Protective Effect of Surfactant Protein D in Pulmonary Vaccinia Virus Infection: Implication of A27 Viral Protein
Abstract
:1. Introduction

2. Results and Discussion
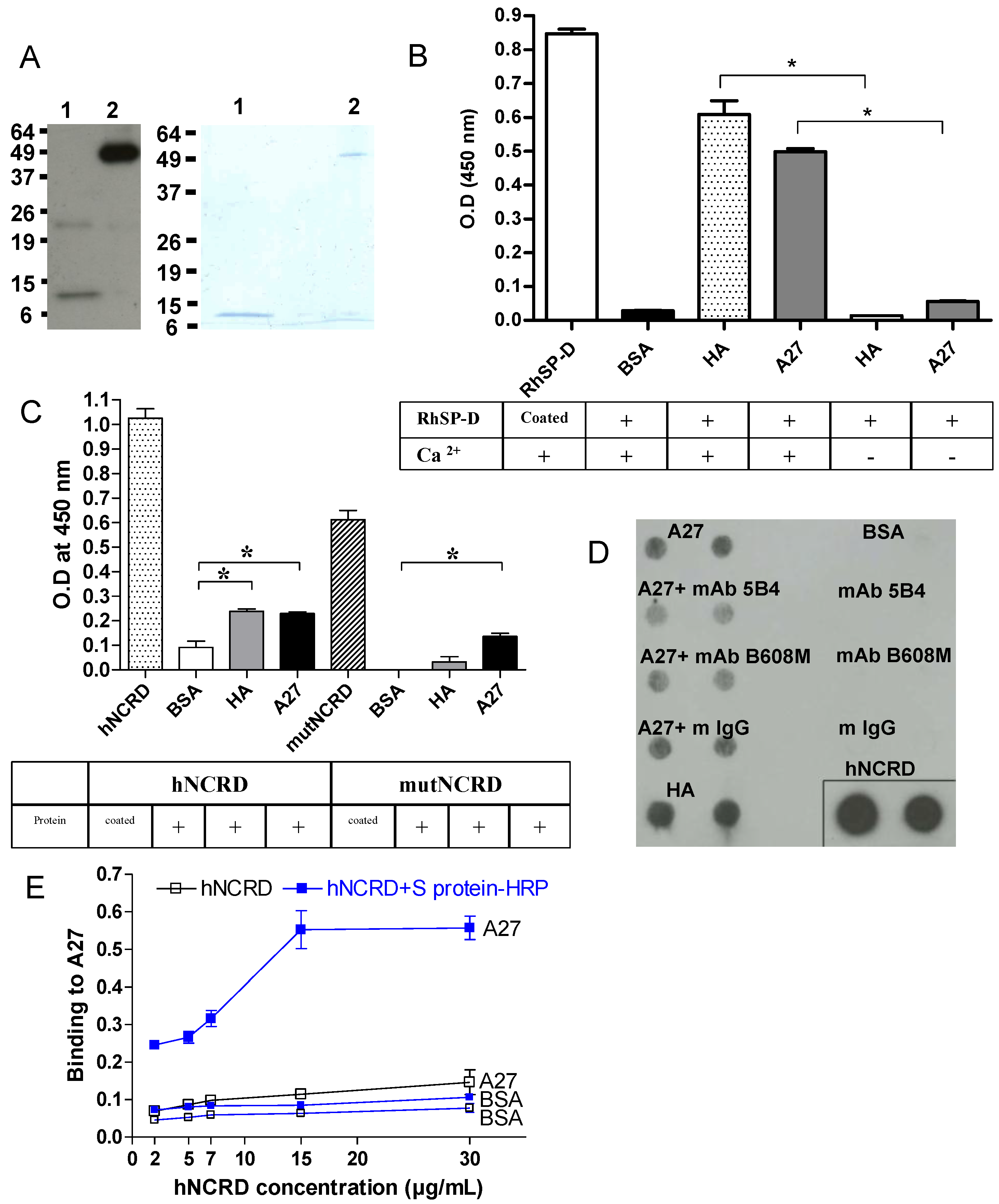
2.1. Vaccinia virus interaction with surfactant protein D

2.2. SP-D inhibits VACV infectivity
2.3. Multimeric SP-D but not NCRD domains inhibits VACV infectivity
 , VACV-WR+RhSP-D; □, VACV-WR+BSA. (C) VACV-WR was incubated with 0.6 µg/mL of RhSP-D and 2 µg/mL of NCRD domain of SP-D from human (hNCRD), rat (rNCRD), or human E31K mutant (mutNCRD) for 1 h.
, VACV-WR+RhSP-D; □, VACV-WR+BSA. (C) VACV-WR was incubated with 0.6 µg/mL of RhSP-D and 2 µg/mL of NCRD domain of SP-D from human (hNCRD), rat (rNCRD), or human E31K mutant (mutNCRD) for 1 h.
 , VACV-WR+RhSP-D; □, VACV-WR+BSA. (C) VACV-WR was incubated with 0.6 µg/mL of RhSP-D and 2 µg/mL of NCRD domain of SP-D from human (hNCRD), rat (rNCRD), or human E31K mutant (mutNCRD) for 1 h.
, VACV-WR+RhSP-D; □, VACV-WR+BSA. (C) VACV-WR was incubated with 0.6 µg/mL of RhSP-D and 2 µg/mL of NCRD domain of SP-D from human (hNCRD), rat (rNCRD), or human E31K mutant (mutNCRD) for 1 h. 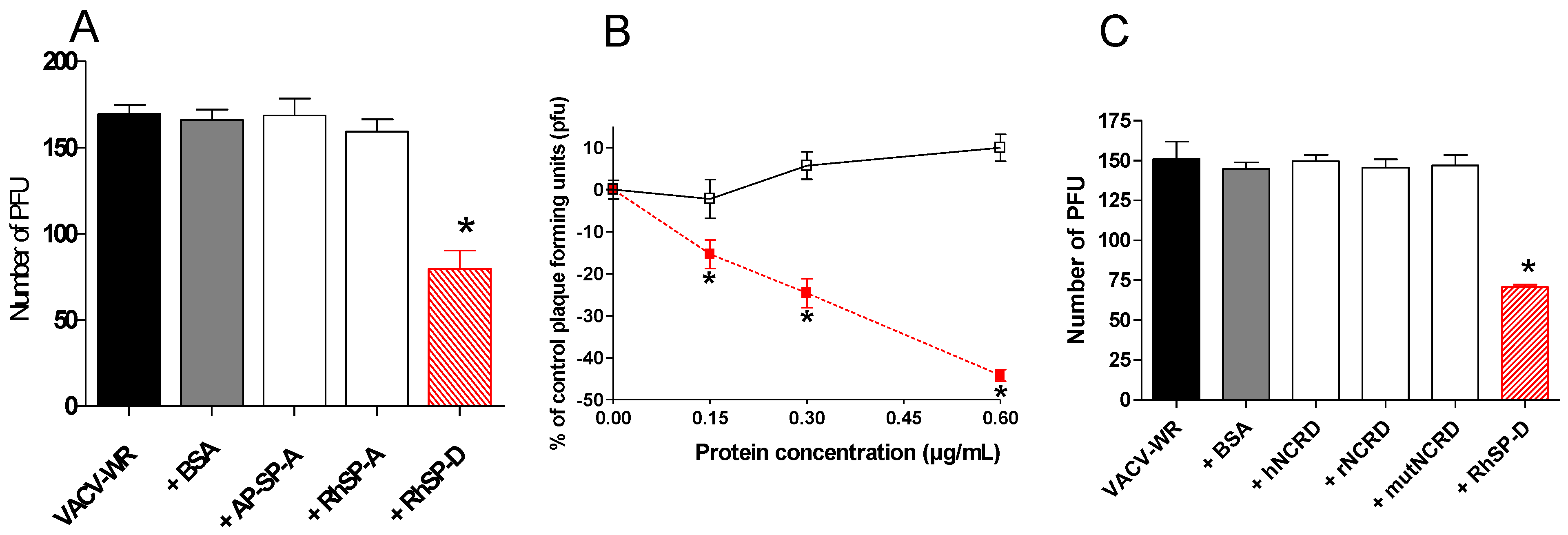
2.4. Comparison of infection inhibiting activity of SP-D with other human defense lectins
2.6. Visualizationof the interaction of SP-D to VACV particle by electron microscopy (EM)

2.7. Analysis of SP-D binding to VACV by surface plasmon resonance (SPR) spectroscopy
2.7.2. Characterization of the interaction between SP-D and recombinant A27
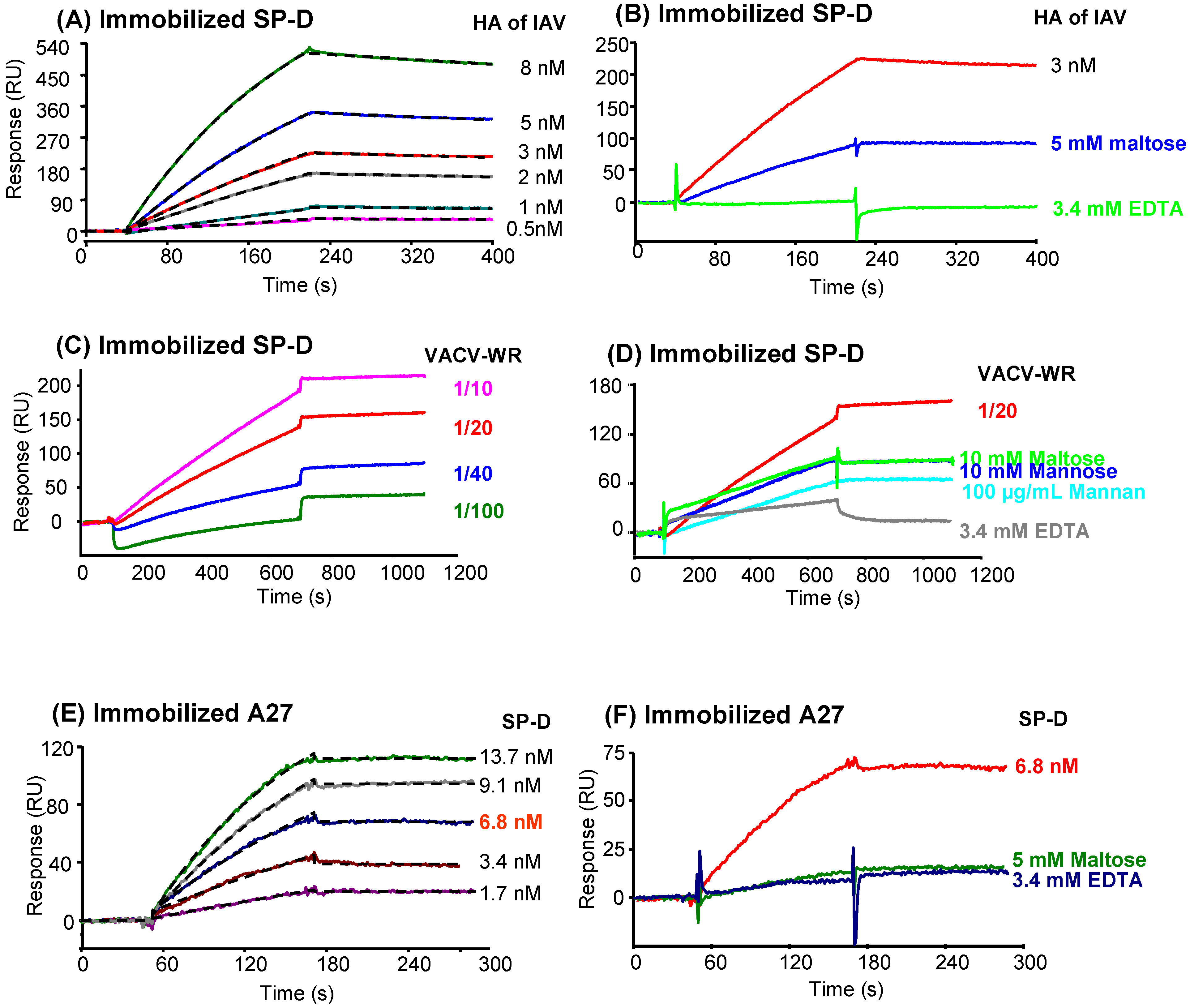
2.7.3. Characterization of the interaction of A27 with other proteins of the defense collagen family
2.8. Protective role of SP-D in VACV infection in mice model
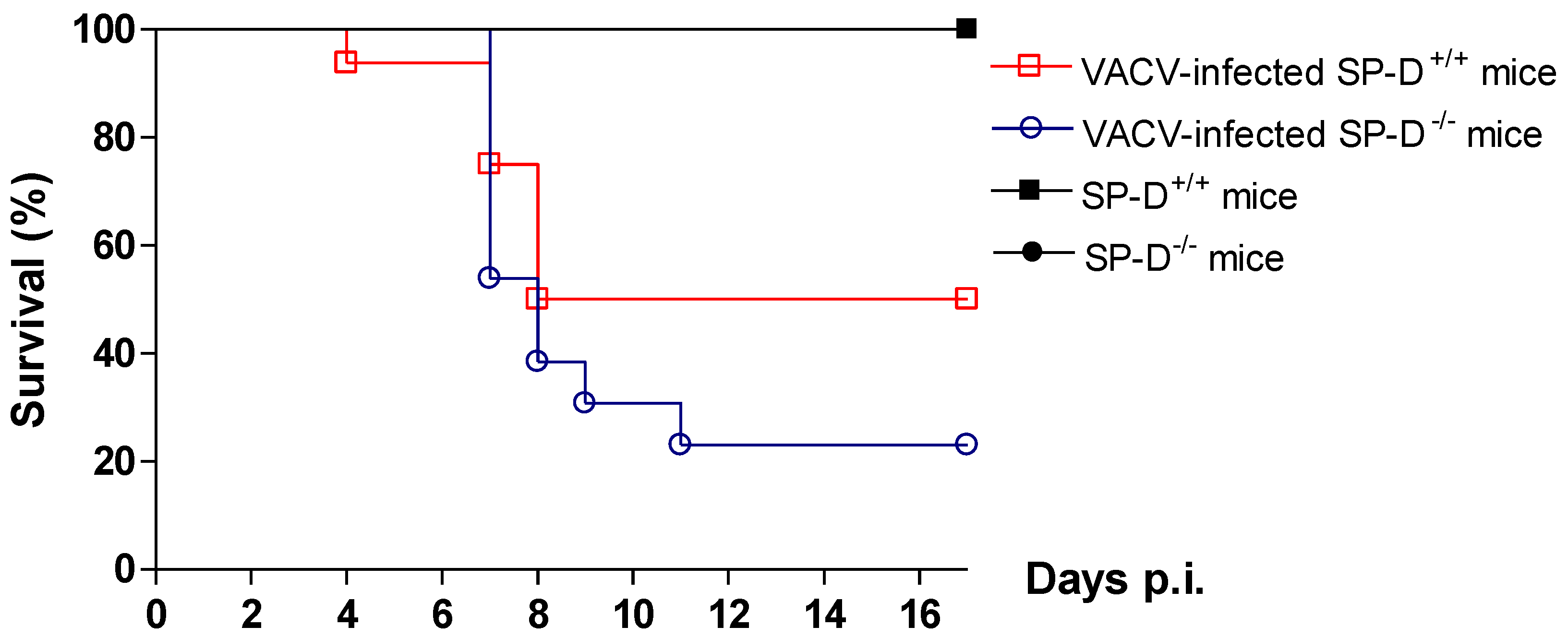
3. Experimental Section
3.1. Cells and viruses
3.2. Reagents
3.3. Recombinant surfactant proteins
3.4. Other recombinant proteins
3.6. A27 SDS-PAGE and western-blot analyses
3.7. Surfactant protein interaction with VACV strains by overlay assay
3.8. hNCRD interaction with A27 by overlay assay
3.9. Lectins-virus infection inhibition assay
3.10. Binding of SP-D to A27 recombinant VACV protein
3.11. ELISA assay for measurement of binding of NCRDs to A27 recombinant VACV protein
3.12. Electron Microscopy
3.13. Surface plasmon resonance analyses on immobilized SP-D
3.15. Surface Plasmon resonance data evaluation
3.16. Ethics Statement
3.17. Animals
3.18. VACV-WR intranasal infection model
3.19. Statistical analysis
4. Conclusions
Abbreviations
| A27 | vaccinia virus A27 membrane protein (14-kDa fusion protein) |
| CRD | carbohydrate recognition domain |
| HA | hemagglutinin protein of IAV |
| IAV | Influenza A virus |
| MBL | mannan-binding lectin |
| NCRD | recombinant trimeric neck+CRDs |
| SP-A | surfactant protein A |
| SP-D | surfactant protein D |
| RSV | respiratory syncycial virus |
| VACV | vaccinia virus |
| VARV | variola virus |
Acknowledgments
Financial support
Conflict of Interest
References and Notes
- Breman, J.G.; Arita, I. The confirmation and maintenance of smallpox eradication. N. Engl. J. Med. 1980, 303, 1263–1273. [Google Scholar] [CrossRef]
- Sherman, I. W. Twelve diseases that changed our world. Emerg. Infect. Dis. 2008, 14, 866. [Google Scholar]
- Dudgeon, J. A. Virus infections of the upper respiratory tract (abridged). Proc. R. Soc. Med. 1969, 62, 1–2. [Google Scholar]
- Fenner, F.; Henderson, D.A.; Arita, I.; Jezek, Z.; Ladnyi, I.D. Smallpox and its eradication; WHO: Geneva, 1988; p. 1460. [Google Scholar]
- Buller, R.M.; Palumbo, G.J. Poxvirus pathogenesis. Microbiol. Rev. 1991, 55, 80–122. [Google Scholar]
- Andreeva, A.V.; Kutuzov, M.A.; Voyno-Yasenetskaya, T.A. Regulation of surfactant secretion in alveolar type II cells. Am. J. Physiol. Lung Cell Mol. Physiol. 2007, 293, 259–271. [Google Scholar] [CrossRef]
- Wustneck, R.; Perez-Gil, J.; Wustneck, N.; Cruz, A.; Fainerman, V.B.; Pison, U. Interfacial properties of pulmonary surfactant layers. Adv. Colloid. Interface. Sci. 2005, 117, 33–58. [Google Scholar]
- Haagsman, H.P.; Hogenkamp, A.; van Eijk, M.; Veldhuizen, E.J. Surfactant collectins and innate immunity. Neonatology 2008, 93, 288–294. [Google Scholar] [CrossRef]
- Haczku, A. Protective role of the lung collectins surfactant protein A and surfactant protein D in airway inflammation. J. Allergy. Clin. Immunol. 2008, 122, 861–879; quiz 880–881. [Google Scholar] [CrossRef]
- Kishore, U.; Greenhough, T.J.; Waters, P.; Shrive, A.K.; Ghai, R.; Kamran, M.F.; Bernal, A.L.; Reid, K.B.; Madan, T.; Chakraborty, T. Surfactant proteins SP-A and SP-D: structure, function and receptors. Mol. Immunol. 2006, 43, 1293–1315. [Google Scholar] [CrossRef]
- Holmskov, U.; Malhotra, R.; Sim, R.B.; Jensenius, J.C. Collectins: collagenous C-type lectins of the innate immune defense system. Immunol. Today 1994, 15, 67–74. [Google Scholar] [CrossRef]
- Sahly, H.; Keisari, Y.; Crouch, E.; Sharon, N.; Ofek, I. Recognition of bacterial surface polysaccharides by lectins of the innate immune system and its contribution to defense against infection: the case of pulmonary pathogens. Infect. Immun. 2008, 76, 1322–1332. [Google Scholar] [CrossRef]
- Reading, P.C.; Pickett, D.L.; Tate, M.D.; Whitney, P.G.; Job, E.R.; Brooks, A.G. Loss of a single N-linked glycan from the hemagglutinin of influenza virus is associated with resistance to collectins and increased virulence in mice. Respir. Res. 2009, 10, 117. [Google Scholar] [CrossRef]
- Crouch, E.; Hartshorn, K.; Ofek, I. Collectins and pulmonary innate immunity. Immunol. Rev. 2000, 173, 52–65. [Google Scholar]
- Epstein, J.; Eichbaum, Q.; Sheriff, S.; Ezekowitz, R.A. The collectins in innate immunity. Curr. Opin. Immunol. 1996, 8, 29–35. [Google Scholar]
- Reading, P.C.; Morey, L.S.; Crouch, E.C.; Anders, E.M. Collectin-mediated antiviral host defense of the lung: evidence from influenza virus infection of mice. J. Virol. 1997, 71, 8204–8212. [Google Scholar]
- Seaton, B.A.; Crouch, E.C.; McCormack, F.X.; Head, J.F.; Hartshorn, K.L.; Mendelsohn, R. Review: Structural determinants of pattern recognition by lung collectins. Innate. Immun. 2010, 16, 143–150. [Google Scholar] [CrossRef]
- Hakansson, K.; Reid, K.B. Collectin structure: a review. Protein Sci. 2000, 9, 1607–1617. [Google Scholar] [CrossRef]
- Ohya, M.; Nishitani, C.; Sano, H.; Yamada, C.; Mitsuzawa, H.; Shimizu, T.; Saito, T.; Smith, K.; Crouch, E.; Kuroki, Y. Human pulmonary surfactant protein D binds the extracellular domains of Toll-like receptors 2 and 4 through the carbohydrate recognition domain by a mechanism different from its binding to phosphatidylinositol and lipopolysaccharide. Biochemistry 2006, 45, 8657–8664. [Google Scholar] [CrossRef]
- Nie, X.; Nishitani, C.; Yamazoe, M.; Ariki, S.; Takahashi, M.; Shimizu, T.; Mitsuzawa, H.; Sawada, K.; Smith, K.; Crouch, E.; Nagae, H.; Takahashi, H.; Kuroki, Y. Pulmonary surfactant protein D binds MD-2 through the carbohydrate recognition domain. Biochemistry 2008, 47, 12878–12885. [Google Scholar] [CrossRef]
- DeSilva, N.S.; Ofek, I.; Crouch, E.C. Interactions of surfactant protein D with fatty acids. Am. J. Respir. Cell Mol. Biol. 2003, 29, 757–770. [Google Scholar] [CrossRef]
- Grubor, B.; Meyerholz, D.K.; Ackermann, M.R. Collectins and cationic antimicrobial peptides of the respiratory epithelia. Vet. Pathol. 2006, 43, 595–612. [Google Scholar] [CrossRef]
- Hartshorn, K.L.; Crouch, E.C.; White, M.R.; Eggleton, P.; Tauber, A.I.; Chang, D.; Sastry, K. Evidence for a protective role of pulmonary surfactant protein D (SP-D) against influenza A viruses. J. Clin. Invest. 1994, 94, 311–319. [Google Scholar] [CrossRef]
- Hickling, T.P.; Bright, H.; Wing, K.; Gower, D.; Martin, S.L.; Sim, R.B.; Malhotra, R. A recombinant trimeric surfactant protein D carbohydrate recognition domain inhibits respiratory syncytial virus infection in vitro and in vivo. Eur. J. Immunol. 1999, 29, 3478–3484. [Google Scholar] [CrossRef]
- Griese, M. Respiratory syncytial virus and pulmonary surfactant. Viral. Immunol. 2002, 15, 357–363. [Google Scholar] [CrossRef]
- LeVine, A.M.; Elliott, J.; Whitsett, J.A.; Srikiatkhachorn, A.; Crouch, E.; DeSilva, N.; Korfhagen, T. Surfactant protein-d enhances phagocytosis and pulmonary clearance of respiratory syncytial virus. Am. J. Respir. Cell Mol. Biol. 2004, 31, 193–199. [Google Scholar] [CrossRef]
- Hartshorn, K.L.; White, M.R.; Tecle, T.; Holmskov, U.; Crouch, E.C. Innate defense against influenza A virus: activity of human neutrophil defensins and interactions of defensins with surfactant protein D. J. Immunol. 2006, 176, 6962–6972. [Google Scholar]
- White, M.R.; Boland, P.; Tecle, T.; Gantz, D.; Sorenson, G.; Tornoe, I.; Holmskov, U.; McDonald, B.; Crouch, E.C.; Hartshorn, K.L. Enhancement of antiviral activity of collectin trimers through cross-linking and mutagenesis of the carbohydrate recognition domain. J. Innate. Immun. 2010, 2, 267–279. [Google Scholar] [CrossRef]
- Qi, L.; Kash, J.C.; Dugan, V.G.; Jagger, B.W.; Lau, Y.F.; Sheng, Z.M.; Crouch, E.C.; Hartshorn, K.L.; Taubenberger, J.K. The ability of pandemic influenza virus hemagglutinins to induce lower respiratory pathology is associated with decreased surfactant protein D binding. Virology 2011, 412, 426–434. [Google Scholar] [CrossRef]
- Kongchanagul, A.; Suptawiwat, O.; Boonarkart, C.; Kitphati, R.; Puthavathana, P.; Uiprasertkul, M.; Auewarakul, P. Decreased expression of surfactant protein D mRNA in human lungs in fatal cases of H5N1 avian influenza. J. Med. Virol. 2011, 83, 1410–1417. [Google Scholar] [CrossRef]
- Hartshorn, K.L.; White, M.R.; Voelker, D.R.; Coburn, J.; Zaner, K.; Crouch, E.C. Mechanism of binding of surfactant protein D to influenza A viruses: importance of binding to haemagglutinin to antiviral activity. Biochem. J. 2000, 351, 449–458. [Google Scholar] [CrossRef]
- Vigerust, D.J.; Ulett, K.B.; Boyd, K.L.; Madsen, J.; Hawgood, S.; McCullers, J.A. N-linked glycosylation attenuates H3N2 influenza viruses. J. Virol. 2007, 81, 8593–8600. [Google Scholar] [CrossRef]
- Hartshorn, K.L.; Webby, R.; White, M.R.; Tecle, T.; Pan, C.; Boucher, S.; Moreland, R.J.; Crouch, E.C.; Scheule, R.K. Role of viral hemagglutinin glycosylation in anti-influenza activities of recombinant surfactant protein D. Respir. Res. 2008, 9, 65. [Google Scholar] [CrossRef]
- Job, E.R.; Deng, Y.M.; Tate, M.D.; Bottazzi, B.; Crouch, E.C.; Dean, M.M.; Mantovani, A.; Brooks, A.G.; Reading, P.C. Pandemic H1N1 influenza A viruses are resistant to the antiviral activities of innate immune proteins of the collectin and pentraxin superfamilies. J. Immunol. 2010, 185, 4284–4291. [Google Scholar]
- Rodriguez, J.F.; Esteban, M. Mapping and nucleotide sequence of the vaccinia virus gene that encodes a 14-kilodalton fusion protein. J. Virol. 1987, 61, 3550–3554. [Google Scholar]
- Doms, R.W.; Blumenthal, R.; Moss, B. Fusion of intra- and extracellular forms of vaccinia virus with the cell membrane. J. Virol. 1990, 64, 4884–4892. [Google Scholar]
- Gong, S.C.; Lai, C.F.; Esteban, M. Vaccinia virus induces cell fusion at acid pH and this activity is mediated by the N-terminus of the 14-kDa virus envelope protein. Virology 1990, 178, 81–91. [Google Scholar] [CrossRef]
- Kochan, G.; Escors, D.; Gonzalez, J.M.; Casasnovas, J.M.; Esteban, M. Membrane cell fusion activity of the vaccinia virus A17-A27 protein complex. Cell Microbiol. 2008, 10, 149–164. [Google Scholar]
- Meyer, H.; Osterrieder, N.; Czerny, C.P. Identification of binding sites for neutralizing monoclonal antibodies on the 14-kDa fusion protein of orthopox viruses. Virology 1994, 200, 778–783. [Google Scholar] [CrossRef]
- Chung, C.S.; Hsiao, J.C.; Chang, Y.S.; Chang, W. A27L protein mediates vaccinia virus interaction with cell surface heparan sulfate. J. Virol. 1998, 72, 1577–1585. [Google Scholar]
- Hsiao, J.C.; Chung, C.S.; Chang, W. Cell surface proteoglycans are necessary for A27L protein-mediated cell fusion: identification of the N-terminal region of A27L protein as the glycosaminoglycan-binding domain. J. Virol. 1998, 72, 8374–8379. [Google Scholar]
- Perino, J.; Foo, C.H.; Spehner, D.; Cohen, G.H.; Eisenberg, R.J.; Crance, J.M.; Favier, A.L. Role of sulfatide in vaccinia virus infection. Biol. Cell 2011, 103, 319–331. [Google Scholar] [CrossRef]
- Ho, Y.; Hsiao, J.C.; Yang, M.H.; Chung, C.S.; Peng, Y.C.; Lin, T.H.; Chang, W.; Tzou, D.L. The Oligomeric Structure of Vaccinia Viral Envelope Protein A27L is Essential for Binding to Heparin and Heparan Sulfates on Cell Surfaces: A Structural and Functional Approach Using Site-specific Mutagenesis. J. Mol. Biol. 2005, 349, 1060–1071. [Google Scholar] [CrossRef]
- Vazquez, M.I.; Rivas, G.; Cregut, D.; Serrano, L.; Esteban, M. The vaccinia virus 14-kilodalton (A27L) fusion protein forms a triple coiled-coil structure and interacts with the 21-kilodalton (A17L) virus membrane protein through a C-terminal alpha-helix. J. Virol. 1998, 72, 10126–10137. [Google Scholar]
- Vazquez, M.I.; Esteban, M. Identification of functional domains in the 14-kilodalton envelope protein (A27L) of vaccinia virus. J. Virol. 1999, 73, 9098–9109. [Google Scholar]
- Cross, K.J.; Langley, W.A.; Russell, R.J.; Skehel, J.J.; Steinhauer, D.A. Composition and functions of the influenza fusion peptide. Protein Pept. Letter. 2009, 16, 766–778. [Google Scholar] [CrossRef]
- Hartshorn, K.; Chang, D.; Rust, K.; White, M.; Heuser, J.; Crouch, E. Interactions of recombinant human pulmonary surfactant protein D and SP-D multimers with influenza A. Am. J. Physiol. 1996, 271, 753–762. [Google Scholar]
- LeVine, A.M.; Whitsett, J.A.; Hartshorn, K.L.; Crouch, E.C.; Korfhagen, T.R. Surfactant protein D enhances clearance of influenza A virus from the lung in vivo. J. Immunol. 2001, 167, 5868–5873. [Google Scholar]
- Hawgood, S.; Brown, C.; Edmondson, J.; Stumbaugh, A.; Allen, L.; Goerke, J.; Clark, H.; Poulain, F. Pulmonary collectins modulate strain-specific influenza a virus infection and host responses. J. Virol. 2004, 78, 8565–8572. [Google Scholar] [CrossRef]
- Carlson, T.K.; Torrelles, J.B.; Smith, K.; Horlacher, T.; Castelli, R.; Seeberger, P.H.; Crouch, E.C.; Schlesinger, L.S. Critical role of amino acid position 343 of surfactant protein-D in the selective binding of glycolipids from Mycobacterium tuberculosis. Glycobiology 2009, 19, 1473–1484. [Google Scholar] [CrossRef]
- Tecle, T.; White, M.R.; Sorensen, G.; Gantz, D.; Kacak, N.; Holmskov, U.; Smith, K.; Crouch, E.C.; Hartshorn, K.L. Critical role for cross-linking of trimeric lectin domains of surfactant protein D in antiviral activity against influenza A virus. Biochem. J. 2008, 412, 323–329. [Google Scholar] [CrossRef]
- Lai, C.F.; Gong, S.C.; Esteban, M. Structural and functional properties of the 14-kDa envelope protein of vaccinia virus synthesized in Escherichia coli. J. Biol. Chem. 1990, 265, 22174–22180. [Google Scholar]
- Crouch, E.; Tu, Y.; Briner, D.; McDonald, B.; Smith, K.; Holmskov, U.; Hartshorn, K. Ligand specificity of human surfactant protein D: expression of a mutant trimeric collectin that shows enhanced interactions with influenza A virus. J. Biol. Chem. 2005, 280, 17046–17056. [Google Scholar]
- Crouch, E.; Persson, A.; Chang, D.; Heuser, J. Molecular structure of pulmonary surfactant protein D (SP-D). J. Biol. Chem. 1994, 269, 17311–17319. [Google Scholar]
- Crouch, E.; Chang, D.; Rust, K.; Persson, A.; Heuser, J. Recombinant pulmonary surfactant protein D. Post-translational modification and molecular assembly. J. Biol. Chem. 1994, 269, 15808–15813. [Google Scholar]
- Fenner, F. The biological characters of several strains of vaccinia, cowpox and rabbitpox viruses. Virology 1958, 5, 502–529. [Google Scholar] [CrossRef]
- Crouch, E.C.; Smith, K.; McDonald, B.; Briner, D.; Linders, B.; McDonald, J.; Holmskov, U.; Head, J.; Hartshorn, K. Species differences in the carbohydrate binding preferences of surfactant protein D. Am. J. Respir. Cell Mol. Biol. 2006, 35, 84–94. [Google Scholar] [CrossRef]
- Malhotra, R.; Haurum, J.S.; Thiel, S.; Sim, R.B. Binding of human collectins (SP-A and MBP) to influenza virus. Biochem. J. 1994, 304, 455–461. [Google Scholar]
- Hartshorn, K.L.; White, M.R.; Tecle, T.; Sorensen, G.; Holmskov, U.; Crouch, E.C. Viral aggregating and opsonizing activity in collectin trimers. Am. J. Physiol. Lung Cell Mol. Physiol. 2010, 298, 79–88. [Google Scholar] [CrossRef] [Green Version]
- Hartshorn, K.L.; White, M.R.; Rynkiewicz, M.; Sorensen, G.; Holmskov, U.; Head, J.; Crouch, E.C. Monoclonal antibody-assisted structure-function analysis of the carbohydrate recognition domain of surfactant protein D. Am. J. Physiol. Lung. Cell Mol. Physiol. 2010, 299, 384–392. [Google Scholar]
- Crouch, E.; Nikolaidis, N.; McCormack, F.X.; McDonald, B.; Allen, K.; Rynkiewicz, M.J.; Cafarella, T.M.; White, M.; Lewnard, K.; Leymarie, N.; Zaia, J.; Seaton, B.A.; Hartshorn, K.L. Mutagenesis of surfactant protein D informed by evolution and x-ray crystallography enhances defenses against influenza A virus in vivo. J. Biol. Chem. 2011, 286, 40681–40692. [Google Scholar] [CrossRef]
- Ji, X.; Olinger, G.G.; Aris, S.; Chen, Y.; Gewurz, H.; Spear, G.T. Mannose-binding lectin binds to Ebola and Marburg envelope glycoproteins, resulting in blocking of virus interaction with DC-SIGN and complement-mediated virus neutralization. J. Gen. Virol. 2005, 86, 2535–2542. [Google Scholar] [CrossRef]
- Anders, E.M.; Hartley, C.A.; Reading, P.C.; Ezekowitz, R.A. Complement-dependent neutralization of influenza virus by a serum mannose-binding lectin. J. Gen. Virol. 1994, 75, 615–622. [Google Scholar] [CrossRef]
- Lemos, M.P.; McKinney, J.; Rhee, K.Y. Dispensability of surfactant proteins A and D in immune control of Mycobacterium tuberculosis infection following aerosol challenge of mice. Infect. Immun. 2011, 79, 1077–1085. [Google Scholar] [CrossRef]
- Lee, H.J.; Essani, K.; Smith, G.L. The genome sequence of Yaba-like disease virus, a yatapoxvirus. Virology 2001, 281, 170–192. [Google Scholar] [CrossRef]
- Tsung, K.; Yim, J.H.; Marti, W.; Buller, R.M.; Norton, J.A. Gene expression and cytopathic effect of vaccinia virus inactivated by psoralen and long-wave UV light. J. Virol. 1996, 70, 165–171. [Google Scholar]
- Gout, E.; Garlatti, V.; Smith, D.F.; Lacroix, M.; Dumestre-Perard, C.; Lunardi, T.; Martin, L.; Cesbron, J.Y.; Arlaud, G.J.; Gaboriaud, C.; Thielens, N.M. Carbohydrate recognition properties of human ficolins: glycan array screening reveals the sialic acid binding specificity of M-ficolin. J. Biol. Chem. 2010, 285, 6612–6622. [Google Scholar] [CrossRef]
- Teillet, F.; Gaboriaud, C.; Lacroix, M.; Martin, L.; Arlaud, G.J.; Thielens, N.M. Crystal structure of the CUB1-EGF-CUB2 domain of human MASP-1/3 and identification of its interaction sites with mannan-binding lectin and ficolins. J. Biol. Chem. 2008, 283, 25715–25724. [Google Scholar]
- Teillet, F.; Lacroix, M.; Thiel, S.; Weilguny, D.; Agger, T.; Arlaud, G.J.; Thielens, N.M. Identification of the site of human mannan-binding lectin involved in the interaction with its partner serine proteases: the essential role of Lys55. J. Immunol. 2007, 178, 5710–5716. [Google Scholar]
- Lacroix, M.; Dumestre-Perard, C.; Schoehn, G.; Houen, G.; Cesbron, J.Y.; Arlaud, G.J.; Thielens, N.M. Residue Lys57 in the collagen-like region of human L-ficolin and its counterpart Lys47 in H-ficolin play a key role in the interaction with the mannan-binding lectin-associated serine proteases and the collectin receptor calreticulin. J. Immunol. 2009, 182, 456–465. [Google Scholar]
- Korfhagen, T.R.; Sheftelyevich, V.; Burhans, M.S.; Bruno, M.D.; Ross, G.F.; Wert, S.E.; Stahlman, M.T.; Jobe, A.H.; Ikegami, M.; Whitsett, J.A.; Fisher, J.H. Surfactant protein-D regulates surfactant phospholipid homeostasis in vivo. J. Biol. Chem. 1998, 273, 28438–28443. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Perino, J.; Thielens, N.M.; Crouch, E.; Spehner, D.; Crance, J.-M.; Favier, A.-L. Protective Effect of Surfactant Protein D in Pulmonary Vaccinia Virus Infection: Implication of A27 Viral Protein. Viruses 2013, 5, 928-953. https://doi.org/10.3390/v5030928
Perino J, Thielens NM, Crouch E, Spehner D, Crance J-M, Favier A-L. Protective Effect of Surfactant Protein D in Pulmonary Vaccinia Virus Infection: Implication of A27 Viral Protein. Viruses. 2013; 5(3):928-953. https://doi.org/10.3390/v5030928
Chicago/Turabian StylePerino, Julien, Nicole M. Thielens, Erika Crouch, Danièle Spehner, Jean-Marc Crance, and Anne-Laure Favier. 2013. "Protective Effect of Surfactant Protein D in Pulmonary Vaccinia Virus Infection: Implication of A27 Viral Protein" Viruses 5, no. 3: 928-953. https://doi.org/10.3390/v5030928