| TOMM34 | translocase of outer mitochondrial membrane 34 | GO:0006626~protein targeting to mitochondrion, |
| YTHDF1 | YTH N6-methyladenosine RNA binding protein 1 | GO:0045948~positive regulation of translational initiation, |
| RAE1 | ribonucleic acid export 1 | GO:0000972~transcription-dependent tethering of RNA polymerase II gene DNA at nuclear periphery, GO:0006406~mRNA export from nucleus, GO:0006409~tRNA export from nucleus, GO:0006606~protein import into nucleus, GO:0007077~mitotic nuclear envelope disassembly, GO:0010827~regulation of glucose transport, GO:0016032~viral process, GO:0016925~protein sumoylation, GO:0019083~viral transcription, GO:0031047~gene silencing by RNA, GO:0071407~cellular response to organic cyclic compound, GO:0075733~intracellular transport of virus, GO:1900034~regulation of cellular response to heat, |
| PARD6B | par-6 family cell polarity regulator beta | GO:0006461~protein complex assembly, GO:0007043~cell-cell junction assembly, GO:0007049~cell cycle, GO:0007163~establishment or maintenance of cell polarity, GO:0007409~axonogenesis, GO:0030334~regulation of cell migration, GO:0051301~cell division, GO:0070830~bicellular tight junction assembly, |
| MRGBP | MRG domain binding protein | GO:0006351~transcription, DNA-templated, GO:0006357~regulation of transcription from RNA polymerase II promoter, GO:0016573~histone acetylation, GO:0040008~regulation of growth, |
| ID | Focus Molecules | Top Diseases and Functions |
|---|---|---|
| 1 | 35 | Cancer, Gastrointestinal Disease, Organismal Injury and Abnormalities |
| 2 | 35 | Amino Acid Metabolism, Molecular Transport, Small Molecule Biochemistry |
| 3 | 34 | Cardiovascular Disease, Gene Expression, Protein Synthesis |
| 4 | 34 | Developmental Disorder, Hereditary Disorder, Neurological Disease |
| 5 | 34 | Dental Disease, Dermatological Diseases and Conditions, Post-Translational Modification |
| 6 | 34 | Hereditary Disorder, Infectious Diseases, RNA Post-Transcriptional Modification |
| 7 | 34 | Carbohydrate Metabolism, Lipid Metabolism, Post-Translational Modification |
| 8 | 34 | Connective Tissue Disorders, Developmental Disorder, Hereditary Disorder |
| 9 | 34 | Cell Cycle, Molecular Transport, Protein Trafficking |
| 10 | 33 | Connective Tissue Disorders, Dermatological Diseases and Conditions, Developmental Disorder |
| 11 | 33 | Cell Morphology, Cellular Assembly and Organization, Cellular Function and Maintenance |
| 12 | 33 | Gene Expression, Post-Translational Modification, RNA Damage and Repair |
| 13 | 33 | Cell Cycle, Cellular Growth and Proliferation, Reproductive System Development and Function |
| 14 | 32 | Infectious Diseases, Molecular Transport, Post-Translational Modification |
| 15 | 32 | Cell Cycle, Cellular Assembly and Organization, DNA Replication, Recombination, and Repair |
| 16 | 32 | Developmental Disorder, Hereditary Disorder, Molecular Transport |
| 17 | 32 | Carbohydrate Metabolism, Nucleic Acid Metabolism, Small Molecule Biochemistry |
| 18 | 31 | Cellular Assembly and Organization, Cellular Response to Therapeutics, DNA Replication, Recombination, and Repair |
| 19 | 31 | Developmental Disorder, Lipid Metabolism, Small Molecule Biochemistry |
| 20 | 31 | Cell Morphology, Cellular Assembly and Organization, Skeletal and Muscular System Development and Function |
| 21 | 31 | Cancer, Cellular Assembly and Organization, Skeletal and Muscular Disorders |
| 22 | 31 | Cell Cycle, Cellular Assembly and Organization, Cellular Compromise |
| 23 | 31 | Molecular Transport, RNA Post-Transcriptional Modification, RNA Trafficking |
| 24 | 31 | Nervous System Development and Function, Neurological Disease, Organ Morphology |
| 25 | 31 | Gene Expression, Neurological Disease, Organismal Functions |
| ID | Regulators | Target Total | Diseases & Functions |
|---|---|---|---|
| 1 | AREG, BNIP3L, CHEK1, E2f, E2F3, EIF4G1, Irgm1, LIN9, MED1, miR-21-5p (and other miRNAs w/seed AGCUUAU), mir-290, NLRP3, PTGER2, RABL6, UXT, YAP1 | 94 | Hepatocellular carcinoma, Oral tumor |
| 2 | AREG, ERG, KDM5B, MIR17HG, TFDP1, YAP1 | 123 | Hepatocellular carcinoma, Intestinal cancer, Large intestine neoplasm |
| 3 | AREG, KDM5B, miR-21-5p (and other miRNAs w/seed AGCUUAU), mir-290, MIR17HG, PTGER2, SMARCB1, TCF3, UXT, YAP1 | 70 | Hepatocellular carcinoma |
| 4 | AREG, CSF2, DYRK1A, E2F2, KDM1A, let-7a-5p (and other miRNAs w/seed GAGGUAG), MED1, NLRP3, TBX2, YAP1 | 200 | Gastrointestinal tract cancer, Hepatocellular carcinoma, Large intestine neoplasm, Oral tumor |
| 5 | MYCN | 3 | Cell death of osteosarcoma cells |
| 6 | EGFR, ERBB2, HRAS, miR-205-5p (and other miRNAs w/seed CCUUCAU), tanespimycin, tazemetostat, YAP1 | 57 | Oral tumor |
| 7 | calcitriol, medroxyprogesterone acetate | 112 | Gastrointestinal adenocarcinoma, Intestinal carcinoma |
| 8 | TP53 | 298 | Gastrointestinal carcinoma |
| 9 | 5-fluorouracil | 28 | Liver tumor |
| 10 | TAL1 | 31 | Liver tumor |
| 11 | NUPR1 | 25 | Hepatocellular carcinoma |
| 12 | MITF | 20 | Hepatocellular carcinoma |
| 13 | 26s Proteasome | 23 | Liver tumor |
| 14 | EP400 | 19 | Liver tumor |
| 15 | CDKN2A | 69 | Intestinal cancer, Large intestine neoplasm |
| 16 | FOXO1 | 45 | Hepatobiliary system cancer |
| 17 | E2F1 | 47 | Hepatocellular carcinoma |
| 18 | HGF | 35 | Hepatocellular carcinoma |
| 19 | arsenic trioxide | 32 | Liver tumor |
| 20 | let-7 | 27 | Hepatocellular carcinoma |
| 21 | TP73 | 36 | Hepatobiliary system cancer |
| 22 | mir-21 | 13 | Oral tumor |
| 23 | valproic acid | 12 | Cell death of osteosarcoma cells |
| ID | Regulators | Target Total | Diseases & Functions |
|---|---|---|---|
| 1 | ACTB, AREG, BRD4, CCND1, CDKN1A, DYRK1A, E2f, E2F3, EIF4G1, EWSR1, FOXM1, GATA1, gentamicin, imipramine blue, LIN9, MED1, MYCN, NLRP3, NTRK2, phenethyl isothiocyanate, Rb, RB1, RBL2, TCF3, TFDP1 | 276 | Female genital neoplasm, Gonadal tumor, Oral tumor, Tumorigenesis of reproductive tract |
| 2 | ATF4, ATF6, BNIP3L, E2f, EIF4G1, epothilone B, ERG, FOXM1, GATA1, gentamicin, imipramine blue, Irgm1, KDM5B, let-7, miR-24-3p (and other miRNAs w/seed GGCUCAG), NLRP3, phenethyl isothiocyanate, RABL6, Rb, RB1, RBL1, RBL2, SMARCB1, ZNF281 | 231 | Cell death of osteosarcoma cells, Female genital neoplasm, Gonadal tumor, Tumorigenesis of reproductive tract |
| 3 | alvespimycin, decitabine, EGFR, EWSR1, gentamicin, KAT6A, miR-34a-5p (and other miRNAs w/seed GGCAGUG), phenethyl isothiocyanate, SYVN1, tazemetostat, YAP1 | 67 | Oral tumor |
| 4 | alvespimycin, calcitriol, decitabine, E2F2, EGFR, ERBB2, estrogen, EWSR1, mir-181, phenethyl isothiocyanate, tazemetostat, Vegf, YAP1 | 210 | Oral tumor, Prostatic carcinoma |
| 5 | CCND1, DDIT3, HDAC1 | 140 | Digestive system cancer, Oral tumor, Prostatic carcinoma |
| 6 | ATF4, ATF6, EIF4G1, EP400, FOXM1, gentamicin, Irgm1, MYC, NELFA, NELFCD, NELFE, NLRP3, ZNF281 | 94 | Ovarian tumor |
| 7 | KDM4C, UXT | 18 | Frequency of tumor, Incidence of tumor |
| 8 | CSF2, DDIT3, ERG, ESR1, miR-291a-3p (and other miRNAs w/seed AAGUGCU) | 131 | Prostatic carcinoma |
| 9 | TP53 | 131 | Prostatic carcinoma |
| 10 | E2F1 | 87 | Tumorigenesis of reproductive tract |
| 11 | HGF | 67 | Female genital neoplasm |
| 12 | TBX2 | 38 | Digestive system cancer |
| 13 | SMARCB1 | 39 | Abdominal carcinoma |
| 14 | TP63 | 42 | Tumorigenesis of reproductive tract |
| 15 | PTGER2 | 40 | Abdominal carcinoma |
| 16 | MITF | 33 | Female genital neoplasm |
| 17 | mir-21 | 27 | Prostatic carcinoma |
| 18 | NFE2L2 | 23 | Gonadal tumor |
| 19 | CD3 | 19 | Oral tumor |
| 20 | DNMT3B | 18 | Female genital neoplasm |
| 21 | cephaloridine | 18 | Tumorigenesis of reproductive tract |
| 22 | CDKN2A | 47 | Tumorigenesis of reproductive tract |
| 23 | NFE2L1 | 6 | Cell death of osteosarcoma cells |
| 24 | EIF4E | 11 | Ovarian tumor |
| 25 | 5-fluorouracil | 5 | Cell death of osteosarcoma cells |
| 26 | mibolerone | 9 | Ovarian tumor |
| 27 | KDM1A | 25 | Female genital neoplasm, Tumorigenesis of reproductive tract |
| 28 | TRAP1 | 6 | Oral tumor |
| 29 | fulvestrant | 45 | Female genital neoplasm |
| 30 | NCOA3 | 15 | Tumorigenesis of reproductive tract |
| 31 | MEF2D | 9 | Female genital neoplasm, Tumorigenesis of reproductive tract |
| ID | Regulators | Target Molecules in Dataset | Diseases & Functions | Known Regulator-Disease/Function Relationship |
|---|---|---|---|---|
| 1 | miR-205-5p (and other miRNAs w/seed CCUUCAU), miR-21-5p (and other miRNAs w/seed AGCUUAU), mir-290 | ABCC2, ATP1A1, BCL2L1, CDH5, CDK2, ERBB3, IRAK1, MSH2, NFIB, PIK3R1, PRKACB, PTEN, RECK, SOX2, TGFBR2, TIMP3, VEGFA, ZEB2 | Hepatobiliary carcinoma, Hepatobiliary system cancer, Liver tumor, Oral tumor | 42% (5/12) |
| 2 | let-7a-5p (and other miRNAs w/seed GAGGUAG) | ADGRG1, BCL2L1, CCND1, CCNE1, CDKN2A, IGF2BP1, IGF2BP3, TYMS, VIM | Hepatocellular carcinoma | 100% (1/1) |
| 3 | let-7 | AGO2, APC, AURKA, BCL2L1, BRCA1, BRCA2, BUB1, BUB1B, CCNA2, CCNB1, CCND1, CCNE2, CDC6, CDCA8, CKS1B, DLC1, E2F5, E2F8, IGF2BP1, MCM2, ORC6, RFC4, RRM1, RRM2, SMAD4, SOX9, VIM | Hepatocellular carcinoma | 100% (1/1) |
| 4 | mir-21 | BCL2, CCND1, CDH5, CDKN2A, DLGAP5, IRAK1, KNTC1, LEPR, PTEN, STAT1, TACC3, TIMP3, TOP2A | Oral tumor | 0% (0/1) |
| ID | Regulators | Target Molecules in Dataset | Diseases & Functions | Known Regulator-Disease/Function Relationship |
|---|---|---|---|---|
| 1 | let-7, miR-24-3p (and other miRNAs w/seed GGCUCAG) | ACVR1B, APC, AURKA, AURKB, BCL2L1, BRCA1, BRCA2, BUB1, BUB1B, CCNA2, CCNB1, CCND1, CCNE2, CDC20, CDC25A, CDC6, CDK1, CDK4, CDKN2A, CKS1B, DBF4, DLC1, E2F4, E2F8, FANCD2, FBL, FEN1, HMGA1, IGF2BP1, MCM10, MCM2, MCM7, MCM8, NOLC1, NUF2, PLAGL2, RFC4, RFC5, RRM1, RRM2, SALL4, SLC25A13, SMAD4, SOX9, TARBP2, VIM, XPO5 | Female genital neoplasm, Gonadal tumor, Tumorigenesis of reproductive tract | 50% (3/6) |
| 2 | mir-181, miR-291a-3p (and other miRNAs w/seed AAGUGCU), miR-34a-5p (and other miRNAs w/seed GGCAGUG) | ADCY9, ARHGEF3, BCL2, BIRC5, CCND1, CD46, CDK4, CDKN2A, CENPF, E2F3, E2F5, FAM13B, KIF23, MCM10, NIN, PRC1, PRKACB, PTEN, SOX2, TFAP4, TIMP3, VEGFA, ZEB2 | Oral tumor, Prostatic carcinoma | 0% (0/6) |
| 3 | mir-21 | ANLN, ARL6IP1, ASPM, ATAD2, BCL2, CCNB1, CCND1, CDH5, CDKN2A, CKAP5, CSE1L, KIF23, KNTC1, LEPR, MKI67, NCAPD2, NUSAP1, PIP4K2A, PRC1, PTEN, SOX2, STAT1, TAP1, TBC1D1, TOP2A, YY1, ZWILCH | Prostatic carcinoma | 0% (0/1) |
| 4 | mir-21 | ANLN, ARL6IP1, ASPM, ATAD2, BCL2, C1R, CACYBP, CANX, CCNA2, CCNB1, CCND1, CDC25A, CDH5, CDKN2A, CKAP5, CKS2, CLPB, CSE1L, ECT2, FUBP1, GTSE1, HNRNPA2B1, IFI16, IRAK1, KIF23, KIF4A, KIFC1, KNTC1, LEPR, MKI67, MSH2, NCAPD2, NME1, NPAS2, NUSAP1, PIP4K2A, PRC1, PTEN, RACGAP1, RAD51AP1, RECK, SMC2, SOX2, STAT1, STMN1, TACC3, TAP1, TBC1D1, TCF21, TIMP3, TLR1, TMEM97, TOP2A, TP53RK, UBA7, VRK1, YWHAB, YY1, ZW10, ZWILCH | Frequency of tumor | 100% (1/1) |
| 5 | mir-21 | ANLN, ARL6IP1, ASPM, ATAD2, BCL2, C1R, CACYBP, CANX, CCNA2, CCNB1, CCND1, CDC25A, CDH5, CDKN2A, CKAP5, CKS2, CLPB, CSE1L, ECT2, FUBP1, GTSE1, HNRNPA2B1, IFI16, IRAK1, KIF23, KIF4A, KIFC1, KNTC1, LEPR, MKI67, MSH2, NCAPD2, NME1, NPAS2, NUSAP1, PIP4K2A, PRC1, PTEN, RACGAP1, RAD51AP1, RECK, SMC2, SOX2, STAT1, STMN1, TACC3, TAP1, TBC1D1, TCF21, TIMP3, TLR1, TMEM97, TOP2A, TP53RK, UBA7, VRK1, YWHAB, YY1, ZW10, ZWILCH | Incidence of tumor | 100% (1/1) |
| Upstream Regulators | TCGA CIN | TCGA GS |
|---|---|---|
| NUPR1 | −4.457 | 6.685 |
| CSF2 | 4.849 | −6.057 |
| PTGER2 | 4.427 | −5.06 |
| TP53 | −4.044 | 5.394 |
| EGFR | 3.75 | −5.207 |
| let-7 | −3.031 | 5.836 |
| ERBB2 | 2.986 | −5.804 |
| calcitriol | −3.349 | 5.194 |
| RABL6 | 3.28 | −5.154 |
| MITF | 2.927 | −5.436 |
| E2F1 | 2.141 | −5.933 |
| CDKN2A | −2.944 | 5 |
| KDM1A | 3.328 | −4.551 |
| E2F3 | 2.496 | −5.334 |
| EP400 | 3.183 | −4.482 |
| BNIP3L | −3.714 | 3.571 |
| YAP1 | 3.103 | −4.161 |
| MYCN | 4.044 | −2.997 |
| MYC | 1.087 | −5.862 |
| HGF | 2.874 | −4.014 |
| E2f | 2.984 | −3.881 |
| AREG | 3.525 | −3.213 |
| TBX2 | 2.619 | −4.104 |
| KDM5B | −4.075 | 2.537 |
| Term | Count |
|---|---|
| GO:0051301~cell division | 121 |
| GO:0007062~sister chromatid cohesion | 54 |
| GO:0007067~mitotic nuclear division | 91 |
| GO:0006260~DNA replication | 67 |
| GO:0031145~anaphase-promoting complex-dependent catabolic process | 40 |
| GO:0051436~negative regulation of ubiquitin-protein ligase activity involved in mitotic cell cycle | 37 |
| GO:0000082~G1/S transition of mitotic cell cycle | 44 |
| GO:0051437~positive regulation of ubiquitin-protein ligase activity involved in regulation of mitotic cell cycle transition | 36 |
| GO:0006281~DNA repair | 74 |
| GO:0006521~regulation of cellular amino acid metabolic process | 27 |
| GO:0006270~DNA replication initiation | 20 |
| GO:0043488~regulation of mRNA stability | 39 |
| GO:0006364~rRNA processing | 62 |
| GO:0007059~chromosome segregation | 29 |
| GO:0031047~gene silencing by RNA | 39 |
| GO:0038061~NIK/NF-kappaB signaling | 27 |
| GO:0060071~Wnt signaling pathway, planar cell polarity pathway | 33 |
| GO:0002479~antigen processing and presentation of exogenous peptide antigen via MHC class I, TAP-dependent | 26 |
| GO:0007077~mitotic nuclear envelope disassembly | 21 |
| GO:0000398~mRNA splicing, via spliceosome | 60 |
| GO:0070125~mitochondrial translational elongation | 31 |
| Genes | |||
|---|---|---|---|
| AGPAT2 | BCAT2 | HPRT1 | PAFAH1B1 |
| HACD3 | CERS2 | IMPDH1 | PDGFRA |
| HACD4 | CHDH | ITPA | PNPT1 |
| BDH2 | CHKA | ITPKA | PRIM1 |
| ATIC | CDO1 | ITPKB | PRIM2 |
| ADPRM | CYP1B1 | IDH1 | PTGES2 |
| AKT3 | CYP2U1 | IDH3B | PTGES3 |
| CTPS1 | DGUOK | LTC4S | PTGS1 |
| CTPS2 | DTYMK | LPIN3 | PRKCB |
| DNMT3B | DGAT2 | LIPT2 | PRUNE1 |
| POLA2 | DMGDH | MDH2 | PNPO |
| POLD2 | ENTPD1 | MARS2 | PYCR1 |
| POLE2 | ENTPD6 | MARS | PYCR2 |
| POLE3 | ENOPH1 | MMAB | PC |
| HDDC3 | ERBB2 | MAPK10 | RAC3 |
| NANP | FAM213B | NPR1 | RRM1 |
| NFS1 | FASN | NPR2 | RRM2 |
| NME1 | FGFR1 | NEU1 | RPIA |
| POLR1B | FGFR3 | NIT2 | RPE |
| POLR1C | FLAD1 | NUDT5 | SEPHS1 |
| POLR2C | FTCD | PTEN | SELENOI |
| POLR2H | FAH | PCYT2 | SRR |
| POLR2J | GAL3ST1 | PEMT | SLC1A5 |
| POLR3C | GGCT | PIK3CB | SLC44A5 |
| POLR3E | GGT5 | PIP5K1A | SLC7A5 |
| POLR3F | GGT7 | PIP5K1C | SORD |
| POLR3GL | GNPDA1 | PIP4K2A | SRM |
| POLR3K | G6PC3 | PIP4K2C | SMOX |
| TWISTNB | G6PD | PTDSS1 | SMPD4 |
| UGT8 | GANC | PDE11A | TAZ |
| UXS1 | GBA | PDE1A | TXNDC12 |
| WASF2 | GUSB | PDE2A | TXNRD1 |
| ACACB | GAD1 | PDE4B | TXNRD2 |
| ACSS1 | GCLM | PDE5A | TK1 |
| ACYP1 | EPRS | PDE7B | TYMS |
| AHCY | EARS2 | PGP | TPO |
| ADCY4 | GSTM5 | PIK3R1 | TALDO1 |
| ADCY9 | GSTO2 | PLPP1 | UMPS |
| AK2 | GSS | PPAT | UCKL1 |
| ADH1B | GCSH | PSAT1 | ZNRD1 |
| ALDH1A1 | GLO1 | PSPH | |
| ASPA | GMPS | PAFAH1B3 | |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tanabe, S.; Quader, S.; Ono, R.; Cabral, H.; Aoyagi, K.; Hirose, A.; Yokozaki, H.; Sasaki, H. Molecular Network Profiling in Intestinal- and Diffuse-Type Gastric Cancer. Cancers 2020, 12, 3833. https://doi.org/10.3390/cancers12123833
Tanabe S, Quader S, Ono R, Cabral H, Aoyagi K, Hirose A, Yokozaki H, Sasaki H. Molecular Network Profiling in Intestinal- and Diffuse-Type Gastric Cancer. Cancers. 2020; 12(12):3833. https://doi.org/10.3390/cancers12123833
Chicago/Turabian StyleTanabe, Shihori, Sabina Quader, Ryuichi Ono, Horacio Cabral, Kazuhiko Aoyagi, Akihiko Hirose, Hiroshi Yokozaki, and Hiroki Sasaki. 2020. "Molecular Network Profiling in Intestinal- and Diffuse-Type Gastric Cancer" Cancers 12, no. 12: 3833. https://doi.org/10.3390/cancers12123833