Henipavirus Mediated Membrane Fusion, Virus Entry and Targeted Therapeutics
Abstract
:1. Introduction
2. Attachment Glycoprotein (G)
2.1. Oligomerization of G Glycoprotein
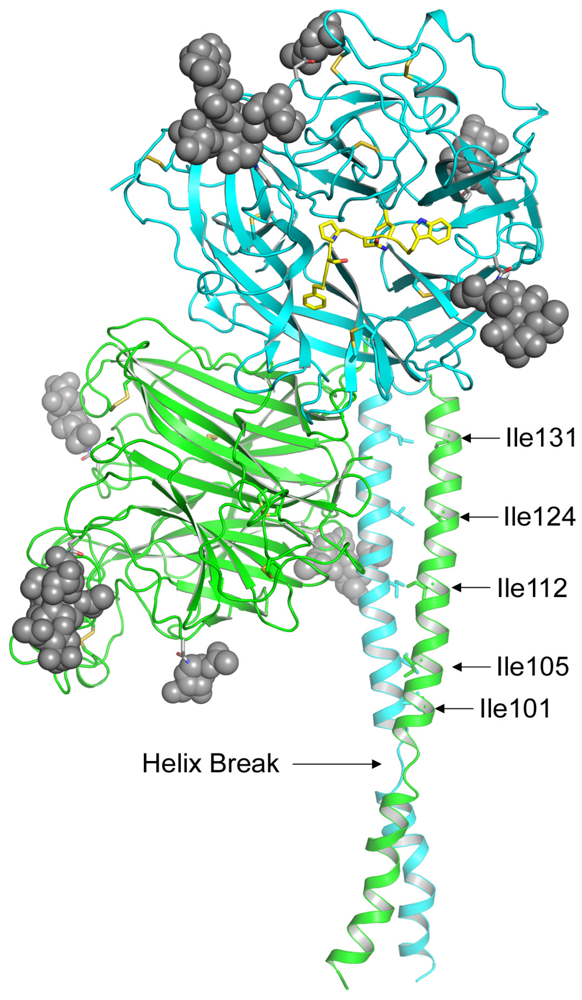
3. Receptors Ephrin-B2 and Ephrin-B3
3.1. Interaction of Henipavirus G with Ephrin Receptors
| Binding Pocket for loop residues | HeV-G glycoprotein residues | NiV-G glycoprotein residues |
|---|---|---|
| F/Y | C240, N557, A558, Q559, E579, I580, Y581, I588, R589 | C240, N557, A558, Q559, E579, I580, Y581, I588, R589 |
| P | P488, G489, Q490, E505, G506, T507, Q530, T531, A532 | P488, G489, Q490, E505, G506, V507, Q530, T531, A532 |
| L | Y458, W504, E505, G506 | F458, W504, E505, G506 |
| W | L305, V401, N402, W504 | L305, I401, R402, W504 |
3.2. Mutations in the G Glycoprotein That Impact Function
| Effect | Location | Group | Subgroup | Residues | Binds | Notes |
|---|---|---|---|---|---|---|
| Decrease | Globular Head—Distal | Group 1 | D257, D260 | Disrupts dimer 1 | ||
| G449, D468 | Disrupts stalk-head interaction 1 | |||||
| K443, K465 | Disrupts hydrogen bonding 1 | |||||
| Globular Head—Proximal | Group 2 | Subgroup 2a | Q530, T531, A532, N557 | Unknown | Decreased receptor binding 2 | |
| Subgroup 2b | N402, E501, E505, G506, E533, Y581, I588 | Ephrin-B2 | No disruption in receptor binding 2 | |||
| Subgroup 2c | Q490, E501, W504, E505, G506, E533, Y581 | Ephrin-B3 | Decreased receptor binding 2 | |||
| N402, I588 | No disruption in receptor binding 2 | |||||
| Stalk | Group 3 | I101, I105, I112, I120, I124, I131, I138, I155, I160 | No disruption in receptor binding 2 | |||
| Increase | Globular Head | Group e1 | Subgroup e1 | D564, D470 | Unknown | No disruption in receptor binding 2 |
| G439 | Decreased receptor binding 2 | |||||
| Subgroup e2 | V401, Q490, Q559 | Ephrin-B2 | No disruption in receptor binding 2 | |||
| W504 | Decreased receptor binding 2 | |||||
| Subgroup e3 | V401, Q559 | Ephrin-B3 | No disruption in receptor binding 2 |
4. Fusion Glycoprotein (F)
4.1. The Fusion Mechanism
5. G and F Glycoprotein Interaction
6. Model of Henipavirus Fusion
7. Therapeutics
7.1. Peptide Fusion Inhibitors and Premature Fusion Triggering
7.2. Monoclonal Antibodies

8. Concluding Remarks
Acknowledgments
Conflict of Interest
References and Notes
- Selvey, L.A.; Wells, R.M.; McCormack, J.G.; Ansford, A.J.; Murray, K.; Rogers, R.J.; Lavercombe, P.S.; Selleck, P.; Sheridan, J.W. Infection of humans and horses by a newly described morbillivirus [see comments]. Med. J. Aust. 1995, 162, 642–645. [Google Scholar]
- Chua, K.B.; Goh, K.J.; Wong, K.T.; Kamarulzaman, A.; Tan, P.S.; Ksiazek, T.G.; Zaki, S.R.; Paul, G.; Lam, S.K.; Tan, C.T. Fatal encephalitis due to Nipah virus among pig-farmers in Malaysia [see comments]. Lancet 1999, 354, 1257–1259. [Google Scholar]
- Chua, K.B.; Bellini, W.J.; Rota, P.A.; Harcourt, B.H.; Tamin, A.; Lam, S.K.; Ksiazek, T.G.; Rollin, P.E.; Zaki, S.R.; Shieh, W.; et al. Nipah virus: A recently emergent deadly paramyxovirus. Science 2000, 288, 1432–1435. [Google Scholar]
- Eaton, B.T.; Broder, C.C.; Middleton, D.; Wang, L.F. Hendra and Nipah viruses: Different and dangerous. Nat. Rev. Microbiol. 2006, 4, 23–35. [Google Scholar]
- Chong, H.T.; Kunjapan, S.R.; Thayaparan, T.; Tong, J.; Petharunam, V.; Jusoh, M.R.; Tan, C.T. Nipah encephalitis outbreak in Malaysia, clinical features in patients from Seremban. Can. J. Neurol. Sci. 2002, 29, 83–87. [Google Scholar]
- Chua, K.B. Nipah virus outbreak in Malaysia. J. Clin. Virol. 2003, 26, 265–275. [Google Scholar]
- Goh, K.J.; Tan, C.T.; Chew, N.K.; Tan, P.S.; Kamarulzaman, A.; Sarji, S.A.; Wong, K.T.; Abdullah, B.J.; Chua, K.B.; Lam, S.K. Clinical features of Nipah virus encephalitis among pig farmers in Malaysia [see comments]. N. Engl. J. Med. 2000, 342, 1229–1235. [Google Scholar]
- Playford, E.G.; McCall, B.; Smith, G.; Slinko, V.; Allen, G.; Smith, I.; Moore, F.; Taylor, C.; Kung, Y.H.; Field, H. Human Hendra virus encephalitis associated with equine outbreak, Australia, 2008. Emerg. Infect. Dis. 2010, 16, 219–223. [Google Scholar]
- Hossain, M.J.; Gurley, E.S.; Montgomery, J.M.; Bell, M.; Carroll, D.S.; Hsu, V.P.; Formenty, P.; Croisier, A.; Bertherat, E.; Faiz, M.A.; et al. Clinical presentation of Nipah virus infection in Bangladesh. Clin. Infect. Dis. 2008, 46, 977–984. [Google Scholar]
- Wong, K.T.; Shieh, W.J.; Kumar, S.; Norain, K.; Abdullah, W.; Guarner, J.; Goldsmith, C.S.; Chua, K.B.; Lam, S.K.; Tan, C.T.; et al. Nipah virus infection: Pathology and pathogenesis of an emerging paramyxoviral zoonosis. Am. J. Pathol. 2002, 161, 2153–2167. [Google Scholar]
- Chong, H.T.; Tan, C.T. Relapsed and late-onset Nipah encephalitis, a report of three cases. Neurol. J. Southeast Asia 2003, 8, 109–112. [Google Scholar]
- Tan, C.T.; Goh, K.J.; Wong, K.T.; Sarji, S.A.; Chua, K.B.; Chew, N.K.; Murugasu, P.; Loh, Y.L.; Chong, H.T.; Tan, K.S.; et al. Relapsed and late-onset Nipah encephalitis. Ann. Neurol. 2002, 51, 703–708. [Google Scholar]
- Wong, K.T.; Robertson, T.; Ong, B.B.; Chong, J.W.; Yaiw, K.C.; Wang, L.F.; Ansford, A.J.; Tannenberg, A. Human Hendra virus infection causes acute and relapsing encephalitis. Neuropathol. Appl. Neurobiol. 2009, 35, 296–305. [Google Scholar]
- Field, H.E.; Mackenzie, J.S.; Daszak, P. Henipaviruses: Emerging paramyxoviruses associated with fruit bats. Curr. Top. Microbiol. Immunol. 2007, 315, 133–159. [Google Scholar]
- Gurley, E.S.; Montgomery, J.M.; Hossain, M.J.; Bell, M.; Azad, A.K.; Islam, M.R.; Molla, M.A.; Carroll, D.S.; Ksiazek, T.G.; Rota, P.A.; et al. Person-to-person transmission of Nipah virus in a Bangladeshi community. Emerg. Infect. Dis. 2007, 13, 1031–1037. [Google Scholar]
- Homaira, N.; Rahman, M.; Hossain, M.J.; Epstein, J.H.; Sultana, R.; Khan, M.S.; Podder, G.; Nahar, K.; Ahmed, B.; Gurley, E.S.; et al. Nipah virus outbreak with person-to-person transmission in a district of Bangladesh, 2007. Epidemiol. Infect. 2010, 138, 1630–1636. [Google Scholar]
- Luby, S.P.; Gurley, E.S.; Hossain, M.J. Transmission of human infection with Nipah virus. Clin. Infect. Dis. 2009, 49, 1743–1748. [Google Scholar]
- Field, H.; Schaaf, K.; Kung, N.; Simon, C.; Waltisbuhl, D.; Hobert, H.; Moore, F.; Middleton, D.; Crook, A.; Smith, G.; et al. Hendra virus outbreak with novel clinical features, Australia. Emerg. Infect. Dis. 2010, 16, 338–340. [Google Scholar]
- Bishop, K.A.; Broder, C.C. Hendra and Nipah: Lethal Zoonotic Paramyxoviruses. In Emerging Infections; Scheld, W.M., Hammer, S.M., Hughes, J.M., Eds.; American Society for Microbiology: Washington, DC, USA, 2008; pp. 155–187. [Google Scholar]
- Hendra Virus, Human, Equine—Australia (04): Queensland Fatal. Pro-MED-mail, Archive No. 20090903.3098. International Society for Infectious Diseases: Brookline, MA, USA, 3 September 2009. Available online: http://www.promedmail.org/ (accessed on 8 February 2012).
- Hendra Virus, Equine—Australia (28): (Queensland, New South Wales). Pro-MED-mail, Archive No. 20111013.3061. International Society for Infectious Diseases: Brookline, MA, USA, 12 October 2011. Available online: http://www.promedmail.org/ (accessed on 8 February 2012).
- Smith, I.; Broos, A.; de Jong, C.; Zeddeman, A.; Smith, C.; Smith, G.; Moore, F.; Barr, J.; Crameri, G.; Marsh, G.; et al. Identifying Hendra virus diversity in pteropid bats. PLoS One 2011, 6. [Google Scholar]
- Pallister, J.; Middleton, D.; Broder, C.C.; Wang, L.F. Henipavirus vaccine development. J. Bioterr. Biodef. 2011, S1. [Google Scholar]
- Luby, S.P.; Hossain, M.J.; Gurley, E.S.; Ahmed, B.N.; Banu, S.; Khan, S.U.; Homaira, N.; Rota, P.A.; Rollin, P.E.; Comer, J.A.; et al. Recurrent zoonotic transmission of Nipah virus into humans, Bangladesh, 2001–2007. Emerg. Infect. Dis. 2009, 15, 1229–1235. [Google Scholar]
- Stone, R. Epidemiology. Breaking the chain in Bangladesh. Science 2011, 331, 1128–1131. [Google Scholar] [CrossRef]
- Nahar, N.; Sultana, R.; Gurley, E.S.; Hossain, M.J.; Luby, S.P. Date palm sap collection: Exploring opportunities to prevent Nipah transmission. Ecohealth 2010, 7, 196–203. [Google Scholar]
- Balzer, M. Hendra vaccine success announced. Aust. Vet. J. 2011, 89, N2–N3. [Google Scholar]
- Sendow, I.; Field, H.E.; Adjid, A.; Ratnawati, A.; Breed, A.C.; Darminto; Morrissy, C.; Daniels, P. Screening for Nipah virus infection in West Kalimantan province, Indonesia. Zoonoses Public Health 2010, 57, 499–503. [Google Scholar] [CrossRef]
- Sendow, I.; Field, H.E.; Curran, J.; Darminto; Morrissy, C.; Meehan, G.; Buick, T.; Daniels, P. Henipavirus in Pteropus vampyrus bats, Indonesia. Emerg. Infect. Dis. 2006, 12, 711–712. [Google Scholar]
- Wacharapluesadee, S.; Lumlertdacha, B.; Boongird, K.; Wanghongsa, S.; Chanhome, L.; Rollin, P.; Stockton, P.; Rupprecht, C.E.; Ksiazek, T.G.; Hemachudha, T. Bat Nipah virus, Thailand. Emerg. Infect. Dis. 2005, 11, 1949–1951. [Google Scholar]
- Drexler, J.F.; Corman, V.M.; Gloza-Rausch, F.; Seebens, A.; Annan, A.; Ipsen, A.; Kruppa, T.; Muller, M.A.; Kalko, E.K.; Adu-Sarkodie, Y.; et al. Henipavirus RNA in African bats. PLoS One 2009, 4. [Google Scholar]
- Hayman, D.T.; Suu-Ire, R.; Breed, A.C.; McEachern, J.A.; Wang, L.; Wood, J.L.; Cunningham, A.A. Evidence of Henipavirus infection in West African fruit bats. PLoS One 2008, 3. [Google Scholar]
- Iehle, C.; Razafitrimo, G.; Razainirina, J.; Andriaholinirina, N.; Goodman, S.M.; Faure, C.; Georges-Courbot, M.C.; Rousset, D.; Reynes, J.M. Henipavirus and Tioman virus antibodies in pteropodid bats, Madagascar. Emerg. Infect. Dis. 2007, 13, 159–161. [Google Scholar]
- Li, Y.; Wang, J.; Hickey, A.C.; Zhang, Y.; Wu, Y.; Zhang, H.; Yuan, J.; Han, Z.; McEachern, J.; Broder, C.C.; et al. Antibodies to Nipah or Nipah-like viruses in bats, China. Emerg. Infect. Dis. 2008, 14, 1974–1976. [Google Scholar]
- Hayman, D.T.; Wang, L.F.; Barr, J.; Baker, K.S.; Suu-Ire, R.; Broder, C.C.; Cunningham, A.A.; Wood, J.L. Antibodies to henipavirus or henipa-like viruses in domestic pigs in Ghana, West Africa. PLoS One 2011, 6. [Google Scholar]
- Lamb, R.A.; Collins, P.L.; Kolakofsky, D.; Melero, J.A.; Nagai, Y.; Oldstone, M.B.A.; Pringle, C.R.; Rima, B.K. Family Paramyxoviridae. In Virus Taxonomy: The Classification and Nomenclature of Viruses. The Eighth Report of the International Committee in Taxonomy of Viruses; Fauquet, C.M., Mayo, M.A., Maniloff, J., Desselberger, U., Ball, L.A., Eds.; Elsevier Academic Press: San Diego, CA, USA, 2005; pp. 655–668. [Google Scholar]
- Harcourt, B.H.; Tamin, A.; Ksiazek, T.G.; Rollin, P.E.; Anderson, L.J.; Bellini, W.J.; Rota, P.A. Molecular characterization of Nipah virus, a newly emergent paramyxovirus. Virology 2000, 271, 334–349. [Google Scholar]
- Bossart, K.N.; Broder, C.C. Paramyxovirus Entry. In Viral Entry into Host Cells; Pöhlmann, S., Simmons, G., Eds.; Landes Bioscience: Austin, TX, USA, 2009. [Google Scholar]
- Lee, B.; Ataman, Z.A. Modes of paramyxovirus fusion: A Henipavirus perspective. Trends Microbiol. 2011, 19, 389–399. [Google Scholar]
- Muhlebach, M.D.; Mateo, M.; Sinn, P.L.; Prufer, S.; Uhlig, K.M.; Leonard, V.H.; Navaratnarajah, C.K.; Frenzke, M.; Wong, X.X.; Sawatsky, B.; et al. Adherens junction protein nectin-4 is the epithelial receptor for measles virus. Nature 2011. [Google Scholar]
- Noyce, R.S.; Bondre, D.G.; Ha, M.N.; Lin, L.T.; Sisson, G.; Tsao, M.S.; Richardson, C.D. Tumor cell marker PVRL4 (nectin 4) is an epithelial cell receptor for measles virus. PLoS Pathog. 2011, 7. [Google Scholar]
- Bowden, T.A.; Aricescu, A.R.; Gilbert, R.J.; Grimes, J.M.; Jones, E.Y.; Stuart, D.I. Structural basis of Nipah and Hendra virus attachment to their cell-surface receptor ephrin-B2. Nat. Struct. Mol. Biol. 2008, 15, 567–572. [Google Scholar]
- Xu, K.; Rajashankar, K.R.; Chan, Y.P.; Himanen, J.P.; Broder, C.C.; Nikolov, D.B. Host cell recognition by the henipaviruses: Crystal structures of the Nipah G attachment glycoprotein and its complex with ephrin-B3. Proc. Natl. Acad. Sci. U. S. A. 2008, 105, 9953–9958. [Google Scholar]
- Bowden, T.A.; Crispin, M.; Harvey, D.J.; Jones, E.Y.; Stuart, D.I. Dimeric architecture of the Hendra virus attachment glycoprotein: Evidence for a conserved mode of assembly. J. Virol. 2010, 84, 6208–6217. [Google Scholar]
- Bowden, T.A.; Crispin, M.; Harvey, D.J.; Aricescu, A.R.; Grimes, J.M.; Jones, E.Y.; Stuart, D.I. Crystal structure and carbohydrate analysis of Nipah virus attachment glycoprotein: A template for antiviral and vaccine design. J. Virol. 2008, 82, 11628–11636. [Google Scholar]
- Xu, K.; Broder, C.C.; Nikolov, D.B. 2012. Memorial Sloan-Kettering Cancer Center, New York, NY, USA; Uniformed Services University, Bethesda, MD, USA. Unpublished work.
- Colgrave, M.L.; Snelling, H.J.; Shiell, B.J.; Feng, Y.R.; Chan, Y.P.; Bossart, K.N.; Xu, K.; Nikolov, D.B.; Broder, C.C.; Michalski, W.P. Site occupancy and glycan compositional analysis of two soluble recombinant forms of the attachment glycoprotein of Hendra virus. Glycobiology 2011. [Google Scholar]
- Bossart, K.N.; Crameri, G.; Dimitrov, A.S.; Mungall, B.A.; Feng, Y.R.; Patch, J.R.; Choudhary, A.; Wang, L.F.; Eaton, B.T.; Broder, C.C. Receptor binding, fusion inhibition, and induction of cross-reactive neutralizing antibodies by a soluble G glycoprotein of Hendra virus. J. Virol. 2005, 79, 6690–6702. [Google Scholar]
- Yuan, P.; Swanson, K.A.; Leser, G.P.; Paterson, R.G.; Lamb, R.A.; Jardetzky, T.S. Structure of the Newcastle disease virus hemagglutinin-neuraminidase (HN) ectodomain reveals a four-helix bundle stalk. Proc. Natl. Acad. Sci. U. S. A. 2011, 108, 14920–14925. [Google Scholar]
- Kelley, L.A.; Sternberg, M.J. Protein structure prediction on the Web: A case study using the Phyre server. Nat. Protoc. 2009, 4, 363–371. [Google Scholar]
- Bishop, K.A.; Hickey, A.C.; Khetawat, D.; Patch, J.R.; Bossart, K.N.; Zhu, Z.; Wang, L.F.; Dimitrov, D.S.; Broder, C.C. Residues in the stalk domain of the Hendra virus G glycoprotein modulate conformational changes associated with receptor binding. J. Virol. 2008, 82, 11398–11409. [Google Scholar]
- Yuan, P.; Thompson, T.B.; Wurzburg, B.A.; Paterson, R.G.; Lamb, R.A.; Jardetzky, T.S. Structural studies of the parainfluenza virus 5 hemagglutinin-neuraminidase tetramer in complex with its receptor, sialyllactose. Structure 2005, 13, 803–815. [Google Scholar]
- Bonaparte, M.I.; Dimitrov, A.S.; Bossart, K.N.; Crameri, G.; Mungall, B.A.; Bishop, K.A.; Choudhry, V.; Dimitrov, D.S.; Wang, L.F.; Eaton, B.T.; et al. Ephrin-B2 ligand is a functional receptor for Hendra virus and Nipah virus. Proc. Natl. Acad. Sci. U. S. A. 2005, 102, 10652–10657. [Google Scholar]
- Negrete, O.A.; Levroney, E.L.; Aguilar, H.C.; Bertolotti-Ciarlet, A.; Nazarian, R.; Tajyar, S.; Lee, B. EphrinB2 is the entry receptor for Nipah virus, an emergent deadly paramyxovirus. Nature 2005, 436, 401–405. [Google Scholar]
- Negrete, O.A.; Wolf, M.C.; Aguilar, H.C.; Enterlein, S.; Wang, W.; Muhlberger, E.; Su, S.V.; Bertolotti-Ciarlet, A.; Flick, R.; Lee, B. Two key residues in ephrinB3 are critical for its use as an alternative receptor for Nipah virus. PLoS Pathog. 2006, 2. [Google Scholar]
- Bishop, K.A.; Stantchev, T.S.; Hickey, A.C.; Khetawat, D.; Bossart, K.N.; Krasnoperov, V.; Gill, P.; Feng, Y.R.; Wang, L.; Eaton, B.T.; et al. Identification of Hendra virus G glycoprotein residues that are critical for receptor binding. J. Virol. 2007, 81, 5893–5901. [Google Scholar]
- Pasquale, E.B. Eph receptors and ephrins in cancer: Bidirectional signalling and beyond. Nat. Rev. Cancer 2010, 10, 165–180. [Google Scholar]
- Lackmann, M.; Boyd, A.W. Eph, a protein family coming of age: More confusion, insight, or complexity? Sci. Signal. 2008, 1. [Google Scholar]
- Bossart, K.N.; Tachedjian, M.; McEachern, J.A.; Crameri, G.; Zhu, Z.; Dimitrov, D.S.; Broder, C.C.; Wang, L.F. Functional studies of host-specific ephrin-B ligands as Henipavirus receptors. Virology 2008, 372, 357–371. [Google Scholar]
- Gale, N.W.; Baluk, P.; Pan, L.; Kwan, M.; Holash, J.; DeChiara, T.M.; McDonald, D.M.; Yancopoulos, G.D. Ephrin-B2 selectively marks arterial vessels and neovascularization sites in the adult, with expression in both endothelial and smooth-muscle cells. Dev. Biol. 2001, 230, 151–160. [Google Scholar]
- Poliakov, A.; Cotrina, M.; Wilkinson, D.G. Diverse roles of eph receptors and ephrins in the regulation of cell migration and tissue assembly. Dev. Cell 2004, 7, 465–480. [Google Scholar]
- Pasquale, E.B. Eph-ephrin bidirectional signaling in physiology and disease. Cell 2008, 133, 38–52. [Google Scholar]
- Hooper, P.; Zaki, S.; Daniels, P.; Middleton, D. Comparative pathology of the diseases caused by Hendra and Nipah viruses. Microbes Infect. 2001, 3, 315–322. [Google Scholar]
- Wong, K.T. Emerging epidemic viral encephalitides with a special focus on henipaviruses. Acta Neuropathol. 2010, 120, 317–325. [Google Scholar]
- Bossart, K.N.; McEachern, J.A.; Hickey, A.C.; Choudhry, V.; Dimitrov, D.S.; Eaton, B.T.; Wang, L.F. Neutralization assays for differential Henipavirus serology using Bio-Plex Protein Array Systems. J. Virol. Methods 2007, 142, 29–40. [Google Scholar]
- Negrete, O.A.; Chu, D.; Aguilar, H.C.; Lee, B. Single amino acid changes in the Nipah and Hendra virus attachment glycoproteins distinguish ephrinB2 from ephrinB3 Usage. J. Virol. 2007, 81, 10804–10814. [Google Scholar]
- Toth, J.; Cutforth, T.; Gelinas, A.D.; Bethoney, K.A.; Bard, J.; Harrison, C.J. Crystal structure of an ephrin ectodomain. Dev. Cell 2001, 1, 83–92. [Google Scholar]
- Koolpe, M.; Burgess, R.; Dail, M.; Pasquale, E.B. EphB receptor-binding peptides identified by phage display enable design of an antagonist with ephrin-like affinity. J. Biol. Chem. 2005, 280, 17301–17311. [Google Scholar]
- Nikolov, D.B.; Li, C.; Barton, W.A.; Himanen, J.P. Crystal structure of the ephrin-B1 ectodomain: Implications for receptor recognition and signaling. Biochemistry 2005, 44, 10947–10953. [Google Scholar]
- Navaratnarajah, C.K.; Oezguen, N.; Rupp, L.; Kay, L.; Leonard, V.H.; Braun, W.; Cattaneo, R. The heads of the measles virus attachment protein move to transmit the fusion-triggering signal. Nat. Struct. Mol. Biol. 2011, 18, 128–134. [Google Scholar]
- Guillaume, V.; Aslan, H.; Ainouze, M.; Guerbois, M.; Wild, T.F.; Buckland, R.; Langedijk, J.P. Evidence of a potential receptor-binding site on the Nipah virus G protein (NiV-G): Identification of globular head residues with a role in fusion promotion and their localization on an NiV-G structural model. J. Virol. 2006, 80, 7546–7554. [Google Scholar]
- Yuan, J.; Marsh, G.; Khetawat, D.; Broder, C.C.; Wang, L.F.; Shi, Z. Mutations in the G-H loop region of ephrin-B2 can enhance Nipah virus binding and infection. J. Gen. Virol. 2011, 92, 2142–2152. [Google Scholar]
- Corey, E.A.; Iorio, R.M. Mutations in the stalk of the measles virus hemagglutinin protein decrease fusion but do not interfere with virus-specific interaction with the homologous fusion protein. J. Virol. 2007, 81, 9900–9910. [Google Scholar]
- Brindley, M.A.; Plemper, R.K. Blue native PAGE and biomolecular complementation reveal a tetrameric or higher-order oligomer organization of the physiological measles virus attachment protein H. J. Virol. 2010, 84, 12174–12184. [Google Scholar]
- Pager, C.T.; Dutch, R.E. Cathepsin L is involved in proteolytic processing of the Hendra virus fusion protein. J. Virol. 2005, 79, 12714–12720. [Google Scholar]
- Meulendyke, K.A.; Wurth, M.A.; McCann, R.O.; Dutch, R.E. Endocytosis plays a critical role in proteolytic processing of the Hendra virus fusion protein. J. Virol. 2005, 79, 12643–12649. [Google Scholar]
- Vogt, C.; Eickmann, M.; Diederich, S.; Moll, M.; Maisner, A. Endocytosis of the Nipah virus glycoproteins. J. Virol. 2005, 79, 3865–3872. [Google Scholar]
- Moll, M.; Diederich, S.; Klenk, H.D.; Czub, M.; Maisner, A. Ubiquitous activation of the Nipah virus fusion protein does not require a basic amino Acid at the cleavage site. J. Virol. 2004, 78, 9705–9712. [Google Scholar]
- Lou, Z.; Xu, Y.; Xiang, K.; Su, N.; Qin, L.; Li, X.; Gao, G.F.; Bartlam, M.; Rao, Z. Crystal structures of Nipah and Hendra virus fusion core proteins. FEBS J. 2006, 273, 4538–4547. [Google Scholar]
- Popa, A.; Pager, C.T.; Dutch, R.E. C-terminal tyrosine residues modulate the fusion activity of the Hendra virus fusion protein. Biochemistry 2011, 50, 945–952. [Google Scholar]
- Weise, C.; Erbar, S.; Lamp, B.; Vogt, C.; Diederich, S.; Maisner, A. Tyrosine residues in the cytoplasmic domains affect sorting and fusion activity of the Nipah virus glycoproteins in polarized epithelial cells. J. Virol. 2010, 84, 7634–7641. [Google Scholar]
- Yin, H.S.; Wen, X.; Paterson, R.G.; Lamb, R.A.; Jardetzky, T.S. Structure of the parainfluenza virus 5 F protein in its metastable, prefusion conformation. Nature 2006, 439, 38–44. [Google Scholar]
- Yin, H.S.; Paterson, R.G.; Wen, X.; Lamb, R.A.; Jardetzky, T.S. Structure of the uncleaved ectodomain of the paramyxovirus (hPIV3) fusion protein. Proc. Natl. Acad. Sci. U. S. A. 2005, 102, 9288–9293. [Google Scholar]
- Lamb, R.A.; Jardetzky, T.S. Structural basis of viral invasion: Lessons from paramyxovirus F. Curr. Opin. Struct. Biol. 2007, 17, 427–436. [Google Scholar]
- Smith, E.C.; Popa, A.; Chang, A.; Masante, C.; Dutch, R.E. Viral entry mechanisms: The increasing diversity of paramyxovirus entry. FEBS J. 2009, 276, 7217–7227. [Google Scholar]
- Karron, R.A.; Buonagurio, D.A.; Georgiu, A.F.; Whitehead, S.S.; Adamus, J.E.; Clements-Mann, M.L.; Harris, D.O.; Randolph, V.B.; Udem, S.A.; Murphy, B.R.; et al. Respiratory syncytial virus (RSV) SH and G proteins are not essential for viral replication in vitro: Clinical evaluation and molecular characterization of a cold-passaged, attenuated RSV subgroup B mutant. Proc. Natl. Acad. Sci. U. S. A. 1997, 94, 13961–13966. [Google Scholar]
- Techaarpornkul, S.; Barretto, N.; Peeples, M.E. Functional analysis of recombinant respiratory syncytial virus deletion mutants lacking the small hydrophobic and/or attachment glycoprotein gene. J. Virol. 2001, 75, 6825–6834. [Google Scholar]
- Dutch, R.E.; Joshi, S.B.; Lamb, R.A. Membrane fusion promoted by increasing surface densities of the paramyxovirus F and HN proteins: Comparison of fusion reactions mediated by simian virus 5 F, human parainfluenza virus type 3 F, and influenza virus HA. J. Virol. 1998, 72, 7745–7753. [Google Scholar]
- Aguilar, H.C.; Matreyek, K.A.; Choi, D.Y.; Filone, C.M.; Young, S.; Lee, B. Polybasic KKR motif in the cytoplasmic tail of Nipah virus fusion protein modulates membrane fusion by inside-out signaling. J. Virol. 2007, 81, 4520–4532. [Google Scholar]
- Aguilar, H.C.; Ataman, Z.A.; Aspericueta, V.; Fang, A.Q.; Stroud, M.; Negrete, O.A.; Kammerer, R.A.; Lee, B. A novel receptor-induced activation site in the Nipah virus attachment glycoprotein (G) involved in triggering the fusion glycoprotein (F). J. Biol. Chem. 2009, 284, 1628–1635. [Google Scholar]
- Whitman, S.D.; Smith, E.C.; Dutch, R.E. Differential rates of protein folding and cellular trafficking for the Hendra virus F and G proteins: Implications for F-G complex formation. J. Virol. 2009, 83, 8998–9001. [Google Scholar]
- Plemper, R.K.; Hammond, A.L.; Cattaneo, R. Measles virus envelope glycoproteins hetero-oligomerize in the endoplasmic reticulum. J. Biol. Chem. 2001, 276, 44239–44246. [Google Scholar]
- Stone-Hulslander, J.; Morrison, T.G. Detection of an interaction between the HN and F proteins in Newcastle disease virus-infected cells. J. Virol. 1997, 71, 6287–6295. [Google Scholar]
- Tong, S.; Compans, R.W. Alternative mechanisms of interaction between homotypic and heterotypic parainfluenza virus HN and F proteins. J. Gen. Virol. 1999, 80, 107–115. [Google Scholar]
- Paterson, R.G.; Johnson, M.L.; Lamb, R.A. Paramyxovirus fusion (F) protein and hemagglutinin-neuraminidase (HN) protein interactions: Intracellular retention of F and HN does not affect transport of the homotypic HN or F protein. Virology 1997, 237, 1–9. [Google Scholar]
- Whitman, S.D.; Dutch, R.E. Surface density of the Hendra G protein modulates Hendra F protein-promoted membrane fusion: Role for Hendra G protein trafficking and degradation. Virology 2007, 363, 419–429. [Google Scholar]
- Sergel, T.; McGinnes, L.W.; Peeples, M.E.; Morrison, T.G. The attachment function of the Newcastle disease virus hemagglutinin-neuraminidase protein can be separated from fusion promotion by mutation. Virology 1993, 193, 717–726. [Google Scholar]
- Melanson, V.R.; Iorio, R.M. Amino acid substitutions in the F-specific domain in the stalk of the newcastle disease virus HN protein modulate fusion and interfere with its interaction with the F protein. J. Virol. 2004, 78, 13053–13061. [Google Scholar]
- Li, J.; Quinlan, E.; Mirza, A.; Iorio, R.M. Mutated form of the Newcastle disease virus hemagglutinin-neuraminidase interacts with the homologous fusion protein despite deficiencies in both receptor recognition and fusion promotion. J. Virol. 2004, 78, 5299–5310. [Google Scholar]
- Connolly, S.A.; Leser, G.P.; Jardetzky, T.S.; Lamb, R.A. Bimolecular complementation of paramyxovirus fusion and hemagglutinin-neuraminidase proteins enhances fusion: Implications for the mechanism of fusion triggering. J. Virol. 2009, 83, 10857–10868. [Google Scholar]
- Plemper, R.K.; Hammond, A.L.; Gerlier, D.; Fielding, A.K.; Cattaneo, R. Strength of envelope protein interaction modulates cytopathicity of measles virus. J. Virol. 2002, 76, 5051–5061. [Google Scholar]
- Plemper, R.K.; Brindley, M.A.; Iorio, R.M. Structural and mechanistic studies of measles virus illuminate paramyxovirus entry. PLoS Pathog. 2011, 7. [Google Scholar]
- Dutch, R.E. Entry and fusion of emerging paramyxoviruses. PLoS Pathog. 2010, 6. [Google Scholar]
- Aguilar, H.C.; Matreyek, K.A.; Filone, C.M.; Hashimi, S.T.; Levroney, E.L.; Negrete, O.A.; Bertolotti-Ciarlet, A.; Choi, D.Y.; McHardy, I.; Fulcher, J.A.; et al. N-Glycans on Nipah virus fusion protein protect against neutralization but reduce membrane fusion and viral entry. J. Virol. 2006, 80, 4878–4889. [Google Scholar]
- Corey, E.A.; Iorio, R.M. Measles virus attachment proteins with impaired ability to bind CD46 interact more efficiently with the homologous fusion protein. Virology 2009, 383, 1–5. [Google Scholar]
- Chan, Y.P.; Broder, C.C. 2012; Department of Microbiology and Immunology, Uniformed Services University, Bethesda, MD, USA. Unpublished work.
- Porotto, M.; Devito, I.; Palmer, S.G.; Jurgens, E.M.; Yee, J.L.; Yokoyama, C.C.; Pessi, A.; Moscona, A. Spring-loaded model revisited: Paramyxovirus fusion requires engagement of a receptor binding protein beyond initial triggering of the fusion protein. J. Virol. 2011, 85, 12867–12880. [Google Scholar]
- Porotto, M.; Palmer, S.G.; Palermo, L.M.; Moscona, A. Mechanism of fusion triggering by human parainfluenza virus type III: Communication between viral glycoproteins during entry. J. Biol. Chem. 2011, 287, 778–793. [Google Scholar]
- Chong, H.T.; Kamarulzaman, A.; Tan, C.T.; Goh, K.J.; Thayaparan, T.; Kunjapan, S.R.; Chew, N.K.; Chua, K.B.; Lam, S.K. Treatment of acute Nipah encephalitis with ribavirin. Ann. Neurol. 2001, 49, 810–813. [Google Scholar]
- Pager, C.T.; Wurth, M.A.; Dutch, R.E. Subcellular localization and calcium and pH requirements for proteolytic processing of the Hendra virus fusion protein. J. Virol. 2004, 78, 9154–9163. [Google Scholar]
- Porotto, M.; Orefice, G.; Yokoyama, C.C.; Mungall, B.A.; Realubit, R.; Sganga, M.L.; Aljofan, M.; Whitt, M.; Glickman, F.; Moscona, A. Simulating henipavirus multicycle replication in a screening assay leads to identification of a promising candidate for therapy. J. Virol. 2009, 83, 5148–5155. [Google Scholar]
- Hendra Virus, Human, Equine—Australia (05): Queensland. Pro-MED-mail, Archive No. 20090910.3189. International Society for Infectious Diseases: Brookline, MA, USA, 10 September 2009. Available online: http://www.promedmail.org/ (accessed on 8 February 2012).
- Freiberg, A.N.; Worthy, M.N.; Lee, B.; Holbrook, M.R. Combined chloroquine and ribavirin treatment does not prevent death in a hamster model of Nipah and Hendra virus infection. J. Gen. Virol. 2010, 91, 765–772. [Google Scholar]
- Georges-Courbot, M.C.; Contamin, H.; Faure, C.; Loth, P.; Baize, S.; Leyssen, P.; Neyts, J.; Deubel, V. Poly(I)-poly(C12U) but not ribavirin prevents death in a hamster model of Nipah virus infection. Antimicrob. Agents Chemother. 2006, 50, 1768–1772. [Google Scholar]
- Rockx, B.; Bossart, K.N.; Feldmann, F.; Geisbert, J.B.; Hickey, A.C.; Brining, D.; Callison, J.; Safronetz, D.; Marzi, A.; Kercher, L.; et al. A novel model of lethal Hendra virus infection in African green monkeys and the effectiveness of ribavirin treatment. J. Virol. 2010, 84, 9831–9839. [Google Scholar]
- Pallister, J.; Middleton, D.; Crameri, G.; Yamada, M.; Klein, R.; Hancock, T.J.; Foord, A.; Shiell, B.; Michalski, W.; Broder, C.C.; et al. Chloroquine administration does not prevent Nipah virus infection and disease in ferrets. J. Virol. 2009, 83, 11979–11982. [Google Scholar]
- Bossart, K.N.; Broder, C.C. Developments towards effective treatments for Nipah and Hendra virus infection. Expert Rev. Anti Infect. Ther. 2006, 4, 43–55. [Google Scholar]
- Bossart, K.N.; Wang, L.F.; Eaton, B.T.; Broder, C.C. Functional expression and membrane fusion tropism of the envelope glycoproteins of Hendra virus. Virology 2001, 290, 121–135. [Google Scholar]
- Bossart, K.N.; Mungall, B.A.; Crameri, G.; Wang, L.F.; Eaton, B.T.; Broder, C.C. Inhibition of Henipavirus fusion and infection by heptad-derived peptides of the Nipah virus fusion glycoprotein. Virol. J. 2005, 2. [Google Scholar]
- Geisbert, T.W.; Broder, C.C. 2012; Galveston National Laboratory and Department of Microbiology and Immunology, University of Texas Medical Branch, Galveston, TX 77550, USA; Uniformed Services University, Bethesda, MD, USA. Unpublished work.
- Porotto, M.; Carta, P.; Deng, Y.; Kellogg, G.E.; Whitt, M.; Lu, M.; Mungall, B.A.; Moscona, A. Molecular determinants of antiviral potency of paramyxovirus entry inhibitors. J. Virol. 2007, 81, 10567–10574. [Google Scholar]
- Porotto, M.; Doctor, L.; Carta, P.; Fornabaio, M.; Greengard, O.; Kellogg, G.E.; Moscona, A. Inhibition of Hendra virus fusion. J. Virol. 2006, 80, 9837–9849. [Google Scholar]
- Porotto, M.; Rockx, B.; Yokoyama, C.C.; Talekar, A.; Devito, I.; Palermo, L.M.; Liu, J.; Cortese, R.; Lu, M.; Feldmann, H.; et al. Inhibition of Nipah virus infection in vivo: Targeting an early stage of paramyxovirus fusion activation during viral entry. PLoS Pathog. 2010, 6. [Google Scholar]
- Porotto, M.; Yi, F.; Moscona, A.; LaVan, D.A. Synthetic protocells interact with viral nanomachinery and inactivate pathogenic human virus. PLoS One 2011, 6. [Google Scholar]
- Farzan, S.F.; Palermo, L.M.; Yokoyama, C.C.; Orefice, G.; Fornabaio, M.; Sarkar, A.; Kellogg, G.E.; Greengard, O.; Porotto, M.; Moscona, A. Premature activation of the paramyxovirus fusion protein before target cell attachment with corruption of the viral fusion machinery. J. Biol. Chem. 2011, 286, 37945–37954. [Google Scholar]
- Broder, C.C. Therapeutics and Vaccines against Hendra and Nipah Viruses. In New Generation Vaccines, 4th; Levine, M.M., Dougan, G., Good, M.F., Liu, M.A., Nabel, G.J., Nataro, J.P., Rappuoli, R., Eds.; Informa Healthcare USA: New York, NY, USA, 2010; pp. 885–894. [Google Scholar]
- Guillaume, V.; Contamin, H.; Loth, P.; Georges-Courbot, M.C.; Lefeuvre, A.; Marianneau, P.; Chua, K.B.; Lam, S.K.; Buckland, R.; Deubel, V.; et al. Nipah virus: Vaccination and passive protection studies in a hamster model. J. Virol. 2004, 78, 834–840. [Google Scholar]
- Guillaume, V.; Contamin, H.; Loth, P.; Grosjean, I.; Courbot, M.C.; Deubel, V.; Buckland, R.; Wild, T.F. Antibody prophylaxis and therapy against Nipah virus infection in hamsters. J. Virol. 2006, 80, 1972–1978. [Google Scholar]
- Guillaume, V.; Wong, K.T.; Looi, R.Y.; Georges-Courbot, M.C.; Barrot, L.; Buckland, R.; Wild, T.F.; Horvat, B. Acute Hendra virus infection: Analysis of the pathogenesis and passive antibody protection in the hamster model. Virology 2009, 387, 459–465. [Google Scholar]
- Hayden, M.S.; Gilliland, L.K.; Ledbetter, J.A. Antibody engineering. Curr. Opin. Immunol. 1997, 9, 201–212. [Google Scholar]
- Rader, C.; Barbas, C.F., 3rd. Phage display of combinatorial antibody libraries. Curr. Opin. Biotechnol. 1997, 8, 503–508. [Google Scholar]
- Zhu, Z.; Bossart, K.N.; Bishop, K.A.; Crameri, G.; Dimitrov, A.S.; McEachern, J.A.; Feng, Y.; Middleton, D.; Wang, L.F.; Broder, C.C.; et al. Exceptionally potent cross-reactive neutralization of Nipah and Hendra viruses by a human monoclonal antibody. J. Infect. Dis. 2008, 197, 846–853. [Google Scholar]
- Zhu, Z.; Dimitrov, A.S.; Bossart, K.N.; Crameri, G.; Bishop, K.A.; Choudhry, V.; Mungall, B.A.; Feng, Y.R.; Choudhary, A.; Zhang, M.Y.; et al. Potent neutralization of Hendra and Nipah viruses by human monoclonal antibodies. J. Virol. 2006, 80, 891–899. [Google Scholar]
- Bossart, K.N.; Zhu, Z.; Middleton, D.; Klippel, J.; Crameri, G.; Bingham, J.; McEachern, J.A.; Green, D.; Hancock, T.J.; Chan, Y.P.; et al. A neutralizing human monoclonal antibody protects against lethal disease in a new ferret model of acute Nipah virus infection. PLoS Pathog. 2009, 5. [Google Scholar]
- Pallister, J.; Middleton, D.; Broder, C.C. 2012; CSIRO Livestock Industries, Australian Animal Health Laboratory, Geelong, VIC, Australia; Department of Microbiology and Immunology, Uniformed Services University, Bethesda, MD, USA. Unpublished work.
- Bossart, K.N.; Geisbert, T.W.; Feldmann, H.; Zhu, Z.; Feldmann, F.; Geisbert, J.B.; Yan, L.; Feng, Y.R.; Brining, D.; Scott, D.; et al. A neutralizing human monoclonal antibody protects African green monkeys from Hendra virus challenge. Sci. Transl. Med. 2011, 3. [Google Scholar]
- Playford, G. Pathology Queensland, Herston, Queensland, Austral. Personal Communication., 2009. [Google Scholar]
- Hendra Virus, Equine—Australia (05): (Queensland) Human Exposure. Pro-MED-mail, Archive No. 20100527.1761. International Society for Infectious Diseases: Brookline, MA, USA, 27 May 2010. Available online: http://www.promedmail.org/ (accessed on 8 February 2012).
- Xu, K.; Broder, C.C.; Nikolov, D.B. 2012; Memorial Sloan-Kettering Cancer Center, New York, NY, USA; Uniformed Services University, Bethesda, MD, USA. Unpublished work.
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Steffen, D.L.; Xu, K.; Nikolov, D.B.; Broder, C.C. Henipavirus Mediated Membrane Fusion, Virus Entry and Targeted Therapeutics. Viruses 2012, 4, 280-308. https://doi.org/10.3390/v4020280
Steffen DL, Xu K, Nikolov DB, Broder CC. Henipavirus Mediated Membrane Fusion, Virus Entry and Targeted Therapeutics. Viruses. 2012; 4(2):280-308. https://doi.org/10.3390/v4020280
Chicago/Turabian StyleSteffen, Deborah L., Kai Xu, Dimitar B. Nikolov, and Christopher C. Broder. 2012. "Henipavirus Mediated Membrane Fusion, Virus Entry and Targeted Therapeutics" Viruses 4, no. 2: 280-308. https://doi.org/10.3390/v4020280





