RNA-Seq Analysis for Assessing the Early Response to DSP Toxins in Mytilus galloprovincialis Digestive Gland and Gill
Abstract
:1. Introduction
2. Results
2.1. Toxin Accumulation
2.2. Transcriptome Sequencing and De Novo Assembly
2.3. DEGs Among Samples
2.4. Gene Functional Annotations
2.5. Real-Time Quantitative PCR (qPCR) Validation
3. Discussion
4. Conclusions
5. Materials and Methods
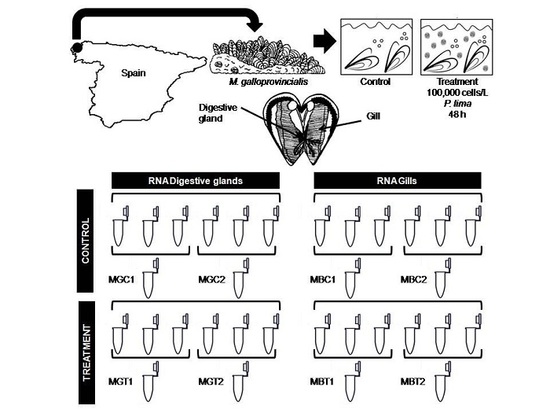
5.1. Sample Collection and Experimental Design
5.2. RNA Extraction
5.3. Library Preparation and Sequencing
5.4. De Novo Assembly
5.5. Differential Expression, Functional Annotation and Functional Enrichment Analysis of DEGs
5.6. Real-Time Quantitative PCR Validation
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Anderson, D.M.; Andersen, P.; Bricelj, V.M.; Cullen, J.J.; Rensel, J.E.J. Monitoring and Management Strategies for Harmful Algal Blooms in Coastal Waters; Unesco: Paris, France, 2001. [Google Scholar]
- Wells, M.L.; Trainer, V.L.; Smayda, T.J.; Karlson, B.S.O.; Trick, C.G.; Kudela, R.M.; Ishikawa, A.; Bernard, S.; Wulff, A.; Anderson, D.M.; et al. Harmful algal blooms and climate change: Learning from the past and present to forecast the future. Harmful Algae 2015, 49, 68–93. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Visciano, P.; Schirone, M.; Berti, M.; Milandri, A.; Tofalo, R.; Suzzi, G. Marine biotoxins: Occurrence, toxicity, regulatory limits and reference methods. Front. Microbiol. 2016, 7, 1051. [Google Scholar] [CrossRef] [PubMed]
- Bialojan, C.; Takai, A. Inhibitory effect of a marine-sponge toxin, okadaic acid, on protein phosphatases. Specificity and kinetics. Biochem. J. 1988, 256, 283–290. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Prado-Alvarez, M.; Flórez-Barrós, F.; Sexto-Iglesias, A.; Méndez, J.; Fernandez-Tajes, J. Effects of okadaic acid on haemocytes from Mytilus galloprovincialis: A comparison between field and laboratory studies. Mar. Environ. Res. 2012, 81, 90–93. [Google Scholar] [CrossRef] [PubMed]
- Prego-Faraldo, M.V.; Valdiglesias, V.; Méndez, J.; Eirín-López, J.M. Okadaic acid meet and greet: An insight into detection methods, response strategies and genotoxic effects in marine invertebrates. Mar. Drugs 2013, 11, 2829–2845. [Google Scholar] [CrossRef] [PubMed]
- Reguera, B.; Velo-Suárez, L.; Raine, R.; Park, M.G. Harmful dinophysis species: A review. Harmful Algae 2012, 14, 87–106. [Google Scholar] [CrossRef]
- Valdiglesias, V.; Prego-Faraldo, M.V.; Pásaro, E.; Méndez, J.; Laffon, B. Okadaic acid: More than a diarrheic toxin. Mar. Drugs 2013, 11, 4328–4349. [Google Scholar] [CrossRef] [PubMed]
- Munday, R. Is protein phosphatase inhibition responsible for the toxic effects of okadaic acid in animals? Toxins 2013, 5, 267–285. [Google Scholar] [CrossRef] [PubMed]
- Prego-Faraldo, M.; Vieira, L.; Eirin-Lopez, J.; Méndez, J.; Guilhermino, L. Transcriptional and biochemical analysis of antioxidant enzymes in the mussel Mytilus galloprovincialis during experimental exposures to the toxic dinoflagellate Prorocentrum lima. Mar. Environ. Res. 2017, 129, 304–315. [Google Scholar] [CrossRef] [PubMed]
- Prego-Faraldo, M.V.; Valdiglesias, V.; Laffon, B.; Eirín-López, J.M.; Méndez, J. In vitro analysis of early genotoxic and cytotoxic effects of okadaic acid in different cell types of the mussel Mytilus galloprovincialis. J. Toxicol. Environ. Health A 2015, 78, 814–824. [Google Scholar] [CrossRef] [PubMed]
- Prego-Faraldo, M.V.; Valdiglesias, V.; Laffon, B.; Mendez, J.; Eirin-Lopez, J.M. Early genotoxic and cytotoxic effects of the toxic dinoflagellate Prorocentrum lima in the mussel Mytilus galloprovincialis. Toxins 2016, 8, 159. [Google Scholar] [CrossRef] [PubMed]
- Gerdol, M.; De Moro, G.; Manfrin, C.; Milandri, A.; Riccardi, E.; Beran, A.; Venier, P.; Pallavicini, A. RNA sequencing and de novo assembly of the digestive gland transcriptome in Mytilus galloprovincialis fed with toxinogenic and non-toxic strains of Alexandrium minutum. BMC Res. Notes 2014, 7, 722. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Moreira, R.; Pereiro, P.; Canchaya, C.; Posada, D.; Figueras, A.; Novoa, B. RNA-Seq in Mytilus galloprovincialis: Comparative transcriptomics and expression profiles among different tissues. BMC Genom. 2015, 16, 728. [Google Scholar] [CrossRef] [PubMed]
- Pazos, A.J.; Ventoso, P.; Martínez-Escauriaza, R.; Pérez-Parallé, M.L.; Blanco, J.; Triviño, J.C.; Sánchez, J.L. Transcriptional response after exposure to domoic acid-producing Pseudo-nitzschia in the digestive gland of the mussel Mytilus galloprovincialis. Toxicon 2017, 140, 60–71. [Google Scholar] [CrossRef] [PubMed]
- Rosani, U.; Varotto, L.; Rossi, A.; Roch, P.; Novoa, B.; Figueras, A.; Pallavicini, A.; Venier, P. Massively parallel amplicon sequencing reveals isotype-specific variability of antimicrobial peptide transcripts in Mytilus galloprovincialis. PLoS ONE 2011, 6, e26680. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Suarez-Ulloa, V.; Fernandez-Tajes, J.; Aguiar-Pulido, V.; Prego-Faraldo, M.V.; Florez-Barros, F.; Sexto-Iglesias, A.; Mendez, J.; Eirin-Lopez, J.M. Unbiased high-throughput characterization of mussel transcriptomic responses to sublethal concentrations of the biotoxin okadaic acid. PeerJ 2015, 3, e1429. [Google Scholar] [CrossRef] [PubMed]
- Suárez-Ulloa, V.; Fernández-Tajes, J.; Aguiar-Pulido, V.; Rivera-Casas, C.; González-Romero, R.; Ausio, J.; Méndez, J.; Dorado, J.; Eirín-López, J.M. The CHROMEVALOA database: A resource for the evaluation of okadaic acid contamination in the marine environment based on the chromatin-associated transcriptome of the mussel Mytilus galloprovincialis. Mar. Drugs 2013, 11, 830–841. [Google Scholar] [CrossRef] [PubMed]
- Manfrin, C.; Dreos, R.; Battistella, S.; Beran, A.; Gerdol, M.; Varotto, L.; Lanfranchi, G.; Venier, P.; Pallavicini, A. Mediterranean mussel gene expression profile induced by okadaic acid exposure. Environ. Sci. Technol. 2010, 44, 8276–8283. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57. [Google Scholar] [CrossRef] [PubMed]
- Blanco, J.; Mariño, C.; Martín, H.; Acosta, C.P. Anatomical distribution of diarrhetic shellfish poisoning (DSP) toxins in the mussel Mytilus galloprovincialis. Toxicon 2007, 50, 1011–1018. [Google Scholar] [CrossRef] [PubMed]
- Moroño, A.; Arévalo, F.; Fernández, M.; Maneiro, J.; Pazos, Y.; Salgado, C.; Blanco, J. Accumulation and transformation of DSP toxins in mussels Mytilus galloprovincialis during a toxic episode caused by Dinophysis acuminata. Aquat. Toxicol. 2003, 62, 269–280. [Google Scholar] [CrossRef]
- Manfrin, C.; De Moro, G.; Torboli, V.; Venier, P.; Pallavicini, A.; Gerdol, M. Physiological and molecular responses of bivalves to toxic dinoflagellates. Invertebr. Surv. J. 2012, 9, 184–199. [Google Scholar]
- Beyer, J.; Green, N.W.; Brooks, S.; Allan, I.J.; Ruus, A.; Gomes, T.; Bråte, I.L.N.; Schøyen, M. Blue mussels (Mytilus edulis spp.) as sentinel organisms in coastal pollution monitoring: A review. Mar. Environ. Res. 2017. [Google Scholar] [CrossRef] [PubMed]
- Romero-Geraldo, R.d.J.; García-Lagunas, N.; Hernandez-Saavedra, N.Y. Effects of in vitro exposure to diarrheic toxin producer Prorocentrum lima on gene expressions related to cell cycle regulation and immune response in Crassostrea gigas. PLoS ONE 2014, 9, e97181. [Google Scholar] [CrossRef]
- Romero-Geraldo, R.d.J.; Hernández-Saavedra, N.Y. Stress gene expression in Crassostrea gigas (Thunberg, 1793) in response to experimental exposure to the toxic dinoflagellate Prorocentrum lima (Ehrenberg) Dodge, 1975. Aquac. Res. 2014, 45, 1512–1522. [Google Scholar] [CrossRef]
- Díaz, P.A.; Reguera, B.; Ruiz-Villarreal, M.; Pazos, Y.; Velo-Suárez, L.; Berger, H.; Sourisseau, M. Climate variability and oceanographic settings associated with interannual variability in the initiation of Dinophysis acuminata blooms. Mar. Drugs 2013, 11, 2964–2981. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.; Chen, Y.; Lu, W.; Wu, B.; Qi, P. Transcriptome analysis of Mytilus coruscus hemocytes in response to Vibrio alginnolyficus infection. Fish Shellfish Immunol. 2017, 70, 560–567. [Google Scholar] [CrossRef] [PubMed]
- Ventoso, P.; Martínez-Escauriaza, R.; Sánchez, J.; Pérez-Parallé, M.; Blanco, J.; Triviño, J.; Pazos, A. In Sequencing and de novo assembly of the digestive gland transcriptome in Mytilus galloprovincialis and analysis of differentially expressed genes in response to domoic acid. In Proceedings of the International Symposium on Genetics in Aquaculture XII, Santiago de Compostela, Spain, 21–27 June 2015; p. 93, 229. [Google Scholar]
- Chi, C.; Giri, S.; Jun, J.; Kim, S.; Kim, H.; Kang, J.; Park, S. Detoxification- and immune-related transcriptomic analysis of gills from bay scallops (Argopecten irradians) in response to algal toxin okadaic acid. Toxins 2018, 10, 308. [Google Scholar] [CrossRef] [PubMed]
- Huang, L.; Zou, Y.; Weng, H.-W.; Li, H.-Y.; Liu, J.-S.; Yang, W.-D. Proteomic profile in Perna viridis after exposed to Prorocentrum lima, a dinoflagellate producing DSP toxins. Environ. Pollut. 2015, 196, 350–357. [Google Scholar] [CrossRef] [PubMed]
- Guo, F.; An, T.; Rein, K.S. The algal hepatoxoxin okadaic acid is a substrate for human cytochromes CYP3A4 and CYP3A5. Toxicon 2010, 55, 325–332. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Miao, J.; Chi, L.; Pan, L.; Song, Y. Generally detected genes in comparative transcriptomics in bivalves: Toward the identification of molecular markers of cellular stress response. Environ. Toxicol. Pharmacol. 2015, 39, 475–481. [Google Scholar] [CrossRef] [PubMed]
- Yamashita, M.; Yamashita, Y.; Suzuki, T.; Kani, Y.; Mizusawa, N.; Imamura, S.; Takemoto, K.; Hara, T.; Hossain, M.A.; Yabu, T. Selenoneine, a novel selenium-containing compound, mediates detoxification mechanisms against methylmercury accumulation and toxicity in zebrafish embryo. Mar. Biotechnol. 2013, 15, 559–570. [Google Scholar] [CrossRef] [PubMed]
- Creppy, E.E.; Traoré, A.; Baudrimont, I.; Cascante, M.; Carratú, M.-R. Recent advances in the study of epigenetic effects induced by the phycotoxin okadaic acid. Toxicology 2002, 181, 433–439. [Google Scholar] [CrossRef]
- Murgarella, M.; Puiu, D.; Novoa, B.; Figueras, A.; Posada, D.; Canchaya, C. A first insight into the genome of the filter-feeder mussel Mytilus galloprovincialis. PLoS ONE 2016, 11, e0151561. [Google Scholar] [CrossRef]
- Zhang, G.; Fang, X.; Guo, X.; Li, L.; Luo, R.; Xu, F.; Yang, P.; Zhang, L.; Wang, X.; Qi, H. The oyster genome reveals stress adaptation and complexity of shell formation. Nature 2012, 490, 49–54. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Ilott, N.E.; Ponting, C.P. Predicting long non-coding RNAs using RNA sequencing. Methods 2013, 63, 50–59. [Google Scholar] [CrossRef] [PubMed]
- Milan, M.; Coppe, A.; Reinhardt, R.; Cancela, L.M.; Leite, R.B.; Saavedra, C.; Ciofi, C.; Chelazzi, G.; Patarnello, T.; Bortoluzzi, S. Transcriptome sequencing and microarray development for the manila clam, Ruditapes philippinarum: Genomic tools for environmental monitoring. BMC Genom. 2011, 12, 234. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lozano, V.; Martínez-Escauriaza, R.; Pérez-Parallé, M.; Pazos, A.; Sánchez, J. Two novel multidrug resistance associated protein (MRP/ABCC) from the mediterranean mussel (Mytilus galloprovincialis): Characterization and expression patterns in detoxifying tissues. Can. J. Zool. 2015, 93, 567–578. [Google Scholar] [CrossRef]
- Rosa, R.D.; Santini, A.; Fievet, J.; Bulet, P.; Destoumieux-Garzón, D.; Bachère, E. Big defensins, a diverse family of antimicrobial peptides that follows different patterns of expression in hemocytes of the oyster Crassostrea gigas. PLoS ONE 2011, 6, e25594. [Google Scholar] [CrossRef] [PubMed]
- Bettencourt, R.; Pinheiro, M.; Egas, C.; Gomes, P.; Afonso, M.; Shank, T.; Santos, R.S. High-throughput sequencing and analysis of the gill tissue transcriptome from the deep-sea hydrothermal vent mussel Bathymodiolus azoricus. BMC Genom. 2010, 11, 559. [Google Scholar] [CrossRef] [PubMed]
- Mello, D.F.; da Silva, P.M.; Barracco, M.A.; Soudant, P.; Hégaret, H. Effects of the dinoflagellate Alexandrium minutum and its toxin (saxitoxin) on the functional activity and gene expression of Crassostrea gigas hemocytes. Harmful Algae 2013, 26, 45–51. [Google Scholar] [CrossRef]
- Chi, C.; Giri, S.S.; Jun, J.W.; Kim, H.J.; Kim, S.W.; Yun, S.; Park, S.C. Effects of algal toxin okadaic acid on the non-specific immune and antioxidant response of bay scallop (Argopecten irradians). Fish Shellfish Immunol. 2017, 65, 111–117. [Google Scholar] [CrossRef] [PubMed]
- Gerdol, M.; Venier, P. An updated molecular basis for mussel immunity. Fish Shellfish Immunol. 2015, 46, 17–38. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Dunin-Horkawicz, S.; Feder, M.; Bujnicki, J.M. Phylogenomic analysis of the GIY−YIG nuclease superfamily. BMC Genom. 2006, 7, 98. [Google Scholar] [CrossRef] [PubMed]
- Biscotti, M.A.; Gerdol, M.; Canapa, A.; Forconi, M.; Olmo, E.; Pallavicini, A.; Barucca, M.; Schartl, M. The lungfish transcriptome: A glimpse into molecular evolution events at the transition from water to land. Sci. Rep. 2016, 6, 21571. [Google Scholar] [CrossRef] [PubMed]
- Dittrich, C.M.; Kratz, K.; Sendoel, A.; Gruenbaum, Y.; Jiricny, J.; Hengartner, M.O. LEM−3–A LEM domain containing nuclease involved in the DNA damage response in C. elegans. PLoS ONE 2012, 7, e24555. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Balsinde, J. Phospholipase A2; Cellular Regulation, Function, and Inhibition 2016. Available online: http://www.balsinde.org/publists/engplasic.pdf (accessed on 24 May 2018).
- Korbecki, J.; Baranowska-Bosiacka, I.; Gutowska, I.; Chlubek, D. The effect of reactive oxygen species on the synthesis of prostanoids from arachidonic acid. J. Physiol. Pharmacol. 2013, 64, 409–421. [Google Scholar] [PubMed]
- Kim, C.; Kim, J.-Y.; Kim, J.-H. Cytosolic phospholipase A2, lipoxygenase metabolites, and reactive oxygen species. BMB Rep. 2008, 41, 555–559. [Google Scholar] [CrossRef] [PubMed]
- Bierer, T.L.; Bui, L.M. Improvement of arthritic signs in dogs fed green-lipped mussel (Perna canaliculus). J. Nutr. 2002, 132, 1634S–1636S. [Google Scholar] [CrossRef] [PubMed]
- Zhao, M.; Brunk, U.T.; Eaton, J.W. Delayed oxidant-induced cell death involves activation of phospholipase A2. FEBS Lett. 2001, 509, 399–404. [Google Scholar] [CrossRef] [Green Version]
- Mukherjee, A.; Ghosal, S.; Maity, C. Lysosomal membrane stabilization by α-tocopherol against the damaging action of Vipera russelli venom phospholipase A2. Cell. Mol. Life Sci. 1997, 53, 152–155. [Google Scholar] [CrossRef] [PubMed]
- Romero-Geraldo, R.d.J.; García-Lagunas, N.; Hernández-Saavedra, N.Y. Crassostrea gigas exposure to the dinoflagellate Prorocentrum lima: Histological and gene expression effects on the digestive gland. Mar. Environ. Res. 2016, 120, 93–102. [Google Scholar] [CrossRef] [PubMed]
- Viarengo, A.; Marro, A.; Marchi, B.; Burlando, B. Single and combined effects of heavy metals and hormones on lysosomes of haemolymph cells from the mussel Mytilus galloprovincialis. Mar. Biol. 2000, 137, 907–912. [Google Scholar] [CrossRef]
- Marchi, B.; Burlando, B.; Moore, M.; Viarengo, A. Mercury- and copper-induced lysosomal membrane destabilisation depends on [Ca2+]i dependent phospholipase A2 activation. Aquat. Toxicol. 2004, 66, 197–204. [Google Scholar] [CrossRef] [PubMed]
- Núñez-Acuña, G.; Aballay, A.E.; Hégaret, H.; Astuya, A.P.; Gallardo-Escárate, C. Transcriptional responses of Mytilus chilensis exposed in vivo to saxitoxin (STX). J. Mollus. Stud. 2013, 79, 323–331. [Google Scholar] [CrossRef]
- Estrada, N.; de Jesús Romero, M.; Campa-Córdova, A.; Luna, A.; Ascencio, F. Effects of the toxic dinoflagellate, Gymnodinium catenatum on hydrolytic and antioxidant enzymes, in tissues of the giant lions-paw scallop Nodipecten subnodosus. Comp. Biochem. Phys. C Toxicol. Pharmacol. 2007, 146, 502–510. [Google Scholar] [CrossRef] [PubMed]
- Artigaud, S.; Lacroix, C.; Richard, J.; Flye-Sainte-Marie, J.; Bargelloni, L.; Pichereau, V. Proteomic responses to hypoxia at different temperatures in the great scallop (Pecten maximus). PeerJ 2015, 3, e871. [Google Scholar] [CrossRef] [PubMed]
- Detree, C.; Núñez-Acuña, G.; Roberts, S.; Gallardo-Escárate, C. Uncovering the complex transcriptome response of Mytilus chilensis against saxitoxin: Implications of harmful algal blooms on mussel populations. PLoS ONE 2016, 11, e0165231. [Google Scholar] [CrossRef] [PubMed]
- Venier, P.; Varotto, L.; Rosani, U.; Millino, C.; Celegato, B.; Bernante, F.; Lanfranchi, G.; Novoa, B.; Roch, P.; Figueras, A. Insights into the innate immunity of the mediterranean mussel Mytilus galloprovincialis. BMC Genom. 2011, 12, 69. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Domeneghetti, S.; Manfrin, C.; Varotto, L.; Rosani, U.; Gerdol, M.; De Moro, G.; Pallavicini, A.; Venier, P. How gene expression profiles disclose vital processes and immune responses in Mytilus spp. Invertebr. Surv. J. 2011, 8, 179–189. [Google Scholar]
- Romero, A.; Dios, S.; Poisa-Beiro, L.; Costa, M.M.; Posada, D.; Figueras, A.; Novoa, B. Individual sequence variability and functional activities of fibrinogen-related proteins (FREPs) in the mediterranean mussel (Mytilus galloprovincialis) suggest ancient and complex immune recognition models in invertebrates. Dev. Comp. Immunol. 2011, 35, 334–344. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cheng, C. Physico-Immunological Characterizations of Exogenous Substances (Palmitoleic Acid and Okadaic Acid) in Bivalves. Ph.D. Thesis, The Graduate School of Seoul National University, Seoul, Korea, 2017. [Google Scholar]
- Zhang, X.-J.; Qin, G.-M.; Yan, B.-L.; Xu, J.; Bi, K.-R.; Qin, L. Phenotypic and molecular characterization of pathogenic Listonella anguillarum isolated from half-smooth tongue sole Cynoglossus semilaevis. Acta Oceanol. Sin. 2009, 5, 012. [Google Scholar]
- Intecmar, Xunta de Galicia. Available online: http://www.intecmar.gal/ (accessed on 4 January 2015).
- EU-Harmonised Standard Operating Procedure for Determination of Lipophilic Marine Biotoxins in Molluscs by LC-MS/MS. Available online: http://www.aecosan.msssi.gob.es/CRLMB/docs/docs/metodos_analiticos_de_desarrollo/EU-Harmonised-SOP-LIPO-LCMSMS_Version5.pdf (accessed on 21 September 2018).
- Andrews, S. FastQC: A Quality Control Tool for High throughput Sequence Data. 2010, unpublished. Available online: https://www.bioinformatics.babraham.ac.uk/projects/fastqc/ (accessed on 7 October 2015).
- Schmieder, R.; Edwards, R. Quality control and preprocessing of metagenomic datasets. Bioinformatics 2011, 27, 863–864. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gordon, A.; Hannon, G. Fastx-Toolkit. FASTQ/A Short-Reads Preprocessing Tools. 2010, unpublished. Available online: http://hannonlab.cshl.edu/fastx_toolkit/ (accessed on 16 October 2015).
- Love, M.I.; Huber, W.; Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [PubMed]
- Team, R. RStudio: Integrated Development for R; RStudio, Inc.: Boston, MA, USA, 2015; Available online: http://www.rstudio.com (accessed on 27 October 2015).
- Benjamini, Y.; Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. B Methodol. 1995, 57, 289–300. [Google Scholar]
- Conesa, A.; Götz, S.; García-Gómez, J.M.; Terol, J.; Talón, M.; Robles, M. Blast2GO: A universal tool for annotation, visualization and analysis in functional genomics research. Bioinformatics 2005, 21, 3674–3676. [Google Scholar] [CrossRef] [PubMed]
- Götz, S.; García-Gómez, J.M.; Terol, J.; Williams, T.D.; Nagaraj, S.H.; Nueda, M.J.; Robles, M.; Talón, M.; Dopazo, J.; Conesa, A. High-throughput functional annotation and data mining with the Blast2GO suite. Nucleic Acids Res. 2008, 36, 3420–3435. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Finn, R.D.; Mistry, J.; Schuster-Böckler, B.; Griffiths-Jones, S.; Hollich, V.; Lassmann, T.; Moxon, S.; Marshall, M.; Khanna, A.; Durbin, R. Pfam: Clans, web tools and services. Nucleic Acids Res. 2006, 34, D247–D251. [Google Scholar] [CrossRef] [PubMed]
- CIMminer. Available online: http://discover.nci.nih.gov/cimminer (accessed on 19 June 2018).
- Cubero-Leon, E.; Ciocan, C.M.; Minier, C.; Rotchell, J.M. Reference gene selection for qPCR in mussel, Mytilus edulis, during gametogenesis and exogenous estrogen exposure. Environ. Sci. Pollut. Res. Int. 2012, 19, 2728–2733. [Google Scholar] [CrossRef] [PubMed]
- Universal ProbeLibrary. Available online: https://lifescience.roche.com/en_es/brands/universal-probe-library.html#assay-design-centre (accessed on 20 July 2018).
- Pfaffl, M.W.; Tichopad, A.; Prgomet, C.; Neuvians, T.P. Determination of stable housekeeping genes, differentially regulated target genes and sample integrity: Bestkeeper–Excel-based tool using pair-wise correlations. Biotechnol. Lett. 2004, 26, 509–515. [Google Scholar] [CrossRef] [PubMed]
- Xie, F.; Xiao, P.; Chen, D.; Xu, L.; Zhang, B. miRDeepFinder: A miRNA analysis tool for deep sequencing of plant small RNAs. Plant Mol. Biol. 2012, 80, 75–84. [Google Scholar] [CrossRef] [PubMed]
- Hellemans, J.; Mortier, G.; De Paepe, A.; Speleman, F.; Vandesompele, J. qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol. 2007, 8, R19. [Google Scholar] [CrossRef] [PubMed]
- Ruijter, J.; Ramakers, C.; Hoogaars, W.; Karlen, Y.; Bakker, O.; Van den Hoff, M.; Moorman, A. Amplification efficiency: Linking baseline and bias in the analysis of quantitative PRC data. Nucleic Acids Res. 2009, 37, e45. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W. A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res. 2001, 29, e45. [Google Scholar] [CrossRef] [PubMed]
- Pfaffl, M.W.; Horgan, G.W.; Dempfle, L. Relative expression software tool (REST©) for group-wise comparison and statistical analysis of relative expression results in real-time PCR. Nucleic Acids Res. 2002, 30, e36. [Google Scholar] [CrossRef] [PubMed]







| Total number of contigs | 95,702 | L25 | 1682 bp |
| Total length | 71,623.079 Kb | N50 | 21,152 |
| Maximum contig length | 16,082 Kb | L50 | 1062 bp |
| Minimum contig length | 102 pb | N75 | 42,376 |
| Average contig length | 748 bp | L75 | 668 bp |
| N25 | 7537 | %GC | 33.20% |
| Sequence ID | Description | Length (bp) | baseMean | Log2FC | FC | p-Value | Adjusted p-Value |
|---|---|---|---|---|---|---|---|
| ci|000006456|Bact|Sample_MGT2|2 | cytochrome c oxidase subunit 1, partial | 910 | 23,389.22 | 7.09 | 136.29 | 1.96 × 10112 | 1.70 × 10−107 |
| ci|000001182|Bact|Sample_MBT2|2 | * ATP-synt_A | 578 | 3975.18 | 7.21 | 147.76 | 6.65 × 10 −66 | 1.92 × 10−61 |
| Contig39610 | ribosomal protein L23a, partial | 1166 | 2205.27 | 7.92 | 243.02 | 3.89 × 10−43 | 6.71 × 10−39 |
| ci|000005084|Bact|Sample_MGT2|2 | cytochrome c oxidase subunit I | 1848 | 12,611.71 | 6.23 | 75.08 | 1.91 × 10−37 | 2.75 × 10−33 |
| Contig34888 | NA | 529 | 608.03 | 8.47 | 355.72 | 4.04 × 10−35 | 4.98 × 10−31 |
| ci|000015505|Bact|Sample_MGT1|2 | NA | 588 | 1046.84 | 5.05 | 33.22 | 9.05 × 10−35 | 9.78 × 10−31 |
| ci|000014133|Bact|Sample_MGT2|2 | * Glyco_hydro_16 | 949 | 411.92 | 6.95 | 124.04 | 3.09 × 10−34 | 2.97 × 10−30 |
| Contig22742 | NA | 1165 | 944.38 | 6.61 | 97.63 | 4.80 × 10−33 | 4.14 × 10−29 |
| Contig33832 | Kazal-like serine protease inhibitor domain-containing protein | 507 | 514.55 | 6.70 | 103.84 | 6.63 × 10−33 | 5.20 × 10−29 |
| ci|000004031|Bact|Sample_MGT1|2 | * Porin_3 | 1024 | 888.45 | 5.19 | 36.44 | 3.14 × 10−31 | 2.09 × 10−27 |
| ci|000016700|Bact|Sample_MGT1|2 | † COX1_MYTED | 750 | 12,157.03 | 6.86 | 116.52 | 2.32 × 10−30 | 1.43 × 10−26 |
| ci|000022316|Bact|Sample_MBT1|2 | * Ribosomal_L23 | 340 | 280.82 | 6.68 | 102.35 | 2.26 × 10−28 | 1.15 × 10−24 |
| Contig17884 | PREDICTED: 60 kDa SS-A/Ro ribonucleoprotein | 1726 | 227.65 | 5.51 | 45.57 | 6.21 × 10−25 | 2.68 × 10−21 |
| ci|000012420|Bact|Sample_MGT2|2 | NA | 560 | 761.81 | 9.97 | 1004.61 | 2.04 × 10−24 | 8.01 × 10−21 |
| ci|000010593|Bact|Sample_MBC1|2 | * Ribosomal_L7Ae | 458 | 403.47 | 8.29 | 313.91 | 6.26 × 10−23 | 2.16 × 10−19 |
| ci|000001186|Bact|Sample_MGT2|2 | NA | 316 | 209.87 | 6.26 | 76.55 | 7.11 × 10−23 | 2.36 × 10−19 |
| ci|000001089|Bact|Sample_MGT2|2 | NA | 1106 | 224.57 | 5.35 | 40.77 | 1.03 × 10−20 | 3.08 × 10−17 |
| Contig35276 | NA | 420 | 188.91 | 7.45 | 174.32 | 6.35 × 10−20 | 1.77 × 10−16 |
| Contig38903 | * Myticin-prepro | 506 | 510.91 | 9.28 | 623.48 | 1.02 × 10−19 | 2.68 × 10−16 |
| ci|000022507|Bact|Sample_MBT2|2 | * Ribosomal_S9 | 1568 | 918.24 | 5.20 | 36.82 | 3.34 × 10−19 | 7.79 × 10−16 |
| ci|000000480|Bact|Sample_MGT2|2 | * Astacin | 858 | 563.33 | 6.67 | 101.71 | 4.63 × 10−19 | 1.00 × 10−15 |
| ci|000018470|Bact|Sample_MGT1|2 | * Lectin_C | 633 | 186.73 | 7.90 | 238.40 | 5.47 × 10−19 | 1.13 × 10−15 |
| ci|000004710|Bact|Sample_MBC2|2 | NA | 1395 | 219.14 | 6.06 | 66.62 | 1.35 × 10−18 | 2.66 × 10−15 |
| ci|000008308|Bact|Sample_MGT1|2 | NA | 464 | 637.37 | 5.04 | 32.92 | 1.82 × 10−18 | 3.49 × 10−15 |
| ci|000004147|Bact|Sample_MGT2|2 | NA | 622 | 507.89 | 6.81 | 112.02 | 2.47 × 10−18 | 4.65 × 10−15 |
| Sequence ID | Description | Length (bp) | baseMean | Log2FC | FC | p-Value | Adjusted p-Value |
|---|---|---|---|---|---|---|---|
| ci|000007816|Bact|Sample_MGC1|2 | NA | 539 | 7514.26 | −7.48 | −178.70 | 1.66 × 10−82 | 7.17 × 10−78 |
| Contig22552 | NA | 622 | 131,251.75 | −6.93 | −121.52 | 5.49 × 10−60 | 1.18 × 10−55 |
| Contig26868 | NADH dehydrogenase subunit 5, partial | 719 | 1916.17 | −6.96 | −124.62 | 2.16 × 10−32 | 1.56 × 10−28 |
| Contig28135 | 40S ribosomal protein S10-like | 559 | 406.63 | −5.12 | −34.81 | 5.46 × 10−29 | 3.14 × 10−25 |
| Contig30578 | * DUF1082 | 529 | 3132.77 | −9.97 | −1005.57 | 2.05 × 10−28 | 1.11 × 10−24 |
| Contig28105 | * SRCR | 1419 | 329.84 | −7.74 | −213.17 | 2.60 × 10−26 | 1.25 × 10−22 |
| ci|000000372|Bact|Sample_MGC1|2 | NA | 723 | 851.06 | −6.15 | −70.81 | 7.19 × 10−26 | 3.27 × 10−22 |
| ci|000009048|Bact|Sample_MBC1|2 | NA | 703 | 359.37 | −8.39 | −334.64 | 1.14 × 10−24 | 4.67 × 10−21 |
| Contig26906 | NA | 530 | 286.75 | −8.12 | −279.10 | 1.47 × 10−23 | 5.52 × 10−20 |
| ci|000018684|Bact|Sample_MBC2|2 | * Cytochrom_B_N_2 | 643 | 2708.10 | −7.79 | −221.90 | 5.93 × 10−23 | 2.13 × 10−19 |
| ci|000000728|Bact|Sample_MGC2|2 | NA | 768 | 510.67 | −9.66 | −810.78 | 8.67 × 10−23 | 2.77 × 10−19 |
| Contig29976 | uncharacterized protein LOC567525 isoform X1/* Fibrinogen_C | 1089 | 167.09 | −6.83 | −114.10 | 6.31 × 10−21 | 1.95 × 10−17 |
| ci|000000734|Bact|Sample_MGC2|2 | * Zona_pellucida | 1185 | 255.85 | −9.02 | −518.06 | 4.16 × 10−20 | 1.20 × 10−16 |
| Contig26843 | NA | 984 | 230.17 | −4.23 | −18.76 | 7.68 × 10−20 | 2.07 × 10−16 |
| ci|000002253|Bact|Sample_MGC2|2 | PREDICTED: GTPase IMAP family member 7/* AIG1 | 1188 | 255.05 | −7.83 | −228.11 | 1.33 × 10−19 | 3.37 × 10−16 |
| ci|000003979|Bact|Sample_MGC1|2 | NA | 1136 | 233.95 | −8.88 | −471.60 | 2.94 ×10−19 | 7.25 × 10−16 |
| ci|000021317|Bact|Sample_MBC1|2 | NA | 834 | 1818.89 | −6.97 | −125.28 | 3.04 × 10−19 | 7.29 × 10−16 |
| ci|000008655|Bact|Sample_MGC2|2 | NA | 374 | 1501.18 | −8.07 | −267.93 | 4.48 × 10−19 | 1.00 × 10−15 |
| ci|000004674|Bact|Sample_MGC2|2 | Perlucin | 660 | 174.13 | −5.61 | −48.70 | 4.57 × 10−19 | 1.00 × 10−15 |
| ci|000023153|Bact|Sample_MBT1|2 | * COX1 | 605 | 4253.61 | −6.85 | −115.03 | 5.47 × 10−19 | 1.13 × 10−15 |
| ci|000001983|Bact|Sample_MBC2|2 | * KOW | 607 | 4924.53 | −2.78 | −6.88 | 8.86 × 10−19 | 1.78 × 10−15 |
| ci|000005149|Bact|Sample_MGC1|2 | * TIG | 612 | 148.86 | −6.00 | −64.19 | 2.90 × 10−17 | 5.11 × 10−14 |
| ci|000009215|Bact|Sample_MGC2|2 | * Glyco_hydro_10 | 946 | 136.75 | −7.11 | −138.62 | 4.08 × 10−17 | 7.06 × 10−14 |
| Contig28020 | NA | 570 | 166.31 | −8.45 | −350.56 | 4.96 × 10−17 | 8.24 × 10−14 |
| ci|000015516|Bact|Sample_MBT2|2 | * Ribosomal_L22 | 1338 | 1356.05 | −4.92 | −30.37 | 1.19 × 10−16 | 1.87 × 10−13 |
| Sequence ID | Description | Length (bp) | baseMean | Log2FC | FC | p-Value | Adjusted p-Value |
|---|---|---|---|---|---|---|---|
| ci|000029194|Bact|Sample_MBT1|2 | * EF-hand_1 and 7 | 508 | 3570.44 | 9.76 | 868.30 | 7.36 × 10−98 | 6.48 × 10−93 |
| ci|000006043|Bact|Sample_MBT1|2 | NA | 471 | 4432.18 | 6.05 | 66.47 | 4.32 × 10−60 | 1.90 × 10−55 |
| ci|000001929|Bact|Sample_MBT2|2 | NA | 690 | 803.53 | 7.61 | 195.15 | 4.55 × 10−49 | 1.00 × 10−44 |
| ci|000002899|Bact|Sample_MBT1|2 | NA | 779 | 1066.24 | 9.48 | 715.09 | 9.38 × 10−40 | 1.65 × 10−35 |
| Contig35833 | NA | 944 | 520.96 | 8.45 | 350.51 | 6.42 × 10−29 | 6.28 × 10−25 |
| ci|000022507|Bact|Sample_MBT2|2 | NADH dehydrogenase subunit 6 | 1568 | 475.01 | 3.90 | 14.88 | 1.10 × 10−28 | 9.70 × 10−25 |
| ci|000017597|Bact|Sample_MBT1|2 | * Antistasin | 795 | 266.99 | 7.13 | 140.36 | 1.22 × 10−28 | 9.79 × 10−25 |
| ci|000007496|Bact|Sample_MBT2|2 | NA | 745 | 425.52 | 6.39 | 84.07 | 4.01 × 10−28 | 2.94 × 10−24 |
| ci|000020755|Bact|Sample_MBT2|2 | † NU4M_MYTED | 1483 | 849.56 | 5.15 | 35.44 | 1.44 × 10−26 | 9.05 × 10−23 |
| ci|000025759|Bact|Sample_MBT2|2 | NA | 2007 | 257.03 | 7.56 | 188.10 | 1.81 × 10−26 | 1.06 × 10−22 |
| Contig39610 | * Ribosomal_L23 | 1166 | 453.85 | 6.46 | 87.80 | 3.94 × 10−26 | 2.16 × 10−22 |
| Contig15942 | NA | 482 | 916.44 | 5.84 | 57.29 | 6.24 × 10−26 | 3.23 × 10−22 |
| ci|000001411|Bact|Sample_MGC2|2 | * HSBP1 | 580 | 421.11 | 4.55 | 23.41 | 1.82 × 10−25 | 8.92 × 10−22 |
| ci|000005084|Bact|Sample_MGT2|2 | * COX1 | 1848 | 3923.98 | 3.98 | 15.75 | 3.81 × 10−25 | 1.77 × 10−21 |
| ci|000003417|Bact|Sample_MBT2|2 | * Phospholip_A2_1 | 562 | 203.18 | 6.08 | 67.55 | 1.10 × 10−23 | 4.05 × 10−20 |
| Contig20144 | NA | 2258 | 205.66 | 3.81 | 14.01 | 4.48 × 10−23 | 1.58 × 10−19 |
| ci|000019916|Bact|Sample_MBT1|2 | NA | 768 | 241.64 | 7.15 | 141.88 | 1.42 × 10−22 | 4.45 × 10−19 |
| ci|000001302|Bact|Sample_MBT1|2 | NA | 675 | 281.74 | 6.61 | 97.65 | 1.36 × 10−21 | 4.00 × 10−18 |
| ci|000018492|Bact|Sample_MBT1|2 | NA | 1552 | 159.56 | 5.50 | 45.19 | 1.59 × 10−21 | 4.36 × 10−18 |
| Contig13066 | Calcyphosin-like protein | 2325 | 853.76 | 3.68 | 12.85 | 2.01 × 10−21 | 5.35 × 10−18 |
| ci|000018122|Bact|Sample_MBT2|2 | * HYR and TMEM154 | 3321 | 2042.07 | 4.34 | 20.27 | 2.30 × 10−21 | 5.95 ×10−18 |
| Contig12937 | † RS27L_HUMAN | 2183 | 321.15 | 7.71 | 210.04 | 3.66 × 10−21 | 9.21 × 10−18 |
| ci|000000451|Bact|Sample_MGT2|2 | NA | 655 | 587.43 | 2.92 | 7.57 | 1.42 × 10−20 | 3.38 × 10−17 |
| ci|000003122|Bact|Sample_MBT2|2 | NA | 1513 | 215.45 | 7.54 | 185.87 | 4.85 × 10−20 | 1.12 × 10−16 |
| Contig40138 | NA | 584 | 136.63 | 6.17 | 72.15 | 7.64 × 10−20 | 1.72 × 10−16 |
| Sequence ID | Description | Length (bp) | baseMean | Log2FC | FC | p-Value | Adjusted p-Value |
|---|---|---|---|---|---|---|---|
| ci|000007038|Bact|Sample_MBC2|2 | low-density lipoprotein receptor-related protein 8 isoform X1 | 689 | 1321.99 | −9.45 | −700.32 | 1.06 × 10−53 | 3.11 × 10−49 |
| Contig3681 | NA | 895 | 1561.88 | −10.25 | −1216.97 | 9.98 × 10−37 | 1.46 × 10−32 |
| Contig11592 | NA | 798 | 652.29 | −9.55 | −752.01 | 4.28 × 10−31 | 5.38 × 10−27 |
| Contig1183 | NA | 581 | 802.66 | −4.76 | −27.10 | 9.28 × 10−30 | 1.02 × 10−25 |
| Contig8105 | NA | 1717 | 199.06 | −6.43 | −86.51 | 3.62 × 10−27 | 2.45 × 10−23 |
| ci|000015242|Bact|Sample_MGT2|2 | NA | 663 | 429.13 | −7.24 | −151.36 | 4.36 × 10−25 | 1.92 × 10−21 |
| ci|000005973|Bact|Sample_MBC1|2 | NA | 1860 | 257.90 | −4.59 | −24.07 | 5.34 × 10−25 | 2.24 × 10−21 |
| Contig10936 | * Oxidored_q1 | 2797 | 18,810.53 | −1.67 | −3.18 | 3.68 × 10−24 | 1.47 × 10−20 |
| ci|000000312|Bact|Sample_MBC1|2 | NA | 972 | 208.97 | −7.32 | −159.95 | 4.63 × 10−24 | 1.77 × 10−20 |
| Contig6277 | NA | 702 | 2009.78 | −10.35 | −1303.40 | 6.01 × 10−23 | 2.04 × 10−19 |
| ci|000016192|Bact|Sample_MBC1|2 | * Ldl_recept_a and PRKCSH-like | 946 | 4008.01 | −2.65 | −6.26 | 9.71 × 10−23 | 3.17 × 10−19 |
| Contig3876 | Predicted protein | 536 | 295.82 | −8.43 | −344.34 | 5.01 × 10−22 | 1.52 × 10−18 |
| Contig4774 | neurocalcin homolog | 1267 | 339.45 | −4.83 | −28.39 | 1.49 × 10−21 | 4.22 × 10−18 |
| ci|000000823|Bact|Sample_MBC2|2 | NA | 803 | 325.92 | −3.99 | −15.91 | 1.06 × 10−20 | 2.59 × 10−17 |
| Contig6059 | NA | 486 | 5331.78 | −1.93 | −3.82 | 8.13 × 10−20 | 1.76 × 10−16 |
| Contig7283 | cytochrome c oxidase subunit I | 2879 | 132.24 | −5.09 | −34.06 | 9.51 × 10−20 | 1.99 × 10−16 |
| ci|000015433|Bact|Sample_MBC1|2 | cytochrome c oxidase subunit I | 1136 | 17,057.57 | −3.52 | −11.49 | 3.00 × 10−19 | 5.74 × 10−16 |
| ci|000004320|Bact|Sample_MBC1|2 | * Lipoxygenase | 1950 | 564.59 | −9.46 | −703.80 | 3.48 × 10−19 | 6.24 × 10−16 |
| ci|000001144|Bact|Sample_MBC2|2 | NA | 434 | 221.23 | −8.61 | −389.88 | 4.06 × 10−19 | 7.15 × 10−16 |
| ci|000008127|Bact|Sample_MBC2|2 | NA | 2503 | 364.67 | −10.03 | −1043.76 | 7.20 × 10−19 | 1.22 × 10−15 |
| ci|000005247|Bact|Sample_MBC1|2 | NA | 1246 | 153.48 | −6.35 | −81.80 | 1.26 × 10−18 | 2.09 × 10−15 |
| ci|000001610|Bact|Sample_MBC1|2 | NA | 1057 | 341.22 | −9.94 | −982.30 | 1.93 × 10−18 | 3.11 × 10−15 |
| ci|000000874|Bact|Sample_MBC2|2 | NA | 697 | 153.63 | −5.02 | −32.42 | 5.21 × 10−18 | 7.77 × 10−15 |
| ci|000002263|Bact|Sample_MBC1|2 | * Pfam-B_5682 | 1222 | 151.49 | −4.77 | −27.37 | 6.23 × 10−18 | 8.99 × 10−15 |
| ci|000003990|Bact|Sample_MGT2|2 | NA | 521 | 310.73 | −9.76 | −866.01 | 2.46 × 10−17 | 3.23 × 10−14 |
| Category | Number of Genes | p-Value |
|---|---|---|
| PF04548.11//AIG1 | 26 | 0.00248912 |
| PF01926.18//MMR_HSR1 | 25 | 0.0029543 |
| PF00059.16//Lectin_C | 21 | 0.01366918 |
| PF00100.18//Zona_pellucida | 16 | 0.00403746 |
| PF13499.1//EF-hand_7 | 14 | 0.00221868 |
| PF13405.1//EF-hand_6 | 14 | 0.00355134 |
| PF00036.27//EF-hand_1 | 13 | 0.00065889 |
| PF13202.1//EF-hand_5 | 13 | 0.02872925 |
| PF13833.1//EF-hand_8 | 12 | 0.02032022 |
| PF00361.15//Oxidored_q1 | 10 | 0.00489835 |
| PF00119.15//ATP-synt_A | 8 | 0.00023995 |
| PF10690.4//Myticin-prepro | 8 | 0.02237525 |
| PF07679.11//I-set | 7 | 0.04744078 |
| Category | Number of Genes | p-Value |
|---|---|---|
| PF00386.16//C1q | 36 | 5.2 × 10−8 |
| PF00036.27//EF-hand_1 | 31 | 0.00035296 |
| PF13499.1//EF-hand_7 | 29 | 0.00014495 |
| PF13405.1//EF-hand_6 | 27 | 8.61 × 10−5 |
| PF00147.13//Fibrinogen_C | 25 | 0.01665835 |
| PF13202.1//EF-hand_5 | 23 | 0.00079724 |
| PF13833.1//EF-hand_8 | 20 | 0.00015502 |
| PF10690.4//Myticin-prepro | 13 | 0.01435613 |
| PF00361.15//Oxidored_q1 | 13 | 0.03222834 |
| PF07679.11//I-set | 9 | 0.00010238 |
| PF09458.5//H_lectin | 9 | 0.00621282 |
| PF01607.19//CBM_14 | 9 | 0.02592731 |
| PB002965//Pfam-B_2965 | 9 | 0.03289021 |
| PF13895.1//Ig_2 | 8 | 0.00039907 |
| PF00119.15//ATP-synt_A | 8 | 0.01065053 |
| PF13927.1//Ig_3 | 7 | 0.00090571 |
| PF00092.23//VWA | 7 | 0.00272518 |
| PF07686.12//V-set | 7 | 0.01729404 |
| PF03281.9//Mab-21 | 7 | 0.03056683 |
| Gene Name | Abbreviation | Reference | E | Amplicon Size (bp) | Tm (°C) | Primers 5′→3′ |
|---|---|---|---|---|---|---|
| Tropomyosin | TPM | ab000907.1 | 1.90 | 67 | F-55.3 R-57.1 | F-GATGCTGAAAATCGTGCAAC R-CGGTCTACTTCTTTTTGCAACTT |
| Ribosomal proteins S4 | rpS4 | Lozano et al. (2015) | 1.83 | 138 | F-58.8 R-60.3 | F-TGGGTTATCGAGGGCGTAG R-TCCCTTAGTTTGTTGAGGACCTG |
| 18S ribosomal RNA | 18S | L33452.1 | 1.86 | 60 | F-58.3 R-55.9 | F-CCTGGAAAGGTCGGGTAAC R-AATTACAAGCCCCAATCCCTA |
| 18S ribosomal RNA | 18S-L33448 | Cubero-Leon et al. (2012) | 1.79 | 114 | F-56.3 R-56.0 | F-CATTAGTCAAGAACGAAAGTCAGAG R-GCCTGCCGAGTCATTGAAG |
| Glyceraldehyde 3-phosphate-dehydrogenase | GAPDH | Lozano et al. (2015) | 1.92 | 114 | F-59.4 R-58.4 | F-AGGAATGGCCTTCAGGG R-TCAGATGCTGCTTTAATGGCTG |
| Elongation Factor 1 | EF1 | Suarez-Ulloa et al. (2013) | 1.89 | 106 | F-55.8 R-57.0 | F-CCTCCCACCATCAAGACCTA R-GGCTGGAGCAAAGGTAACAA |
| Big defensin 2 | BD2 | Contig37896 | 1.83 | 110 | F-60.3 R-59.3 | F-TCTGAGCAGGGAGTATCAACAG R-TGGACAAAACAGCTACTAACAAGG |
| NADH dehidrogenase subunit 5 | NADH5 | Contig24266 | 1.86 | 90 | F-53.7 R-56.5 | F-GCAGTCATGCGCAAAAAG R-ACCCGGTACAAATATGGCTAAA |
| Dynactin-subunit-6-like | DYNA | Contig14551 | 1.89 | 60 | F-58.9 R-58.9 | F-AGTATTCTCAGGCATGGTTTCTG R-GGTTGTATAATTGGAGGCATGTG |
| GIY-YIG domain containing protein | GIY-YIG DC | ci|000000744|Bact|Sample_MBC1|2 | 1.83 | 70 | F-57.6 R-55.3 | F-AATCTACCAATTGCTTGTCTGTCA R-CGAAACGTAGTGTGCGAAAA |
| KAZAL domain containing protein | KAZAL DC | Contig33832 | 1.91 | 60 | F-53.2 R-60.3 | F-ATAATCGGCAGTGCAAAACA R-TTCCTTACTGAGTCAGTCG |
| Cytosolic phospholipase A-2 like | CPLA2 | ci|000001655|Bact|Sample_MBT1|2 | 1.80 | 73 | F-61.6 R-57.1 | F-CCTGTACTGTGAGATTAGGTTATTGC R-CAGAAGGTTATTGACCGAAAGAA |
| Arachidonate 15-lipoxygenase B-like | ALOX15B | ci|000023941|Bact|Sample_MBT2|2 | 1.81 | 94 | F-58.5 R-55.9 | F-TGTTGTGAGTGAAGCAATAACTCTAA R-CGGAATAAATCG AGAGAACCA |
| Alpha-L-fucosidase-like | FUCA | ci|000010451|Bact|Sample_MBT1|2 | 1.87 | 74 | F-61.0 R-55.3 | F-GGAATTCCAGTAGGAATCAGTAGC R-TGGTAAATGCATACAAACCTGAA |
| H_Lectin domain containing protein | H_Lectin DC | Contig19341 | 1.85 | 73 | F-56.5 R-55.3 | F-CCCTTCTTTGCTTTAGATGCTT R-TTGATGGCCAGATTACGACA |
| Fibrinogen_C domain containing protein | Fibrinogen_C DC | ci|000024772|Bact|Sample_MBC1|2 | 1.86 | 67 | F-57.3 R-59.4 | F-AAGGTTGTCTCCAGCGTTTC R-CGGTGATGCCTCTACCAACT |
| Rank | Normfinder | Stability | BestKeeper | SD | r |
|---|---|---|---|---|---|
| 1 | rpS4 | 0.07 | rpS4 | 0.46 | 0.732 |
| 2 | TPM | 0.17 | TPM | 0.50 | 0.448 |
| 3 | GAPDH | 0.20 | GAPDH | 0.64 | 0.669 |
| 4 | 18S | 0.37 | 18S | 0.71 | 0.827 |
| 5 | 18S-L33448 | 0.76 | 18S-L33448 | 1.08 | |
| 6 | EF1 | 1.78 | EF1 | 2.91 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Prego-Faraldo, M.V.; Martínez, L.; Méndez, J. RNA-Seq Analysis for Assessing the Early Response to DSP Toxins in Mytilus galloprovincialis Digestive Gland and Gill. Toxins 2018, 10, 417. https://doi.org/10.3390/toxins10100417
Prego-Faraldo MV, Martínez L, Méndez J. RNA-Seq Analysis for Assessing the Early Response to DSP Toxins in Mytilus galloprovincialis Digestive Gland and Gill. Toxins. 2018; 10(10):417. https://doi.org/10.3390/toxins10100417
Chicago/Turabian StylePrego-Faraldo, María Verónica, Luisa Martínez, and Josefina Méndez. 2018. "RNA-Seq Analysis for Assessing the Early Response to DSP Toxins in Mytilus galloprovincialis Digestive Gland and Gill" Toxins 10, no. 10: 417. https://doi.org/10.3390/toxins10100417