Functional Characterization of Cucumis metuliferus Proteinase Inhibitor Gene (CmSPI) in Potyviruses Resistance
Abstract
:1. Introduction
2. Materials and Methods
2.1. Cloning CmSPI Gene cDNA and Full Length Genomic Fragment of Cucumis metuliferus
| Primer Name | Oligonucleotides |
|---|---|
| 5′ and 3′ RACE reaction | |
| HMK2007-14 | 5′-AAT TCC AAC ACA AAT CAT CAT CTT-3′ |
| HMK2007-15 | 5′-TAA CAA ACA ACC AAA CTC GAT CAC-3′ |
| Full length genome walking | |
| HMK2010-35 | 5′-CTA ATA CgA CTC ACT ATA ggg CTC gAg Cgg CCg CCC ggg CAg gT-3′ |
| HMK2010-36 | 5′-ACC TgC CC-3′ |
| HMK2010-37 | 5′-ggA TCC TAA TAC gAC TCA CTA TAg ggC-3′ |
| HMK2010-38 | 5′-ACT CAC TAT Agg gCT CgA gCg ggC-3′ |
| HMK2011-118 | 5′-TAT TCC AAC AAG TTC CGG CCA CTG-3′ |
| HMK2011-119 | 5′-GAA AGT TGT TGA AAT TCC GAA GGT TG-3′ |
| HMK2011-127 | 5′-gct gga act gga acc act aaa ga-3′ |
| HMK2011-128 | 5′-cca taa atg gaa aca tac cag gag-3′ |
| CmSPI RNAi construction | |
| HMK2007-93 | 5′-ggA TCC CCA Tgg CCC ggg CgA ATT CCA AgC TT-3′ |
| HMK2007-94 | 5′-TgT ACA CTC gAg TAg Agg ggA TCC AgA TCT-3′ |
| Full length genomic CmSPI construction | |
| HMK2012-6 | 5′-CAA ATA GGA GAA GAT GTT CTC G-3′ |
| HMK2012-23 | 5′-CAG GTG ACA CAT GCG TAT AAC-3′ |
| Genomic PCR and probe synthesis | |
| HMK2011-102 | 5′-gaa ctt tct gga tct act tta ttt g-3′ |
| HMK2011-103 | 5′-aca caa act tca tct aac ctt aaa c-3′ |
| RT-PCR and probe synthesis | |
| HMK2013-60 | 5′-acg cGA ATT CGT CGA CCT CGA GAA CCT TCG GAA TTT CAA CAA C-3′ |
| HMK2013-61 | 5′-acg cGG ATC CGA GCT CAA GCT TAT GGC TGA TAT TTG TCC TCC T-3′ |
2.2. Sequence Analysis and Construction of Phylogenetic Tree
2.3. Construction of Binary Vector and Bacterial Strain
2.4. Plant Material and Transformation
2.5. Inoculation of Plant Viruses
2.6. Gene Expression Detection
2.6.1. Genomic PCR
2.6.2. Southern Hybridization
2.6.3. RT-PCR (Reverse Transcription PCR)
2.6.4. Northern Hybridization
2.6.5. Western Hybridization
2.6.6. Detection of Small Interfering RNA
3. Results
3.1. Cloning and Functional Characterization of C. metuliferus CmSPI Gene
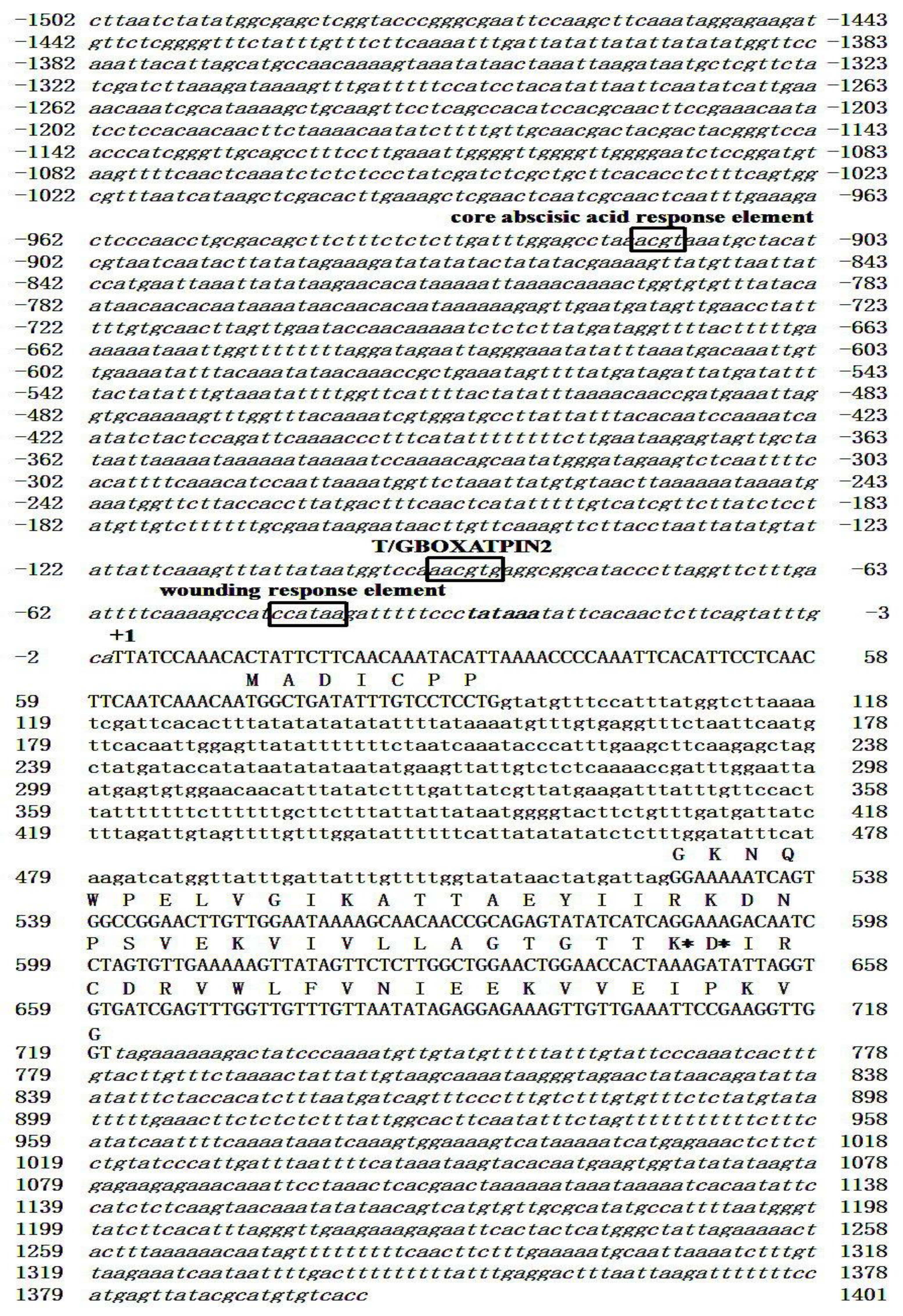
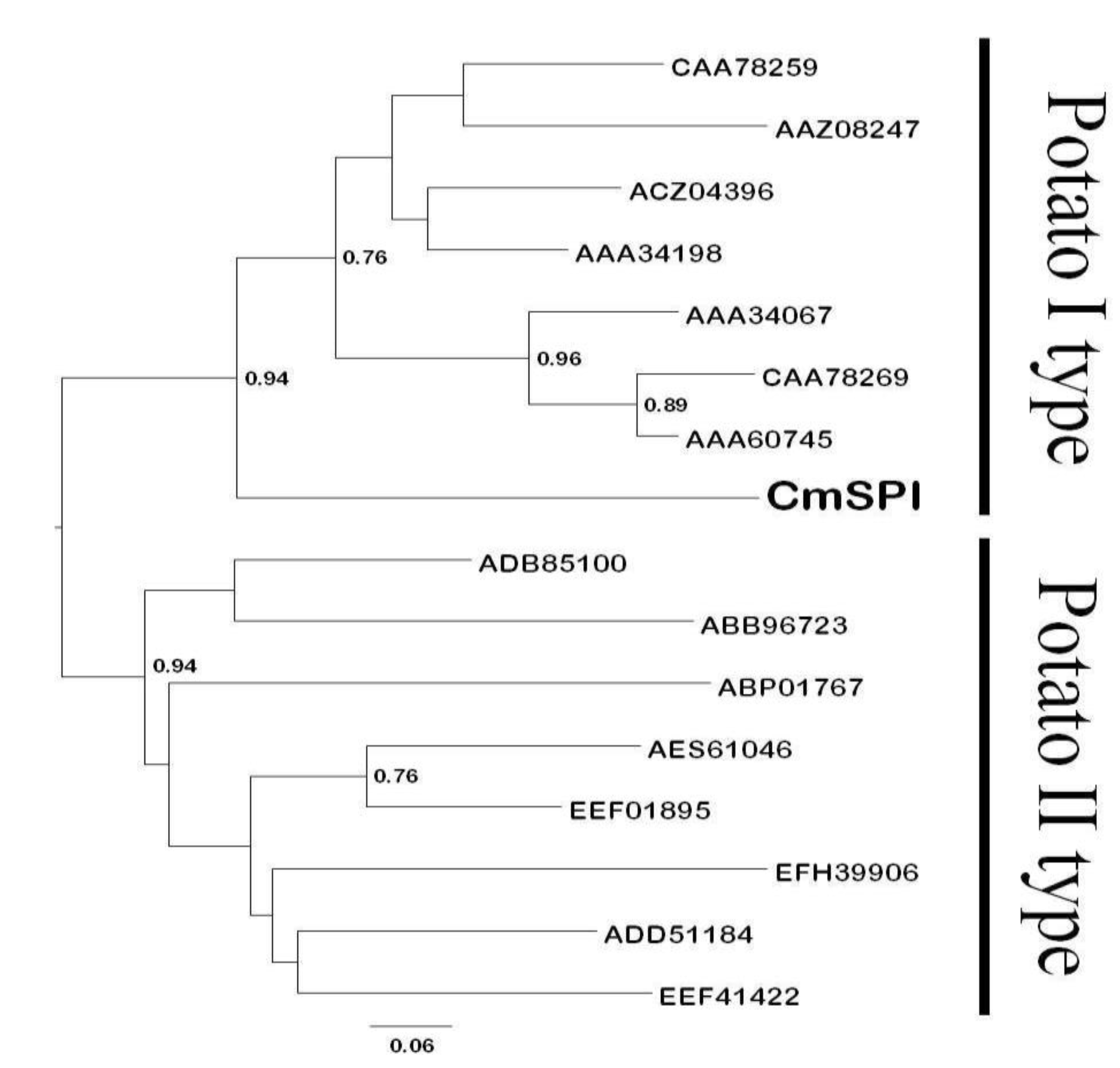
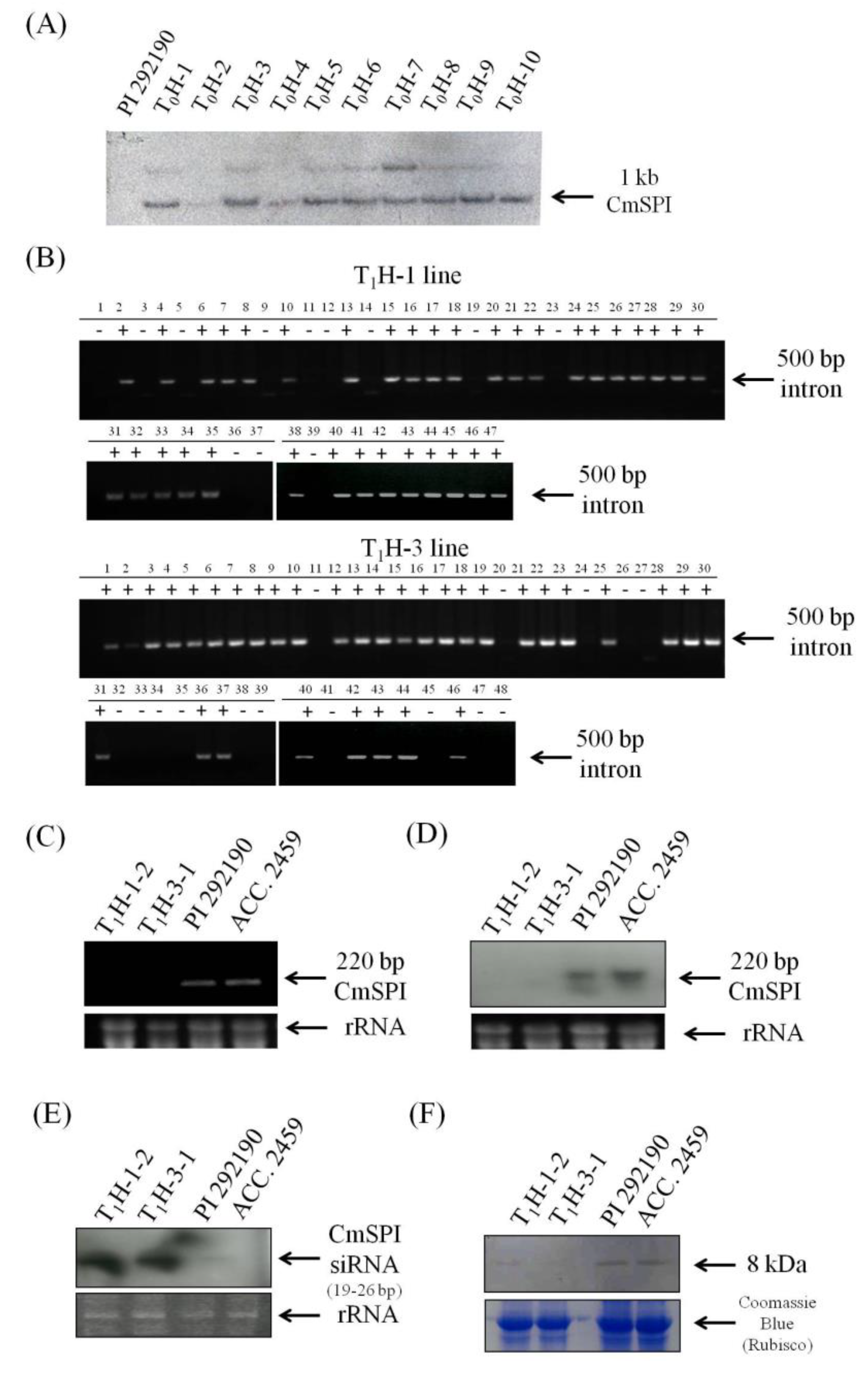
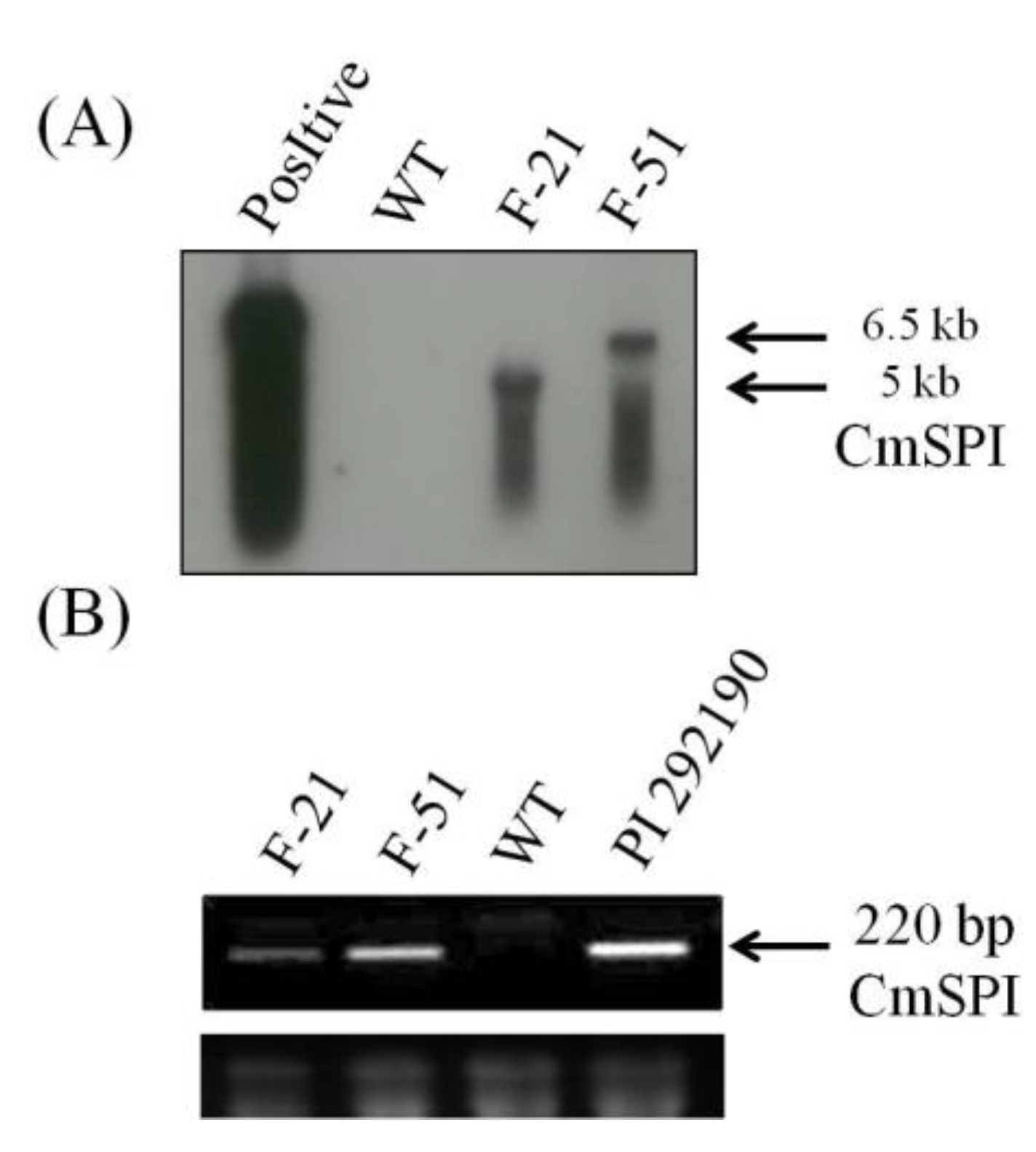
3.2. Generation of CmSPI RNAi Transgenic Plant


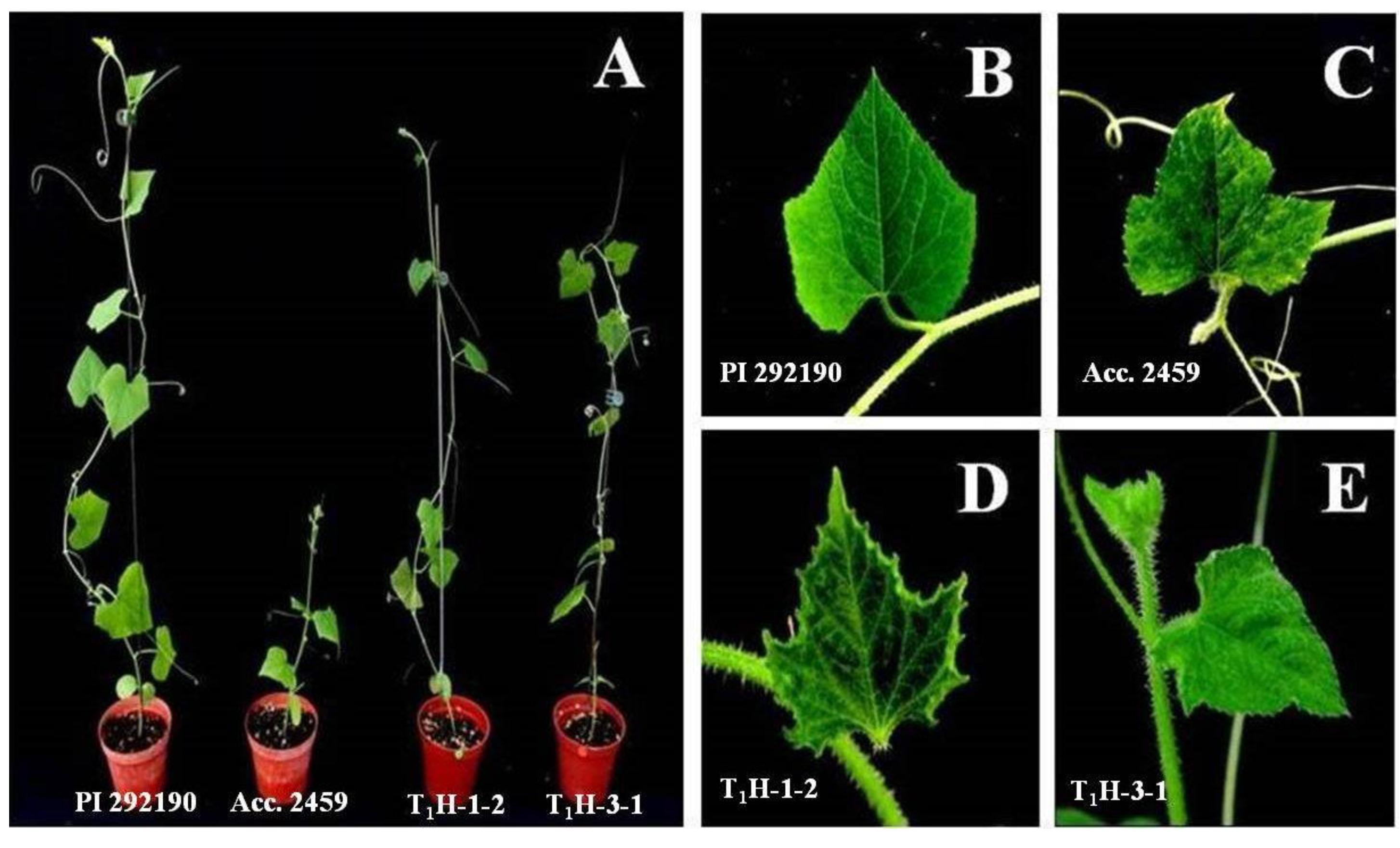
3.3. Alteration of Virus Resistance in CmSPI RNAi Transgenic C. metuliferus

3.4. Resistance to Virus in Transgenic N. benthamiana
| Transgenic Lines | Symptomatic Plants/Total of Inoculated Plants |
|---|---|
| T1F-3 | 28/28 |
| T1F-5 | 25/25 |
| T1F-14 | 21/22 |
| T1F-21 | 18/30 |
| T1F-51 | 17/30 |
| T1F-52 | 27/30 |
| T1F-53 | 29/29 |
| T1F-54 | 30/30 |
| T1F-55 | 27/28 |
| Non-transformed control | 27/27 |

| Transgenic Lines | Symptomatic Plants/Total of Inoculated Plants |
|---|---|
| T2F-21-2 | 33/44 |
| T2F-51-10 | 2/44 |
| Non-transformed control | 15/15 |
4. Discussion
4.1. CmSPI Is Involved in Plant Physiological Responses and Plant Defense
4.2. Silencing of CmSPI Alters PRSV Resistance in C. metuliferus
4.3. Expression of CmSPI in N. benthamiana Plants Can Supply Resistance for PVY
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- De Leo, F.; Volpicella, M.; Licciulli, F.; Liuni, S.; Gallerani, R.; Ceci, L.R. Plant-PIs: A database for plant protease inhibitors and their genes. Nucleic Acids Res. 2002, 30, 347–348. [Google Scholar] [CrossRef] [PubMed]
- Rawlings, N.D.; Barrett, A.J. Evolutionary families of peptidases. Biochem. J. 1993, 299, 205–218. [Google Scholar]
- Rawlings, N.D.; Tolle, D.P.; Barrett, A.J. MEROPS: The peptidase database. Nucleic Acid Res. 2004, 32, 160–164. [Google Scholar] [CrossRef] [PubMed]
- Kidric, M.; Kos, J.; Sabotic, J. Protease and their endogenous inhibitors in the plant response to abiotic stress. Botanica Serbica 2014, 38, 139–158. [Google Scholar]
- Ryan, C.A. Proteinase inhibitors in plants: Genes for improving defenses against insects and pathogens. Annu. Rev. Phytopathol. 1990, 28, 425–449. [Google Scholar] [CrossRef]
- Habib, H.; Fazili, K.M. Plant protease inhibitors: A defense strategy in plants. Biltechnol. Mol. Biol. 2007, 2, 68–85. [Google Scholar]
- Haq, S.K.; Atif, S.M.; Khan, R.H. Protein proteinase inhibitor genes in combat against insects, pests, and pathogens: Natural and engineered phytoprotection. Arch. Biochem. Biophys. 2004, 43, 145–159. [Google Scholar] [CrossRef] [PubMed]
- Falco, M.C.; Silva-Filho, M.C. Expression of soybean proteinases inhibitor in transgenic sugarcane plants: Effects on natural defense against Diatraea saccharalis. Plant Physiol. Biochem. 2003, 41, 761–766. [Google Scholar] [CrossRef]
- Urwin, P.E.; Levesley, A.; McPherson, M.J.; Atkinson, H.J. Transgenic resistance to the nematode Rotylenchulus reniformis conferred by Arabidopsis thaliana plants expressing proteinase inhibitors. Mol. Breed. 2000, 6, 257–264. [Google Scholar] [CrossRef]
- Gutierrez-Campos, R.; Torres-Acosta, J.A.; Saucedo-Arias, L.J.; Gomez-Lim, M.A. The use of cysteine proteinase inhibitors to engineer resistance against potyviruses in transgenic tobacco plants. Nat. Biotechnol. 1999, 17, 1223–1226. [Google Scholar] [PubMed]
- Provvidenti, R.; Robinson, R.W. Inheritance of resistance to Watermelon mosaic virus 1 in Cucumis metulifetrus (Naud.) May. J. Hered. 1977, 68, 56–57. [Google Scholar]
- Provvidenti, R.; Gonsalves, D. Resistance to Papaya ringspot virus in Cucumis metuliferus and its relationship to resistance to Watermelon mosaic virus 1. J. Hered. 1982, 73, 239–240. [Google Scholar]
- Yeh, S.D.; Chen, Y.H. Use of resistant Cucumis metuliferus for selection of nitrous-acid induced attenuated strains of papaya ringspot virus. Phytophathology 1989, 79, 1257–1261. [Google Scholar] [CrossRef]
- Lin, Y.T.; Jan, F.J.; Lin, C.W.; Chung, C.H.; Chen, J.C.; Yeh, S.D.; Ku, H.M. Differential gene expression in response to Papaya ringspot virus infection in Cucumis metuliferus using cDNA-amplified fragment length polymorphism analysis. PLoS ONE 2013, 8, e68749. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fire, A.; Xu, S.; Montgomery, M.K.; Kostas, S.A.; Driver, S.E.; Mello, C.C. Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 1998, 391, 806–811. [Google Scholar] [CrossRef] [PubMed]
- Axtell, M.J. Classification and comparison of small RNAs from plants. Annu. Rev. Plant Biol. 2013, 64, 137–159. [Google Scholar] [CrossRef] [PubMed]
- Waterhouse, P.M.; Helliwell, C.A. Exploring plant genomes by RNA-induced gene silencing. Nat. Rev. Genet. 2003, 4, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Wesley, S.V.; Helliwell, C.A.; Smith, N.A.; Wang, M.B.; Rouse, D.T.; Liu, Q.; Gooding, P.S.; Singh, S.P.; Abbott, D.; Stoutjesdijk, P.A.; et al. Construct design for efficient, effective and highthroughput gene silencing in plants. Plant J. 2001, 27, 581–590. [Google Scholar] [CrossRef] [PubMed]
- Sibert, P.D.; Chenchik, A.; Kellogg, D.E.; Lukyanov, K.A.; Lukyanov, S.A. An improved PCR method for walking in uncloned genomic DNA. Nucleic Acids Res. 1995, 23, 1087–1088. [Google Scholar] [CrossRef]
- Napoli, C.; Lemieux, C.; Jorgensen, R. Introduction of a chimeric chalcone synthase gene into petunia results in reversible co-suppression of homologous genes in trans. Plant Cell 1990, 2, 279–289. [Google Scholar] [CrossRef] [PubMed]
- Higo, K.; Ugawa, Y.; Iwamoto, M.; Korenaga, T. Plant cis-acting regulatory DNA elements (PLACE) database: 1999. Nucleic Acids Res. 1999, 27, 297–300. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Tamura, K.; Jakobsen, I.B.; Nei, M. MEGA2: Molecular evolutionary genetics analysis software. Bioinformatics 2001, 17, 1244–1245. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Hasegawa, M.; Kishino, H.; Yano, T. Dating of the human-ape splitting by a molecular clock of mitochondrial DNA. J. Mol. Evol. 1985, 22, 160–174. [Google Scholar] [CrossRef] [PubMed]
- Jan, F.J. Roles of nontarget DNA and viral gene length in influencing multi-virus resistance through homology-dependent gene silencing. Ph. D. Thesis, Department of Plant Pathology, Cornell University, Ithaca, NY, USA, 1998. [Google Scholar]
- Lin, Y.T.; Lin, C.W.; Chung, C.H.; Su, M.H.; Ho, H.Y.; Yeh, S.D.; Jan, F.J.; Ku, H.M. In Vitro Regeneration and genetic transformation of Cucumis metuliferus through cotyledon organogenesis. HortScience 2011, 46, 616–621. [Google Scholar]
- Horsch, R.B.; Fry, J.E.; Hoffmann, N.L.; Eichholtz, D.; Rogers, S.G.; Fraley, R.T. A simple method for transferring genes into plants. Science 1985, 227, 1229–1231. [Google Scholar]
- Fulton, T.M.; Chunwongse, J.; Tanksley, S.D. Microprep protocol for extraction of DNA from tomato and other herbaceous plants. Plant Mol. Biol. Rep. 1995, 13, 207–209. [Google Scholar] [CrossRef]
- Hattori, T.; Totsuka, M.; Hobo, T.; Kagaya, Y.; Yamamoto-Toyoda, A. Experimentally determined sequence requirement of ACGT-containing abscisic acid response element. Plant Cell Physiol. 2002, 43, 136–140. [Google Scholar] [CrossRef] [PubMed]
- Ma, S.; Bohnert, H.J. Integration of Arabidopsis thaliana stress-related transcript profiles, promoter structures, and cell-specific expression. Genome Biol. 2007, 8, eR49. [Google Scholar] [CrossRef] [PubMed]
- An, G.; Costa, M.A.; Ha, S.B. Nopaline synthase promoter is wound inducible and auxin lnducible. Plant Cell 1990, 2, 225–233. [Google Scholar] [CrossRef] [PubMed]
- Birk, Y. Protein proteinase inhibitors in legume seeds-overview. Arch. Latinoam. Nutr. 1996, 44, 26–30. [Google Scholar]
- Sugawara, H.; Shibuya, K.; Yoshioka, T.; Hashiba, T.; Satoh, S. Is a cysteine proteinase inhibitor involved in the regulation of petal wilting in senescing carnation (Dianthus caryophyllus L.) flowers? J. Exp. Bot. 2002, 53, 407–413. [Google Scholar] [CrossRef] [PubMed]
- Mosolov, V.V.; Valueva, T.A. Proteinase inhibitors and their function in plants: A review. Prikl. Biokhimiia Mikrobiol. 2005, 41, 261–282. [Google Scholar] [CrossRef]
- Huang, Y.; Xiao, B.; Xiong, L. Characterization of a stress responsive proteinase inhibitor gene with positive effect in improving drought resistance in rice. Planta 2007, 226, 73–85. [Google Scholar] [CrossRef] [PubMed]
- Sin, S.F.; Chye, M.L. Expression of proteinase inhibitor II proteins during floral development in Solanum americanum. Planta 2004, 219, 1010–1022. [Google Scholar] [CrossRef] [PubMed]
- Sin, S.F.; Yeung, E.C.; Chye, M.L. Downregulation of Solanum americanum genes encoding proteinase inhibitor II causes defective seed development. Plant J. 2006, 45, 58–70. [Google Scholar] [CrossRef] [PubMed]
- Drame, K.N.; Passaquet, C.; Repellin, A.; Zuily-Fodil, Y. Cloning, characterization and differential expression of a Bowman-Birk inhibitor during progressive water deficit and subsequent recovery in peanut (Arachis hypogaea) leaves. J. Plant Physiol. 2013, 170, 225–229. [Google Scholar] [CrossRef] [PubMed]
- Botella, M.A.; Xu, Y.; Prabha, T.N.; Zhao, Y.; Narasimhan, M.L.; Wilson, K.A.; Nielsen, S.S.; Bressan, R.A.; Hasegawa, P.M. Differential expression of soybean cysteine proteinase inhibitor genes during development and in response to wounding and methyl jasmonate. Plant Physiol. 1996, 112, 1201–1210. [Google Scholar] [CrossRef] [PubMed]
- Kasschau, K.D.; Xie, Z.; Allen, E.; Llave, C.; Chapman, E.J.; Krizan, K.A.; Carrington, J.C. P1/HC-Pro, a Viral Suppressor of RNA Silencing, Interferes with Arabidopsis development and miRNA function. Dev. Cell 2003, 4, 205–217. [Google Scholar] [CrossRef]
- Dasgupta, I.; Malathi, V.G.; Mukherjee, S.K. Genetic engineering for virus resistance. Curr. Sci. 2003, 84, 341–354. [Google Scholar]
- Yeh, S.D.; Jan, F.J.; Chiang, C.H.; Doong, T.J.; Chen, M.C.; Chung, P.H.; Bau, H.J. Complete nucleotide sequence and genetic organization of Papaya ringspot virus RNA. J. Gen. Virol. 1992, 73, 2531–2541. [Google Scholar] [CrossRef] [PubMed]
- Verchot, J.; Koonon, E.V.; Carrington, J.C. The 35S-kDa protein from the N-terminus of the potyviral polyprotein functions as a third virus-encoded proteinase. Virology 1991, 185, 527–538. [Google Scholar] [CrossRef]
- Jan, F.J.; Fagoaga, C.; Pang, S.Z.; Gonsalves, D. A single chimeric transgene derived from two distinct viruses confers multi-virus resistance in transgenic plants through homology-dependent gene silencing. J. Gen. Virol. 2000, 81, 2103–2109. [Google Scholar] [PubMed]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lin, C.-W.; Su, M.-H.; Lin, Y.-T.; Chung, C.-H.; Ku, H.-M. Functional Characterization of Cucumis metuliferus Proteinase Inhibitor Gene (CmSPI) in Potyviruses Resistance. Viruses 2015, 7, 3816-3834. https://doi.org/10.3390/v7072799
Lin C-W, Su M-H, Lin Y-T, Chung C-H, Ku H-M. Functional Characterization of Cucumis metuliferus Proteinase Inhibitor Gene (CmSPI) in Potyviruses Resistance. Viruses. 2015; 7(7):3816-3834. https://doi.org/10.3390/v7072799
Chicago/Turabian StyleLin, Chia-Wei, Mei-Hsiu Su, Yu-Tsung Lin, Chien-Hung Chung, and Hsin-Mei Ku. 2015. "Functional Characterization of Cucumis metuliferus Proteinase Inhibitor Gene (CmSPI) in Potyviruses Resistance" Viruses 7, no. 7: 3816-3834. https://doi.org/10.3390/v7072799






