Changes of Scots Pine Phyllosphere and Soil Fungal Communities during Outbreaks of Defoliating Insects
Abstract
:1. Introduction
2. Materials and Methods
2.1. Study Design and Site Characteristics
2.2. DNA Extraction from Soil and Needles
2.3. PCR-DGGE
2.4. Silver Staining and Gel Drying
2.5. Selection of Soil DGGE Gels and Profiles for Sequencing
2.6. Band Excision, Reamplification, and Sequencing
2.7. Data Analysis of DGGE Profiles
3. Results
3.1. Bacterial Communities
3.2. Fungal Communities
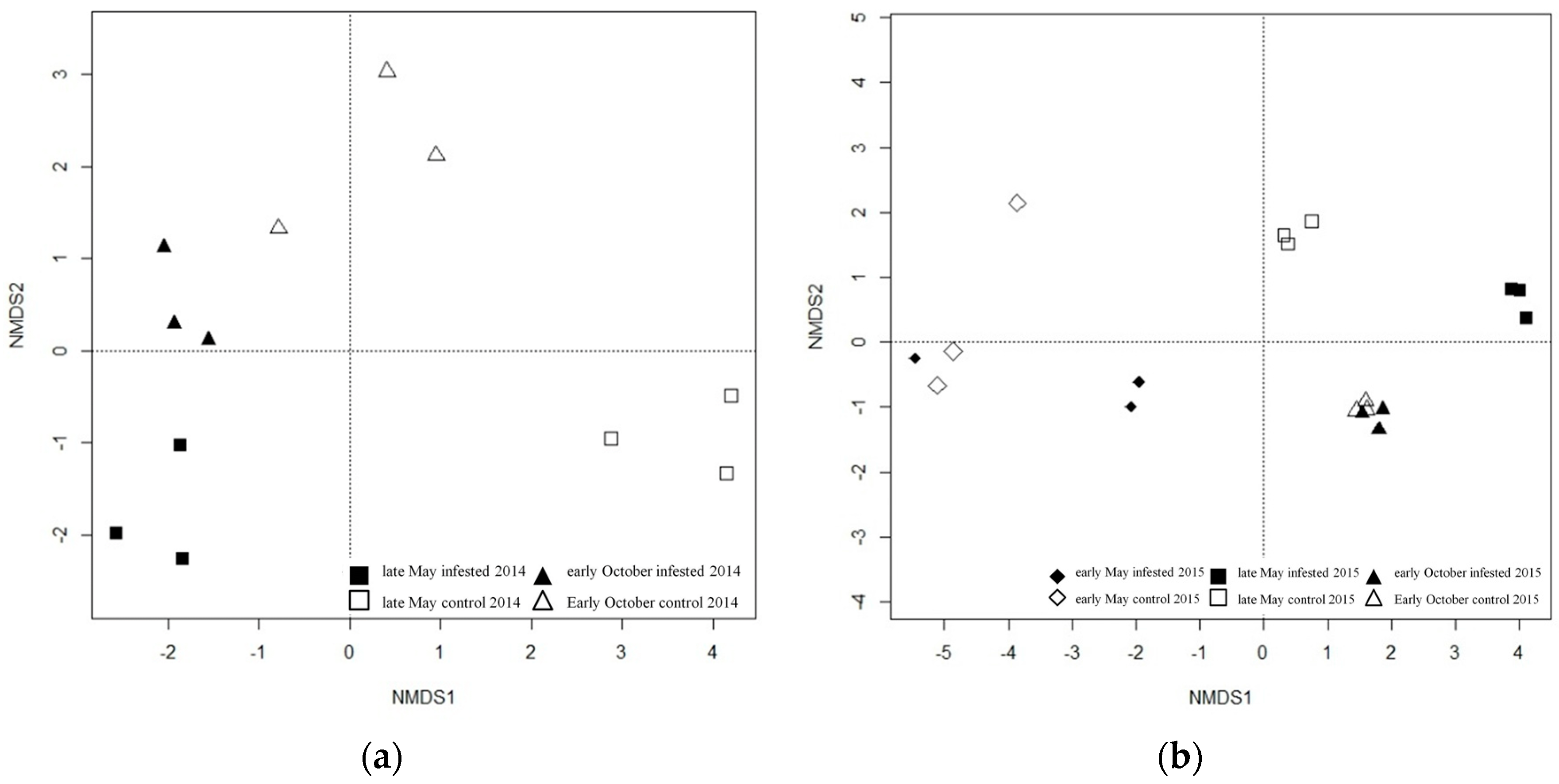
3.2.1. Phyllosphere Fungi
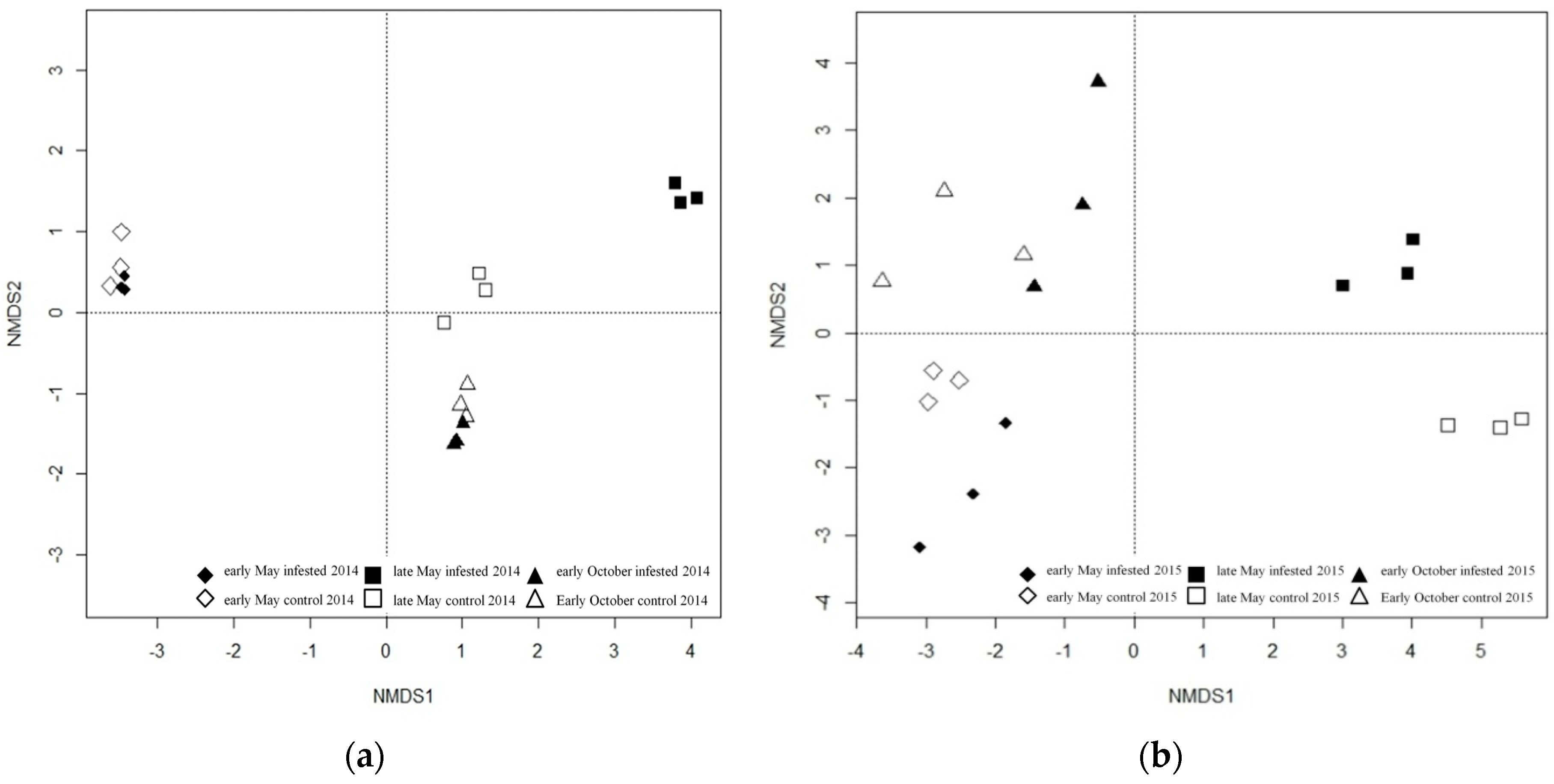
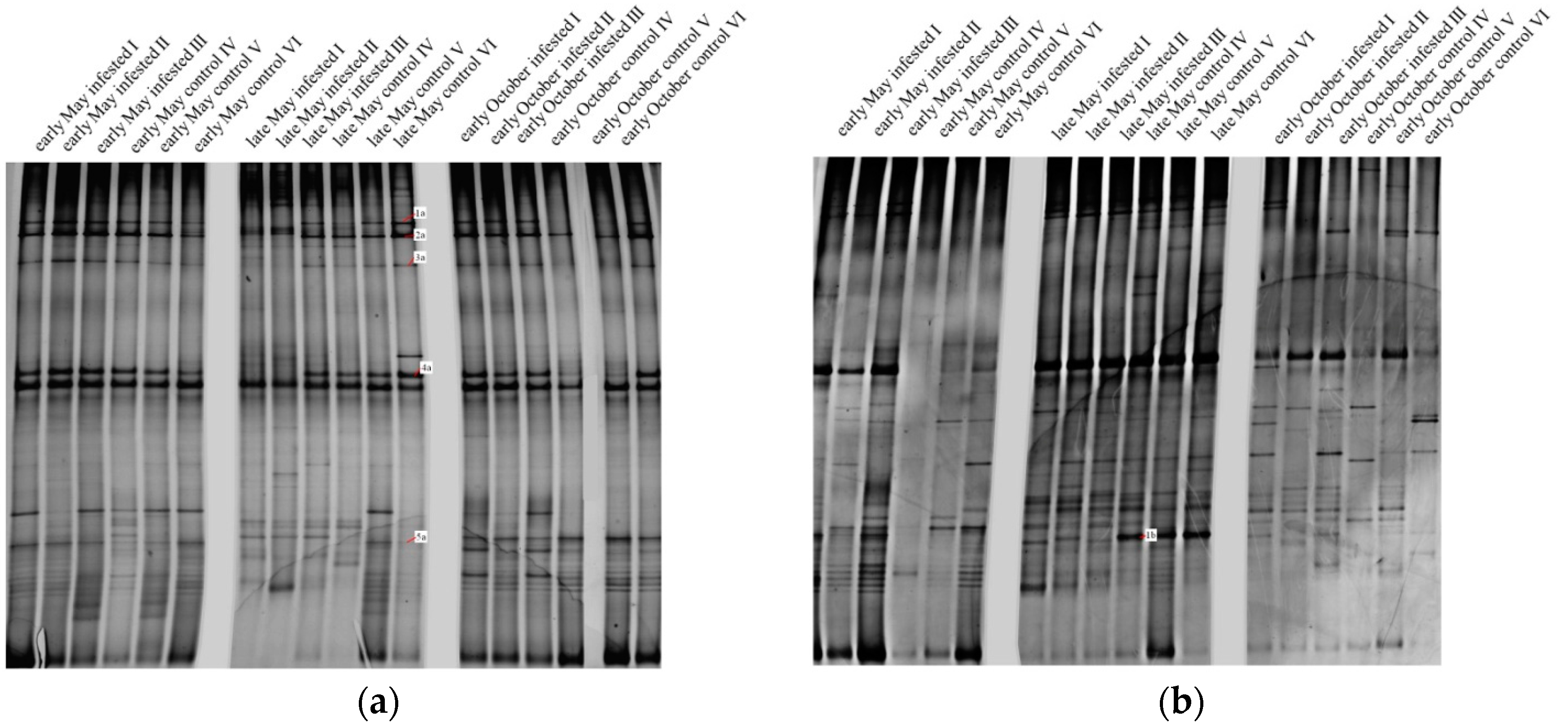
3.2.2. Soil Fungal Community
4. Discussion
4.1. Bacterial Community of Pine Needles and Soil
4.2. Fungal Community of Pine Needles
4.3. Soil Fungal Community
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Van Lierop, P.; Lindquist, E.; Sathyapala, S.; Franceschini, G. Global forest area disturbance from fire, insect pests, diseases and severe weather events. For. Ecol. Manag. 2015, 352, 78–88. [Google Scholar] [CrossRef]
- Dale, V.H.; Joyce, L.A.; McNulty, S.; Neilson, R.P.; Ayres, M.P.; Flannigan, M.D.; Hanson, P.J.; Irland, L.C.; Lugo, A.E.; Peterson, C.J.; et al. Climate change and forest disturbances. BioScience 2001, 51, 723–734. [Google Scholar] [CrossRef]
- Majunke, C.; Möller, K.; Funke, M. Die Nonne. Waldschutz-Merkblatt 52, 3rd ed.; Umweltschutz und Raumordnung des Landes Brandenburg; Landesforstanstalt Eberswalde und Ministerium für Landwirtschaft: Eberswalde, Germany, 2008; p. 10.
- Möller, K.; Heydeck, P. Situationsbericht zum Auftreten von Schaderregern und Schäden im Land Brandenburg; Aktuelle Waldschutzsituation 07/2013; Ministerium für Infrastruktur und Landwirtschaft des Landes Brandenburg: Eberswalde, Germany, 2013; p. 3.
- Langström, B.; Annila, E.; Hellqvist, C.; Varama, M.; Niemelä, P. Tree mortality, needle biomass recovery and growth losses in Scots pine following defoliation by Diprion pini (L.) and subsequent attack by Tomicus piniperda (L.). Scand. J. For. Res. 2001, 16, 342–353. [Google Scholar] [CrossRef]
- Armour, H.; Straw, N.; Day, K. Interactions between growth, herbivory and long-term foliar dynamics of Scots pine. Trees Struct. Funct. 2003, 17, 70–80. [Google Scholar] [CrossRef]
- Hicke, J.A.; Allen, C.D.; Desai, A.R.; Dietze, M.C.; Hall, R.J.; Kashian, D.M.; Moore, D.; Raffa, K.F.; Sturrock, R.N.; Vogelmann, J. Effects of biotic disturbances on forest carbon cycling in the United States and Canada. Glob. Chang. Biol. 2012, 18, 7–34. [Google Scholar] [CrossRef]
- Vorholt, J.A. Microbial life in the phyllosphere. Nat. Rev. Microbiol. 2012, 10, 828–840. [Google Scholar] [CrossRef] [PubMed]
- Knief, C. Analysis of plant microbe interactions in the era of next generation sequencing technologies. Front. Plant Sci. 2014, 5, 216. [Google Scholar] [CrossRef] [PubMed]
- Nakatsu, C.H.; Torsvik, V.; Øvreås, L. Soil community analysis using DGGE of 16S rDNA polymerase chain reaction products. Soil Sci. Soc. Am. J. 2000, 64, 1382–1388. [Google Scholar] [CrossRef]
- Dı́ez, B.; Pedrós-Alió, C.; Marsh, T.L.; Massana, R. Application of denaturing gradient gel electrophoresis (DGGE) to study the diversity of marine picoeukaryotic assemblages and comparison of DGGE with other molecular techniques. Appl. Environ. Microbiol. 2001, 67, 2942–2951. [Google Scholar] [CrossRef] [PubMed]
- González-Toril, E.; Llobet-Brossa, E.; Casamayor, E.O.; Amann, R.; Amils, R. Microbial ecology of an extreme acidic environment, the Tinto River. Appl. Environ. Microbiol. 2003, 69, 4853–4865. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.Y.; He, L.M.; Wu, J.; Jiang, Q. Bacterial community diversity associated with four marine sponges from the South China Sea based on 16S rDNA-DGGE fingerprinting. J. Exp. Mar. Biol. Ecol. 2006, 329, 75–85. [Google Scholar] [CrossRef]
- Brons, J.K.; Van Elsas, J.D. Analysis of bacterial communities in soil by use of denaturing gradient gel electrophoresis and clone libraries, as influenced by different reverse primers. Appl. Environ. Microbiol. 2008, 74, 2717–2727. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.H.; Crowley, D.E.; Borneman, J.; Keen, N.T. Microbial phyllosphere populations are more complex than previously realized. Proc. Natl. Acad. Sci. USA 2001, 98, 3889–3894. [Google Scholar] [CrossRef] [PubMed]
- Dik, A.J.; van Pelt, J.A. Interaction between phyllosphere yeasts, aphid honeydew and fungicide effectiveness in wheat under field conditions. Plant Pathol. 1992, 41, 661–675. [Google Scholar] [CrossRef]
- Stadler, B.; Müller, T. Aphid honeydew and its effect on the phyllosphere microflora of Picea abies (L.) Karst. Oecologia 1996, 108, 771–776. [Google Scholar] [CrossRef] [PubMed]
- Kimmins, J.P. Relative contributions of leaching, litter-fall and defoliation by Neodiprion sertifer (Hymenoptera) to the removal of cesium-134 from red pine. Oikos 1972, 226–234. [Google Scholar] [CrossRef]
- Schowalter, T.D.; Hargrove, W.; Crossley, D.A., Jr. Herbivory in forested ecosystems. Ann. Rev. Entomol. 1986, 31, 177–196. [Google Scholar] [CrossRef]
- Stadler, B.; Müller, T. Effects of aphids and moth caterpillars on epiphytic microorganisms in canopies of forest trees. Can. J. For. Res. 2000, 30, 631–638. [Google Scholar] [CrossRef]
- Müller, T.; Müller, M.; Behrendt, U.; Stadler, B. Diversity of culturable phyllosphere bacteria on beech and oak: The effects of lepidopterous larvae. Microbiol. Res. 2003, 158, 291–297. [Google Scholar] [CrossRef] [PubMed]
- Le Mellec, A.; Habermann, M.; Michalzik, B. Canopy herbivory altering C to N ratios and soil input patterns of different organic matter fractions in a Scots pine forest. Plant Soil 2009, 325, 255–262. [Google Scholar] [CrossRef]
- L-M-Arnold, A.; Grüning, M.; Simon, J.; Reinhardt, A.B.; Lamersdorf, N.; Thies, C. Forest defoliator pests alter carbon and nitrogen cycles. R. Soc. Open Sci. 2016, 3, 160361. [Google Scholar] [CrossRef] [PubMed]
- Stremińska, M.A.; Blaszczyk, M.; Kolk, A. Microbial Abundance and Some of Their Physiological Activities in Soil Organic Horizon of Pine Forest Affected by Insect Herbivory. Pol. J. Environ. Stud. 2006, 15, 905–914. [Google Scholar]
- Kuikka, K.; Härmä, E.; Markkola, A.; Rautio, P.; Roitto, M.; Saikkonen, K.; Ahonen-Jonnarth, U.; Finlay, R.; Tuomi, J. Severe defoliation of Scots pine reduces reproductive investment by ectomycorrhizal symbionts. Ecology 2003, 84, 2051–2061. [Google Scholar] [CrossRef]
- Saravesi, K.; Aikio, S.; Wäli, P.R.; Ruotsalainen, A.L.; Kaukonen, M.; Huusko, K.; Suokas, M.; Brown, S.P.; Jumpponen, A.; Tuomi, J.; et al. Moth outbreaks alter root-associated fungal communities in subarctic mountain birch forests. Microb. Ecol. 2015, 69, 788–797. [Google Scholar] [CrossRef] [PubMed]
- Ferrenberg, S.; Knelman, J.E.; Jones, J.M.; Beals, S.C.; Bowman, W.D.; Nemergut, D.R. Soil bacterial community structure remains stable over a 5-year chronosequence of insect-induced tree mortality. Front. Microbiol. 2014, 5, 681. [Google Scholar] [CrossRef] [PubMed]
- Štursová, M.; Šnajdr, J.; Cajthaml, T.; Bárta, J.; Šantrůčková, H.; Baldrian, P. When the forest dies: The response of forest soil fungi to a bark beetle-induced tree dieback. ISME J. 2014, 8, 1920–1931. [Google Scholar] [CrossRef]
- Treu, R.; Karst, J.; Randall, M.; Pec, G.J.; Cigan, P.W.; Simard, S.W.; Cooke, J.E.K.; Erbilgin, N.; Cahill, J.F. Decline of ectomycorrhizal fungi following a mountain pine beetle epidemic. Ecology 2014, 95, 1096–1103. [Google Scholar] [CrossRef] [PubMed]
- Menkis, A.; Marciulynas, A.; Gedminas, A.; Lynikien, J. High-throughput sequencing reveals drastic changes in fungal communities in the phyllosphere of Norway spruce (Picea abies) following invasion of the spruce bud scale (Physokermes piceae). Microb. Ecol. 2015, 70, 904. [Google Scholar] [CrossRef] [PubMed]
- Abdelfattah, A.; Wisniewski, M.; Nicosia, M.G.L.D.; Cacciola, S.O.; Schena, L. Metagenomic analysis of fungal diversity on strawberry plants and the effect of management practices on the fungal community structure of aerial organs. PLoS ONE 2016, 11, e0160470. [Google Scholar] [CrossRef] [PubMed]
- Pec, G.J.; Karst, J.; Taylor, D.L.; Cigan, P.W.; Erbilgin, N.; Cooke, J.E.; Simard, S.W.; Cahill, J.F. Change in soil fungal community structure driven by a decline in ectomycorrhizal fungi following a mountain pine beetle (Dendroctonus ponderosae) outbreak. New Phytol. 2017, 213, 864–873. [Google Scholar] [CrossRef] [PubMed]
- Mikkelson, K.M.; Lozupone, C.A.; Sharp, J.O. Altered edaphic parameters couple to shifts in terrestrial bacterial community structure associated with insect-induced tree mortality. Soil Biol. Biochem. 2016, 95, 19–29. [Google Scholar] [CrossRef]
- Mikkelson, K.M.; Bokman, C.M.; Sharp, J.O. Rare taxa maintain microbial diversity and contribute to terrestrial community dynamics throughout bark beetle infestation. Appl. Environ. Microbiol. 2016, 82, 6912–6919. [Google Scholar] [CrossRef] [PubMed]
- Brouillard, B.M.; Mikkelson, K.M.; Bokman, C.M.; Berryman, E.M.; Sharp, J.O. Extent of localized tree mortality influences soil biogeochemical response in a beetle-infested coniferous forest. Soil Biol. Biochem. 2017, 114, 309–318. [Google Scholar] [CrossRef]
- Brandfass, C.; Karlovsky, P. Upscaled CTAB-based DNA extraction and real-time PCR assays for Fusarium culmorum and F. graminearum DNA in plant material with reduced sampling error. Int. J. Mol. Sci. 2008, 9, 2306–2321. [Google Scholar] [CrossRef] [PubMed]
- Sambrook, J.; Fritsch, E.F.; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed.; Cold Spring Harbor Laboratory Press: New York, NY, USA, 1989. [Google Scholar]
- Vainio, E.J.; Hantula, J. Direct analysis of wood-inhabiting fungi using denaturing gradient gel electrophoresis of amplified ribosomal DNA. Mycol. Res. 2000, 104, 927–936. [Google Scholar] [CrossRef]
- SILVER SEQUENCE™ DNA Sequencing System. Available online: https://www.promega.de/-/media/files/resources/protocols/technical-manuals/0/silver-sequence-dna-sequencing-system-protocol.pdf (accessed on 8 July 2017).
- Voříšková, J.; Brabcová, V.; Cajthaml, T.; Baldrian, P. Seasonal dynamics of fungal communities in a temperate oak forest soil. New Phytol. 2014, 201, 269–278. [Google Scholar] [CrossRef] [PubMed]
- Basic Local Alignment Search Tool. Available online: https://blast.ncbi.nlm.nih.gov/Blast.cgi (accessed on 8 July 2017).
- Fromin, N.; Hamelin, J.; Tarnawski, S.; Roesti, D.; Jourdain-Miserez, K.; Forestier, N.; Teyssier-Cuvelle, S.; Gillet, F.; Aragna, M.; Rossi, P. Statistical analysis of denaturing gel electrophoresis (DGE) fingerprinting patterns. Environ. Microbiol. 2002, 4, 634–643. [Google Scholar] [CrossRef] [PubMed]
- Fry, J.C.; Webster, G.; Cragg, B.A.; Weightman, A.J.; Parkes, R.J. Analysis of DGGE profiles to explore the relationship between prokaryotic community composition and biogeochemical processes in deep subseafloor sediments from the Peru Margin. FEMS Microbiol. Ecol. 2006, 58, 86–98. [Google Scholar] [CrossRef] [PubMed]
- Oksanen, J.; Blanchet, F.G.; Kindt, R.; Legendre, P.; Minchin, P.R.; O’hara, R.B.; Simspons, G.L.; Solymos, P.; Stevens, M.H.; Szoecs, E.; et al. Community Ecology Package 2013. Version 2.0-4. Available online: https://cran.ism.ac.jp/web/packages/vegan/vegan.pdf (accessed on 8 July 2017).
- Lilley, A.K.; Hails, R.S.; Cory, J.S.; Bailey, M.J. The dispersal and establishment of pseudomonad populations in the phyllosphere of sugar beet by phytophagous caterpillars. FEMS Microbiol. Ecol. 1997, 24, 151–157. [Google Scholar] [CrossRef]
- Stadler, B.; Müller, T.; Orwig, D.; Cobb, R. Hemlock woolly adelgid in New England forests: Canopy impacts transforming ecosystem processes and landscapes. Ecosystems 2005, 8, 233–247. [Google Scholar] [CrossRef]
- Gómez, S.; Orians, C.M.; Preisser, E.L. Exotic herbivores on a shared native host: Tissue quality after individual, simultaneous, and sequential attack. Oecologia 2012, 169, 1015–1024. [Google Scholar] [CrossRef] [PubMed]
- Rubino, L.; Charles, S.; Sirulnik, A.G.; Tuininga, A.R.; Lewis, J.D. Invasive insect effects on nitrogen cycling and host physiology are not tightly linked. Tree Physiol. 2015, 35, 124–133. [Google Scholar] [CrossRef] [PubMed]
- Grüning, M.M.; Simon, J.; Rennenberg, H.; l-M-Arnold, A. Defoliating Insect Mass Outbreak Affects Soil N Fluxes and Tree N Nutrition in Scots Pine Forests. Front. Plant Sci. 2017, 8, 954. [Google Scholar] [CrossRef] [PubMed]
- Vestgarden, L.S. Carbon and nitrogen turnover in the early stage of Scots pine (Pinus sylvestris L.) needle litter decomposition: Effects of internal and external nitrogen. Soil Biol. Biochem. 2001, 33, 465–474. [Google Scholar] [CrossRef]
- Kang, J.C.; Hyde, K.D.; Kong, R.Y. Studies on Amphisphaeriales: The Amphisphaeriaceae (sensu stricto). Mycol. Res. 1999, 103, 53–64. [Google Scholar] [CrossRef]
- Jeewon, R.E.C.Y.; Liew, E.C.Y.; Hyde, K.D. Phylogenetic evaluation of species nomenclature of Pestalotiopsis in relation to host association. Fungal Divers. 2004, 17, 39–55. [Google Scholar]
- Cannon, P.F.; Carmarán, C.C.; Romero, A.I. Studies on biotrophic fungi from Argentina: Microcyclus porlieriae, with a key to South American species of Microcyclus. Mycol. Res. 1995, 99, 353–356. [Google Scholar] [CrossRef]
- Funk, A.; Parker, A.K. Scirrhia pini n. sp., the perfect state of Dothistroma pini Hulbary. Can. J. Bot. 1966, 44, 1171–1176. [Google Scholar] [CrossRef]
- Butin, H. Teleomorph-und anamorph-Entwicklung von Scirrhia pini Funk & Parker auf Nadeln von Pinus nigra Arnold. Sydowia 1985, 38, 20–27. [Google Scholar]
- Dobbeler, P. Ascospore diversity of bryophilous Hypocreales and two new hepaticolous Nectria species. Mycologia 2005, 97, 924–934. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.J.; Geiser, D.M.; Proctor, R.H.; Rooney, A.P.; O’Donnell, K.; Trail, F.; Gardiner, D.M.; Manners, J.M.; Kazan, K. Fusarium pathogenomics. Ann. Rev. Microbiol. 2013, 67, 399–416. [Google Scholar] [CrossRef] [PubMed]
- Berbee, M.L.; Pirseyedi, M.; Hubbard, S. Cochliobolus phylogenetics and the origin of known, highly virulent pathogens, inferred from ITS and glyceraldehyde-3-phosphate dehydrogenase gene sequences. Mycologia 1999, 91, 964–977. [Google Scholar] [CrossRef]
- Zhang, Y.; Schoch, C.L.; Fournier, J.; Crous, P.W.; De Gruyter, J.; Woudenberg, J.H.C.; Hirayama, K.; Tanaka, K.; Pointing, S.B.; Spatafora, J.W.; et al. Multi-locus phylogeny of Pleosporales: A taxonomic, ecological and evolutionary re-evaluation. Stud. Mycol. 2009, 64, 85–102. [Google Scholar] [CrossRef] [PubMed]
- Simon, U.K.; Groenewald, J.Z.; Crous, P.W. Cymadothea trifolii, an obligate biotrophic leaf parasite of Trifolium, belongs to Mycosphaerellaceae as shown by nuclear ribosomal DNA analyses. Persoonia 2009, 22, 49–55. [Google Scholar] [CrossRef] [PubMed]
- Kaur, R.; Shweta, M.M. Onychomycosis due to Rhizomucor in psoriatic patient with HIV infection. Indian J. Dermatol. 2013, 58, 242. [Google Scholar] [CrossRef] [PubMed]
- Battaglia, E.; Visser, L.; Nijssen, A.; van Veluw, G.J.; Wösten, H.A.B.; de Vries, R.P. Analysis of regulation of pentose utilisation in Aspergillus niger reveals evolutionary adaptations in Eurotiales. Stud. Mycol. 2011, 69, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Houbraken, J.; Samson, R.A. Phylogeny of Penicillium and the segregation of Trichocomaceae into three families. Stud. Mycol. 2011, 70, 1–51. [Google Scholar] [CrossRef] [PubMed]
- Nehls, U.; Grunze, N.; Willmann, M.; Reich, M.; Kuester, H. Sugar for my honey: Carbohydrate partitioning in ectomycorrhizal symbiosis. Phytochemistry 2007, 68, 82–91. [Google Scholar] [CrossRef] [PubMed]
- Smith, S.E.; Read, D.J. Mycorrhizal Symbiosis, 3rd ed.; Academic Press: Amsterdam, The Netherlands, 2008. [Google Scholar]
- Gieger, T.; Thomas, F.M. Effects of defoliation and drought stress on biomass partitioning and water relations of Quercus robur and Quercus petraea. Basic Appl. Ecol. 2002, 3, 171–181. [Google Scholar] [CrossRef]
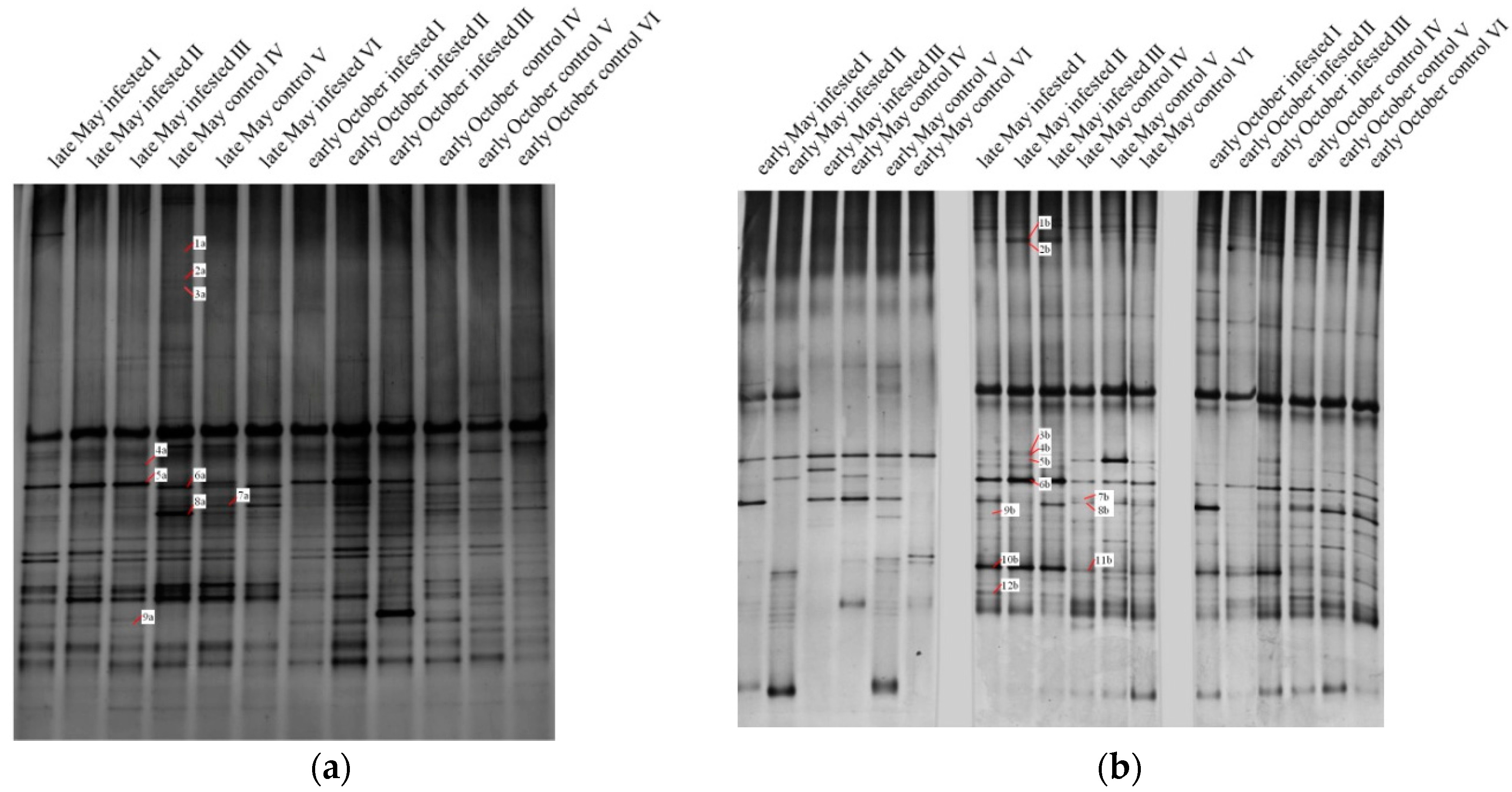
| Band | Phylum | Order | Infested Plots | Control Plots | ||||
|---|---|---|---|---|---|---|---|---|
| I | II | III | IV | V | VI | |||
| 1a | Ascomycota | Capnodiales | − | − | − | + | + | + |
| 2a | Ascomycota | Capnodiales | − | − | − | + | + | + |
| 3a | Ascomycota | Capnodiales | − | − | − | + | + | + |
| 4a | Ascomycota | Dothideales | − | + | + | − | − | − |
| 5a | Ascomycota | Xylariales | +++ | +++ | +++ | + | ++ | − |
| 6a | Ascomycota | Capnodiales | − | − | − | +++ | ++ | +++ |
| 7a | Ascomycota | Dothideales | − | − | + | ++ | ++ | +++ |
| 8a | Ascomycota | Hypocreales | + | + | − | +++ | ++ | + |
| 9a | Ascomycota | Capnodiales | + | + | + | − | − | − |
| Band | Phylum | Order | Infested Plots | Control Plots | ||||
|---|---|---|---|---|---|---|---|---|
| I | II | III | IV | V | VI | |||
| 1b | Ascomycota | Capnodiales | + | ++ | ++ | + | + | + |
| 2b | Ascomycota | Capnodiales | + | ++ | ++ | + | + | + |
| 3b | Ascomycota | Capnodiales | + | + | + | − | − | − |
| 4b | Ascomycota | Capnodiales | + | + | + | − | − | − |
| 5b | Ascomycota | Capnodiales | + | + | + | ++ | +++ | ++ |
| 6b | Ascomycota | Xylariales | +++ | +++ | +++ | ++ | ++ | ++ |
| 7b | Ascomycota | Capnodiales | + | + | + | + | + | + |
| 8b | Ascomycota | Capnodiales | ++ | − | − | + | ++ | + |
| 9b | Ascomycota | Pleosporales | + | + | + | − | − | − |
| 10b | Ascomycota | Pleosporales | +++ | +++ | +++ | − | − | − |
| 11b | Ascomycota | Capnodiales | − | − | − | + | + | + |
| 12b | Ascomycota | Capnodiales | + | + | + | − | − | − |
| Band | Phylum | Family | Infested Plots | Control Plots | ||||
|---|---|---|---|---|---|---|---|---|
| I | II | III | IV | V | VI | |||
| 1a | Zygomycota | Mucoraceae | − | − | ++ | ++ | ++ | +++ |
| 2a | Zygomycota | Mucoraceae | − | − | +++ | +++ | +++ | +++ |
| 3a | Zygomycota | Mucoraceae | + | + | ++ | ++ | ++ | ++ |
| 4a | Zygomycota | Mucoraceae | − | − | +++ | +++ | +++ | +++ |
| 5a | Ascomycota | Trichocomaceae | − | − | − | − | ++ | + |
| Band | Phylum | Genus | Infested Plots | Control Plots | ||||
|---|---|---|---|---|---|---|---|---|
| I | II | III | IV | V | VI | |||
| 1b | Basidiomycota | Russula | − | + | − | +++ | +++ | +++ |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Beule, L.; Grüning, M.M.; Karlovsky, P.; L-M-Arnold, A. Changes of Scots Pine Phyllosphere and Soil Fungal Communities during Outbreaks of Defoliating Insects. Forests 2017, 8, 316. https://doi.org/10.3390/f8090316
Beule L, Grüning MM, Karlovsky P, L-M-Arnold A. Changes of Scots Pine Phyllosphere and Soil Fungal Communities during Outbreaks of Defoliating Insects. Forests. 2017; 8(9):316. https://doi.org/10.3390/f8090316
Chicago/Turabian StyleBeule, Lukas, Maren Marine Grüning, Petr Karlovsky, and Anne L-M-Arnold. 2017. "Changes of Scots Pine Phyllosphere and Soil Fungal Communities during Outbreaks of Defoliating Insects" Forests 8, no. 9: 316. https://doi.org/10.3390/f8090316





