Chemical Screening Method for the Rapid Identification of Microbial Sources of Marine Invertebrate-Associated Metabolites
Abstract
:1. Introduction
2. Results and Discussion
2.1. UPLC/MS Library of EC Secondary Metabolites
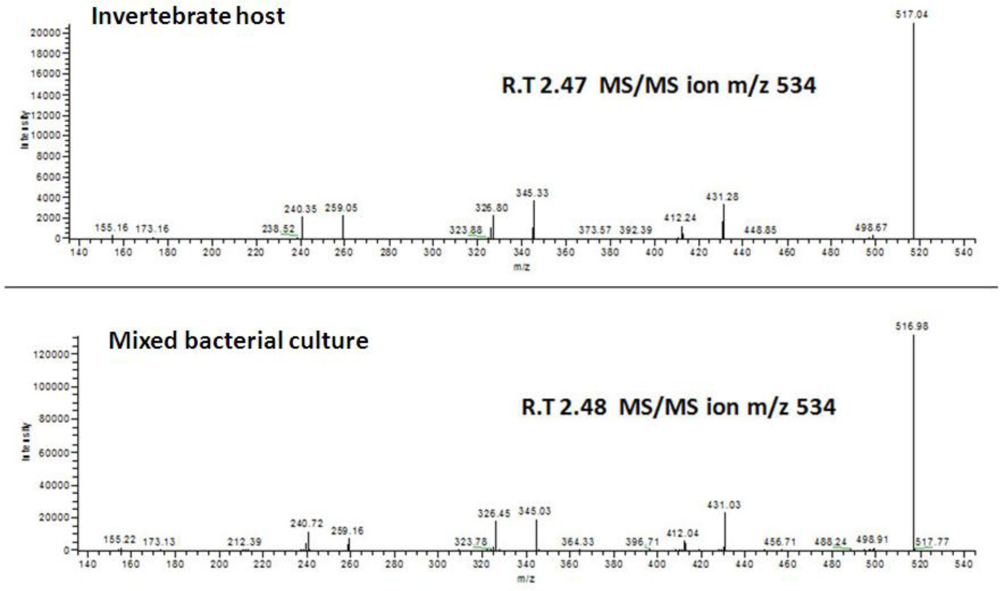
2.2. Screening Mixed Cultures for “EC Metabolites”
2.3. Isolation and Identification of Pure Cultures Producing Target Compounds 41, 47 and 53
2.4. Structure Elucidation of Compounds 41, 47 and 53
2.5. Media Optimization
2.6. Taxonomic Identification of Bacteria Producing 3-Hydroxybutyric Acid Oligomers
3. Conclusions
4. Experimental Section
4.1. Identification of Secondary Metabolites Present in EC
4.2. Chemical Screening of MWP Mixed Cultures
4.3. Chemical Screening of Axenic MWP Cultures
4.4. TB Media Optimization Study
4.5. Structure Determination
4.6. 16S rDNA Sequencing
Acknowledgments
References
- Newman, DJ; Cragg, GM; Snader, KM. Natural Products as Sources of New Drugs over the Period 1981–2002. J. Nat. Prod 2003, 66, 1022–1037. [Google Scholar]
- Newman, DJ; Cragg, GM. Natural Products as Sources of New Drugs over the Last 25 Years. J Nat Prod 2007, 70, 461–477. [Google Scholar]
- Berrue, F; Kerr, RG. Diterpenes from Gorgonian Corals. Nat Prod Rep 2009, 26, 681–710. [Google Scholar]
- Blunt, JW; Copp, BR; Munro, MHG; Northcote, PT; Prinsep, MR. Marine Natural Products. Nat Prod Rep 2010, 27, 165–237. [Google Scholar]
- Marris, E. Marine Natural Products: Drugs from the Deep. Nature 2006, 443, 904–905. [Google Scholar]
- Simmons, TL; Andrianasolo, E; McPhail, K; Flatt, P; Gerwick, WH. Marine Natural Products as Anticancer Drugs. Mol Cancer Ther 2005, 4, 333–342. [Google Scholar]
- Simmons, TL; Coates, RC; Clark, BR; Engene, N; Gonzalez, D; Esquenazi, E; Dorrestein, PC; Gerwick, WH. Biosynthetic Origin of Natural Products Isolated from Marine Microorganism-Invertebrate Assemblages. Proc Natl Acad Sci USA 2008, 105, 4587–4594. [Google Scholar]
- Molinski, TF; Dalisay, DS; Lievens, SL; Saludes, JP. Drug Development from Marine Natural Products. Nat Rev Drug Discov 2009, 8, 69–85. [Google Scholar]
- Piel, J. Bacterial Symbionts: Prospects for the Sustainable Production of Invertebrate-Derived Pharmaceuticals. Curr Med Chem 2006, 13, 39–50. [Google Scholar]
- Hildebrand, M; Waggoner, LE; Lim, GE; Sharp, KH; Ridley, CP; Haygood, MG. Approaches to Identify, Clone, and Express Symbiont Bioactive Metabolite Genes. Nat Prod Rep 2004, 21, 122–142. [Google Scholar]
- Kennedy, J; Marchesi, JR; Dobson, ADW. Metagenomic Approaches to Exploit the Biotechnological Potential of the Microbial Consortia of Marine Sponges. Appl Microbiol Biotechnol 2007, 75, 11–20. [Google Scholar]
- Oclarit, JM; Okada, H; Ohta, S; Kaminura, K; Yamaoka, Y; Iizuka, T; Miyashiro, S; Ikegami, S. Anti-Bacillus Substance in the Marine Sponge, Hyatella Species, Produced by an Associated Vibrio Species Bacterium. Microbios 1994, 78, 7–16. [Google Scholar]
- Shigemori, H; Bae, MA; Yazawa, K; Sasaki, T; Kobayashi, J. Alteramide A, a New Tetracyclic Alkaloid from a Bacterium Alteromonas Sp. Associated with the Marine Sponge Halichondria Okadai. J Org Chem 1992, 57, 4317–4320. [Google Scholar]
- Stierle, AC; Cardellina, JH, II; Singleton, FL. A Marine Micrococcus Produces Metabolites Ascribed to the Sponge Tedania Ignis. Experientia 1988, 44, 1021. [Google Scholar]
- Duetz, WA. Microtiter Plates as Mini-Bioreactors: Miniaturization of Fermentation Methods. Trends Microbiol 2007, 15, 469–475. [Google Scholar]
- Look, SA; Fenical, W; Van, E; Donna; Clardy, J. Erythrolides: Unique Marine Diterpenoids Interrelated by a Naturally Occurring Di-Ï€-Methane Rearrangement. J Am Chem Soc 1984, 106, 5026–5027. [Google Scholar]
- Fenical, W; Pawlik, JR. Defensive Properties of Secondary Metabolites from the Caribbean Gorgonian Coral Erythropodium Caribaeorum. Mar Ecol Prog Ser 1991, 75, 1–8. [Google Scholar]
- Dookran, R; Maharaj, D; Mootoo, BS; Ramsewak, R; McLean, S; Reynolds, WF; Tinto, WF. Diterpenes from the Gorgonian Coral Erythropodium Caribaeorum from the Southern Caribbean. J Nat Prod 1993, 56, 1051–1056. [Google Scholar]
- Cinel, B; Roberge, M; Behrisch, H; van Ofwegen, L; Castro, CB; Andersen, RJ. Antimitotic Diterpenes from Erythropodium Caribaeorum Test Pharmacophore Models for Microtubule Stabilization. Org Lett 2000, 2, 257–260. [Google Scholar]
- Britton, R; Roberge, M; Berisch, H; Andersen, RJ. Antimitotic Diterpenoids from Erythropodium Caribaeorum: Isolation Artifacts and Putative Biosynthetic Intermediates. Tetrahedron Lett 2001, 42, 2953–2956. [Google Scholar]
- Plumb, R; Castro-Perez, J; Granger, J; Beattie, I; Joncour, K; Wright, A. Ultra-Performance Liquid Chromatography Coupled to Quadrupole-Orthogonal Time-of-Flight Mass Spectrometry. Rapid Commun Mass Spectrom 2004, 18, 2331–2337. [Google Scholar]
- Toyo’oka, T. Determination Methods for Biologically Active Compounds by Ultra-Performance Liquid Chromatography Coupled with Mass Spectrometry: Application to the Analyses of Pharmaceuticals, Foods, Plants, Environments, Metabonomics, and Metabolomics. J Chromatogr Sci 2008, 46, 233–247. [Google Scholar]
- Yao, S; Wu, T; Li, X; Tu, B; Song, H. Ten Years of Research into Phytomedicines Analysis—An Era in New Technologies and Methods. Curr Pharm Anal 2010, 6, 269–288. [Google Scholar]
- Altschul, SF; Madden, TL; Schaffer, AA; Zhang, J; Zhang, Z; Miller, W; Lipman, DJ. Gapped BLAST and PSI-BLAST: A New Generation of Protein Database Search Programs. Nucleic Acids Res 1997, 25, 3389–3402. [Google Scholar]
- Sun, W; Teng, K; Meighen, E. Detection of Poly(3-Hydroxybutyrate) Granules by Electron Microscopy of Vibrio Harveyi Stained with Malachite Green. Can J Microbiol 1995, 41, 131–137. [Google Scholar]
- Kimura, B; Hokimoto, S; Takahashi, H; Fujii, T. Photobacterium Histaminum Okuzumi et al. 1994 is a Later Subjective Synonym of Photobacterium Damselae Subsp. Damselae (Love et al. 1981) Smith et al. 1991. Int J Syst Evol Microbiol 2000, 50, 1339–1342. [Google Scholar]
- Smith, SK; Sutton, DC; Fuerst, JA; Reichelt, JL. Evaluation of the Genus Listonella and Reassignment of Listonella Damsela (Love et al.) MacDonell and Colwell to the Genus Photobacterium as Photobacterium Damsela Comb. Nov. with an Emended Description. Int J Syst Bacteriol 1991, 41, 529–534. [Google Scholar]
- Lee, SY. Plastic Bacteria? Progress and Prospects for Polyhydroxyalkanoate Production in Bacteria. Trends Biotechnol 1996, 14, 431–438. [Google Scholar]
- Zinn, M; Witholt, B; Egli, T. Occurrence, Synthesis and Medical Application of Bacterial Polyhydroxyalkanoate. Adv Drug Delivery Rev 2001, 53, 5–21. [Google Scholar]
- Williams, P. Quorum Sensing, Communication and Cross-Kingdom Signalling in the Bacterial World. Microbiology 2007, 153, 3923–3938. [Google Scholar]
- Angell, S; Bench, BJ; Williams, H; Watanabe Coran, MH. Pyocyanin Isolated from a Marine Microbial Population: Synergistic Production between Two Distinct Bacterial Species and Mode of Action. Chem Biol 2006, 13, 1349–1359. [Google Scholar]
- Medsker, LL; Jenkins, D; Thomas, JF; Koch, C. Odorous Compounds in Natural Waters. 2-Exo-Hydroxy-2-Methylbornane, the Major Odorous Compound Produced by several Actinomycetes. Environ Sci Technol 1969, 3, 476–477. [Google Scholar]
- Reasoner, DJ; Geldreich, EE. A New Medium for the Enumeration and Subculture of Bacteria from Potable Water. Appl Environ Microbiol 1985, 49, 1–7. [Google Scholar]
- Shirling, EB; Gottlieb, D. Methods for Characterization of Streptomyces Species. Int J Syst Bacteriol 1966, 16, 313–340. [Google Scholar]
- Sambrook, J; Fritsch, EF; Maniatis, T. Molecular Cloning: A Laboratory Manual, 2nd ed; Cold Spring Harbor Laboratory: Cold Spring Harbor, NY, USA, 1989. [Google Scholar]
- ZoBell, CE. Studies on Marine Bacteria. I. the Cultural Requirements of Heterotrophic Aerobes. J Mar Res 1941, 4, 42–75. [Google Scholar]
- Marshall, RT. Standard Methods for the Examination of Dairy Products, 16th ed; American Public Health Association: Washington, D.C., USA, 1993. [Google Scholar]
- Baker, GC; Smith, JJ; Cowan, DA. Review and Re-Analysis of Domain-Specific 16S Primers. J Microbiol Methods 2003, 55, 541–555. [Google Scholar]

| No Antibiotics | +Penicillin G | +Kanamycin | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 2 | 3 | 4 | 5 | 6 | 7 | 8 | 9 | 10 | 11 | 12 | |
| A | NB | NB pH 5.8 | 1/10 NB pH 5.8 | MB pH 5.8 | NB | NB pH 5.8 | 1/10 NB pH 5.8 | MB pH 5.8 | NB | NB pH 5.8 | 1/10 NB pH 5.8 | MB pH 5.8 |
| B | LB | NB pH6.5 | 1/10 NB pH6.5 | MB pH6.5 | LB | NB pH6.5 | 1/10 NB pH6.5 | MB pH6.5 | LB | NB pH6.5 | 1/10 NB pH6.5 | MB pH6.5 |
| C | MB | NB pH 7.2 | 1/10 NB pH 7.2 | MB pH 7.2 | MB | NB pH 7.2 | 1/10 NB pH 7.2 | MB pH 7.2 | MB | NB pH 7.2 | 1/10 NB pH 7.2 | MB pH 7.2 |
| D | TB | NB pH 8.0 | 1/10 NB pH 8.0 | MB pH 8.0 | TB | NB pH 8.0 | 1/10 NB pH 8.0 | MB pH 8.0 | TB | NB pH 8.0 | 1/10 NB pH 8.0 | MB pH 8.0 |
| E | dH20 | ISP1 10μg/mL NA | NB + 0.5% butanol | 1/10 MB pH 5.8 | dH20 | ISP1 10μg/mL NA | NB + 0.5% butanol | 1/10 MB pH 5.8 | dH20 | ISP1 10μg/mL NA | NB + 0.5% butanol | 1/10 MB pH 5.8 |
| F | M9 glycerol | NB/SW | ASW | 1/10 MB pH6.5 | M9 glycerol | NB/SW | ASW | 1/10 MB pH6.5 | M9 glycerol | NB/SW | ASW | 1/10 MB pH6.5 |
| G | M9 glucose | Emerson | ASW glycerol | 1/10 MB pH 7.2 | M9 glucose | Emerson | ASW glycerol | 1/10 MB pH 7.2 | M9 glucose | Emerson | ASW glycerol | 1/10 MB pH 7.2 |
| H | M9 arabinose | R2 Broth | ASW glucose | 1/10 MB pH 8.0 | M9 arabinose | R2 Broth | ASW glucose | 1/10 MB pH 8.0 | M9 arabinose | R2 Broth | ASW glucose | 1/10 MB pH 8.0 |
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Berrue, F.; Withers, S.T.; Haltli, B.; Withers, J.; Kerr, R.G. Chemical Screening Method for the Rapid Identification of Microbial Sources of Marine Invertebrate-Associated Metabolites. Mar. Drugs 2011, 9, 369-381. https://doi.org/10.3390/md9030369
Berrue F, Withers ST, Haltli B, Withers J, Kerr RG. Chemical Screening Method for the Rapid Identification of Microbial Sources of Marine Invertebrate-Associated Metabolites. Marine Drugs. 2011; 9(3):369-381. https://doi.org/10.3390/md9030369
Chicago/Turabian StyleBerrue, Fabrice, Sydnor T. Withers, Brad Haltli, Jo Withers, and Russell G. Kerr. 2011. "Chemical Screening Method for the Rapid Identification of Microbial Sources of Marine Invertebrate-Associated Metabolites" Marine Drugs 9, no. 3: 369-381. https://doi.org/10.3390/md9030369
APA StyleBerrue, F., Withers, S. T., Haltli, B., Withers, J., & Kerr, R. G. (2011). Chemical Screening Method for the Rapid Identification of Microbial Sources of Marine Invertebrate-Associated Metabolites. Marine Drugs, 9(3), 369-381. https://doi.org/10.3390/md9030369





