Abstract
Kobuviruses comprise three species, the Aichivirus A, Aichivirus B, and Aichivirus C (porcine kobuvirus). Porcine kobuvirus is endemic to pig farms and is not restricted geographically but, rather, is distributed worldwide. The complete genomic sequences of four porcine kobuvirus strains isolated during a diarrhea outbreak in piglets in the Gansu province of China were determined. Two of these strains exhibited variations relative to the traditional strains. The potential 3C/3D cleavage sites of the variant strains were Q/C, which differed from the Q/S in the traditional porcine kobuvirus genome. A 90-nucleotide deletion in the 2B protein and a single nucleotide insertion in the 3′UTR were found in the variant strains. The VP1 regions of all four porcine kobuviruses in our study were highly variable (81%–86%). Ten common amino acid mutations were found specifically at certain positions within the VP1 region. Significant recombination sites were identified using SimPlot scans of whole genome sequences. Porcine kobuviruses were also detected in pig serum, indicating that the virus can escape the gastrointestinal tract and travel to the circulatory system. These findings suggest that mutations and recombination events may have contributed to the high level of genetic diversity of porcine kobuviruses and serve as a driving force in its evolution.
1. Introduction
Picornaviruses (Picornaviridae family) are small, non-enveloped viruses with single-stranded, positive-sense genomic RNA. This family is divided into the following 17 genera: Aphthovirus, Aquamavirus, Avihepatovirus, Cardiovirus, Cosavirus, Dicipivirus, Enterovirus, Erbovirus, Hepatovirus, Kobuvirus, Megrivirus, Parechovirus, Salivirus, Sapelovirus, Senecavirus, Teschovirus, and Tremovirus [1]. To date, three species of kobuviruses have been identified, including Aichivirus A (formerly human Aichi virus), Aichivirus B (formerly bovine kobuvirus), and Aichivirus C (formerly porcine kobuvirus). Kobuviruses are new members of the Picornaviridae family. To date, kobuviruses have been detected in humans [2], cattle [3], pigs [4], sheep [5], bats [6], dogs [7], cats [8], ferrets [9], and goats [10]; this indicates that these viruses have a wide range of host species. The kobuviruses have the same genome organization, which includes a 5′UTR, a leader (L) protein, three structural proteins (VP0, VP3, and VP1), seven non-structural proteins (2A–2C and 3A–3D), and a 3′UTR.
Porcine kobuvirus was first detected in fecal samples from domestic pigs in Hungary, in 2007 [4]. Recently, porcine kobuviruses have also been detected at a high frequency in stool samples from healthy pigs in Hungary, China [11], Japan [12], Thailand [13], Brazil [14], The Netherlands [14], and Korea [15]. In addition, the prevalence of these viruses in pigs with diarrhea was shown to be very high in China during the last two years. Since December, 2010, there has been a large-scale outbreak of diarrheal disease in suckling piglets, characterized by watery diarrhea, dehydration, and vomiting. This outbreak led to 80% morbidity and 60% mortality in affected piglets in several farms in the Gansu Province of China. A recent report suggested that porcine kobuvirus infection correlated with diarrhea in pigs [15]. Porcine kobuviruses are a newly recognized viral agent in animals. Little is known about this virus, and only eight porcine kobuvirus genome sequences are available in GenBank. For this reason, it is important to increase the awareness of this disease and to stimulate further clinical, epidemiological, and molecular studies regarding porcine kobuviruses. In this study, we report four complete genome sequences and describe the detailed genomic organization of two porcine kobuvirus variant strains.
2. Results and Discussion
2.1. Prevalence and Complete Genomic Sequences of Porcine Kobuviruses
Overall, five (31.3%) of 17 fecal samples and two (33.3%) of six serum samples tested positive for kobuvirus infection using Kv primers. A total of 11 overlapping cDNA clones spanning the entire genome of K-11/2012/CH, K-4/2012/CH, GS-1/2012/CH, and GS-2/2012/CH were obtained, and their nucleotide sequences have been deposited into the GenBank database. Their accession numbers are KC414936, KC424638, KC424639, and KC424649, respectively. The complete RNA genomes of the K-11/2012/CH and K-4/2012/CH strains were similar to the sequence of the Hungarian strain, which consists of 8210 nucleotides (nt), excluding the poly (A) tail. A large open reading frame (ORF) of 7467 nt encoding a potential polyprotein precursor of 2488 amino acids (aa) was found. This ORF was located at the 5′ end and was preceded by 576 nt; 167 nt followed the ORF prior to the poly (A) tail. Relative to the genomes of the traditional strains, the genomes of the GS-1/2012/CH and GS-2/2012/CH variant strains contained 8121 nt, excluding the poly (A) tail; a 90-nucleotide deletion in the 2B protein and a single nucleotide insertion in the 3′UTR were also observed (Table 1).

Table 1.
Comparison of the nucleotide and amino acid sequences of the study strains to the sequences of S-1-HUN.
| Region | Length (nt) | Nucleotide identity (%) | Amino acid identity (%) | Predicted N-terminal cleavage site | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| K-11 | K-4 | GS-1 | GS-2 | K-11 | K-4 | GS-1 | GS-2 | K1-1/K-4 | GS-1/GS-2 | S-1-HUN | ||
| 5′UTR | 576 | 97 | 95 | 95 | 95 | - | - | - | - | - | - | - |
| L | 585 | 87 | 87 | 86 | 86 | 93 | 93 | 93 | 93 | - | - | - |
| VP0 | 1,098 | 83 | 87 | 87 | 85 | 90 | 96 | 96 | 93 | Q/G | Q/G | Q/G |
| VP3 | 669 | 87 | 87 | 86 | 77 | 93 | 97 | 95 | 97 | Q/H | Q/H | Q/H |
| VP1 | 762 | 85 | 86 | 82 | 85 | 96 | 96 | 89 | 96 | Q/A | Q/A | Q/A |
| 2A | 408 | 89 | 88 | 88 | 87 | 95 | 93 | 90 | 93 | Q/C | Q/C | Q/C |
| 2B | 585/495 * | 87 | 87 | 76 | 77 | 97 | 93 | 95 | 96 | Q/G | Q/G | Q/G |
| 2C | 1,005 | 92 | 90 | 91 | 91 | 98 | 98 | 98 | 98 | Q/G | Q/G | Q/G |
| 3A | 270 | 87 | 87 | 87 | 88 | 91 | 93 | 93 | 93 | Q/G | Q/G | Q/G |
| 3B | 102 | 96 | 89 | 92 | 91 | 100 | 97 | 100 | 100 | Q/G | Q/G | Q/G |
| 3C | 576 | 86 | 88 | 88 | 88 | 95 | 99 | 97 | 98 | Q/S | Q/C | Q/S |
| 3D | 1,407 | 93 | 93 | 93 | 93 | 98 | 98 | 99 | 99 | Q/S | Q/S | Q/S |
| 3′UTR | 167/168 * | 98 | 97 | 96 | 96 | - | - | - | - | - | - | - |
| Structural (VP0–VP3) | 2,329 | 81 | 86 | 85 | 83 | 93 | 97 | 94 | 95 | - | - | - |
| Non-structural(2A–3D) | 353/4,263 * | 91 | 90 | 89 | 89 | 97 | 97 | 95 | 95 | - | - | - |
| Complete genome | 8,210/8,121 | 86 | 89 | 88 | 87 | 95 | 96 | 94 | 94 | - | - | - |
* Indicates the nucleotide length of the variant strains.
2.2. Analysis of the UTRs and Coding Regions of the Four Strains
An alignment of the sequences of the four genomic sequences with porcine kobuvirus strain S-1-HUN, revealed that the genome sequences of K-4/2012/CH, K-11/2012/CH, GS-1/2012/CH, and GS-2/2012/CH shared 89%, 86%, 88%, and 87% identity to the S-1-HUN at the nucleotide level, respectively (Table 1). The highest nucleotide identities (96% and 98%) between the variant strains and the S-1-HUN strain were observed in the 3′UTR (Table 1), and the lowest nucleotide identities (76% and 87%) between the variant strains and the S-1-HUN strain were observed in the 2B coding region (Table 1). K-4/2012/CH and K-11/2012/CH polyprotein sequences were analyzed for potential cleavage sites, and the sites were identical to those in theS-1-HUN strain (Table 1). However, the potential 3C/3D cleavage site for the GS-1/2012/CH and GS-2/2012/CH variant strains was Q/C, which differed from the Q/S sequence in the traditional porcine kobuvirus strains.
2.3. Recombination Analysis of the Variant Stains
The four complete kobuvirus genome sequences were analyzed using Simplot. The similarity plot displays the consecutive nucleotide identity (%) among the queried strain and parental strains. The Bootscan plots display the likelihood of recombination within the GS-1/2012/CH variant strain; the 481–1,414 and 3,482–4,100 nucleotide regions are similar to regions found in the K-4/2012/CH strain. The other genomic regions of the GS-1/2012/CH variant strain, including 4,100–4,369, 7,143–7,384, and 7,760–8,068, are closely related to the K-11/2012/CH strain (Figure 1A). Significant recombination signals were identified in the 481–774, 1,165–1,414, 3,842–4,369, 4,842–5,121, 7,143–7,384, and 7,760–8,068 nucleotide regions (Figure 1B).
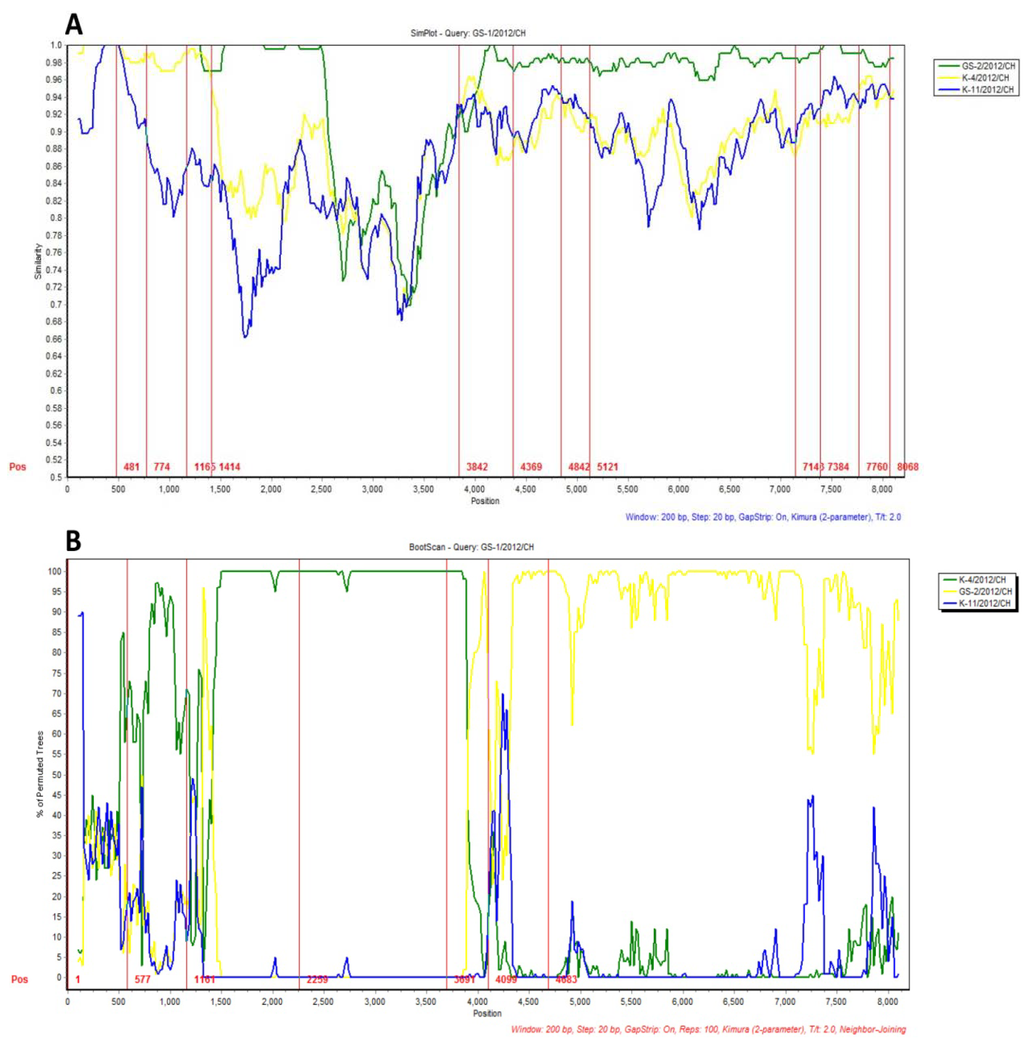
Figure 1.
Nucleotide similarities and recombination among the kobuvirus genomes. (A) Nucleotide similarities among theK-4/2012/CH, K-11/2012/CH, GS-1/2012/CH, and GS-2/2012/CH genomes, as determined using the SimPlot program. The x-axis indicates the nucleotide position along the alignment, and the y-axis shows the similarity; (B) Bootscan plot of the GS-1/2012/CH strain with GS-1/2012/CH, K-4/2012/CH, and K-4/2012/CH reference sequences as out groups. The analysis was performed using pairwise distance, modeled with a window size of 200, step size of 20, and 100 bootstrap replicates.
2.4. Analysis of Coding Region Amino Acid Sequences
The structural and non-structural protein regions of the both K-4/2012/CH and K-11/2012/CH strains contain 2329aa and 4353aa, respectively. The amino acid identity identities among the of structural regions (VP0–VP3) are 90%–97%, and the non-structural regions (2A–3D) share 93%–100% amino acid identity with that of the S-1-HUN strain (Table 2). Compared with the K-4/2012/CH and K-11/2012/CH strains, the variant strain has a 30-animo acid deletion within the 2B protein. The amino acid identities among the structural regions (VP0–VP3) are 89%–97%, and the non-structural regions (2A–3D) share 90%–100% amino acid identity with that of the S-1-HUN strain (Table 2). The 2B amino acid sequence alignment for the 12 strains analyzed in this study and the eight strains from the GenBank database (K-30-HUN/2008 and S-1-HUN are Hungarian strains; others are Chinese strains) is shown in Figure 2A. Ten amino acid mutations were observed at positions 7, 11, 15, 28, 29, 30, 33, 37, 41, and 44. Though the function of the porcine kobuvirus 2B protein is not known, protein 2B has been shown to assume different functions in different kobuviruses [15].

Table 2.
Kobuvirus strains used in this study.
| Accession No. | Virus strain | Kobuvirus species | Host species | Country reported |
|---|---|---|---|---|
| GQ927711 | D/VI2287/2004 | Aichi virus | Human | Germany |
| JX564249 | kvgh99012632/2010 | Aichi virus | Human | Taiwan |
| DQ028632 | Goiania/GO/03/01/Brazil | Aichi virus | Human | Brazil |
| FJ890523 | Chshc7 | Aichi virus | Human | China |
| AB040749 | A846/88 | Aichi virus | Human | Japan |
| HQ650197 | KB101 | Bovine virus | Cattle | South Korea |
| AB084788 | U-1 | Bovine virus | Cattle | Japan |
| HQ650168 | KB12 | Bovine virus | Cattle | South Korea |
| HQ650180 | KB15 | Bovine virus | Cattle | South Korea |
| JF755427 | M-5/USA/2010 | Mouse virus | Mouse | USA |
| JN387133 | AN211D/USA/2009 | Dog virus | Dog | USA |
| KF006985 | MpKoV38 | Ferret virus | Ferret | Netherlands |
| GU245693 | TB3/HUN/2009 | Sheep virus | Sheep | Hungary |
| HQ400969 | 97DA4 | Porcine virus | Sus scrofa | South Korea |
| HQ400970 | 111DA18 | Porcine virus | Sus scrofa | South Korea |
| HQ400968 | 99DA6 | Porcine virus | Sus scrofa | South Korea |
| HQ400967 | 110DA17 | Porcine virus | Sus scrofa | South Korea |
| HQ400963 | 95DA2 | Porcine virus | Sus scrofa | South Korea |
| HQ400966 | 123DB10 | Porcine virus | Sus scrofa | South Korea |
| HQ400962 | 94DA1 | Porcine virus | Sus scrofa | South Korea |
| HQ400948 | 15OA16 | Porcine virus | Sus scrofa | South Korea |
| JF422792 | D11 | Porcine virus | Pig | China |
| JF422788 | D1 | Porcine virus | Pig | China |
| JX401523 | CH/HNXX-4/2012 | Porcine virus | Piglet | China |
| NC_016769 | SH-W-CHN | Porcine virus | Pig | China |
| JX177612 | WB-1-HUN/2011 | Porcine virus | Sus scrofa | Hungary |
| JQ692069 | WHU1 | Porcine virus | Sus scrofa | China |
| GQ249161 | K-30-HUN/2008 | Porcine virus | Sus scrofa | Hungary |
| EU787450 | S-1-HUN | Porcine virus | Sus scrofa | Hungary |
| JX827598 | CH/HZ/2011 | Porcine virus | Pig | China |
| GU298971 | Ch16/2008/CHN | Porcine virus | Pig | China |
| GU298967 | Ch40/2008/CHN | Porcine virus | Pig | China |
| GU292559 | JY-2010a/CHN | Porcine virus | Pig | China |
| KC414936 | K-11/2012/CH | Porcine virus | Sus scrofa | China |
| KC424638 | K-4/2012/CH | Porcine virus | Sus scrofa | China |
| KC424639 | GS-1/2012/CH | Porcine virus | Sus scrofa | China |
| KC424649 | GS-2/2012/CH | Porcine virus | Sus scrofa | China |
| KC204684 | XX | Porcine virus | Pig | China |
VP1 is highly exposed and is the most immunodominant part of the kobuvirus capsid protein [3], and this protein is the most variable among the structural proteins [16]. Eight VP1 regions were analyzed in this study, and more than three amino acid mutations were found specifically at positions 6 (S, N, and D); 22 (E, S, and I); 81 (N, I, and V); 131 (G, S, and T); 132 (G, E, and N); 133 (D, E, and N); 136 (T, D, G, and I); 199 (S, A, N, and T); 209 (Y, M, and L); and 235 (S, V, and F) (Figure 2B).
2.5. Phylogenetic Analysis
We compared the entire genome sequences of K-4/2012/CH, K-11/2012/CH, GS-1/2012/CH, and GS-2/2012/CH with those of other kobuviruses. A phylogenetic tree based on whole genome sequences indicted that all four of the study sequences were placed in the porcine kobuvirus genetic clade, which was separate from the other kobuviruses (Figure 3). Within this clade, the K-4/2012/CH, K-11/2012/CH, GS-1/2012/CH, and GS-2/2012/CH strains form two groups; the GS-1/2012/CH and GS-2/2012/CH variant strains form a separate group with the other porcine kobuviruses.

Figure 2.
Comparison of the deduced protein amino acid sequences of the study strains and the reference strains. (A) Comparison of the deduced 2B protein amino acid sequences. An asterisk (*) indicates an amino acid substitution in the VP1 region; a dash (-) indicates an amino acid deletion in the variant strain; (B) Comparison of the deduced VP1 amino acid sequences. An asterisk (*) indicates no fewer than three amino acid substitutions within the VP1 region.
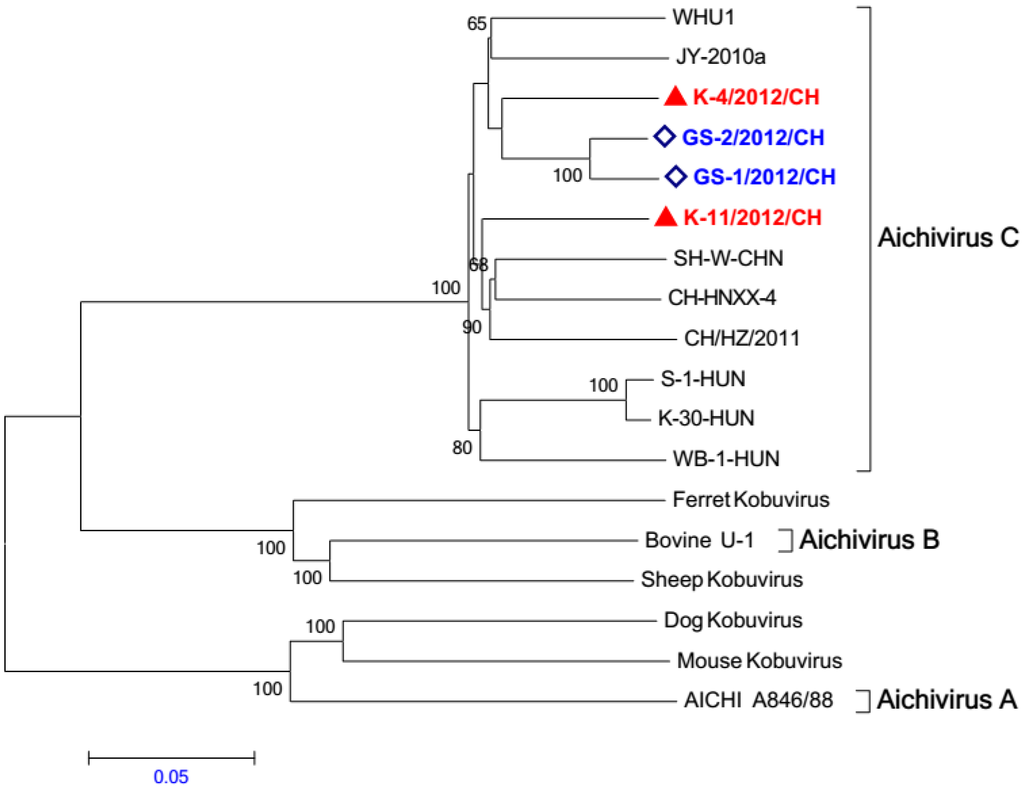
Figure 3.
Relationships among kobuviruses, including porcine kobuvirus and other kobuviruses (human, bovine, sheep, dog, ferret, and mouse) based on whole genome sequences. Bootstrap values (based on 1,000 replicates) for each node are provided when >50%.
The sequences of the 3D region have been particularly useful for placing viruses within species or genera and for comparing viruses of different genera or families [17]. A phylogenetic analysis shows two genetic lineages of porcine kobuviruses, however, these viruses are phylogenetically distinct from both bovine kobuviruses and Aichi viruses based on partial sequences of the 3D region (Figure 4). The four porcine kobuviruses isolated in our study and the Hungarian strains were in the same lineage, and the South Korean strains were placed in another lineage.
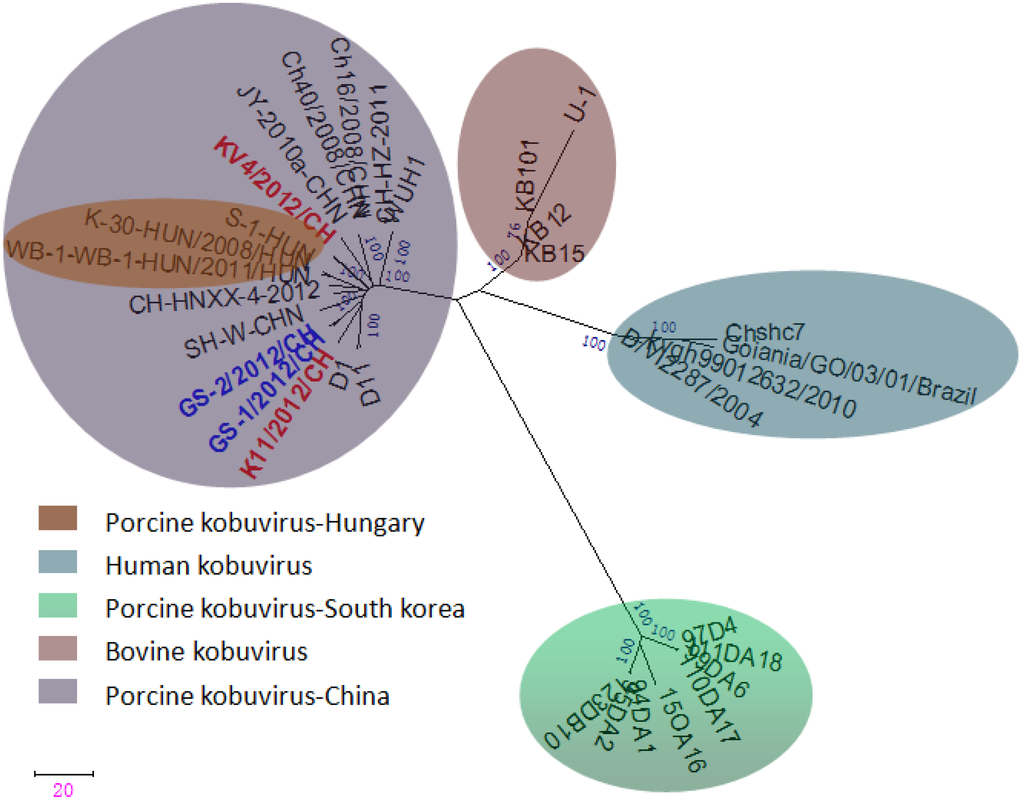
Figure 4.
Phylogenetic tree of porcine kobuviruses based on the sequences from the 3D region (588 nt). The phylogenetic tree was constructed using the neighbor-joining clustering method; distance was calculated using the maximum composite likelihood correction for evolutionary rate in MEGA version 5.0. Bootstrap values (based on 1,000 replicates) for each node are provided when >50%.
2.6. Viral Culture
No cytopathic effects (CPE) were observed in Vero, BHK-21 (Baby Hamster Kidney), or PK-15 (Pig Kidney) cells following serial passage of cultures. Porcine kobuvirus RNA replication were not detected in the cells and supernatants in every passage were collected.
3. Experimental
3.1. Fecal and Serum Samples
Seventeen fecal and serum samples were collected in July, 2012, from dead piglets; these piglets showed signs of watery diarrhea and dehydration for <10 days and were from two farms (located in Jiayuguan and Linxia City) in Gansu Province. Six serum samples from a one-year-old pig in Jiuquan City within Gansu Province were also obtained. Fecal samples were prepared as 10% homogenates in PBS and then centrifuged at 10,000 ×g for 20 min. The resultant supernatants were applied to Vero, PK, or BHK cells prior to RT-PCR analysis.
3.2. RT-PCR, Cloning, and Sequencing
Total RNA was extracted from fecal and serum samples using TRIzol reagent according to the manufacturer’s instructions (Invitrogen Life Technologies, Grand Island, NY, USA). RNA was reverse transcribed with avian myeloblastosis virus reverse transcriptase (Promega, Beijing, China) in the presence of random primersand oligo dT15 (TaKaRa Bio Inc., Otsu, Japan). The cDNA was immediately amplified with primers that were designed based on the sequences of kobuvirus reference strains (Table 3). The 5′ and 3′ ends of the genome were confirmed using a SMARTer rapid amplification of cDNA ends (RACE) cDNA amplification kit (Clontech, Japan). PCR reactions contained 1 µL of FastPfu DNA Polymerase (TransGen Biotech, Beijing, China), 10 µL of 5× FastPfu Buffer, 5 µL of dNTP mixture (2.5 mM), 1 µL of each primer, 2 µL of template, and sterile H2O to yield a final volume of 50 µL. The PCR conditions were as follows: 94 °C for 5 min; 35 cycles of 94 °C for 30 s, 55 °C for 30 s, and 72 °C for 1 min; and a final extension at 72 °C for 10 min. Amplicons were separated in a 1.5% agarosegel.

Table 3.
Primers designed from contigs to acquire the porcine kobuvirus genomes.
| Primer name | Sequence (5'–3') | Size of PCR product (bp) |
|---|---|---|
| Pokv-30F | CCCTCACCCTCTTTTCCG | 369 |
| Pokv-398R | ACCGCAGTCCATGCTCTA | |
| Pokv-302F | AAACTCCTACCCGACAAA | 966 |
| Pokv-1267R | CAGGRCCATCACCAAGRC | |
| Pokv-1134F | YAMCACTCCTTGCCAGATC | 1048 |
| Pokv-2181R | AGAACCAGTWGGRACAGM | |
| Pokv-2048F | ACCTCTAYCAGGGCAAYA | 955 |
| Pokv-3002R | GGTTCAGGGACWGTAGTAGC | |
| Pokv-2894F | TSCGYGGCATCCAAGCAC | 946 |
| Pokv-3839R | AGACCAAGGCGGGAAAGG | |
| Pokv-3723F | CTGTCCAGACGTGCGGRTYT | 1019 |
| Pokv-4741R | CCATGAGCCACTCGGTGTTC | |
| Pokv-4668F | GACGGTTGAACAYCAAGGTG | 928 |
| Pokv-5595R | GAARGAAGGYTGCCAAAGAG | |
| Pokv-5462F | ARCCCTTCGAYCCTGTGGAG | 925 |
| Pokv-6386R | GGAGGACCAGAAAGAGTAGAAAT | |
| Pokv-6219F | YATCGGTCCRGACACCTTTG | 970 |
| Pokv-7188R | GATAGCGTGKAYGGGAGCAG | |
| Pokv-7036F | TTGCCRMYTCCWGAGTTRGA | 695 |
| Pokv-7730R | AAAGTRTCTGTTTTRTTTGCTG | |
| Pokv-7614F | CTAYGGTGATGATGTGATCTATG | 582 |
| Pokv-8195R | AAGTAAAGGACAGCCAGGGA | |
| Kv-F | TCTGGATGCGTTGGCACTTCCAT | 519 |
| Kv-R | CCAGCGGGTCTGAAGGTAAGAGT |
3.3. Sequencing and Phylogenetic Analysis
PCR products were cloned into the pMD-18 T vector (TaKaRa, Otsu, Japan). Competent E. coli DH5a cells (TaKaRa) were transformed with the recombinant plasmid. Three clones were sequenced using the Applied Biosystems (ABI) 3730xl DNA analyzer. Sequences were analyzed using the ClustalX version 2.0 [18], DNASTAR, and MegAlign version 5.0 (DNAStar Inc., Madison, WI, USA) software packages. A dendrogram was constructed using the neighbor-joining clustering method in MEGA 5.0 [18]. Possible recombination events in the coding region were analyzed using Simplot software [20]. All kobuvirus strains used in the study are listed in Table 2.
3.4. Viral Culture
Original fecal samples that tested positive for porcine kobuvirus by RT-PCR were propagated separately for virus isolation. Vero, PK-15, and BHK-21 cells were used as general-purpose cell lines for propagating picornaviruses.
4. Conclusions
The first porcine kobuvirus was identified in Hungary in 2007. Since then, porcine kobuviruses have been reported in several Asian countries. Porcine kobuviruses are not geographically restricted and are widely distributed in pigs worldwide. Porcine kobuviruses were also detected at a high frequency in pigs without clinical diarrhea [16]. Here, we report the complete nucleotide sequences and genetic organization of two traditional strains (K-4/2012/CH and K-11/2012/CH) and two variant strains (GS-1/2012/CH and GS-2/2012/CH).
The complete genome sequences of four strains were compared to the prototype porcine kobuvirus strain (S-1-HUN). The genomes of the K-11/2012/CH and K-4/2012/CH strains were similar to that of the Hungarian strain, which consists of 8210 nucleotides. However, the variant strains, GS-1/2012/CH and GS-2/2012/CH, contain 8121 nucleotides, with a 90-nucleotide deletion within the 2B protein and a single nucleotide insertion within the 3′UTR. Though the function of the porcine kobuvirus 2B protein is unknown, the 2B protein has many different functions in different kobuviruses [21]. In our study, ten amino acid mutations were observed in the 2B protein-coding region, though further studies are required to address whether the variant strains cause serious diarrheal outbreaks in pigs.
The VP1 regions of the four porcine kobuviruses in our study were highly variable [22]. The overall nucleotide identity among the VP1 sequences was 81%–86% (Table 1). Ten amino acid mutations were consistently observed at certain positions within the VP1 region. The mechanisms responsible for the evolution of the viral RNA sequences remain largely unknown. Host, viral, and environmental factors may play roles in this process [23].
Natural recombination has been reported in most picornavirus genera, and it is a key genetic feature of these infectious agents [24]. By exchanging sequences, the virus may induce changes in its genome sequence. This may lead to the acquisition of many key adaptive mutations [25], thus generating genetic variation and even producing new viral variants. One striking difference between porcine kobuvirus and the members of the other two kobuvirus species is that the 5′UTR of porcine kobuvirus contains an IRES related to that of hepatitis C virus [26,27] , whereas the 5′UTRs of the other species contain IRESs that are structurally and mechanistically unrelated [28,29]. In our study, significant recombination signals were identified using SimPlot scans of whole genome sequences. However, kobuvirus recombination cannot be discussed without also considering other evolutionary properties, such as global prevalence and its dynamic epidemiology.
Kobuviruses are transmitted primarily by the fecal-oral route, and other picornaviruses may use a similar mechanism [16]. Porcine kobuviruses are thought to be confined to the intestine, but porcine kobuvirus viremia has also been recently reported [5]. We also detected porcine kobuvirus sequences in pig serum. This finding suggests that porcine kobuvirus can escape the gastrointestinal tract and travel to the circulatory system. In addition to fecal-oral transmission, further experiments are necessary to investigate the possibility of kobuvirus transmission through breastfeeding and via blood-borne infections.
In summary, porcine kobuviruses are a newly recognized viral agent in animals. Porcine kobuviruses maybe persist for long periods because this viral infection can be asymptomatic. The presence of mutations and recombination events may contribute to the high levels of genetic diversity in porcine kobuviruses. The genomic changes and viremia may make this virus a potentially dangerous disease-causing agent in general, particularly in humans.
Acknowledgments
This work was supported by the National Key Technologies R&D Program (No. 2013BAD12B04) and the Special Fund for Agro-scientific Research in the Public Interest (No. 201303042).
Conflicts of Interest
The authors declare no conflict of interest.
References
- International Committee on Taxonomy of Viruses. ICTV official taxonomy: Updates since the 9th report. Available online: http://talk.ictvonline.org/files/ictv_documents/m/msl/4440.aspx (accessed on 14 January 2012).
- Yamashita, T.; Kobayashi, S.; Sakae, K.; Nakata, S.; Chiba, S.; Ishihara, Y.; Isomura, S. Isolation of cytopathic small round viruses with BS-C-1 cells from patients with gastroenteritis. J. Infect. Dis. 1991, 164, 954–957. [Google Scholar] [CrossRef]
- Yamashita, T.; Ito, M.; Kabashima, Y.; Tsuzuki, H.; Fujiura, A.; Sakae, K. Isolation and characterization of a new species of kobuvirus associated with cattle. J. Gen. Virol. 2003, 84, 3069–3077. [Google Scholar] [CrossRef]
- Reuter, G.; Boldizsar, A.; Kiss, I.; Pankovics, P. Candidate new species of Kobuvirus in porcine hosts. Emerging Infect. Dis. 2008, 14, 1968–1970. [Google Scholar] [CrossRef]
- Reuter, G.; Kecskemeti, S.; Pankovics, P. Evolution of porcine kobuvirus infection, Hungary. Emerging Infect. Dis. 2010, 16, 696–698. [Google Scholar] [CrossRef]
- Li, L.; Victoria, J.G.; Wang, C.; Jones, M.; Fellers, G.M.; Kunz, T.H.; Delwart, E. Bat guano virome: Predominance of dietary viruses from insects and plants plus novel mammalian viruses. J. Virol. 2010, 84, 6955–6965. [Google Scholar]
- Li, L.; Pesavento, P.A.; Shan, T.; Leutenegger, C.M.; Wang, C.; Delwart, E. Viruses in diarrhoeic dogs include novel kobuviruses and sapoviruses. J. Gen. Virol. 2011, 92, 2534–2541. [Google Scholar] [CrossRef]
- Chung, J.Y.; Kim, S.H.; Kim, Y.H.; Lee, M.H.; Lee, K.K.; Oem, J.K. Detection and genetic characterization of feline kobuviruses. Virus Genes 2013. [Google Scholar] [CrossRef]
- Smits, S.L.; Raj, V.S.; Oduber, M.D.; Schapendonk, C.M.; Bodewes, R.; Provacia, L.; Stittelaar, K.J.; Osterhaus, A.D.; Haagmans, B.L. Metagenomic analysis of the ferret fecal viral flora. PLoS One 2013, 8, e71595. [Google Scholar]
- Lee, M.H.; Jeoung, H.Y.; Lim, J.A.; Song, J.Y.; Song, D.S.; An, D.J. Kobuvirus in South Korean black goats. Virus Genes 2012, 45, 186–189. [Google Scholar] [CrossRef]
- Yu, J.M.; Jin, M.; Zhang, Q.; Li, H.Y.; Li, D.D.; Xu, Z.Q.; Li, J.S.; Cui, S.X.; Yang, S.H.; Liu, N.; et al. Candidate porcine Kobuvirus, China. Emerging Infect. Dis. 2009, 15, 823–825. [Google Scholar] [CrossRef]
- Khamrin, P.; Maneekarn, N.; Hidaka, S.; Kishikawa, S.; Ushijima, K.; Okitsu, S.; Ushijima, H. Molecular detection of kobuvirus sequences in stool samples collected from healthy pigs in Japan. Infect. Genet. Evol. 2010, 10, 950–954. [Google Scholar] [CrossRef]
- Khamrin, P.; Maneekarn, N.; Kongkaew, A.; Kongkaew, S.; Okitsu, S.; Ushijima, H. Porcine kobuvirus in piglets, Thailand. Emerging Infect. Dis. 2009, 15, 2075–2076. [Google Scholar] [CrossRef]
- Barry, A.F.; Ribeiro, J.; Alfieri, A.F.; van der Poel, W.H.; Alfieri, A.A. First detection of kobuvirus in farm animals in Brazil and The Netherlands. Infect. Genet. Evol. 2011, 11, 1811–1814. [Google Scholar] [CrossRef]
- Park, S.J.; Kim, H.K.; Moon, H.J.; Song, D.S.; Rho, S.M.; Han, J.Y.; Nguyen, V.G.; Park, B.K. Molecular detection of porcine kobuviruses in pigs in Korea and their association with diarrhea. Arch. Virol. 2010, 155, 1803–1811. [Google Scholar] [CrossRef]
- Reuter, G.; Boros, A.; Pankovics, P. Kobuviruses—A comprehensive review. Rev. Med. Virol. 2011, 21, 32–41. [Google Scholar] [CrossRef]
- Oberste, M.S.; Maher, K.; Pallansch, M.A. Genomic evidence that simian virus 2 and six other simian picornaviruses represent a new genus in Picornaviridae. Virology 2003, 314, 283–293. [Google Scholar] [CrossRef]
- Larkin, E.K.; Morris, N.J.; Li, Y.; Nock, N.L.; Stein, C.M. Comparison of affected sibling-pair linkage methods to identify gene x gene interaction in GAW15 simulated data. BMC Proc. 2007, 1, S66. [Google Scholar] [CrossRef]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef]
- Lole, K.S.; Bollinger, R.C.; Paranjape, R.S.; Gadkari, D.; Kulkarni, S.S.; Novak, N.G.; Ingersoll, R.; Sheppard, H.W.; Ray, S.C. Full-length human immunodeficiency virus type 1 genomes from subtype C-infected seroconverters in India, with evidence of intersubtype recombination. J. Virol. 1999, 73, 152–160. [Google Scholar]
- De Jong, A.S.; de Mattia, F.; van Dommelen, M.M.; Lanke, K.; Melchers, W.J.; Willems, P.H.; van Kuppeveld, F.J. Functional analysis of picornavirus 2B proteins: Effects on calcium homeostasis and intracellular protein trafficking. J. Virol. 2008, 82, 3782–3790. [Google Scholar] [CrossRef]
- Okitsu, S.; Khamrin, P.; Thongprachum, A.; Hidaka, S.; Kongkaew, S.; Kongkaew, A.; Maneekarn, N.; Mizuguchi, M.; Hayakawa, S.; Ushijima, H. Sequence analysis of porcine kobuvirus VP1 region detected in pigs in Japan and Thailand. Virus Genes 2012, 44, 253–257. [Google Scholar] [CrossRef]
- Wang, C.; Lan, D.; Cui, L.; Yang, Z.; Yuan, C.; Hua, X. Molecular characterization of a porcine kobuvirus strain in China. Arch. Virol. 2012, 157, 573–578. [Google Scholar] [CrossRef]
- Lukashev, A.N. Recombination among picornaviruses. Rev. Med. Virol. 2010, 20, 327–337. [Google Scholar] [CrossRef]
- Kuiken, T.; Holmes, E.C.; McCauley, J.; Rimmelzwaan, G.F.; Williams, C.S.; Grenfell, B.T. Host species barriers to influenza virus infections. Science 2006, 312, 394–397. [Google Scholar] [CrossRef]
- Hellen, C.U.; de Breyne, S. A distinct group of hepacivirus/pestivirus-like internal ribosomal entry sites in members of diverse picornavirus genera: Evidence for modular exchange of functional noncoding RNA elements by recombination. J. Virol. 2007, 81, 5850–5863. [Google Scholar] [CrossRef]
- Reuter, G.; Boldizsar, A.; Pankovics, P. Complete nucleotide and amino acid sequences and genetic organization of porcine kobuvirus, a member of a new species in the genus Kobuvirus, family Picornavirida. Arch. Virol. 2009, 154, 101–108. [Google Scholar] [CrossRef]
- Yu, Y.; Sweeney, T.R.; Kafasla, P.; Jackson, R.J.; Pestova, T.V.; Hellen, C.U. The mechanism of translation initiation on Aichivirus RNA mediated by a novel type of picornavirus IRES. EMBO J. 2011, 30, 4423–4436. [Google Scholar] [CrossRef]
- Sweeney, T.R.; Dhote, V.; Yu, Y.; Hellen, C.U. A distinct class of internal ribosomal entry site in members of the Kobuvirus and proposed Salivirus and Paraturdivirus genera of the Picornaviridae. J. Virol. 2012, 86, 1468–1486. [Google Scholar]
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).