Identification of Host Micro RNAs That Differentiate HIV-1 and HIV-2 Infection Using Genome Expression Profiling Techniques
Abstract
:1. Introduction
2. Materials and Methods
2.1. Peripheral Blood Mononuclear Blood Cells Isolation and Cell Culture
2.2. Viruses and Infection
2.3. RNA Extraction
2.4. miRNA Profiling Using RT2 MicroRNA PCR Array
2.5. PCR Array Data Analysis
2.6. Validation of Differentially Expressed miRNA
2.7. miRNA Targeted Gene Prediction
2.8. Quantification of Predicted Target Genes by RT-PCR
3. Results
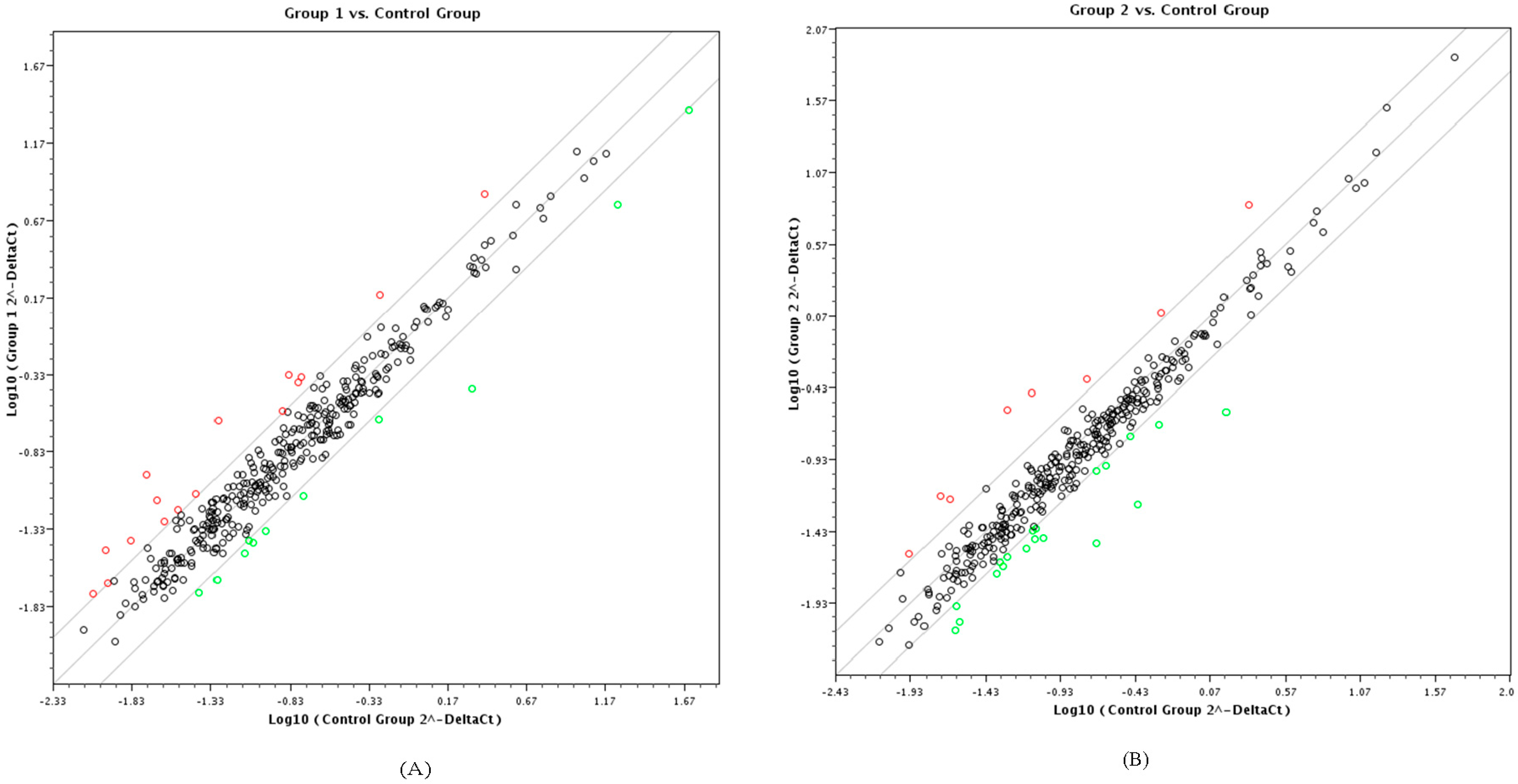
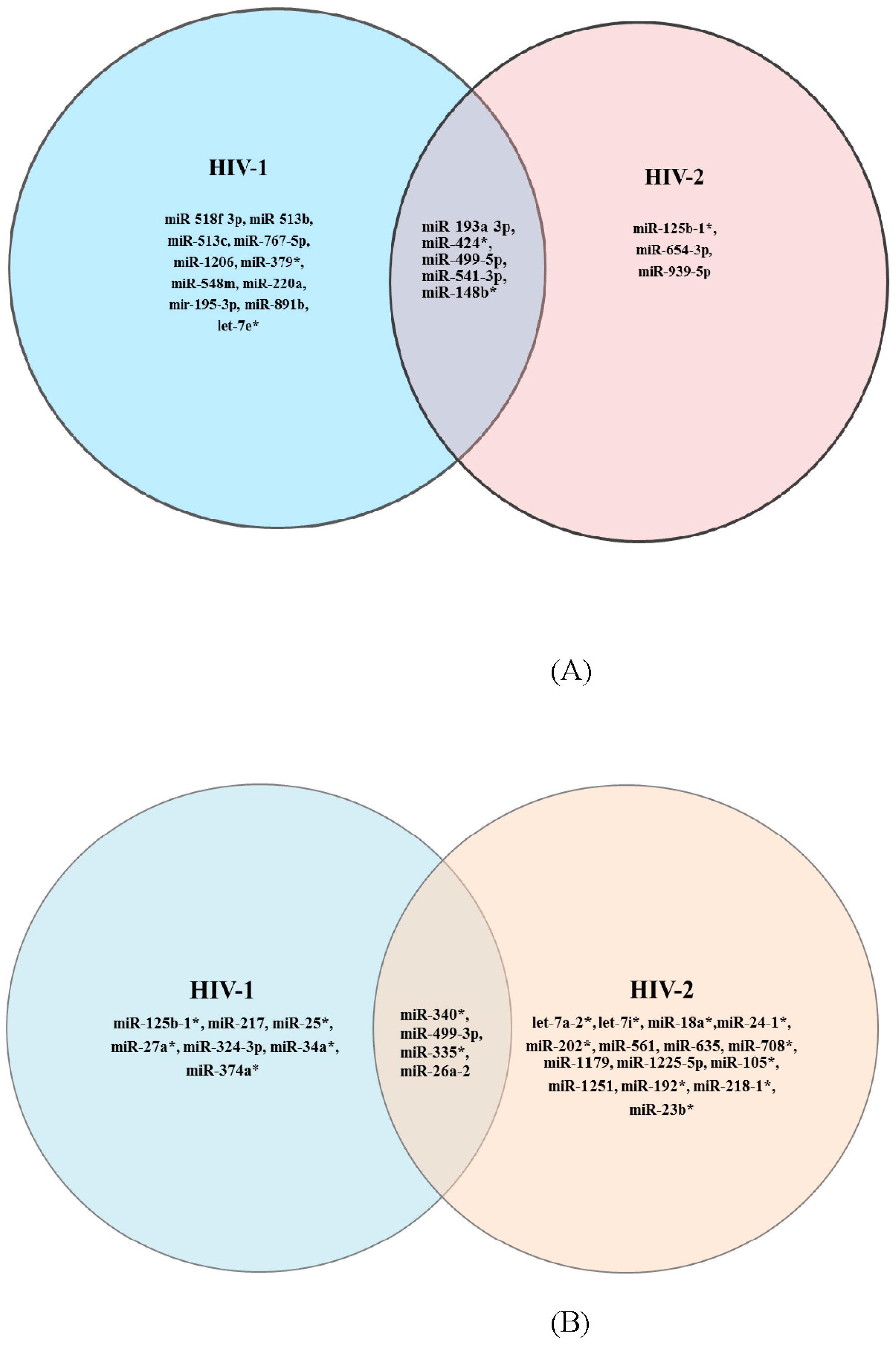
3.1. PCR-Array and Differential miRNA Expression Analysis
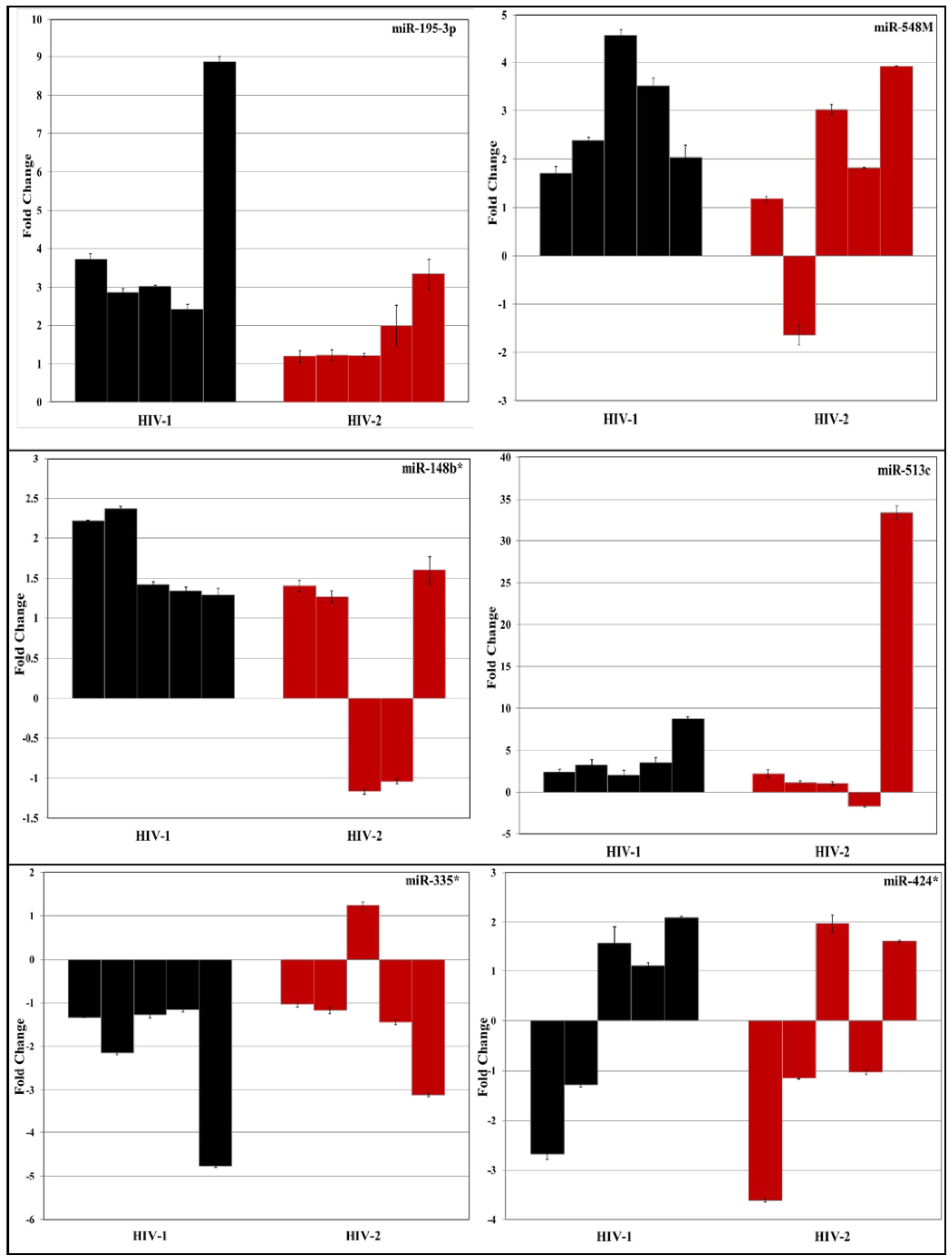
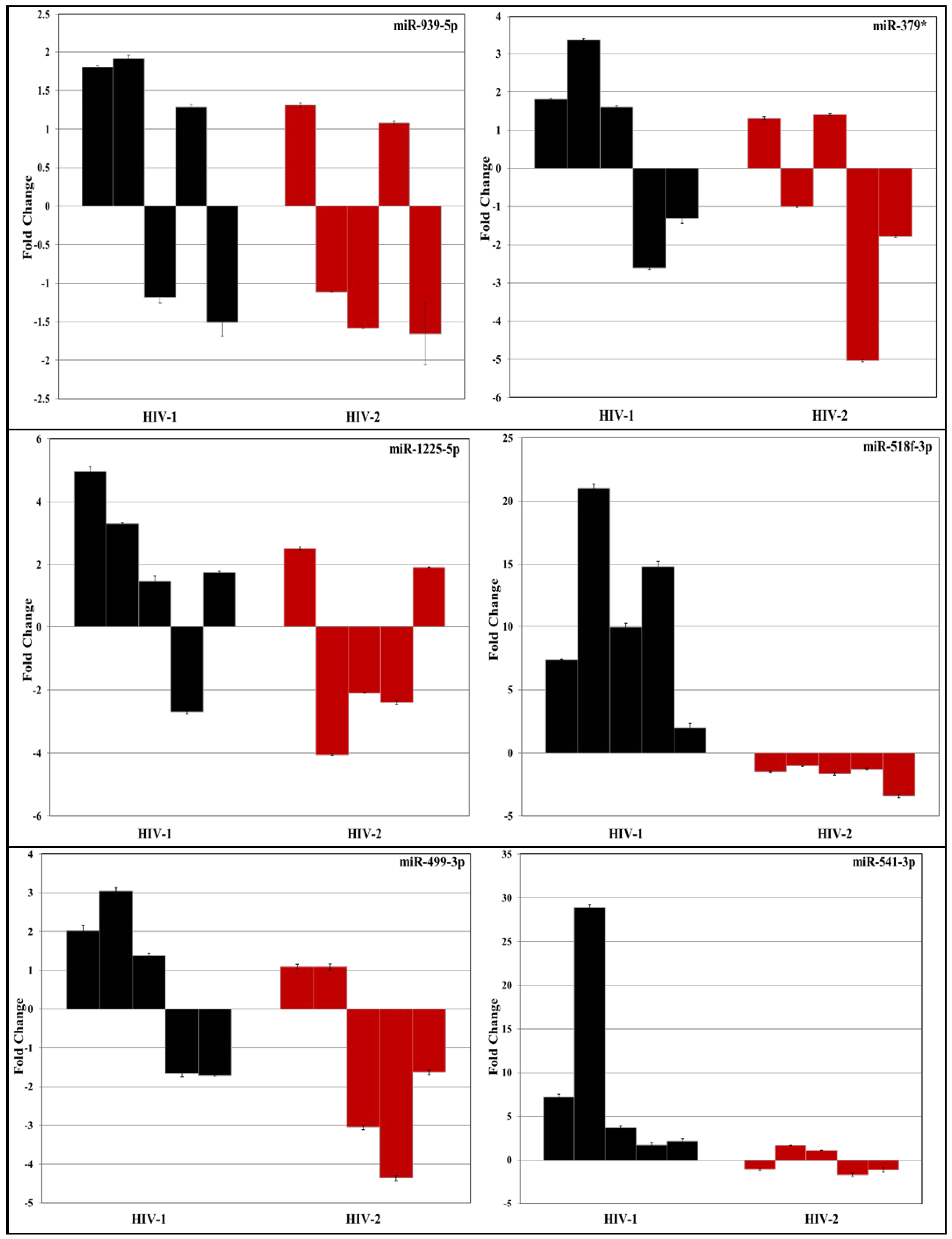
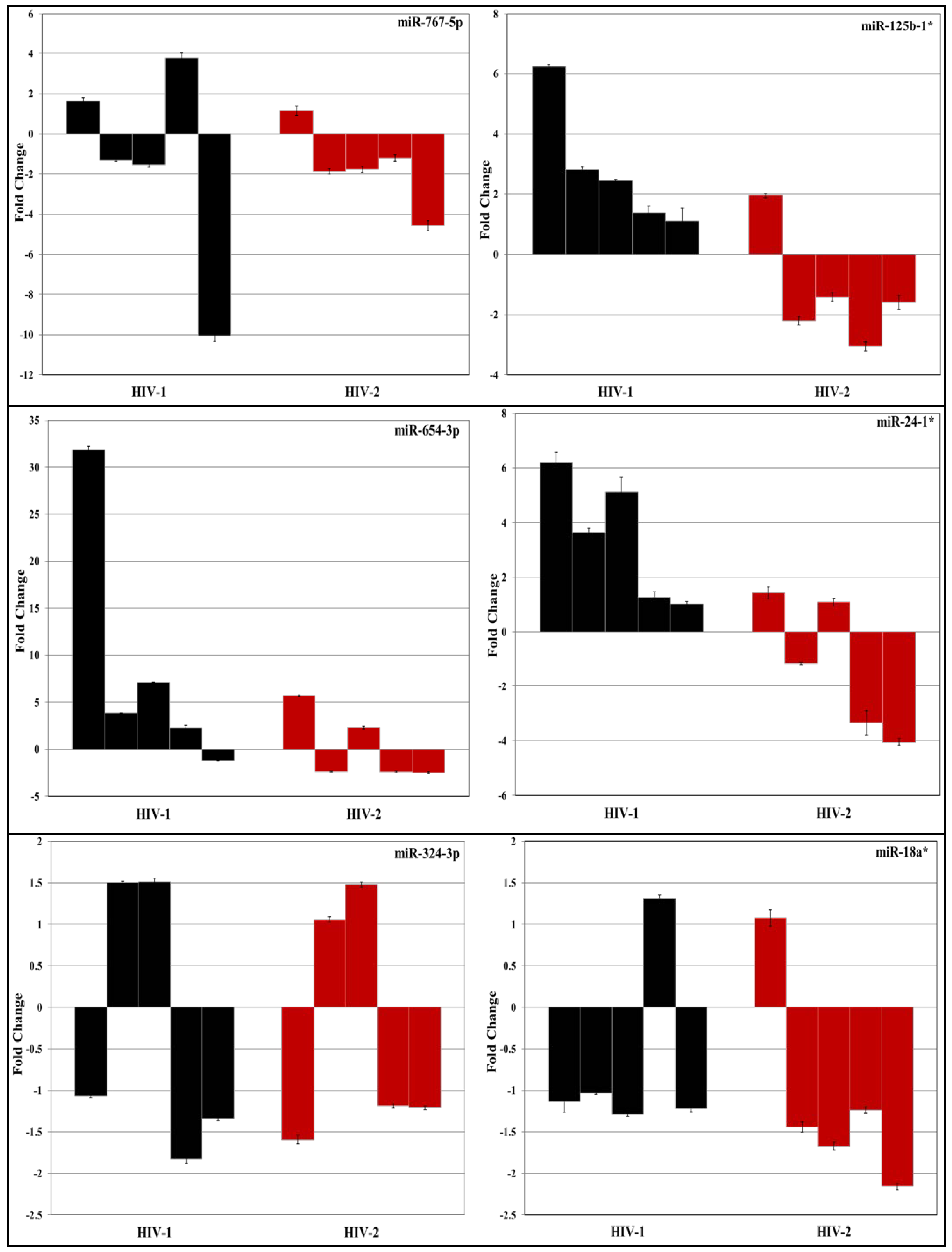
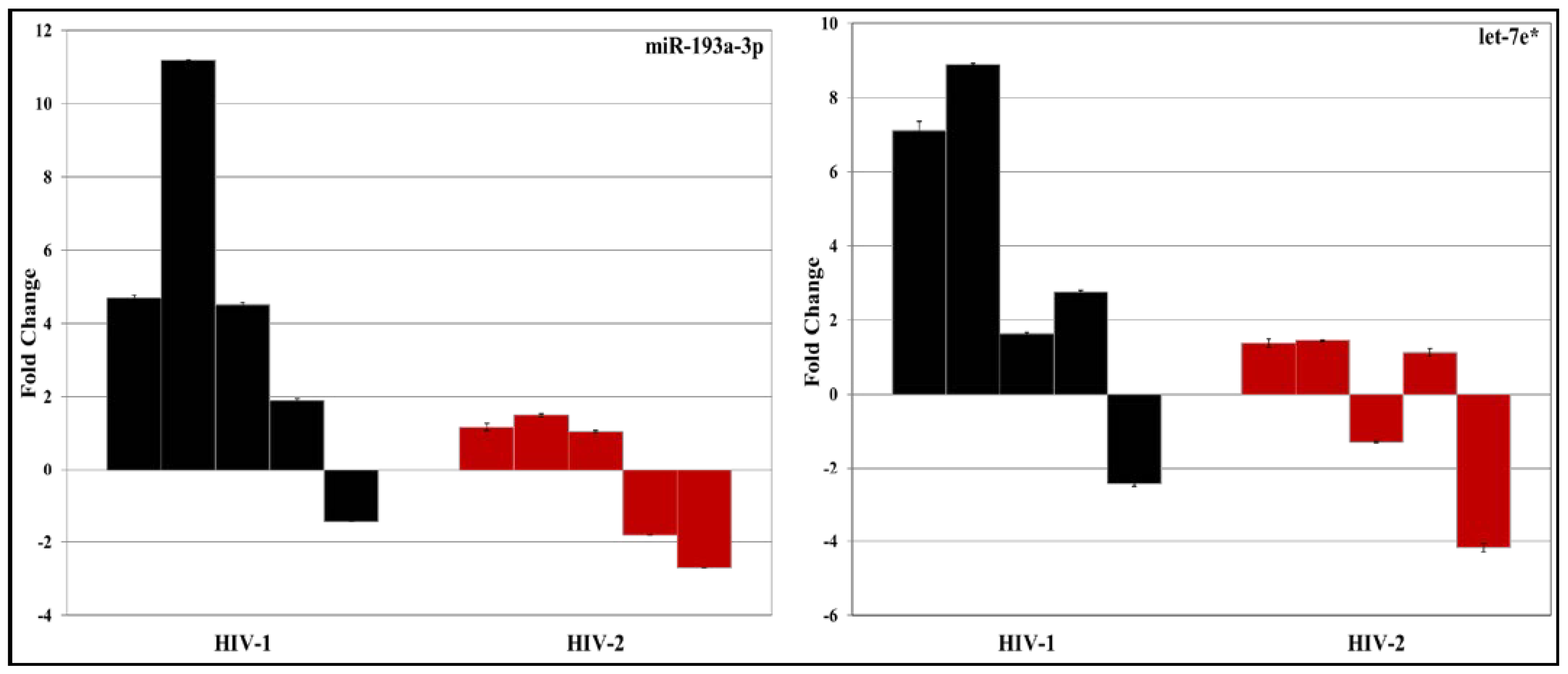
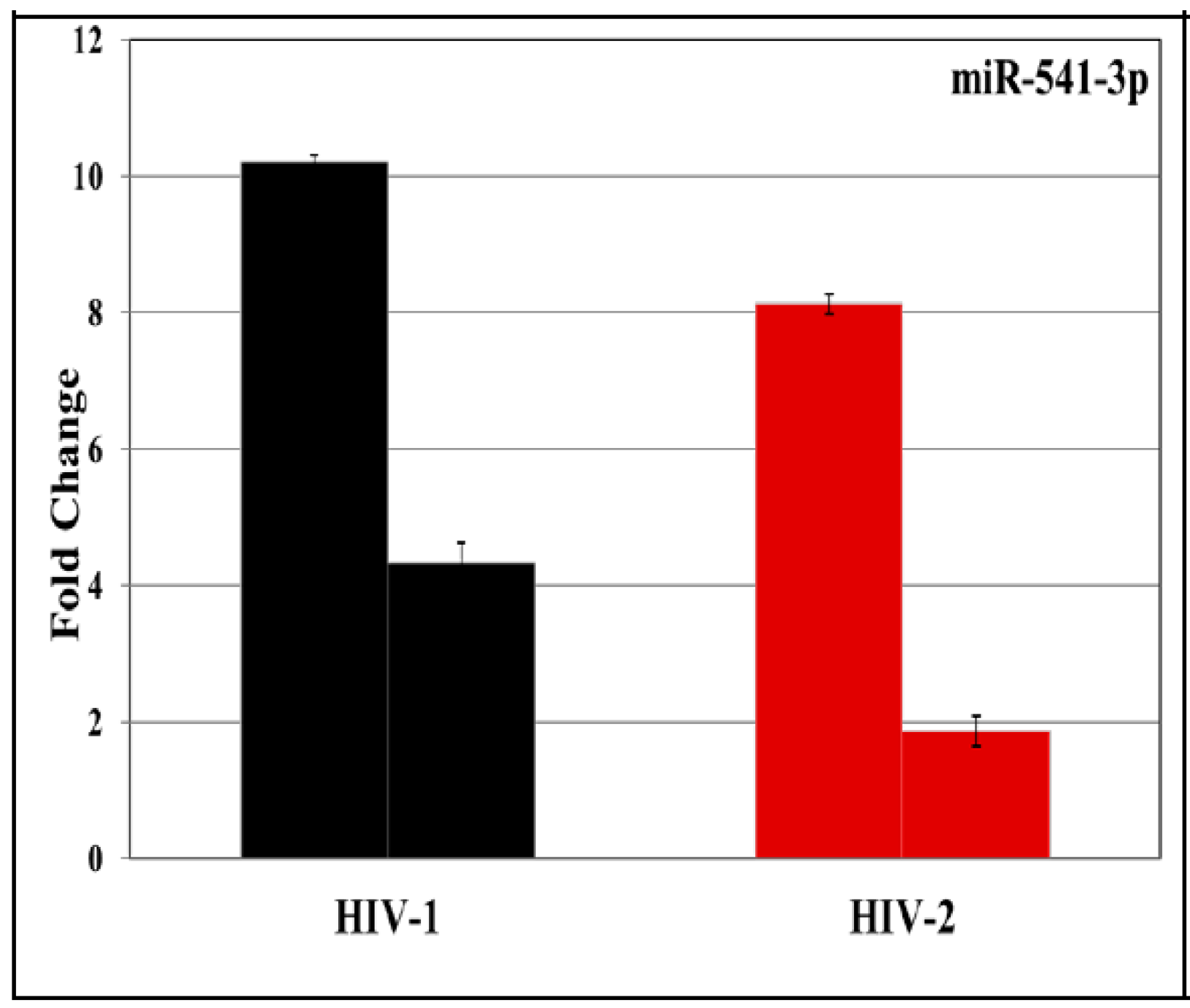
3.2. Validation of miRNA Expression by RT-PCR
3.3. Predicted Target Genes of Differential Expressed miRNA
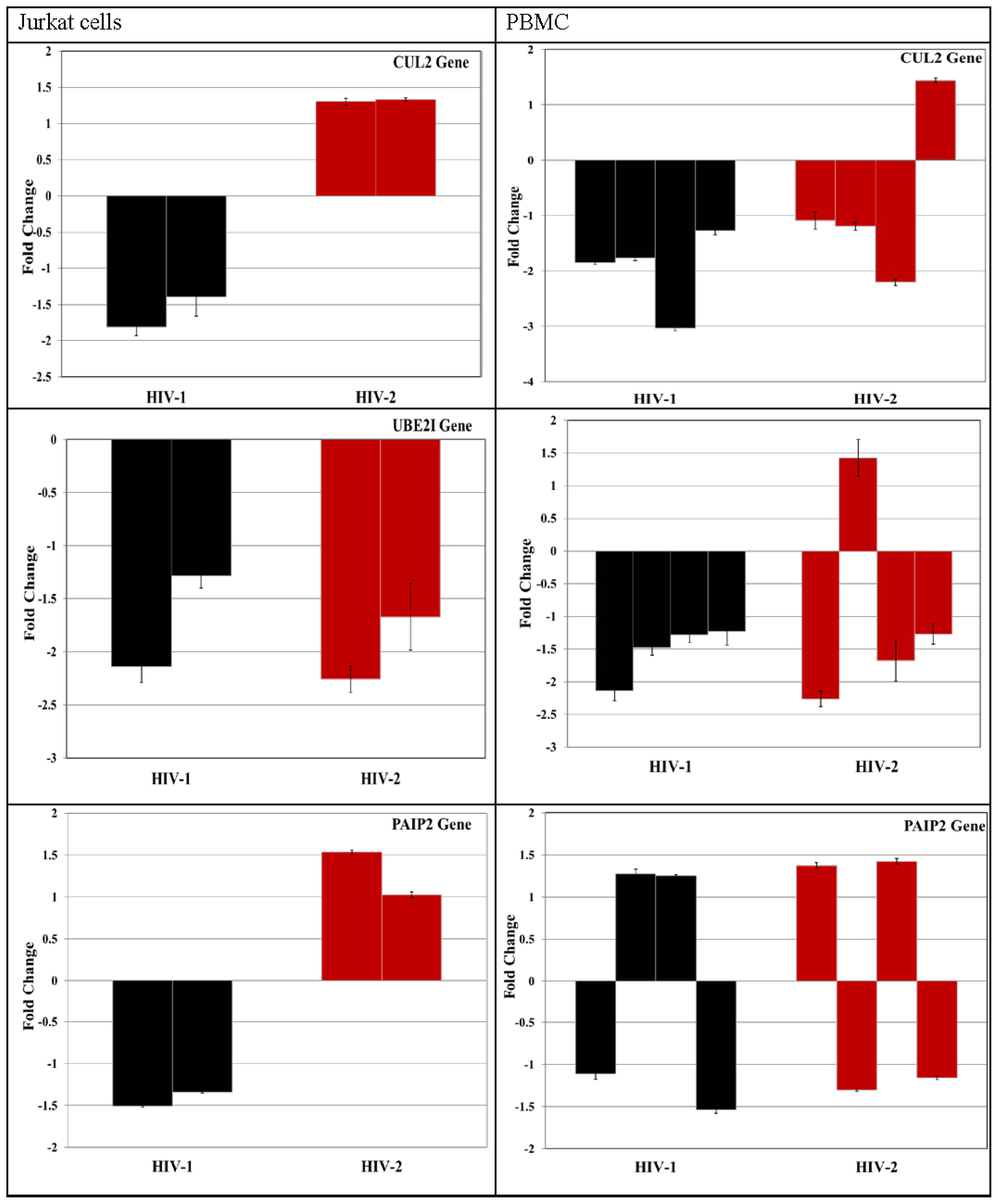
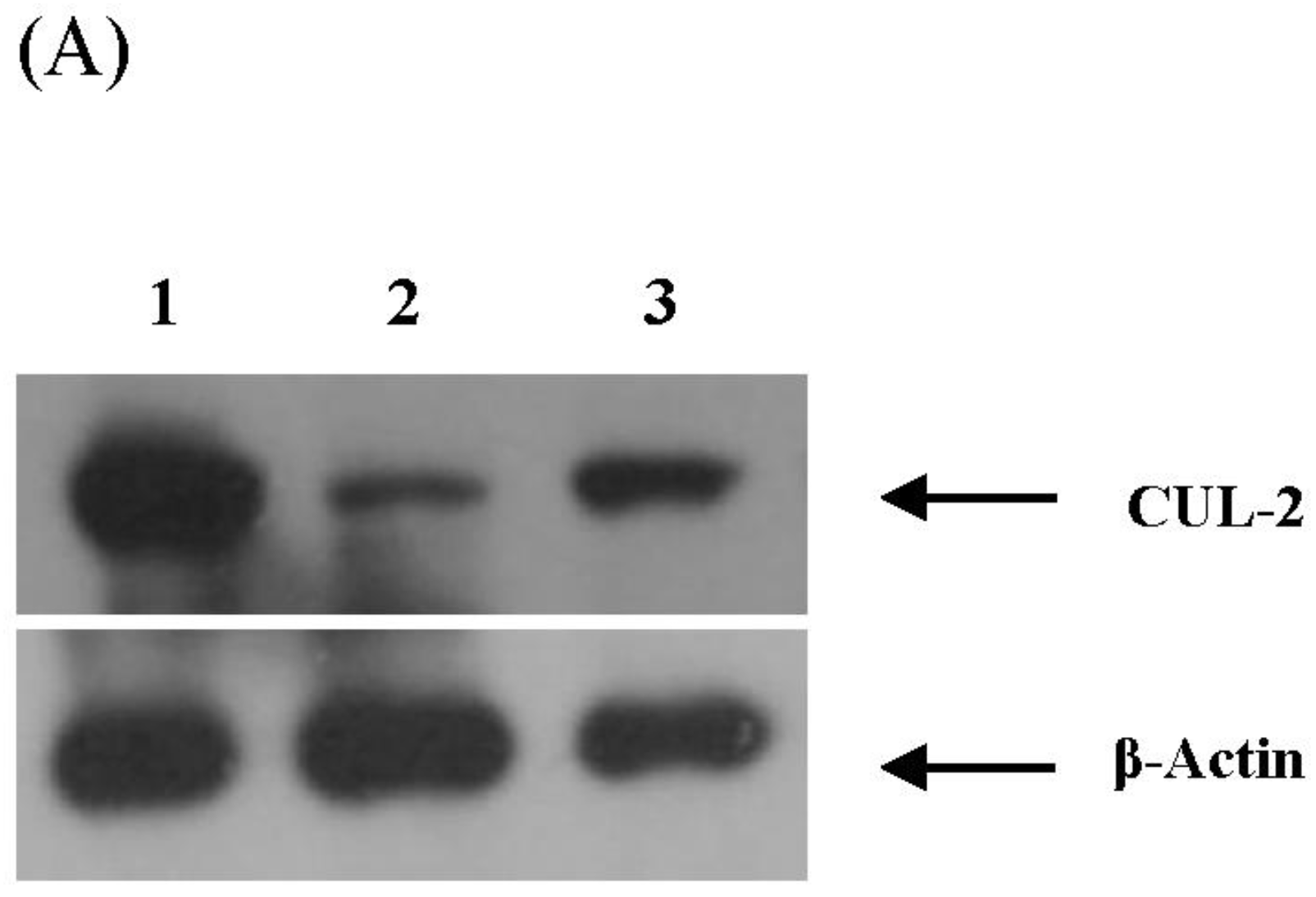
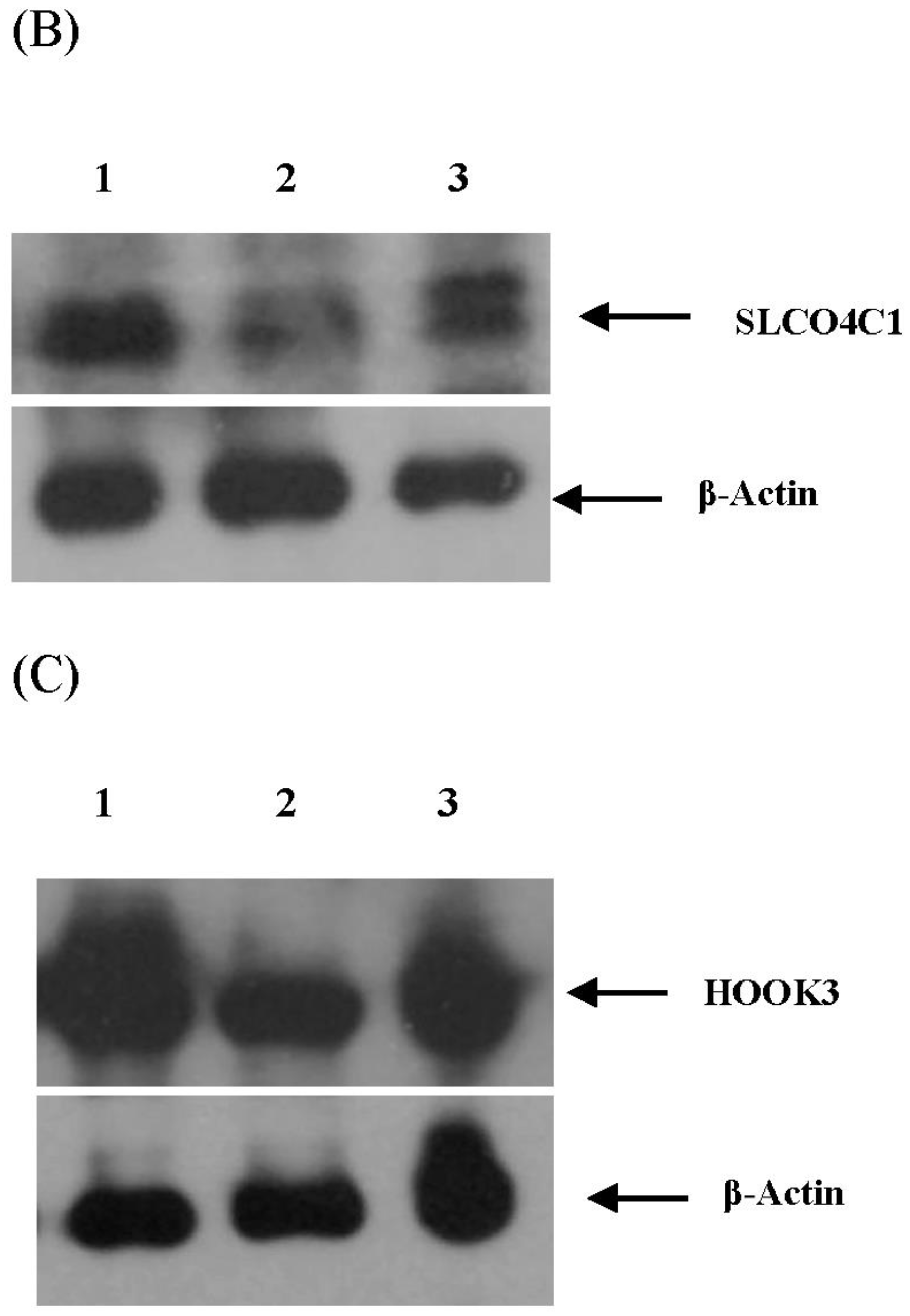
3.4. Validation of miRNA Targets Genes
4. Discussion
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Gao, F.; Yue, L.; Robertson, D.L.; Hill, S.C.; Hui, H.; Biggar, R.J.; Neequaye, A.E.; Whelan, T.M.; Ho, D.D.; Shaw, G.M.; et al. Genetic diversity of human immunodeficiency virus type 2: Evidence for distinct sequence subtypes with differences in virus biology. J. Virol. 1994, 68, 7433–7447. [Google Scholar] [PubMed]
- Poulsen, A.G.; Aaby, P.; Gottschau, A.; Kvinesdal, B.B.; Dias, F.; Molbak, K.; Lauritzen, E. HIV-2 infection in Bissau, West Africa, 1987–1989: Incidence, prevalences, and routes of transmission. J. Acquir. Immune Defic. Syndr. 1993, 6, 941–948. [Google Scholar] [PubMed]
- Sousa, A.E.; Carneiro, J.; Meier-Schellersheim, M.; Grossman, Z.; Victorino, R.M. CD4 T cell depletion is linked directly to immune activation in the pathogenesis of HIV-1 and HIV-2 but only indirectly to the viral load. J. Immunol. 2002, 169, 3400–3406. [Google Scholar] [CrossRef] [PubMed]
- Alimonti, J.B.; Ball, T.B.; Fowke, K.R. Mechanisms of CD4+ T lymphocyte cell death in human immunodeficiency virus infection and AIDS. J. Gen. Virol. 2003, 84, 1649–1661. [Google Scholar] [CrossRef] [PubMed]
- Kanki, P.J.; Travers, K.U.; S, M.B.; Hsieh, C.C.; Marlink, R.G.; Gueye, N.A.; Siby, T.; Thior, I.; Hernandez-Avila, M.; Sankale, J.L.; et al. Slower heterosexual spread of HIV-2 than HIV-1. Lancet 1994, 343, 943–946. [Google Scholar] [CrossRef]
- Hilfinger, J.M.; Clark, N.; Smith, M.; Robinson, K.; Markovitz, D.M. Differential regulation of the human immunodeficiency virus type 2 enhancer in monocytes at various stages of differentiation. J. Virol. 1993, 67, 4448–4453. [Google Scholar] [PubMed]
- Makvandi-Nejad, S.; Rowland-Jones, S. How does the humoral response to HIV-2 infection differ from HIV-1 and can this explain the distinct natural history of infection with these two human retroviruses? Immunol. Lett. 2015, 163, 69–75. [Google Scholar] [CrossRef] [PubMed]
- Popper, S.J.; Sarr, A.D.; Travers, K.U.; Gueye-Ndiaye, A.; Mboup, S.; Essex, M.E.; Kanki, P.J. Lower human immunodeficiency virus (HIV) type 2 viral load reflects the difference in pathogenicity of HIV-1 and HIV-2. J. Infect. Dis. 1999, 180, 1116–1121. [Google Scholar] [CrossRef] [PubMed]
- Popper, S.J.; Sarr, A.D.; Gueye-Ndiaye, A.; Mboup, S.; Essex, M.E.; Kanki, P.J. Low plasma human immunodeficiency virus type 2 viral load is independent of proviral load: Low virus production in vivo. J. Virol. 2000, 74, 1554–1557. [Google Scholar] [CrossRef] [PubMed]
- Leligdowicz, A.; Yindom, L.M.; Onyango, C.; Sarge-Njie, R.; Alabi, A.; Cotten, M.; Vincent, T.; da Costa, C.; Aaby, P.; Jaye, A.; et al. Robust Gag-specific T cell responses characterize viremia control in HIV-2 infection. J. Clin. Investig. 2007, 117, 3067–3074. [Google Scholar] [CrossRef] [PubMed]
- Auld, A.F.; Ekra, K.A.; Shiraishi, R.W.; Tuho, M.Z.; Kouakou, J.S.; Mohamed, F.; Ettiegne-Traore, V.; Sabatier, J.; Essombo, J.; Adjorlolo-Johnson, G.; et al. Temporal trends in treatment outcomes for HIV-1 and HIV-2-infected adults enrolled in Cote d’Ivoire’s national antiretroviral therapy program. PLoS ONE 2014, 9, e98183. [Google Scholar] [CrossRef] [PubMed]
- Sarr, A.D.; Hamel, D.J.; Thior, I.; Kokkotou, E.; Sankale, J.L.; Marlink, R.G.; Coll-Seck, E.M.; Essex, M.E.; Siby, T.; NDoye, I.; et al. HIV-1 and HIV-2 dual infection: Lack of HIV-2 provirus correlates with low CD4+ lymphocyte counts. AIDS 1998, 12, 131–137. [Google Scholar] [CrossRef] [PubMed]
- Koblavi-Deme, S.; Kestens, L.; Hanson, D.; Otten, R.A.; Borget, M.Y.; Bile, C.; Wiktor, S.Z.; Roels, T.H.; Chorba, T.; Nkengasong, J.N. Differences in HIV-2 plasma viral load and immune activation in HIV-1 and HIV-2 dually infected persons and those infected with HIV-2 only in Abidjan, Cote D’Ivoire. AIDS 2004, 18, 413–419. [Google Scholar] [CrossRef] [PubMed]
- Machuca, A.; Ding, L.; Taffs, R.; Lee, S.; Wood, O.; Hu, J.; Hewlett, I. HIV type 2 primary isolates induce a lower degree of apoptosis “in vitro” compared with HIV type 1 primary isolates. AIDS Res. Hum. Retrovir. 2004, 20, 507–512. [Google Scholar] [CrossRef] [PubMed]
- Smoot, M.E.; Ono, K.; Ruscheinski, J.; Wang, P.L.; Ideker, T. Cytoscape 2.8: New features for data integration and network visualization. Bioinformatics 2011, 27, 431–432. [Google Scholar] [CrossRef] [PubMed]
- Ding, Y.; Wu, Z.; Duan, S.; Rou, K.; Yang, Y.; Wang, J.; Gao, M.; Ye, R.; Detels, R.; He, N. Risk factors for incident HSV-2 infections among a prospective cohort of HIV-1-discordant couples in China. Sex. Transm. Infect. 2016, 92, 76–82. [Google Scholar] [CrossRef] [PubMed]
- Devadas, K.; Biswas, S.; Haleyurgirisetty, M.; Wood, O.; Ragupathy, V.; Lee, S.; Hewlett, I. Analysis of host gene expression profile in HIV-1 and HIV-2 infected T-cells. PLoS ONE 2016, 11, e0147421. [Google Scholar] [CrossRef] [PubMed]
- Shalgi, R.; Lieber, D.; Oren, M.; Pilpel, Y. Global and local architecture of the mammalian microRNA-transcription factor regulatory network. PLoS Comput. Biol. 2007, 3, e131. [Google Scholar] [CrossRef] [PubMed]
- Swaminathan, G.; Navas-Martin, S.; Martin-Garcia, J. Interplay between microRNAs, Toll-like receptors, and HIV-1: Potential implications in HIV-1 replication and chronic immune activation. Discov. Med. 2014, 18, 15–27. [Google Scholar] [PubMed]
- Houzet, L.; Yeung, M.L.; de Lame, V.; Desai, D.; Smith, S.M.; Jeang, K.T. MicroRNA profile changes in human immunodeficiency virus type 1 (HIV-1) seropositive individuals. Retrovirology 2008, 5, 118. [Google Scholar] [CrossRef] [PubMed]
- Reynoso, R.; Laufer, N.; Hackl, M.; Skalicky, S.; Monteforte, R.; Turk, G.; Carobene, M.; Quarleri, J.; Cahn, P.; Werner, R.; et al. MicroRNAs differentially present in the plasma of HIV elite controllers reduce HIV infection in vitro. Sci. Rep. 2014, 4, 5915. [Google Scholar] [CrossRef] [PubMed]
- Hubert, A.; Subra, C.; Jenabian, M.A.; Labrecque, P.F.T.; Tremblay, C.; Laffont, B.; Provost, P.; Routy, J.P.; Gilbert, C. Elevated abundance, size, and MicroRNA content of plasma extracellular vesicles in viremic HIV-1+ patients: Correlations with known markers of disease progression. J. Acquir. Immune Defic. Syndr. 2015, 70, 219–227. [Google Scholar] [CrossRef] [PubMed]
- Masotti, A.; Donninelli, G.; Da Sacco, L.; Varano, B.; Del Corno, M.; Gessani, S. HIV-1 gp120 influences the expression of microRNAs in human monocyte-derived dendritic cells via STAT3 activation. BMC Genom. 2015, 16, 480. [Google Scholar] [CrossRef] [PubMed]
- Monteleone, K.; Selvaggi, C.; Cacciotti, G.; Falasca, F.; Mezzaroma, I.; D’Ettorre, G.; Turriziani, O.; Vullo, V.; Antonelli, G.; Scagnolari, C. MicroRNA-29 family expression and its relation to antiviral immune response and viro-immunological markers in HIV-1-infected patients. BMC Infect. Dis. 2015, 15, 51. [Google Scholar] [CrossRef] [PubMed]
- Yang, Z.; Yang, J.; Wang, J.; Lu, X.; Jin, C.; Xie, T.; Wu, N. Identify potential regulators in HIV-1 latency by joint microRNA and mRNA analysis. Cell. Physiol. Biochem. 2015, 36, 569–584. [Google Scholar] [CrossRef] [PubMed]
- Guha, D.; Mancini, A.; Sparks, J.; Ayyavoo, V. HIV-1 infection dysregulates cell cycle regulatory protein p21 in CD4+ T cells through miR-20a and miR-106b regulation. J. Cell. Biochem. 2016. [Google Scholar] [CrossRef] [PubMed]
- Chen, A.K.; Sengupta, P.; Waki, K.; Van Engelenburg, S.B.; Ochiya, T.; Ablan, S.D.; Freed, E.O.; Lippincott-Schwartz, J. MicroRNA binding to the HIV-1 Gag protein inhibits Gag assembly and virus production. Proc. Natl. Acad. Sci. USA 2014, 111, E2676–E2683. [Google Scholar] [CrossRef] [PubMed]
- Roth, W.W.; Huang, M.B.; Addae Konadu, K.; Powell, M.D.; Bond, V.C. Micro RNA in exosomes from HIV-infected macrophages. Int. J. Environ. Res. Public Health 2015, 13. [Google Scholar] [CrossRef] [PubMed]
- Kapoor, R.; Arora, S.; Ponia, S.S.; Kumar, B.; Maddika, S.; Banerjea, A.C. The miRNA miR-34a enhances HIV-1 replication by targeting PNUTS/PPP1R10, which negatively regulates HIV-1 transcriptional complex formation. Biochem. J. 2015, 470, 293–302. [Google Scholar] [CrossRef] [PubMed]
- Sampey, G.C.; Saifuddin, M.; Schwab, A.; Barclay, R.; Punya, S.; Chung, M.C.; Hakami, R.M.; Asad Zadeh, M.; Lepene, B.; Klase, Z.A.; et al. Exosomes from HIV-1-infected cells stimulate production of pro-inflammatory cytokines through trans-activating response (TAR) RNA. J. Biol. Chem. 2016, 291, 1251–1266. [Google Scholar] [CrossRef] [PubMed]
- Kumar, V.; Torben, W.; Kenway, C.S.; Schiro, F.R.; Mohan, M. Longitudinal examination of the intestinal lamina propria cellular compartment of SIV-infected rhesus macaques provides broader and deeper insights into the link between aberrant microRNA expression and persistent immune activation. J. Virol. 2016. [Google Scholar] [CrossRef] [PubMed]
- Klase, Z.; Kale, P.; Winograd, R.; Gupta, M.V.; Heydarian, M.; Berro, R.; McCaffrey, T.; Kashanchi, F. HIV-1 TAR element is processed by Dicer to yield a viral micro-RNA involved in chromatin remodeling of the viral LTR. BMC Mol. Biol. 2007, 8, 63. [Google Scholar] [CrossRef] [PubMed]
- Klase, Z.; Winograd, R.; Davis, J.; Carpio, L.; Hildreth, R.; Heydarian, M.; Fu, S.; McCaffrey, T.; Meiri, E.; Ayash-Rashkovsky, M.; et al. HIV-1 TAR miRNA protects against apoptosis by altering cellular gene expression. Retrovirology 2009, 6, 18. [Google Scholar] [CrossRef] [PubMed]
- Klase, Z.A.; Sampey, G.C.; Kashanchi, F. Retrovirus infected cells contain viral microRNAs. Retrovirology 2013, 10, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Narayanan, A.; Iordanskiy, S.; Das, R.; Van Duyne, R.; Santos, S.; Jaworski, E.; Guendel, I.; Sampey, G.; Dalby, E.; Iglesias-Ussel, M.; et al. Exosomes derived from HIV-1-infected cells contain trans-activation response element RNA. J. Biol. Chem. 2013, 288, 20014–20033. [Google Scholar] [CrossRef] [PubMed]
- Gallo, R.C.; Salahuddin, S.Z.; Popovic, M.; Shearer, G.M.; Kaplan, M.; Haynes, B.F.; Palker, T.J.; Redfield, R.; Oleske, J.; Safai, B.; et al. Frequent detection and isolation of cytopathic retroviruses (HTLV-III) from patients with AIDS and at risk for AIDS. Science 1984, 224, 500–503. [Google Scholar] [CrossRef] [PubMed]
- Shaw, G.M.; Hahn, B.H.; Arya, S.K.; Groopman, J.E.; Gallo, R.C.; Wong-Staal, F. Molecular characterization of human T-cell leukemia (lymphotropic) virus type III in the acquired immune deficiency syndrome. Science 1984, 226, 1165–1171. [Google Scholar] [CrossRef] [PubMed]
- Zhuo, Y.; Gao, G.; Shi, J.A.; Zhou, X.; Wang, X. miRNAs: Biogenesis, origin and evolution, functions on virus-host interaction. Cell. Physiol. Biochem. 2013, 32, 499–510. [Google Scholar] [CrossRef] [PubMed]
- Clark, N.M.; Smith, M.J.; Hilfinger, J.M.; Markovitz, D.M. Activation of the human T-cell leukemia virus type I enhancer is mediated by binding sites for Elf-1 and the pets factor. J. Virol. 1993, 67, 5522–5528. [Google Scholar] [PubMed]
- Li, Z.; Wu, S.; Wang, J.; Li, W.; Lin, Y.; Ji, C.; Xue, J.; Chen, J. Evaluation of the interactions of HIV-1 integrase with small ubiquitin-like modifiers and their conjugation enzyme Ubc9. Int. J. Mol. Med. 2012, 30, 1053–1060. [Google Scholar] [PubMed]
- Chang, S.T.; Thomas, M.J.; Sova, P.; Green, R.R.; Palermo, R.E.; Katze, M.G. Next-generation sequencing of small RNAs from HIV-infected cells identifies phased microrna expression patterns and candidate novel microRNAs differentially expressed upon infection. MBio 2013, 4, e00549-12. [Google Scholar] [CrossRef] [PubMed]
- Egana-Gorrono, L.; Escriba, T.; Boulanger, N.; Guardo, A.C.; Leon, A.; Bargallo, M.E.; Garcia, F.; Gatell, J.M.; Plana, M.; Arnedo, M.; et al. Differential microRNA expression profile between stimulated PBMCs from HIV-1 infected elite controllers and viremic progressors. PLoS ONE 2014, 9, e106360. [Google Scholar]
- Duskova, K.; Nagilla, P.; Le, H.S.; Iyer, P.; Thalamuthu, A.; Martinson, J.; Bar-Joseph, Z.; Buchanan, W.; Rinaldo, C.; Ayyavoo, V. MicroRNA regulation and its effects on cellular transcriptome in human immunodeficiency virus-1 (HIV-1) infected individuals with distinct viral load and CD4 cell counts. BMC Infect. Dis. 2013, 13, 250. [Google Scholar] [CrossRef] [PubMed]
- Witwer, K.W.; Clements, J.E. Evidence for miRNA expression differences of HIV-1-positive, treatment-naive patients and elite suppressors: A re-analysis. Blood 2012, 119, 6395–6396. [Google Scholar] [CrossRef] [PubMed]
- Witwer, K.W.; Watson, A.K.; Blankson, J.N.; Clements, J.E. Relationships of PBMC microRNA expression, plasma viral load, and CD4+ T-cell count in HIV-1-infected elite suppressors and viremic patients. Retrovirology 2012, 9, 5. [Google Scholar] [CrossRef] [PubMed]
- Chiang, K.; Liu, H.; Rice, A.P. miR-132 enhances HIV-1 replication. Virology 2013, 438, 1–4. [Google Scholar] [CrossRef] [PubMed]
- Huang, J.; Wang, F.; Argyris, E.; Chen, K.; Liang, Z.; Tian, H.; Huang, W.; Squires, K.; Verlinghieri, G.; Zhang, H. Cellular microRNAs contribute to HIV-1 latency in resting primary CD4+ T lymphocytes. Nat. Med. 2007, 13, 1241–1247. [Google Scholar] [CrossRef] [PubMed]
- Ahluwalia, J.K.; Khan, S.Z.; Soni, K.; Rawat, P.; Gupta, A.; Hariharan, M.; Scaria, V.; Lalwani, M.; Pillai, B.; Mitra, D.; Brahmachari, S.K. Human cellular microRNA hsa-miR-29a interferes with viral nef protein expression and HIV-1 replication. Retrovirology 2008, 5, 117. [Google Scholar] [CrossRef] [PubMed]
- Sung, T.L.; Rice, A.P. miR-198 inhibits HIV-1 gene expression and replication in monocytes and its mechanism of action appears to involve repression of cyclin T1. PLoS Pathog. 2009, 5, e1000263. [Google Scholar] [CrossRef] [PubMed]
- Swaminathan, S.; Murray, D.D.; Kelleher, A.D. The role of microRNAs in HIV-1 pathogenesis and therapy. AIDS 2012, 26, 1325–1334. [Google Scholar] [CrossRef] [PubMed]
- Triboulet, R.; Mari, B.; Lin, Y.L.; Chable-Bessia, C.; Bennasser, Y.; Lebrigand, K.; Cardinaud, B.; Maurin, T.; Barbry, P.; Baillat, V.; et al. Suppression of microRNA-silencing pathway by HIV-1 during virus replication. Science 2007, 315, 1579–1582. [Google Scholar] [CrossRef] [PubMed]
- Orecchini, E.; Doria, M.; Michienzi, A.; Giuliani, E.; Vassena, L.; Ciafre, S.A.; Farace, M.G.; Galardi, S. The HIV-1 Tat protein modulates CD4 expression in human T cells through the induction of miR-222. RNA Biol. 2014, 11, 334–338. [Google Scholar] [CrossRef] [PubMed]
- Guo, H.; Li, W.; Zheng, T.; Liu, Z. miR-195 targets HDGF to inhibit proliferation and invasion of NSCLC cells. Tumour Biol. J. Int. Soc. Oncodev. Biol. Med. 2014, 35, 8861–8866. [Google Scholar] [CrossRef] [PubMed]
- Farberov, L.; Herzig, E.; Modai, S.; Isakov, O.; Hizi, A.; Shomron, N. MicroRNA-mediated regulation of p21 and TASK1 cellular restriction factors enhances HIV-1 infection. J. Cell Sci. 2015, 128, 1607–1616. [Google Scholar] [CrossRef] [PubMed]
- Swaminathan, G.; Navas-Martin, S.; Martin-Garcia, J. MicroRNAs and HIV-1 infection: Antiviral activities and beyond. J. Mol. Biol. 2014, 426, 1178–1197. [Google Scholar] [CrossRef] [PubMed]
- Swaminathan, G.; Pascual, D.; Rival, G.; Perales-Linares, R.; Martin-Garcia, J.; Navas-Martin, S. Hepatitis C virus core protein enhances HIV-1 replication in human macrophages through TLR2, JNK, and MEK1/2-dependent upregulation of TNF-alpha and IL-6. FEBS Lett. 2014, 588, 3501–3510. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Saini, N. Downregulation of BCL2 by miRNAs augments drug-induced apoptosis—A combined computational and experimental approach. J. Cell Sci. 2012, 125, 1568–1578. [Google Scholar] [CrossRef] [PubMed]
- Itesako, T.; Seki, N.; Yoshino, H.; Chiyomaru, T.; Yamasaki, T.; Hidaka, H.; Yonezawa, T.; Nohata, N.; Kinoshita, T.; Nakagawa, M.; et al. The microRNA expression signature of bladder cancer by deep sequencing: The functional significance of the miR-195/497 cluster. PLoS ONE 2014, 9, e84311. [Google Scholar] [CrossRef] [PubMed]
- Gupta, P.; Cairns, M.J.; Saksena, N.K. Regulation of gene expression by microRNA in HCV infection and HCV-mediated hepatocellular carcinoma. Virol. J. 2014, 11, 64. [Google Scholar] [CrossRef] [PubMed]
- Hrdlickova, R.; Nehyba, J.; Bargmann, W.; Bose, H.R., Jr. Multiple tumor suppressor microRNAs regulate telomerase and TCF7, an important transcriptional regulator of the Wnt pathway. PLoS ONE 2014, 9, e86990. [Google Scholar] [CrossRef] [PubMed]
- Leivonen, S.K.; Sahlberg, K.K.; Makela, R.; Due, E.U.; Kallioniemi, O.; Borresen-Dale, A.L.; Perala, M. High-throughput screens identify microRNAs essential for HER2 positive breast cancer cell growth. Mol. Oncol. 2014, 8, 93–104. [Google Scholar] [CrossRef] [PubMed]
- Dai, C.; Ye, J.; Qin, H.; Zhao, S.; He, P.; Zhou, H. Fabrication of High Energy Li-Ion Cells with Li4Ti5O12 micro-sphere as anode and 0.5Li2MnO3-0.5LiNi0.4Co0.2Mn0.4O2 micro-sphere as cathode. Chem. Asian J. 2016. [Google Scholar] [CrossRef] [PubMed]
- Sarikas, A.; Hartmann, T.; Pan, Z.Q. The cullin protein family. Genome Biol. 2011, 220, 2–12. [Google Scholar] [CrossRef] [PubMed]
- Mousnier, A.; Kubat, N.; Massias-Simon, A.; Segeral, E.; Rain, J.C.; Benarous, R.; Emiliani, S.; Dargemont, C. von Hippel Lindau binding protein 1-mediated degradation of integrase affects HIV-1 gene expression at a postintegration step. Proc. Natl. Acad. Sci. USA 2007, 104, 13615–13620. [Google Scholar] [CrossRef] [PubMed]
- Okumura, F.; Matsuzaki, M.; Nakatsukasa, K.; Kamura, T. The Role of elongin BC-containing ubiquitin ligases. Front. Oncol. 2012, 2, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Jaber, T.; Bohl, C.R.; Lewis, G.L.; Wood, C.; West, J.T., Jr.; Weldon, R.A., Jr. Human Ubc9 contributes to production of fully infectious human immunodeficiency virus type 1 virions. J. Virol. 2009, 83, 10448–10459. [Google Scholar] [CrossRef] [PubMed]
- Bohl, C.R.; Abrahamyan, L.G.; Wood, C. Human Ubc9 is involved in intracellular HIV-1 Env stability after trafficking out of the trans-Golgi network in a Gag dependent manner. PLoS ONE 2013, 8, e69359. [Google Scholar] [CrossRef] [PubMed]




| PBMC | HIV-1 (pg/mL) | HIV-2 (pg/mL) |
|---|---|---|
| Donor 1 | 3148.13 | 28011.78 |
| Donor 2 | 4019.29 | 27121.72 |
| Donor 3 | 1653.06 | 24534.10 |
| No. | miRNA | Fold Change | |
|---|---|---|---|
| HIV-1 MN | HIV-2 ROD | ||
| 1 | hsa-let-7a-2* | −1.2169 | −6.6374 |
| 2 | hsa-let-7e* | 2.0544 | 1.4174 |
| 3 | hsa-let-7i* | −1.3374 | −2.2122 |
| 4 | hsa-miR-18a* | −1.1164 | −6.7632 |
| 5 | hsa-miR-105* | −1.4353 | −2.4315 |
| 6 | hsa-miR-148b* | 2.5483 | 2.4489 |
| 7 | hsa-miR-192* | −1.5557 | −2.0177 |
| 8 | hsa-miR-193a-3p | 2.7988 | 2.2536 |
| 9 | hsa-miR-195-3p | 3.1952 | −1.9213 |
| 10 | hsa-miR-1179 | −1.1205 | −2.7279 |
| 11 | hsa-miR-1206 | 2.1144 | −1.0175 |
| 12 | hsa-miR-1251 | −1.4678 | −2.3088 |
| 13 | hsa-miR-125b-1* | −5.5759 | 3.3442 |
| 14 | hsa-miR-1225-5p | −1.21 | −5.85 |
| 15 | hsa-miR-23b* | −1.0803 | −2.0489 |
| 16 | hsa-miR-24-1* | −1.1715 | −3.0804 |
| 17 | hsa-miR-25* | −2.3575 | 1.2734 |
| 18 | hsa-miR-26a-2* | −2.0392 | −2.0489 |
| 19 | hsa-miR-27a* | −2.9617 | 1.8709 |
| 20 | hsa-miR-202* | −1.1892 | −2.8631 |
| 21 | hsa-miR-217 | −2.1554 | −1.3649 |
| 22 | hsa-miR-218-1* | −1.7084 | −2.3989 |
| 23 | hsa-miR-220a | 2.4943 | 1.5979 |
| 24 | hsa-miR-34a* | −2.2859 | −1.3974 |
| 25 | hsa-miR-324-3p | −2.0879 | 1.4893 |
| 26 | hsa-miR-335* | −2.2532 | −2.5759 |
| 27 | hsa-miR-340* | −2.3827 | −2.0177 |
| 28 | hsa-miR-374a* | −2.2562 | −1.6367 |
| 29 | hsa-miR-379* | 2.1035 | −1.0755 |
| 30 | hsa-miR-424* | 4.5302 | 5.0506 |
| 31 | hsa-miR-499-3p | −2.3432 | −2.0667 |
| 32 | hsa-miR-499-5p | 3.3336 | 2.9536 |
| 33 | hsa-miR-513b | 1.9684 | 1.2096 |
| 34 | hsa-miR-513c | 2.6708 | −1.7721 |
| 35 | hsa-miR-518f-3p | 3.4019 | 1.8917 |
| 36 | hsa-miR-541-3p | 5.6293 | 3.5696 |
| 37 | hsa-miR-548m | 2.1098 | 2.0008 |
| 38 | hsa-miR-635 | 1.1136 | −2.3079 |
| 39 | hsa-miR-654-3p | 1.8715 | 2.2779 |
| 40 | hsa-miR-708* | 1.209 | −2.021 |
| 41 | hsa-miR-767-3p | 1.9017 | −1.0177 |
| 42 | hsa-miR-767-5p | 2.6795 | 1.0528 |
| 43 | hsa-miR-769-5p | 1.9541 | 1.0458 |
| 44 | hsa-miR-891b | 2.2034 | 1.1506 |
| 45 | hsa-miR-939-5p | −1.1593 | 4.6161 |
| 46 | SNORD44 | 1.0339 | 1.0843 |
| Target Score 1 | Gene | HIV-1 Interactions 2 | |
|---|---|---|---|
| Protein | Other | ||
| hsa-miR-195-3p | |||
| 99 | CDK6 (cyclin-dependent kinase 6) | - | - |
| 99 | CUL2 (cullin 2) | Vif; p23 | - |
| 99 | RAB6A [RAB6A, member RAS oncogene family] | gp120; gp41 | Replication |
| 99 | CLTC [clathrin, heavy chain (Hc)] | gp120; Nef; p27; Vpr, p15; matrix; pol | - |
| 99 | DSC3 [desmocollin 3] | gp160 | - |
| 99 | DENND1B [DENN/MADD domain containing 1B] | - | - |
| 98 | CADM2 [cell adhesion molecule 2] | - | - |
| 98 | PAIP2 [poly(A) binding protein interacting protein 2] | - | - |
| 98 | UBE2I [ubiquitin-conjugating enzyme E2I] | gp120; gp41; Pr55(Gag); integrase; p6 | - |
| 98 | MTHFD2L [methylenetetrahydrofolate dehydrogenase (NADP+ dependent) 2-like] | - | - |
| hsa-miR-518f-3p | |||
| 87 | VAPB [VAMP (vesicle-associated membrane protein)-associated protein B and C] | - | - |
| 81 | PRDX6 [peroxiredoxin 6] | - | - |
| 79 | RAP1B [RAP1B, member of RAS oncogene family] | - | Replication |
| 79 | SLCO4C1 [solute carrier organic anion transporter family, member 4C1] | - | - |
| 78 | ZNF282 [zinc finger protein 282] | - | - |
| 75 | TOLLIP [toll interacting protein] | - | - |
| 74 | TSN [translin] | - | - |
| 73 | MYO10 [myosin X] | - | - |
| 71 | PARP11 [poly (ADP-ribose) polymerase family, member 11] | - | - |
| 70 | HMP19 [HMP19 protein] | - | - |
| hsa-miR-541-3p | |||
| 90 | LOC100507421 [transmembrane protein 178-like] | - | - |
| 89 | HOOK3 [hook homolog 3 (Drosophila)] | - | - |
| 89 | TRIM67 [tripartite motif containing 67] | - | - |
| 85 | SLC37A1 [solute carrier family 37 (glycerol-3-phosphate transporter), member 1] | - | - |
| 83 | CACNA1E [calcium channel, voltage-dependent, R type, alpha 1E subunit] | - | - |
| 83 | LSM12 [LSM12 homolog (S. cerevisiae)] | gp 120 | - |
| 82 | CPLX4 [complexin 4] | - | - |
| 82 | PCDH11X [protocadherin 11 X-linked] | Replication | |
| 80 | HAX1 [HCLS1 associated protein X-1] | Rev, p19 | - |
| 78 | DDX17 [DEAD (Asp-Glu-Ala-Asp) box polypeptide 17] | Rev, p19; pol | - |
© 2016 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Devadas, K.; Biswas, S.; Haleyurgirisetty, M.; Ragupathy, V.; Wang, X.; Lee, S.; Hewlett, I. Identification of Host Micro RNAs That Differentiate HIV-1 and HIV-2 Infection Using Genome Expression Profiling Techniques. Viruses 2016, 8, 121. https://doi.org/10.3390/v8050121
Devadas K, Biswas S, Haleyurgirisetty M, Ragupathy V, Wang X, Lee S, Hewlett I. Identification of Host Micro RNAs That Differentiate HIV-1 and HIV-2 Infection Using Genome Expression Profiling Techniques. Viruses. 2016; 8(5):121. https://doi.org/10.3390/v8050121
Chicago/Turabian StyleDevadas, Krishnakumar, Santanu Biswas, Mohan Haleyurgirisetty, Viswanath Ragupathy, Xue Wang, Sherwin Lee, and Indira Hewlett. 2016. "Identification of Host Micro RNAs That Differentiate HIV-1 and HIV-2 Infection Using Genome Expression Profiling Techniques" Viruses 8, no. 5: 121. https://doi.org/10.3390/v8050121






