Lessons in AIDS Vaccine Development Learned from Studies of Equine Infectious, Anemia Virus Infection and Immunity
Abstract
:1. Introduction to EIAV
2. EIAV Antigenic Variation and Env Diversity: Lessons of Viral Evolution and Immune Evasion



3. EIAV Diversity and the Host Immune Response: Lessons on Immune Maturation
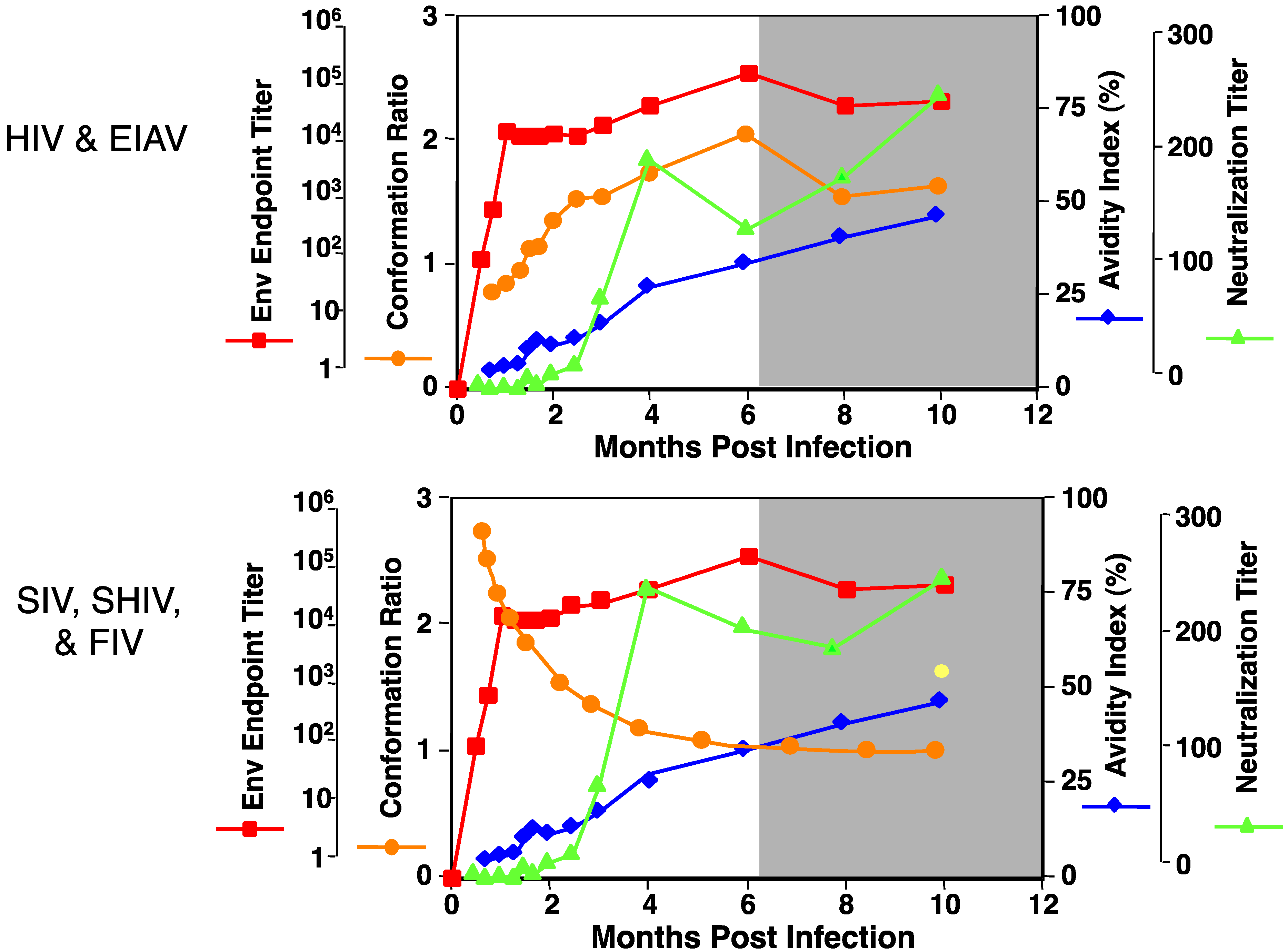
4. Env Diversity and EIAV Vaccines: Lessons on Efficacy and Protection
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Lignée, M. Mémoire et observations sur une maladie de sang, connue sous le nom d’anhémie hydrohémie, cachexie acquise du cheval. Rec. Med. Vet. Ec Alfort. 1843, 20, 30–45. (In French) [Google Scholar]
- Craigo, J.K.; Montelaro, R.C. Equine Infectious Anemia Virus (Retroviridae). In Encyclopedia of Virology, 3rd ed.; Elsevier: Oxford, UK, 2008; Volume 3, pp. 167–174. [Google Scholar]
- Montelaro, R.C.; Ball, J.M.; Rushlow, K. Equine Retroviruses. In The Retroviridae; Levy, J.A., Ed.; Plenum Press: New York, NY, USA, 1993; Volume 2, pp. 257–360. [Google Scholar]
- Clements, J.E.; Zink, M.C.; Narayan, O.; Gabuzda, D. Lentivirus infection of macrophages. Immunol. Ser. 1994, 60, 589–600. [Google Scholar]
- Oaks, J.L.; McGuire, T.C.; Ulibarri, C.; Crawford, T. Equine infectious anemia virus is found in tissue macrophages during subclinical infection. J. Virol. 1998, 72, 7263–7269. [Google Scholar]
- Harrold, S.M.; Cook, S.J.; Cook, R.F.; Rushlow, K.E.; Issel, C.J.; Montelaro, R.C. Tissue sites of persistent infection and active replication of equine infectious anemia virus during acute disease and asymptomatic infection in experimentally infected equids. J. Virol. 2000, 74, 3112–3121. [Google Scholar] [CrossRef]
- Hammond, S.A.; Li, F.; McKeon, B.M., Sr.; Cook, S.J.; Issel, C.J.; Montelaro, R.C. Immune responses and viral replication in long-term inapparent carrier ponies inoculated with equine infectious anemia virus. J. Virol. 2000, 74, 5968–5981. [Google Scholar] [CrossRef]
- Kono, Y.; hirasawa, K.; Fukunaga, Y.; Taniguchi, T. Recrudesence of equine infectious anemia by treatment with immunosuppressive drugs. Nat. Inst. Anim. Hlth. Quart. 1976, 16, 8–15. [Google Scholar]
- Craigo, J.K.; Leroux, C.; Howe, L.; Steckbeck, J.D.; Cook, S.J.; Issel, C.J.; Montelaro, R.C. Transient immune suppression of inapparent carriers infected with a principal neutralizing domain-deficient equine infectious anaemia virus induces neutralizing antibodies and lowers steady-state virus replication. J. Gen. Virol. 2002, 83, 1353–1359. [Google Scholar]
- Pandrea, I.; Onanga, R.; Kornfeld, C.; Rouquet, P.; Bourry, O.; Clifford, S.; Telfer, P.T.; Abernethy, K.; White, L.T.; Ngari, P.; et al. High levels of sivmnd-1 replication in chronically infected mandrillus sphinx. Virology 2003, 317, 119–127. [Google Scholar] [CrossRef]
- Pandrea, I.; Onanga, R.; Souquiere, S.; Mouinga-Ondeme, A.; Bourry, O.; Makuwa, M.; Rouquet, P.; Silvestri, G.; Simon, F.; Roques, P.; et al. Paucity of CD4+ CCR5+ T cells may prevent transmission of simian immunodeficiency virus in natural nonhuman primate hosts by breast-feeding. J. Virol. 2008, 82, 5501–5509. [Google Scholar] [CrossRef]
- VandeWoude, S.; Apetrei, C. Going wild: Lessons from naturally occurring T-lymphotropic lentiviruses. Clin. Microbiol. Rev. 2006, 19, 728–762. [Google Scholar] [CrossRef]
- Leroux, C.; Issel, C.; Montelaro, R.C. Novel and dynamic evolution of equine infectious anemia virus genomic quasispecies associated with sequential disease cycles in an experimentally infected pony. J. Virol. 1997, 71, 9627–9639. [Google Scholar]
- Leroux, C.; Craigo, J.K.; Issel, C.J.; Montelaro, R.C. Equine infectious anemia virus genomic evolution in progressor and nonprogressor ponies. J. Virol. 2001, 75, 4570–4583. [Google Scholar] [CrossRef]
- Lichtenstein, D.L.; Issel, C.J.; Montelaro, R.C. Genomic quasispecies associated with the initiation of infection and disease in ponies experimentally infected with equine infectious anemia virus. J. Virol. 1996, 70, 3346–3354. [Google Scholar]
- Sponseller, B.A.; Sparks, W.O.; Wannemuehler, Y.; Li, Y.; Antons, A.K.; Oaks, J.L.; Carpenter, S. Immune selection of equine infectious anemia virus env variants during the long-term inapparent stage of disease. Virology 2007, 363, 156–165. [Google Scholar] [CrossRef]
- Zheng, Y.H.; Nakaya, T.; Sentsui, H.; Kameoka, M.; Kishi, M.; Hagiwara, K.; Takahashi, H.; Kono, Y.; Ikuta, K. Insertions, duplications and substitutions in restricted gp90 regions of equine infectious anemia virus during febrile episodes in an experimentally infected horse. J. Gen. Virol. 1997, 78, 807–820. [Google Scholar]
- Zheng, Y.H.; Sentsui, H.; Nakaya, T.; Kono, Y.; Ikuta, K. In vivo dynamics of equine infectious anemia viruses emerging during febrile episodes: Insertions/duplications at the principal neutralizing domain. J. Virol. 1997, 71, 5031–5039. [Google Scholar]
- Zheng, Y.H.; Sentsui, H.; Kono, Y.; Ikuta, K. Mutations occurring during serial passage of japanese equine infectious anemia virus in primary horse macrophages. Virus Res. 2000, 68, 93–98. [Google Scholar] [CrossRef]
- Craigo, J.K.; Sturgeon, T.J.; Cook, S.J.; Issel, C.J.; Leroux, C.; Montelaro, R.C. Apparent elimination of EIAV ancestral species in a long-term inapparent carrier. Virology 2006, 344, 340–353. [Google Scholar] [CrossRef]
- Greene, W.K.; Meers, J.; del Fierro, G.; Carnegie, P.R.; Robinson, W.F. Extensive sequence variation of feline immunodeficiency virus env genes in isolates from naturally infected cats. Arch. Virol. 1993, 133, 51–62. [Google Scholar] [CrossRef]
- Leroux, C.; Chastang, J.; Greenland, T.; Mornex, J.F. Genomic heterogeneity of small ruminant lentiviruses: Existence of heterogeneous populations in sheep and of the same lentiviral genotypes in sheep and goats. Arch. Virol. 1997, 142, 1125–1137. [Google Scholar] [CrossRef]
- Simmonds, P.; Balfe, P.; Ludlam, J.; Bishop, O.; Brown, A.J. Analysis of sequence diversity in hypervariable regions of the external glycoprotein of human immunodeficiency virus type 1. J. Virol. 1990, 64, 5840–5850. [Google Scholar]
- Starcich, E.S.; Hahn, B.H.; Shaw, G.M.; McNeely, P.D.; Modrow, S.; Wolf, H.; Parks, E.S.; Parks, W.P.; Josephs, S.F.; Gallo, R.C. Identification and characterization of conserved and variable regions in the envelope gene of HTLV-III/Lav, the retrovirus of AIDS. Cell 1986, 45, 637–648. [Google Scholar] [CrossRef] [Green Version]
- Suarez, D.L.; Whetstone, C.A. Identification of hypervariable and conserved regions in the surface envelope gene in the bovine lentivirus. Virology 1995, 212, 728–733. [Google Scholar] [CrossRef]
- Craigo, J.K.; Barnes, S.; Cook, S.J.; Issel, C.J.; Montelaro, R.C. Divergence, not diversity of an attenuated equine lentivirus vaccine strain correlates with protection from disease. Vaccine 2010, 28, 8095–8104. [Google Scholar] [CrossRef]
- Blankson, J.N.; Persaud, D.; Siliciano, R.F. The challenge of viral reservoirs in HIV-1 infection. Annu. Rev. Med. 2002, 53, 557–593. [Google Scholar] [CrossRef]
- Chun, T.W.; Carruth, L.; Finzi, D.; Shen, X.; DiGiuseppe, J.A.; Taylor, H.; Hermankova, M.; Chadwick, K.; Margolick, J.; Quinn, T.C.; et al. Quantification of latent tissue reservoirs and total body viral load in HIV-1 infection. Nature 1997, 387, 183–188. [Google Scholar] [CrossRef]
- Finzi, D.; Blankson, J.; Siliciano, J.D.; Margolick, J.B.; Chadwick, K.; Pierson, T.; Smith, K.; Lisziewicz, J.; Lori, F.; Flexner, C.; et al. Latent infection of CD4+ T cells provides a mechanism for lifelong persistence of HIV-1, even in patients on effective combination therapy. Nat. Med. 1999, 5, 512–517. [Google Scholar] [CrossRef]
- Persaud, D.; Zhou, Y.; Siliciano, J.M.; Siliciano, R.F. Latency in human immunodeficiency virus type 1 infection: No easy answers. J. Virol. 2003, 77, 1659. [Google Scholar] [CrossRef]
- Han, Y.; Lassen, K.; Monie, D.; Sedaghat, A.R.; Shimoji, S.; Liu, X.; Pierson, T.C.; Margolick, J.B.; Siliciano, R.F.; Siliciano, J.D. Resting CD4+ T cells from human immunodeficiency virus type 1 (HIV-1) -infected individuals carry integrated HIV-1 genomes within actively transcribed host genes. J. Virol. 2004, 78, 6122–6133. [Google Scholar] [CrossRef]
- Lassen, K.G.; Bailey, J.R.; Siliciano, R.F. Analysis of human immunodeficiency virus type 1 transcriptional elongation in resting CD4+ T cells in vivo. J. Virol. 2004, 78, 9105–9114. [Google Scholar] [CrossRef]
- Issel, C.J.; Adams, W.V., Jr.; Meek, L.; Ochoa, R. Transmission of equine infectious anemia virus from horses without clinical signs of disease. J. Am. Vet. Med. Assoc. 1982, 180, 272–275. [Google Scholar]
- McGuire, T.C.; Tumas, D.B.; Byrne, K.M.; Hines, M.T.; Leib, S.R.; Brassfield, A.L.; O'Rourke, K.I.; Perryman, L.E. Major histocompatibility complex-restricted cd8+ cytotoxic T lymphocytes from horses with equine infectious anemia virus recognize env and gag/pr proteins. J. Virol. 1994, 68, 1459–1467. [Google Scholar]
- O'Rourke, K.I.; Perryman, L.E.; McGuire, T.C. Antiviral, antiglycoprotein and neutralizing antibodies in foals with equine infectious anaemia virus. J. Gen. Virol. 1988, 69, 667–674. [Google Scholar] [CrossRef]
- Rwambo, P.M.; Issel, C.J.; Adams, W.V., Jr.; Hussain, K.A.; Miller, M.; Montelaro, R.C. Equine infectious anemia virus (eiav) humoral responses of recipient ponies and antigenic variation during persistent infection. Arch. Virol. 1990, 111, 199–212. [Google Scholar] [CrossRef]
- Montelaro, R.C.; Ball, J.M.; Issel, C.J. Characterization of eiav immunogenicity during persistent infections: Humoral responses and antigen targets. Dev. Biol Stand. 1990, 72, 19–30. [Google Scholar]
- Hammond, S.A.; Cook, S.J.; Lichtenstein, D.L.; Issel, C.J.; Montelaro, R.C. Maturation of the cellular and humoral immune responses to persistent infection in horses by equine infectious anemia virus is a complex and lengthy process. J. Virol. 1997, 71, 3840–3852. [Google Scholar]
- Montelaro, R.C.; Cole, K.S.; Hammond, S.A. Maturation of immune responses to lentivirus infection: Implications for AIDS vaccine development. AIDS Res. Hum. Retrovir. 1998, 14, S255–S259. [Google Scholar] [CrossRef]
- Hussain, K.A.; Issel, C.J.; Schnorr, K.L.; Rwambo, P.M.; West, M.; Montelaro, R.C. Antigenic mapping of the envelope proteins of equine infectious anemia virus: Identification of a neutralization domain and a conserved region on glycoprotein 90. Arch. Virol. 1988, 98, 213–224. [Google Scholar] [CrossRef]
- Payne, S.L.; Fang, F.D.; Liu, C.P.; Dhruva, B.R.; Rwambo, P.; Issel, C.J.; Montelaro, R.C. Antigenic variation and lentivirus persistence: Variations in envelope gene sequences during eiav infection resemble changes reported for sequential isolates of HIV. Virology 1987, 161, 321–331. [Google Scholar] [CrossRef]
- Payne, S.L.; Salinovich, O.; Nauman, S.M.; Issel, C.J.; Montelaro, R.C. Course and extent of variation of equine infectious anemia virus during parallel persistent infections. J. Virol. 1987, 61, 1266–1270. [Google Scholar]
- Payne, S.L.; Rushlow, K.; Dhruva, B.R.; Issel, C.J.; Montelaro, R.C. Localization of conserved and variable antigenic domains of equine infectious anemia virus envelope glycoproteins using recombinant env-encoded protein fragments produced in escherichia coli. Virology 1989, 172, 609–615. [Google Scholar] [CrossRef]
- Perryman, L.E.; O'Rourke, K.I.; McGuire, T.C. Immune responses are required to terminate viremia in equine infectious anemia lentivirus infection. J. Virol. 1988, 62, 3073–3076. [Google Scholar]
- Howe, L.; Leroux, C.; Issel, C.J.; Montelaro, R.C. Equine infectious anemia virus envelope evolution in vivo during persistent infection progressively increases resistance to in vitro serum antibody neutralization as a dominant phenotype. J. Virol. 2002, 76, 10588. [Google Scholar] [CrossRef]
- Howe, L.; Craigo, J.K.; Issel, C.J.; Montelaro, R.C. Specificity of serum neutralizing antibodies induced by transient immune suppression of inapparent carrier ponies infected with a neutralization-resistant equine infectious anemia virus envelope strain. J. Gen. Virol. 2005, 86, 139–149. [Google Scholar] [CrossRef]
- Tagmyer, T.L.; Craigo, J.K.; Cook, S.J.; Issel, C.J.; Montelaro, R.C. Envelope-specific t-helper and cytotoxic t-lymphocyte responses associated with protective immunity to equine infectious anemia virus. J. Gen. Virol. 2007, 88, 1324–1336. [Google Scholar] [CrossRef]
- Tagmyer, T.L.; Craigo, J.K.; Cook, S.J.; Even, D.L.; Issel, C.J.; Montelaro, R.C. Envelope determinants of equine infectious anemia virus vaccine protection and the effects of sequence variation on immune recognition. J. Virol. 2008, 82, 4052–4063. [Google Scholar] [CrossRef]
- Issel, C.J.; Horohov, D.W.; Lea, D.F.; Adams, W.V., Jr.; Hagius, S.D.; McManus, J.M.; Allison, A.C.; Montelaro, R.C. Efficacy of inactivated whole-virus and subunit vaccines in preventing infection and disease caused by equine infectious anemia virus. J. Virol. 1992, 66, 3398–3408. [Google Scholar]
- Wang, S.Z.; Rushlow, K.E.; Issel, C.J.; Cook, R.F.; Cook, S.J.; Raabe, M.L.; Chong, Y.H.; Costa, L.; Montelaro, R.C. Enhancement of eiav replication and disease by immunization with a baculovirus-expressed recombinant envelope surface glycoprotein. Virology 1994, 199, 247–251. [Google Scholar] [CrossRef]
- Grund, C.H.; Lechman, E.R.; Pezzuolo, N.A.; Issel, C.J.; Montelaro, R.C. Fine specificity of equine infectious anaemia virus gp90-specific antibodies associated with protective and enhancing immune responses in experimentally infected and immunized ponies. J. Gen. Virol. 1996, 77, 435–442. [Google Scholar] [CrossRef]
- Raabe, M.L.; Issel, C.J.; Cook, S.J.; Cook, R.F.; Woodson, B.; Montelaro, R.C. Immunization with a recombinant envelope protein (rgp90) of eiav produces a spectrum of vaccine efficacy ranging from lack of clinical disease to severe enhancement. Virology 1998, 245, 151–162. [Google Scholar] [CrossRef]
- Shen, R.X.; Wang, Z. Development and Use of an Equine Infectious Anemia Donkey Leucocyte Attenuated Vaccine. In EIAV: A National Review of Policies, Programs, and Future Objectives; American Quarter Horse Association: Amarillo, TX, USA, 1985; pp. 135–148. [Google Scholar]
- Craigo, J.K.; Zhang, B.; Barnes, S.; Tagmyer, T.L.; Cook, S.J.; Issel, C.J.; Montelaro, R.C. Envelope variation as a primary determinant of lentiviral vaccine efficacy. Proc. Natl. Acad. Sci. USA 2007, 104, 15105–15110. [Google Scholar]
- Cook, R.F.; Cook, S.J.; Bolin, P.S.; Howe, L.J.; Zhou, W.; Montelaro, R.C.; Issel, C.J. Genetic immunization with codon-optimized equine infectious anemia virus (EIAV) surface unit (SU) envelope protein gene sequences stimulates immune responses in ponies. Vet. Microbiol. 2005, 108, 23–37. [Google Scholar] [CrossRef]
- Craigo, J.K.; Durkin, S.; Sturgeon, T.J.; Tagmyer, T.; Cook, S.J.; Issel, C.J.; Montelaro, R.C. Immune suppression of challenged vaccinates as a rigorous assessment of sterile protection by lentiviral vaccines. Vaccine 2007, 25, 834–845. [Google Scholar] [CrossRef]
- Hammond, S.A.; Cook, S.J.; Falo, L.D., Jr.; Issel, C.J.; Montelaro, R.C. A particulate viral protein vaccine reduces viral load and delays progression to disease in immunized ponies challenged with equine infectious anemia virus. Virology 1999, 254, 37–49. [Google Scholar] [CrossRef]
- Li, F.; Craigo, J.K.; Howe, L.; Steckbeck, J.D.; Cook, S.; Issel, C.; Montelaro, R.C. A live attenuated equine infectious anemia virus proviral vaccine with a modified S2 gene provides protection from detectable infection by intravenous virulent virus challenge of experimentally inoculated horses. J. Virol. 2003, 77, 7244–7253. [Google Scholar] [CrossRef]
- Craigo, J.K.; Li, F.; Steckbeck, J.D.; Durkin, S.; Howe, L.; Cook, S.J.; Issel, C.; Montelaro, R.C. Discerning an effective balance between equine infectious anemia virus attenuation and vaccine efficacy. J. Virol. 2005, 79, 2666–2677. [Google Scholar] [CrossRef]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Craigo, J.K.; Montelaro, R.C. Lessons in AIDS Vaccine Development Learned from Studies of Equine Infectious, Anemia Virus Infection and Immunity. Viruses 2013, 5, 2963-2976. https://doi.org/10.3390/v5122963
Craigo JK, Montelaro RC. Lessons in AIDS Vaccine Development Learned from Studies of Equine Infectious, Anemia Virus Infection and Immunity. Viruses. 2013; 5(12):2963-2976. https://doi.org/10.3390/v5122963
Chicago/Turabian StyleCraigo, Jodi K., and Ronald C. Montelaro. 2013. "Lessons in AIDS Vaccine Development Learned from Studies of Equine Infectious, Anemia Virus Infection and Immunity" Viruses 5, no. 12: 2963-2976. https://doi.org/10.3390/v5122963



