Molecular Characterizations of Subcellular Localization Signals in the Nucleocapsid Protein of Porcine Epidemic Diarrhea Virus
Abstract
:1. Introduction
2. Results and Discussion
2.1. Polyclonal Antibody React Specifically with the N Protein of PEDV

2.2. N Protein Can Localize in Nucleolus in PEDV-Infected and Transfected Cell Lines

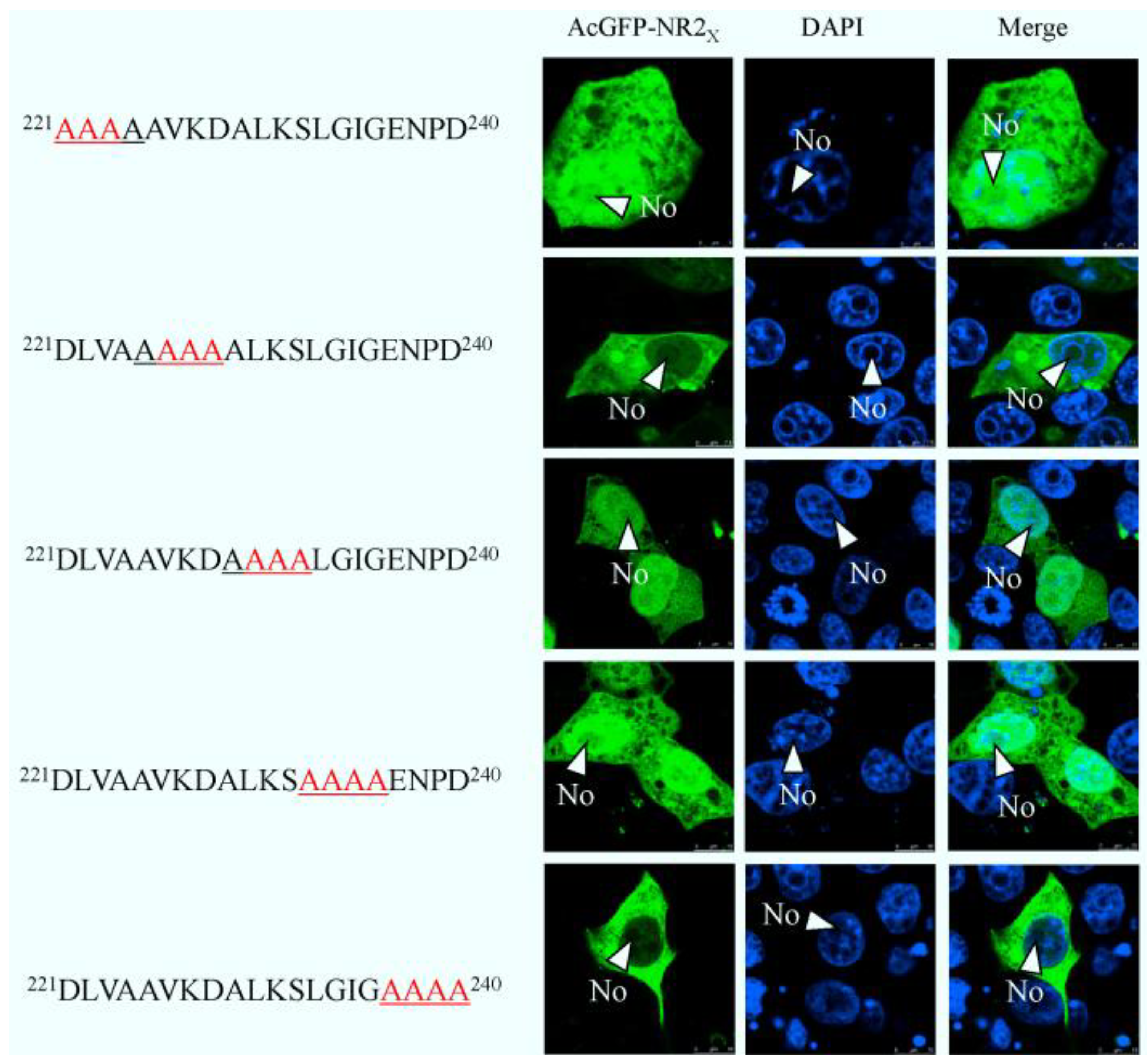
2.3. Bioinformatics Analysis of Subcellular Localization Signals and Delineation of a NoLS in Region 1 of PEDV N Protein
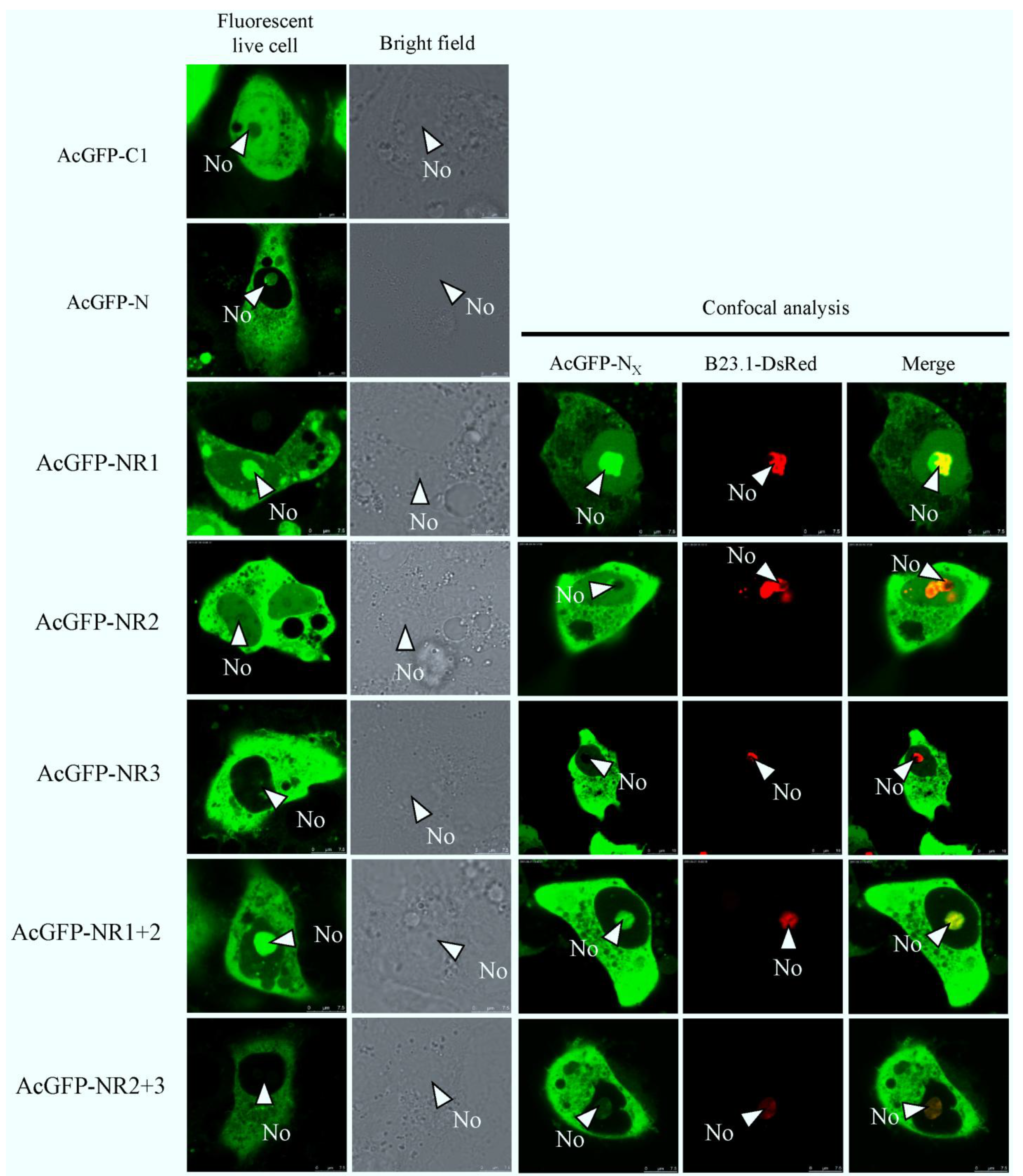
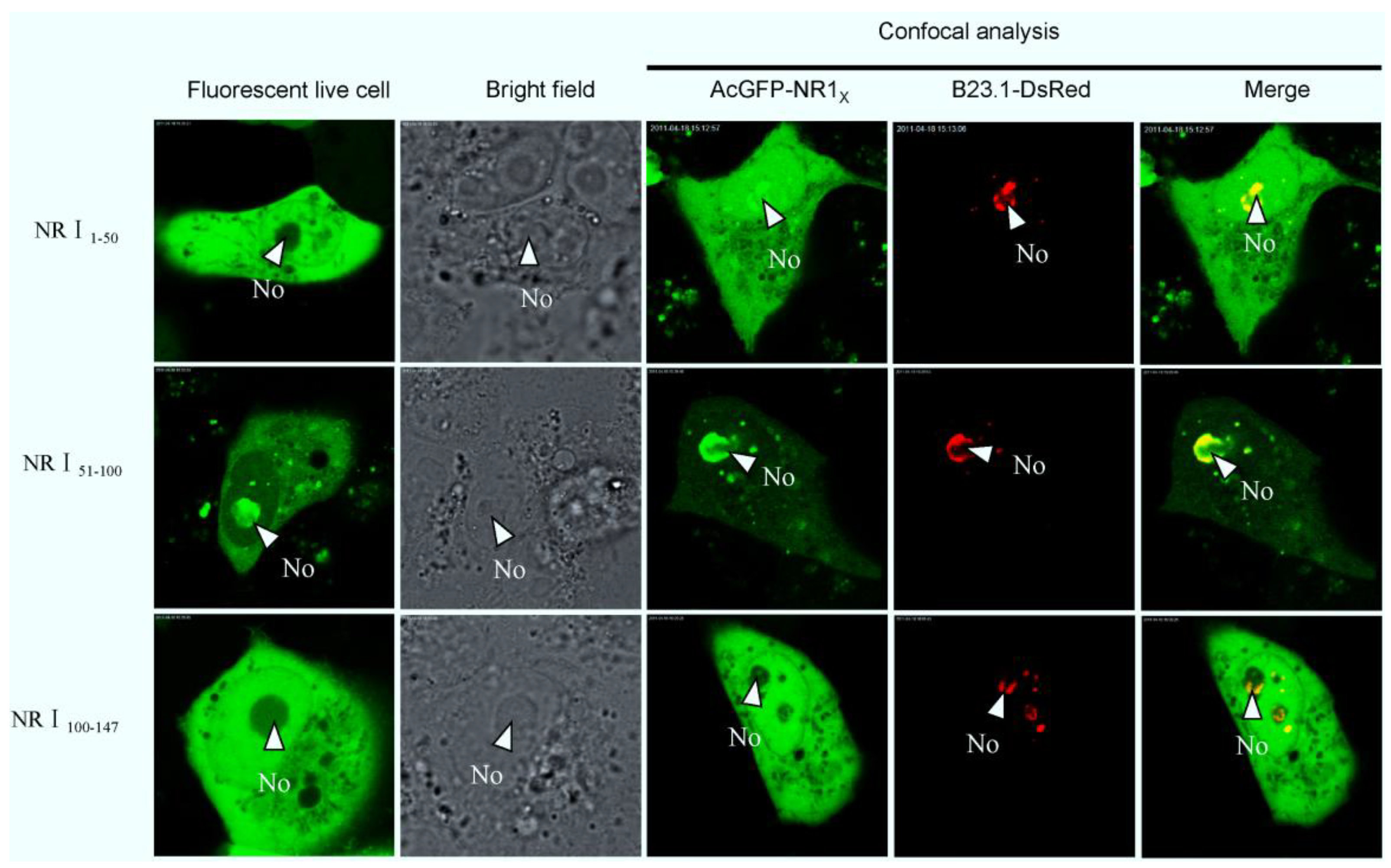
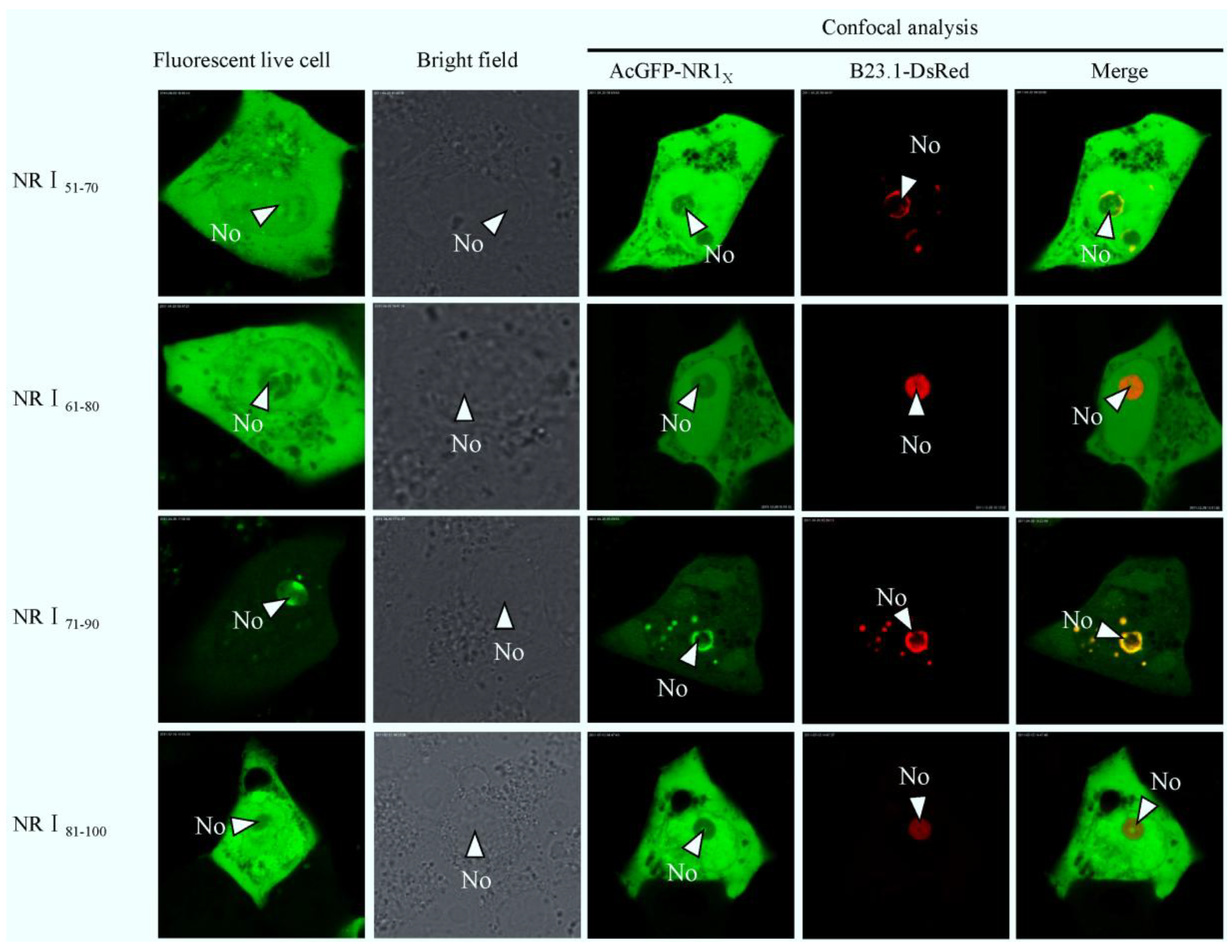

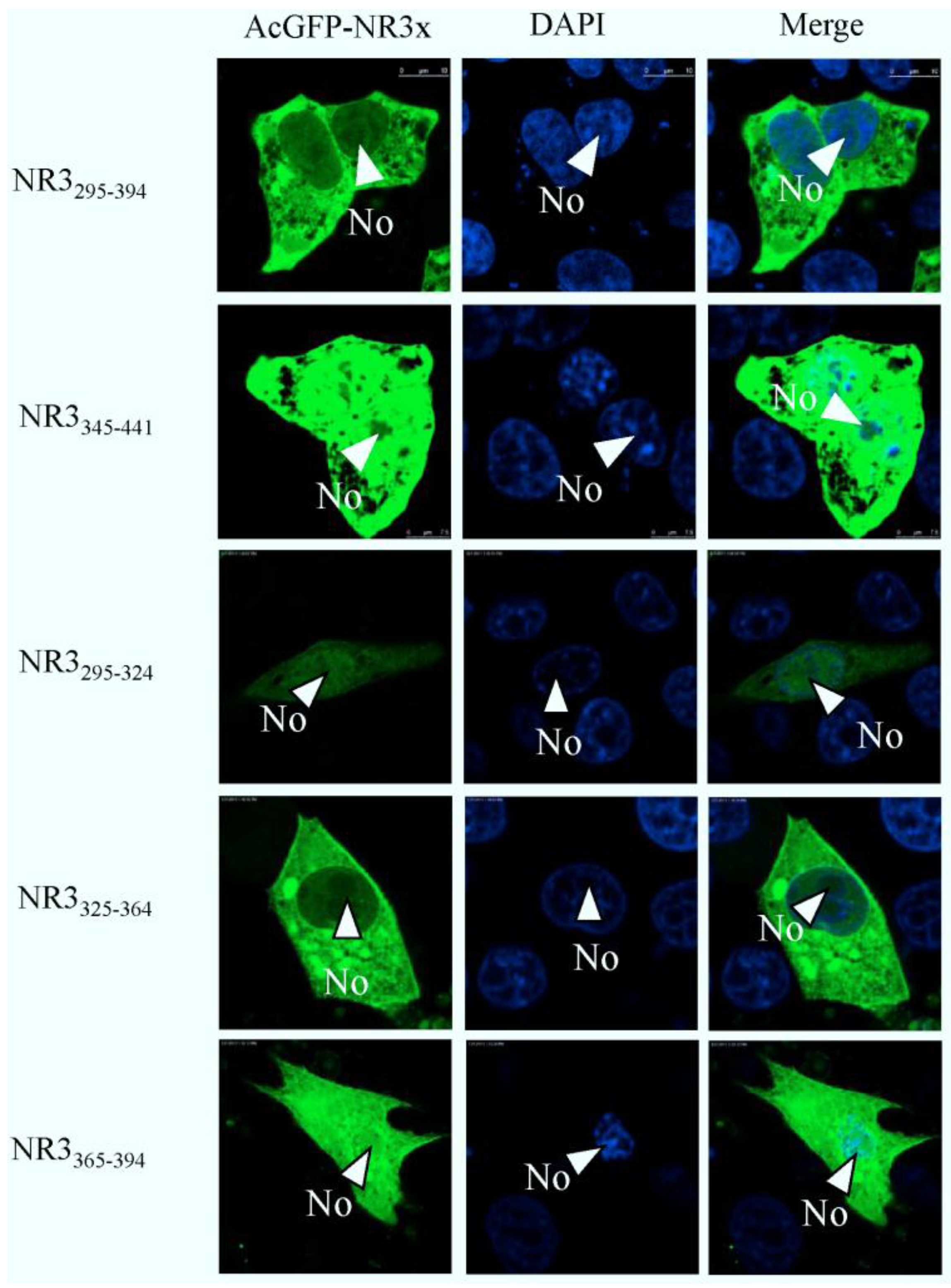
2.4. Mapping a Functional NES in PEDV N Protein Region 3
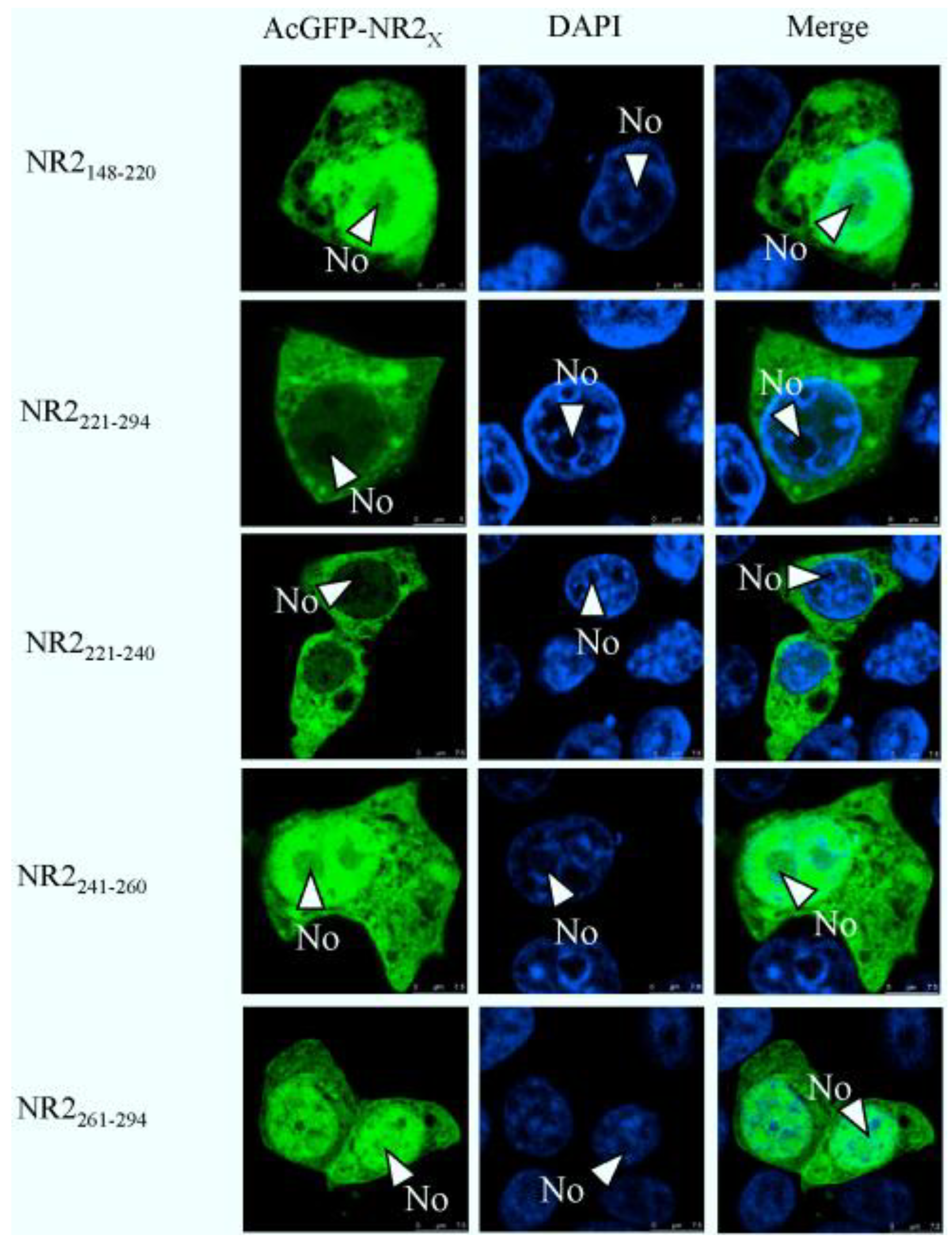

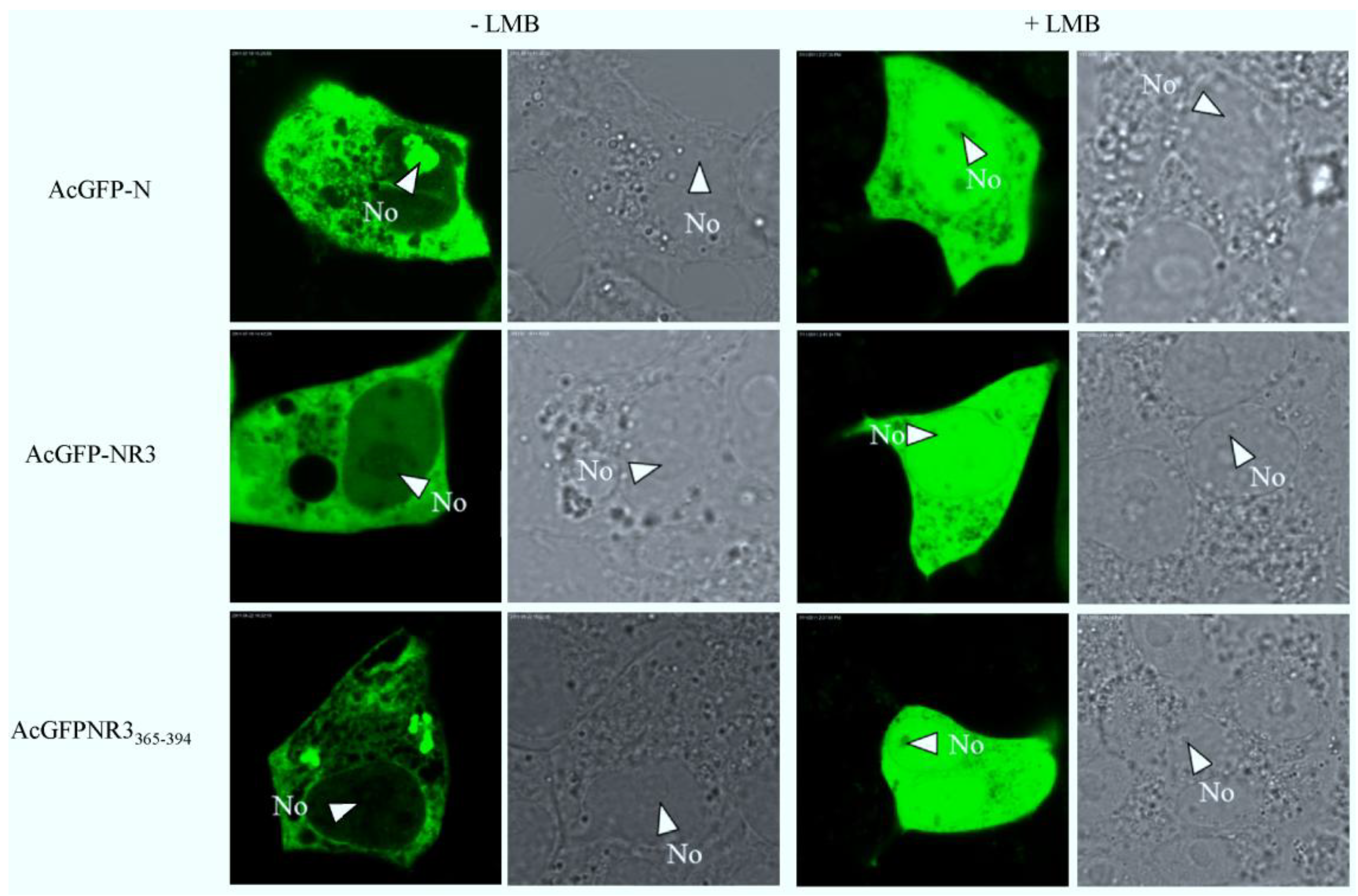
2.5. The Nuclear Export of N Protein Is CRM1 Dependent
3. Experimental Section
3.1. Cells Culture and Virus Infection
3.2. Construction of Plasmids
| Name of plasmids | Forward primer(5'→3') | Reverse primer(5'→3') |
|---|---|---|
| pET30a-N | CAGAGGATCCATGGCTTCTGTCAGC | CAAACTCGAGTTAATTTCCTGTATC |
| pcDNA3.1-N | AGAGGTACCATGGCTTCTGTCAGCTTTCAG | GCGCTCGAGTTAATTTCCTGTATCGAAGAT |
| pAcGFP-NR1+2 | GCACTCGAGCTATGGCTTCTGTCAGCTTTC | GCCCCATCTGGTACCTTAGCAAGCTGCTAC |
| pAcGFP-NR2+3 | TTCACCTCGAGCTATGCGTAGCAGGAGT | GCCGGTACCTTAATTTCCTGTATCGAAGAT |
| pAcGFP-NR1 | GCACTCGAGCTATGGCTTCTGTCAGCTTTC | GCCACGACTGGTACCTTACGAATTTGCACG |
| pAcGFP-NR2 | TTCACCTCGAGCTATGCGTAGCAGGAGT | GCCCCATCTGGTACCTTAGCAAGCTGCTAC |
| pAcGFP-NR3 | TAGCGCTCGAGCTATGTTCGGACCCAGAGG | GCCGGTACCTTAATTTCCTGTATCGAAGAT |
| pAcGFP-NR11–50 | GCACTCGAGCTATGGCTTCTGTCAGCTTTC | ATTGGTACCTTAAATTTGCTGGTCCTTATT |
| pAcGFP-NR151–100 | ACCCTCGAGTTGGGTACTGGAATGAGCAAA | CTTGGTACCTTATTT AGCAACCCAGAAAAC |
| pAcGFP-NR1101–147 | GGGCTCGAGAAGAAGGCGCAAAGACTGAAC | GCCACGACTGGTACCTTACGAATTTGCACG |
| pAcGFP-NR151–70 | ACCCTCGAGTTGGGTACTGGAATGAGCAAA | ATGGGTACCTTAAGGTTGTTCAATTCGCTC |
| pAcGFP-NR161–80 | TTCCTCGAGTTATGCGCCGTGGTGAGCGAA | CGCGGTACCTTATGTTCCGAGGTAGTAGAAATG |
| pAcGFP-NR171–90 | TTGCTCGAGTTTCCAATTGGCATTTCTACT | GCCGGTACCTTAAGTCCTATAACGGAGGTCGCCGTG |
| pAcGFP-NR181–100 | ACCCTCGAGTTGGACCTCACGGCGACCTCC | CTTGGTACCTTATTT AGCAACCCAGAAAAC |
| pAcGFP-NR2148–220 | GCACTCGAGCTCGTAGCAGGAGTCGTGGCAA | GCCGGTACCTTAATCGCGTGATG |
| pAcGFP-NR2221–294 | GCACTCGAGCTGATCTGGTGGCTGCTGTCAA | GCCGGTACCTTAGCAAGCTGCTACGCTATTTTC |
| pAcGFP-NR2221–240 | GCACTCGAGCTGATCTGGTGGCTGCTGTCAA | GCCGGTACCTTAGTCAGGATTTTCTCCA |
| pAcGFP-NR2241–260 | GCACTCGAGCTAGGCATAAGCAACAGCAGAA | GCCGGTACCTTATGTATTTTTGCCGCTGTTGTC |
| pAcGFP-NR2261–294 | GCACTCGAGCTCCTAAGAAGAACAAATCCA | GCCGGTACCTTAGCAAGCTGCTACGCTATTTTC |
| pAcGFP-NR3295–394 | GCACTCGAGCTTTCGGACCCAGAGGGGGCTT | GCCGGTACCTTACGTGGTTTCACGCTTGTTCT |
| pAcGFP-NR3345–441 | GCACTCGAGCTGACTCTTACGAGATTAC | GCCGGTACCTTAATTTCCTGTATCGAAGAT |
| pAcGFP-NR3295–324 | GCACTCGAGCTTTCGGACCCAGAGGGGGCTT | GCGGTACCACTGGCGATCT |
| pAcGFP-NR3325–364 | GCACTCGAGCTTTAGCACCAAAT | GCCGGTACCAACATTTGGATCTGACT |
| pAcGFP-NR3365–394 | GCACTCGAGCTGAGCTTCTTGTTTCACAGG | GCCGGTACCCGTGGTTTCACGCTTGTTCT |
| pAcGFP-NR2221–224DLVA-AAAA | TCGAGCTGCTGCTGCTGCTGCTGTCAAGGATGCACTTAAATCTTTGGGTATTGGAGAAAATCCTGACTAAGGTAC | CTTAGTCAGGATTTTCTCCAATACCCAAAGATTTAAGTGCATCCTTGACAGCAGCAGCAGCAGCAGC |
| pAcGFP-NR2225–228AVKD-AAAA | TCGAGCTGATCTGGTGGCTGCTGCTGCTGCTGCACTTAAATCTTTGGGTATTGGAGAAAATCCTGACTAAGGTAC | CTTAGTCAGGATTTTCTCCAATACCCAAAGATTTAAGTGCAGCAGCAGAAGCAGCCACCAGATCAGC |
| pAcGFP-NR2229–232ALKS-AAAA | TCGAGCTGATCTGGTGGCTGCTGTCAAGGATGCTGCTGCTGCTTTGGGTATTGGAGAAAATCCTGACTAAGGTAC | CTTAGTCAGGATTTTCTCCAATACCCAAAGCAGCAGCAGCATCCTTGACAGCAGCCACCAGATCAGC |
| pAcGFP-NR2233–236LGIG-AAAA | TCGAGCTGATCTGGTGGCTGCTGTCAAGGATGCACTTAAATCTGCTGCTGCTGCTGAAAATCCTGACTAAGGTAC | CTTAGTCAGGATTTTCAGCAGCAGCAGCAGATTTAAGTGCATCCTTGACAGCAGCCACCAGATCAGC |
| pAcGFP-NR2237–240ENPD-AAAA | TCGAGCTGATCTGGTGGCTGCTGTCAAGGATGCACTTAAATCTTTGGGTATTGGAGCTGCTGCTGCTTAAGGTAC | CTTAAGCAGCAGCAGCTCCAATACCCAAAGATTTAAGTGCATCCTTGACAGCAGCCACCAGATCAGC |
| pAcGFP-N ∆221–236 | GCCTCGAGATGGCTTCTGTCAGCTTT(1-U) | ATGCCTGTCAGGATTTTCATCGCGTGATGTCATT(1-L) |
| GGAATGACATCACGCGATGAAAATCCTGACAGGCATAA(2-U) | GCGGTACCTTAATTTCCTGTGTC(2-L) |
3.3. Generation of Polyclonal Antisera in Mouse
3.4. Transfection and Western Blotting
3.5. Indirect Immunofluorescence Assay
3.6. Confocal Microscopic Analysis
3.7. LMB Treatment
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References and Notes
- Egberink, H.F.; Ederveen, J.; Callebaut, P.; Horzinek, M.C. Characterization of the structural proteins of porcine epizootic diarrhea virus, strain CV777. Am. J. Vet. Res. 1988, 49, 1320–1324. [Google Scholar]
- Hiscox, J.A.; Wurm, T.; Wilson, L.; Britton, P.; Cavanagh, D.; Brooks, G. The coronavirus infectious bronchitis virus nucleoprotein localizes to the nucleolus. J. Virol. 2001, 75, 506–512. [Google Scholar] [CrossRef]
- Brian, D.; Baric, R.S. Coronavirus genome structure and replication. In Coronavirus Replication and Reverse Genetics; Enjuanes, L., Ed.; Springer-Verlag: Berlin, Germany, 2005; pp. 2–22. [Google Scholar]
- You, J.; Dove, B.K.; Enjuanes, L.; DeDiego, M.L.; Alvarez, E.; Howell, G.; Heinen, P.; Zambon, M.; Hiscox, J.A. Subcellular localization of the severe acute respiratory syndrome coronavirus nucleocapsid protein. J. Gen. Virol. 2005, 86, 3303–3310. [Google Scholar] [CrossRef]
- You, J.H.; Reed, M.L.; Hiscox, J.A. Trafficking motifs in the SARS coronavirus nucleocapsid protein. Biochem. Biophys. Res. Commun. 2007, 358, 1015–1020. [Google Scholar] [CrossRef]
- Li, F.Q.; Xiao, H.; Tam, J.P.; Liu, D.X. Sumoylation of the nucleocapsid protein of severe acute respiratory syndrome coronavirus. FEBS Lett. 2005, 579, 2387–2396. [Google Scholar] [CrossRef]
- Li, Y.H.; Li, J.; Liu, X.E.; Wang, L.; Li, T.; Zhou, Y.H.; Zhuang, H. Detection of the nucleocapsid protein of severe acute respiratory syndrome coronavirus in serum: Comparison with results of other viral markers. J. Virol. Methods 2005, 130, 45–50. [Google Scholar] [CrossRef]
- Surjit, M.; Liu, B.; Chow, V.T.; Lal, S.K. The nucleocapsid protein of severe acute respiratory syndrome-coronavirus inhibits the activity of cyclincyclin-dependent kinase complex and blocks S phase progression in mammalian cells. J. Biol. Chem. 2006, 281, 10669–10681. [Google Scholar] [CrossRef]
- Spiegel, M.; Pichlmair, A.; Martinez-Sobrido, L.; Cros, J.; García-Sastre, A.; Haller, O.; Weber, F. Inhibition of beta interferon induction by severe acute respiratory syndrome coronavirus suggests a two-step model for activation of interferon regulatory factor-3. J. Virol. 2005, 79, 2079–2086. [Google Scholar] [CrossRef]
- Zheng, B.; He, M.L.; Wong, K.L.; Lum, C.T.; Poon, L.L.; Peng, Y.; Guan, Y.; Lin, M.C.; Kung, H.F. Potent inhibition of SARS associated coronavirus (SCoV) infection and replication by type I interferons (IFN-alpha/beta) but not by type II interferon (IFN-gamma). J. Interferon Cytokine Res. 2004, 24, 388–390. [Google Scholar] [CrossRef] [Green Version]
- Yan, X.; Hao, Q.; Mu, Y.; Timani, K.A.; Ye, L.; Zhu, Y.; Wu, J. Nucleocapsid protein of SARS-CoV activates the expression of cyclooxygenase-2 by binding directly to regulatory elements for nuclear factor-kappa B and CCAAT/enhancer binding protein. Int. J. Biochem. Cell Biol. 2006, 38, 1417–1428. [Google Scholar] [CrossRef]
- Kopecky-Bromberg, S.A.; Martinez-Sobrido, L.; Frieman, M.; Baric, R.A.; Palese, P. Severe acute respiratory syndrome coronavirus open reading frame (ORF) 3b, ORF 6, and nucleocapsid proteins function as interferon antagonists. J. Virol. 2007, 81, 548–557. [Google Scholar] [CrossRef]
- He, R.; Leeson, A.; Andonov, A.; Li, Y.; Bastien, N.; Cao, J.; Osiowy, C.; Dobie, F.; Cutts, T.; Ballantine, M.; et al. Activation of AP-1 signal transduction pathway by SARS coronavirus nucleocapsid protein. Biochem. Biophys. Res. Commun. 2003, 311, 870–876. [Google Scholar] [CrossRef]
- Surjit, M.; Liu, B.; Kumar, P.; Chow, V.T.; Lal, S.K. The nucleocapsid protein of the SARS coronavirus is capable of self-association through a C-terminal 209 amino acid interaction domain. Biochem. Biophys. Res. Commun. 2004, 317, 1030–1036. [Google Scholar] [CrossRef]
- Surjit, M.; Liu, B.; Jameel, S.; Chow, V.T.; Lal, S.K. The SARS coronavirus nucleocapsid protein induces actin reorganization and apoptosis in COS-1 cells in the absence of growth factors. Biochem. J. 2004, 383, 13–18. [Google Scholar] [CrossRef]
- Zhang, L.; Wei, L.; Jiang, D.; Wang, J.; Cong, X.; Fei, R. SARS-CoV nucleocapsid protein induced apoptosis of COS-1 mediated by the mitochondrial pathway. Artif. Cells Blood Substit. Immobil. Biotechnol. 2007, 35, 237–253. [Google Scholar] [CrossRef]
- Luo, C.; Luo, H.; Zheng, S.; Gui, C.; Yue, L.; Yu, C.; Sun, T.; He, P.; Chen, J.; Shen, J.; et al. Nucleocapsid protein of SARS coronavirus tightly binds to human cyclophilin A. Biochem. Biophys. Res. Commun. 2004, 321, 557–565. [Google Scholar] [CrossRef]
- Sonia, Z.; Isabel, S.; Jose, L.M.; Patricia, S.; Juan, P.D.; Luis, E. Coronavirus nucleocapsid protein is an RNA chaperone. Virology 2007, 357, 215–227. [Google Scholar] [CrossRef]
- Lam, Y.W.; Trinkle, M.L.; Lamond, A.I. The nucleolus. J. Cell Sci. 2005, 118, 1335–1337. [Google Scholar] [CrossRef]
- Rowland, R.R.; Yoo, D. Nucleolar-cytoplasmic shuttling of PRRSV nucleocapsid protein: A simple case of molecular mimicry or the complex regulation by nuclear import, nucleolar localization and nuclear export signal sequences. Virus Res. 2003, 95, 23–33. [Google Scholar] [CrossRef]
- Wurm, T.; Chen, H.; Hodgson, T.; Britton, P.; Brooks, G.; Hiscox, J.A. Localization to the nucleolus is a common feature of coronavirus nucleoproteins, and the protein may disrupt host cell division. J. Virol. 2001, 75, 9345–9356. [Google Scholar] [CrossRef]
- Timani, K.A.; Liao, Q.; Ye, L.; Zeng, Y.; Liu, J.; Zheng, Y.; Ye, L.; Yang, X.; Lingbao, K.; Gao, J.; et al. Nuclear/nucleolar localization properties of C-terminal nucleocapsid protein of SARS coronavirus. Virus Res. 2005, 114, 23–34. [Google Scholar] [CrossRef]
- Chen, H.; Wurm, T.; Britton, P.; Brooks, G.; Hiscox, J.A. Interaction of the coronavirus nucleoprotein with nucleolar antigens and the host cell. J. Virol. 2002, 76, 5233–5250. [Google Scholar]
- Yoo, D.; Wootton, S.K.; Li, G.; Song, C.; Rowland, R.R. Colocalization and interaction of the porcine arterivirus nucleocapsid protein with the small nucleolar RNA-associated protein fibrillarin. J. Virol. 2003, 77, 12173–12183. [Google Scholar]
- Zeng, Y.; Ye, L.; Zhu, S.; Zheng, H.; Zhao, P.; Cai, W.; Su, L.; She, Y.; Wu, Z. The nucleocapsid protein of SARS-associated coronavirus inhibits B23 phosphorylation. Biochem. Biophys. Res. Commun. 2008, 369, 287–291. [Google Scholar] [CrossRef]
- Carmo-Fonseca, M.; Mendes-Soares, L.; Campos, I. To be or not to be in the nucleolus. Nat. Cell Biol. 2000, 2, E107–E112. [Google Scholar] [CrossRef]
- Bednenko, J.; Cingolani, G.; Gerace, L. Nucleocytoplasmic transport: Navigating the channel. Traffic. 2003, 4, 127–135. [Google Scholar] [CrossRef]
- Gorlich, D.; Mattaj, I.W. Nucleocytoplasmic transport. Science 1996, 271, 1513–1518. [Google Scholar]
- La Cour, T.; Gupta, R.; Rapacki, K.; Skriver, K.; Poulsen, F.M.; Brunak, S. NESbase version 1.0: A database of nuclear export signals. Nucleic Acids Res. 2003, 31, 393–396. [Google Scholar] [CrossRef]
- Matsuyama, A.; Arai, R.; Yashiroda, Y.; Shirai, A.; Kamata, A.; Sekido, S.; Kobayashi, Y.; Hashimoto, A.; Hamamoto, M.; Hiraoka, Y.; et al. ORFeome cloning and global analysis of protein localization in the fission yeast Schizosaccharomyces pombe. Nat. Biotechnol. 2006, 24, 841–847. [Google Scholar] [CrossRef]
- Fornerod, M.; Ohon, M.; Yoshida, M.; Mattaj, I.W. CRM1 is an export receptor for leucine-rich nuclear export signals. Cell 1997, 9, 1051–1060. [Google Scholar]
- Cokol, M.; Nair, R.; Rost, B. Finding nuclear localization signals. EMBO Rep. 2000, 1, 411–415. [Google Scholar] [CrossRef]
- Nakai, K.; Horton, P. PSORT: A programme for detecting sorting signals in proteins and predicting their subcellular localization. Trends Biochem. Sci. 1999, 24, 34–36. [Google Scholar] [CrossRef]
- Reed, M.L.; Dove, B.K.; Jackson, R.M.; Collins, R.; Brooks, G.; Hiscox, J.A. Delineation and modelling of a nucleolar retention signal in the coronavirus nucleocapsid protein. Traffic 2006, 7, 833–848. [Google Scholar] [CrossRef]
- Siomi, H.; Shida, H.; Nam, S.H.; Nosaka, T.; Maki, M.; Hatanaka, M. Sequence requirements for nucleolar localization of human T cell leukemia virus type I pX protein, which regulates viral RNA processing. Cell 1988, 55, 197–209. [Google Scholar] [CrossRef]
- Cheng, G.; Brett, M.E.; He, B. Signals that dictate nuclear, nucleolar, and cytoplasmic shuttling of the gamma (1) 34.5 protein of herpes simplex virus type 1. J. Virol. 2002, 76, 9434–9445. [Google Scholar] [CrossRef]
- Lohrum, M.A.; Ashcroft, M.; Kubbutat, M.H.; Vousden, K.H. Identification of a cryptic nucleolar-localization signal in MDM2. Nat. Cell Biol. 2000, 2, 179–181. [Google Scholar] [CrossRef]
- Liu, J.L.; Lee, L.F.; Ye, Y.; Qian, Z.; Kung, H.J. Nucleolar and nuclear localization properties of a herpesvirus bZIP oncoprotein, MEQ. J. Virol. 1997, 71, 3188–3196. [Google Scholar]
- Birbach, A.; Bailey, S.T.; Ghosh, S.; Schmid, J.A. Cytosolic, nuclear and nucleolar localization signals determine subcellular distribution and activity of the NF-kappa B inducing kinase NIK. J. Cell Sci. 2004, 117, 3615–3624. [Google Scholar] [CrossRef]
- Nagahama, M.; Hara, Y.; Seki, A.; Yamazoe, T.; Kawate, Y.; Shinohara, T.; Hatsuzawa, K.; Tani, K.; Tagaya, M. NVL2 is a nucleolar AAA-ATPase that interacts with ribosomal protein L5 through its nucleolar localization sequence. Mol. Biol. Cell 2004, 15, 5712–5723. [Google Scholar] [CrossRef]
- Guo, H.; Ding, Q.; Lin, F.; Pan, W.; Lin, J.; Zheng, A.C. Characterization of the nuclear and nucleolar localization signals of bovine herpesvirus-1 infected cell protein 27. Virus Res. 2009, 145, 312–320. [Google Scholar] [CrossRef]
- Valdez, B.C.; Perlaky, L.; Henning, D.; Saijo, Y.; Chan, P.K.; Busch, H. Identification of the nuclear and nucleolar localization signals of the protein p120. J. Biol. Chem. 1994, 269, 23776–23783. [Google Scholar]
- Song, Z.; Wu, M. Identification of a novel nucleolar localization signal and a degradation signal in Survivin-deltaEx3: A potential link between nucleolus and protein degradation. Oncogene 2005, 24, 2723–2734. [Google Scholar] [CrossRef]
- Guo, H.S.; Ding, S.W. A viral protein inhibits the long range signaling activity of the gene silencing signal. EMBO J. 2002, 21, 398–407. [Google Scholar] [CrossRef]
- Kim, H.; Chang, D.J.; Lee, J.A.; Lee, Y.S.; Kaang, B.K. Identification of nuclear/nucleolar localization signal in Aplysia learning associated protein of slug with a molecular mass of 18 kDa homologous protein. Neurosci. Lett. 2003, 343, 134–138. [Google Scholar] [CrossRef]
- Mears, W.E.; Lam, V.; Rice, S.A. Identification of nuclear and nucleolar localization signals in the herpes simplex virus regulatory protein ICP27. J. Virol. 1995, 69, 935–947. [Google Scholar]
- Lixin, R.; Efthymiadis, A.; Henderson, B.; Jans, D.A. Novel properties of the nucleolar targeting signal of human angiogenin. Biochem. Biophys. Res. Commun. 2001, 284, 185–193. [Google Scholar] [CrossRef]
- Sheng, Z.; Lewis, J.A.; Chirico, W.J. Nuclear and nucleolar localization of 18-kDa fibroblast growth factor-2 is controlled by C-terminal signals. J. Biol. Chem. 2004, 279, 40153–40160. [Google Scholar] [CrossRef]
- Thebault, S.; Basbous, J.; Gay, B.; Devaux, C.; Mesnard, J.M. Sequence requirement for the nucleolar localization of human I-mfa domain-containing protein (HIC p40). Eur. J. Cell Biol. 2000, 79, 834–838. [Google Scholar] [CrossRef]
- Cochrane, A.W.; Perkins, A.; Rosen, C.A. Identification of sequences important in the nucleolar localization of human immunodeficiency virus Rev: Relevance of nucleolar localization to function. J. Virol. 1990, 64, 881–885. [Google Scholar]
- Siomi, H.; Shida, H.; Maki, M.; Hatanaka, M. Effects of a highly basic region of human immunodeficiency virus Tat protein on nucleolar localization. J. Virol. 1990, 64, 1803–1807. [Google Scholar]
- Lange, A.; Mills, R.E.; Lange, C.J.; Stewart, M.; Devine, S.E.; Corbett, A.H. Classical nuclear localization signals: Definition, function, and interaction with importin alpha. J. Biol. Chem. 2007, 282, 5101–5105. [Google Scholar]
- Macara, I.G. Transport into and out of the nucleus. Microbiol. Mol. Biol. Rev. 2001, 65, 570–594. [Google Scholar] [CrossRef]
- Kubota, N.; Terauchi, Y.; Miki, H.; Tamemoto, H.; Yamauchi, T.; Komeda, K.; Satoh, S.; Nakano, R.; Ishii, C.; Sugiyama, T.; et al. PPAR γ mediates high-fat diet-induced adipocyte hypertrophy and insulin resistance. Mol. Cell 1999, 4, 597–609. [Google Scholar] [CrossRef]
- Dohner, K.; Wolfstein, A.; Prank, U.; Echeverri, C.; Dujardin, D.; Vallee, R.; Sodeik, B. Function of dynein and dynactin in herpessimplex virus capsid transport. Mol. Biol. Cell 2002, 13, 2795–2809. [Google Scholar] [CrossRef]
- Giannakakou, P.; Sackett, D.L.; Ward, Y.; Webster, K.R.; Blagosklonny, M.V.; Fojo, T. p53 Is associated with cellular microtubules and is transported to the nucleus by dynein. Nat. Cell Biol. 2000, 2, 709–717. [Google Scholar] [CrossRef]
- Hiscox, J.A. RNA viruses: Hijacking the dynamic nucleolus. Nat. Rev. Microbiol. 2007, 5, 119–127. [Google Scholar] [CrossRef]
- Cawood, R.; Harrison, S.M.; Dove, B.; Reed, M.L.; Hiscox, J.A. Cell cycle dependent localization of the coronavirus nucleocapsid protein. Cell Cycle 2007, 6, 863–867. [Google Scholar] [CrossRef]
- Lv, M.J.; Chen, J.F.; Shi, H.Y.; Chen, C.J.; Fan, X.P.; Shen, S.C.; Feng, L. Co-localization analysis between porcine epidemic diarrhea virus nucleocapsid protein and nucleolar phosphoprotein B23.1. Acta Microbiol. Sin. 2011, 51, 643–647. [Google Scholar]
© 2014 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Shi, D.; Lv, M.; Chen, J.; Shi, H.; Zhang, S.; Zhang, X.; Feng, L. Molecular Characterizations of Subcellular Localization Signals in the Nucleocapsid Protein of Porcine Epidemic Diarrhea Virus. Viruses 2014, 6, 1253-1273. https://doi.org/10.3390/v6031253
Shi D, Lv M, Chen J, Shi H, Zhang S, Zhang X, Feng L. Molecular Characterizations of Subcellular Localization Signals in the Nucleocapsid Protein of Porcine Epidemic Diarrhea Virus. Viruses. 2014; 6(3):1253-1273. https://doi.org/10.3390/v6031253
Chicago/Turabian StyleShi, Da, Maojie Lv, Jianfei Chen, Hongyan Shi, Sha Zhang, Xin Zhang, and Li Feng. 2014. "Molecular Characterizations of Subcellular Localization Signals in the Nucleocapsid Protein of Porcine Epidemic Diarrhea Virus" Viruses 6, no. 3: 1253-1273. https://doi.org/10.3390/v6031253




