Isolation and Characterization of a Mn(II)-Oxidizing Bacillus Strain from the Demosponge Suberites domuncula
Abstract
:1. Introduction
2. Results
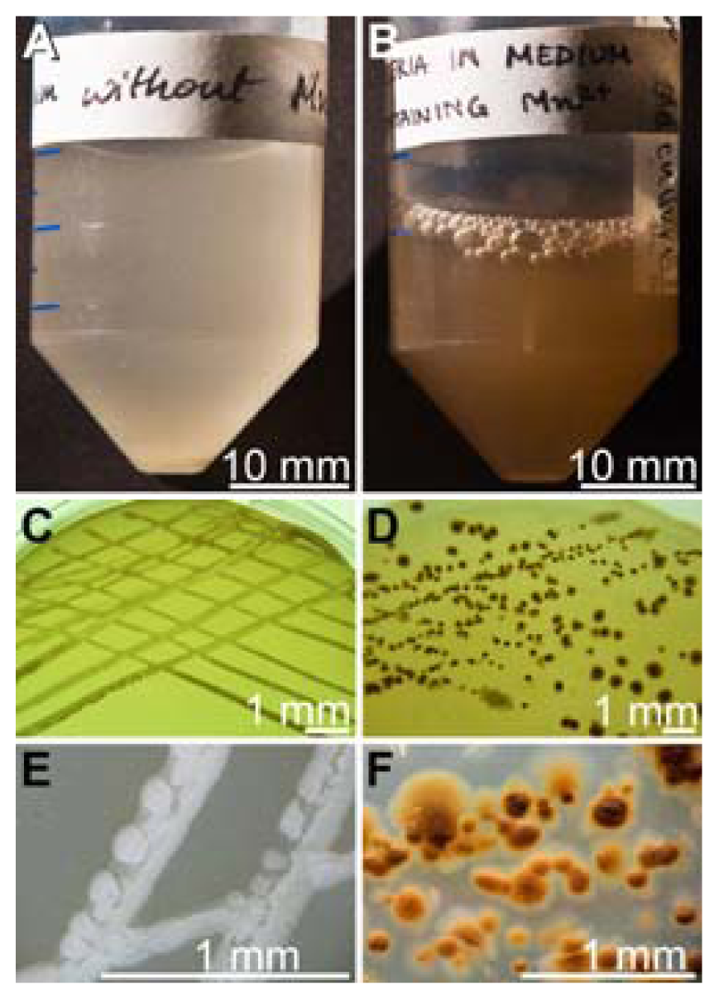
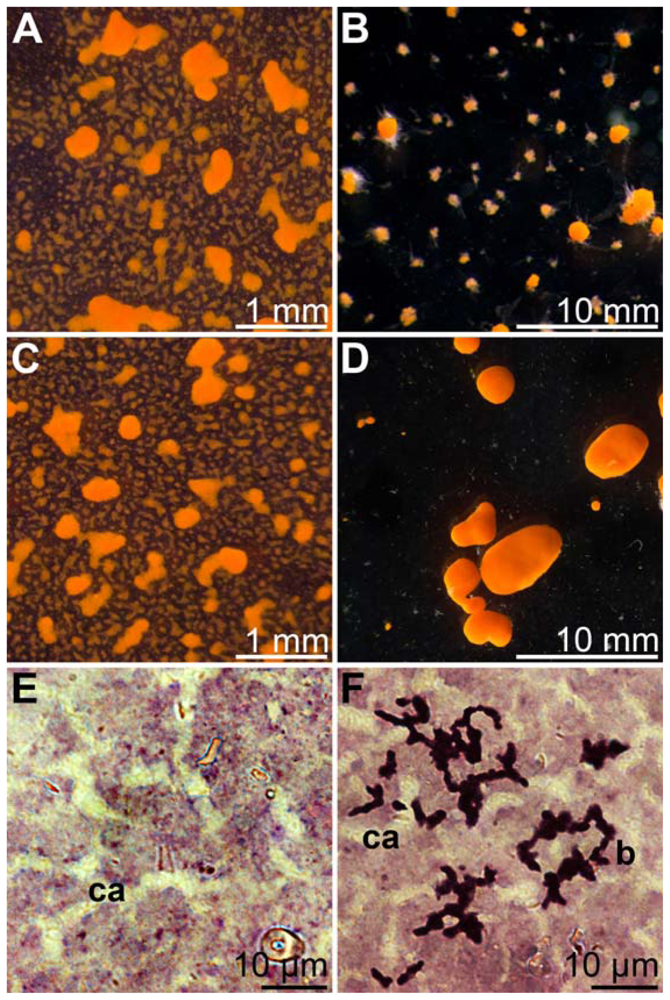
2.1. Isolation of Mn(II)-Oxidizing Bacillus Strain BAC-SubDo-03
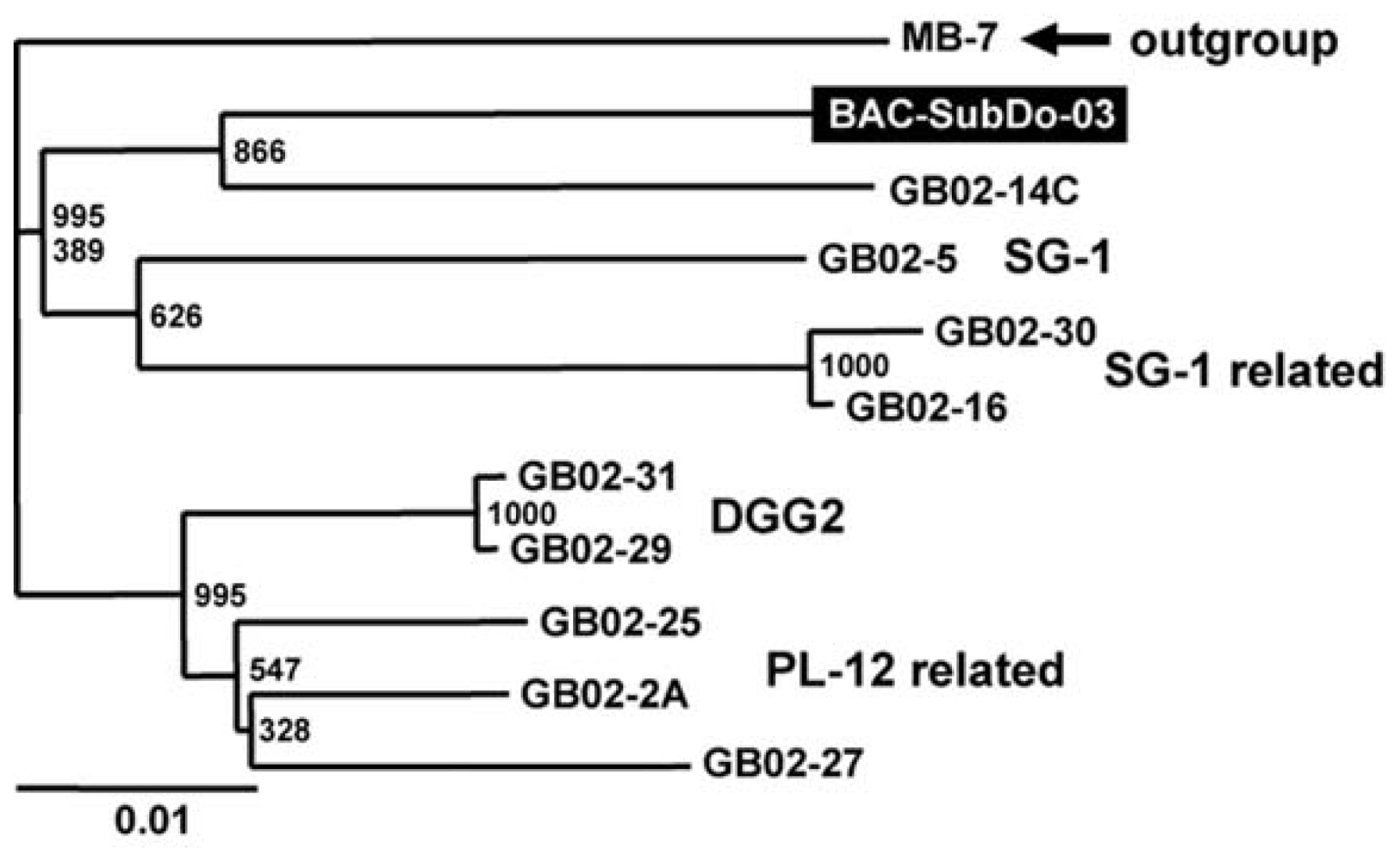
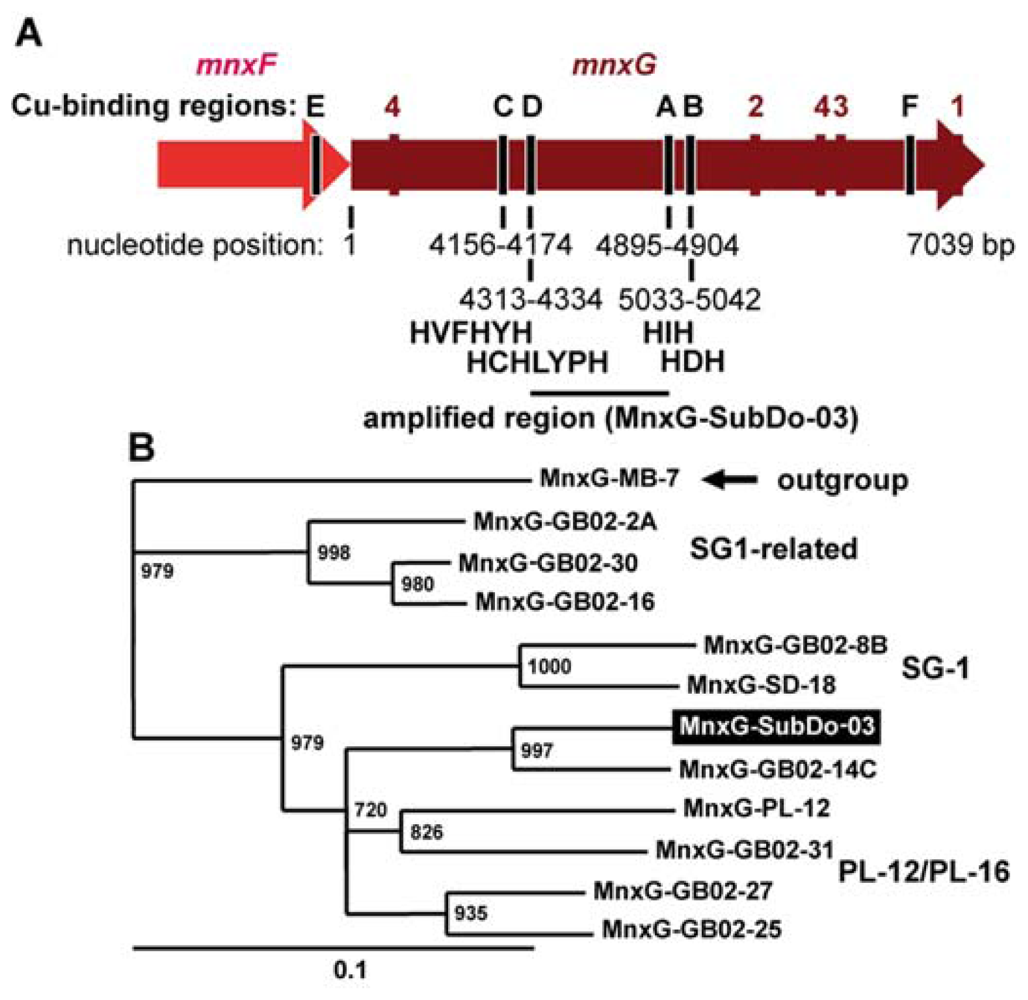
2.2. Phylogenetic Grouping of BAC-SubDo-03
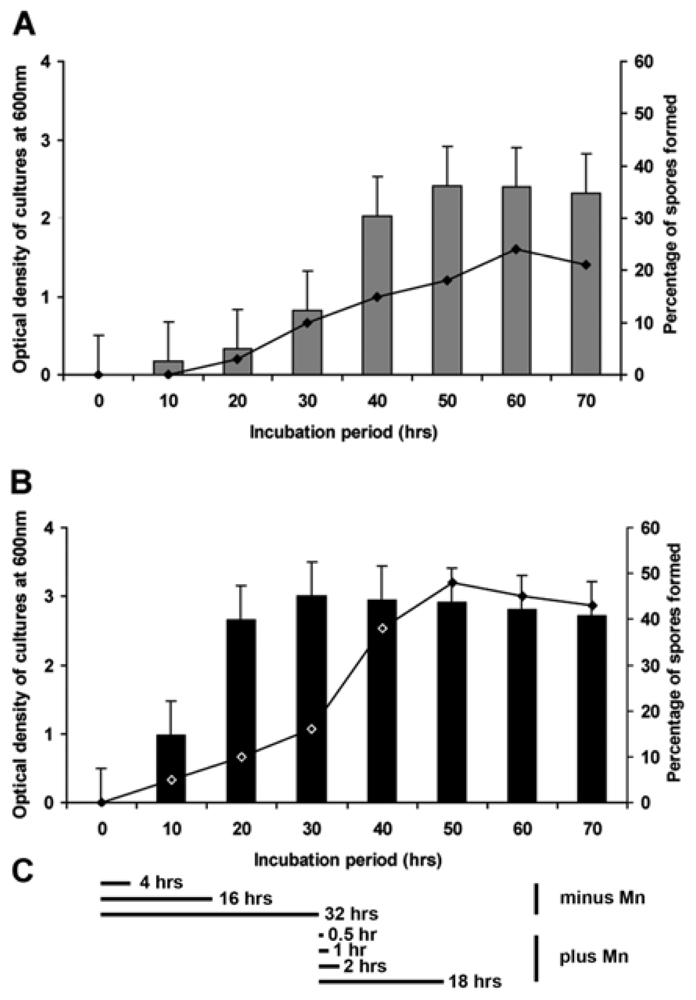
2.3. Formation of Spores
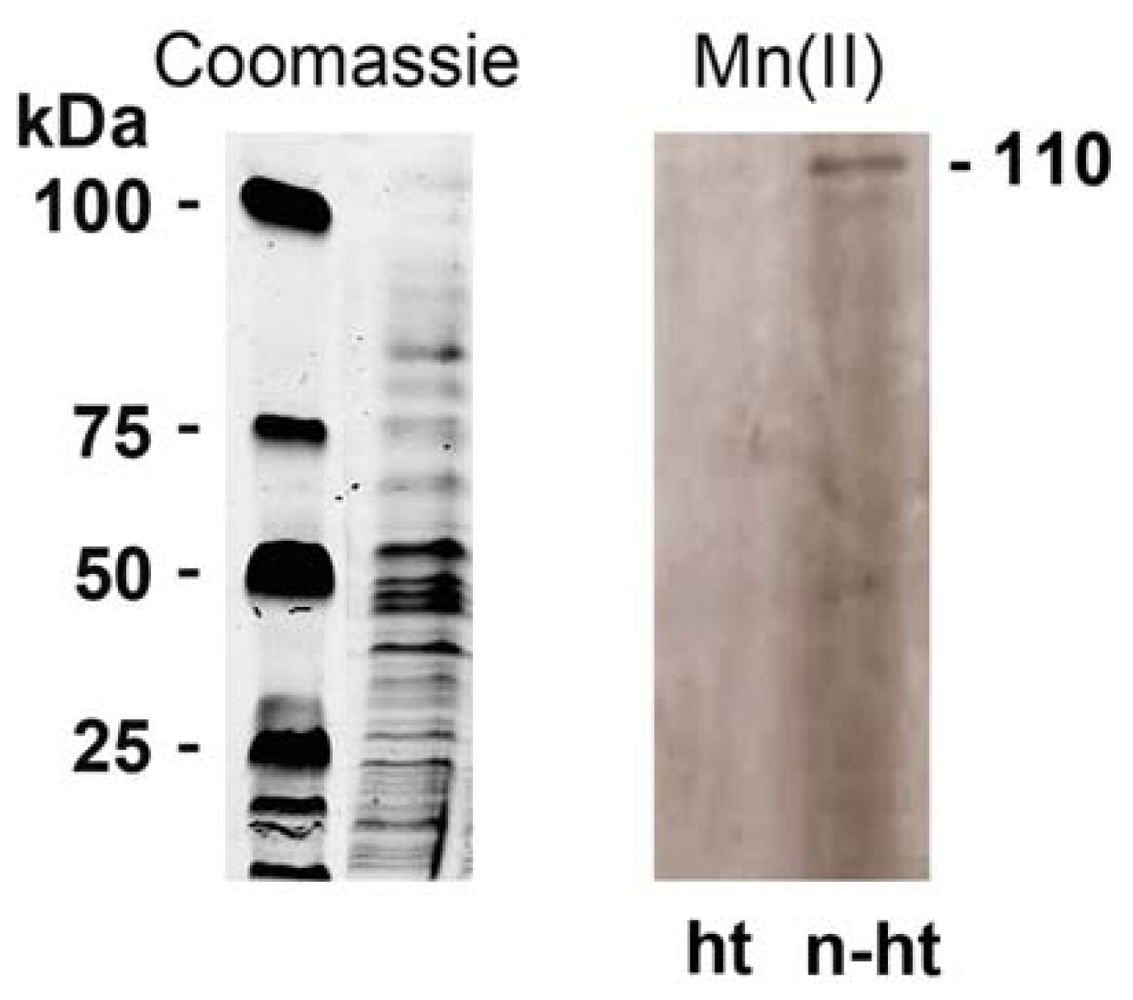
2.4. Detection of Mn-Oxidizing Activity: In-Gel Determination
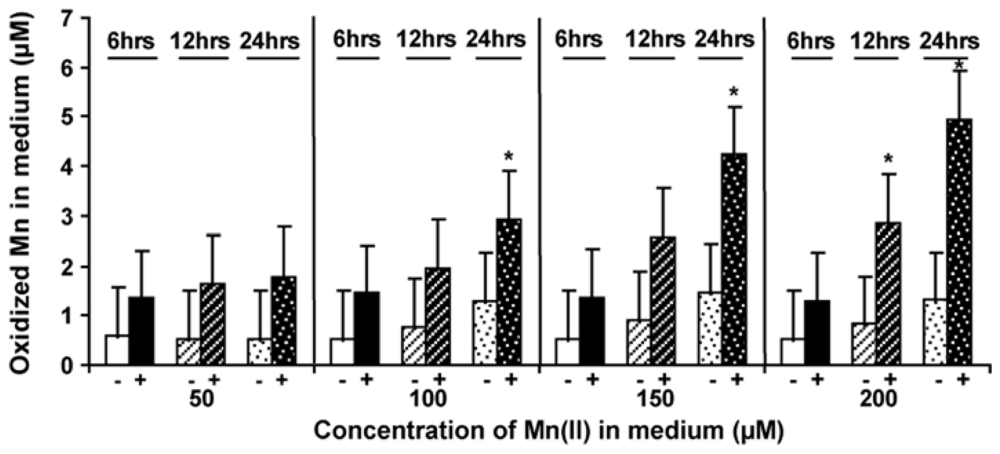
2.5. Level of Mn-Oxidizing Activity Depending on the Presence of Mn in the Medium
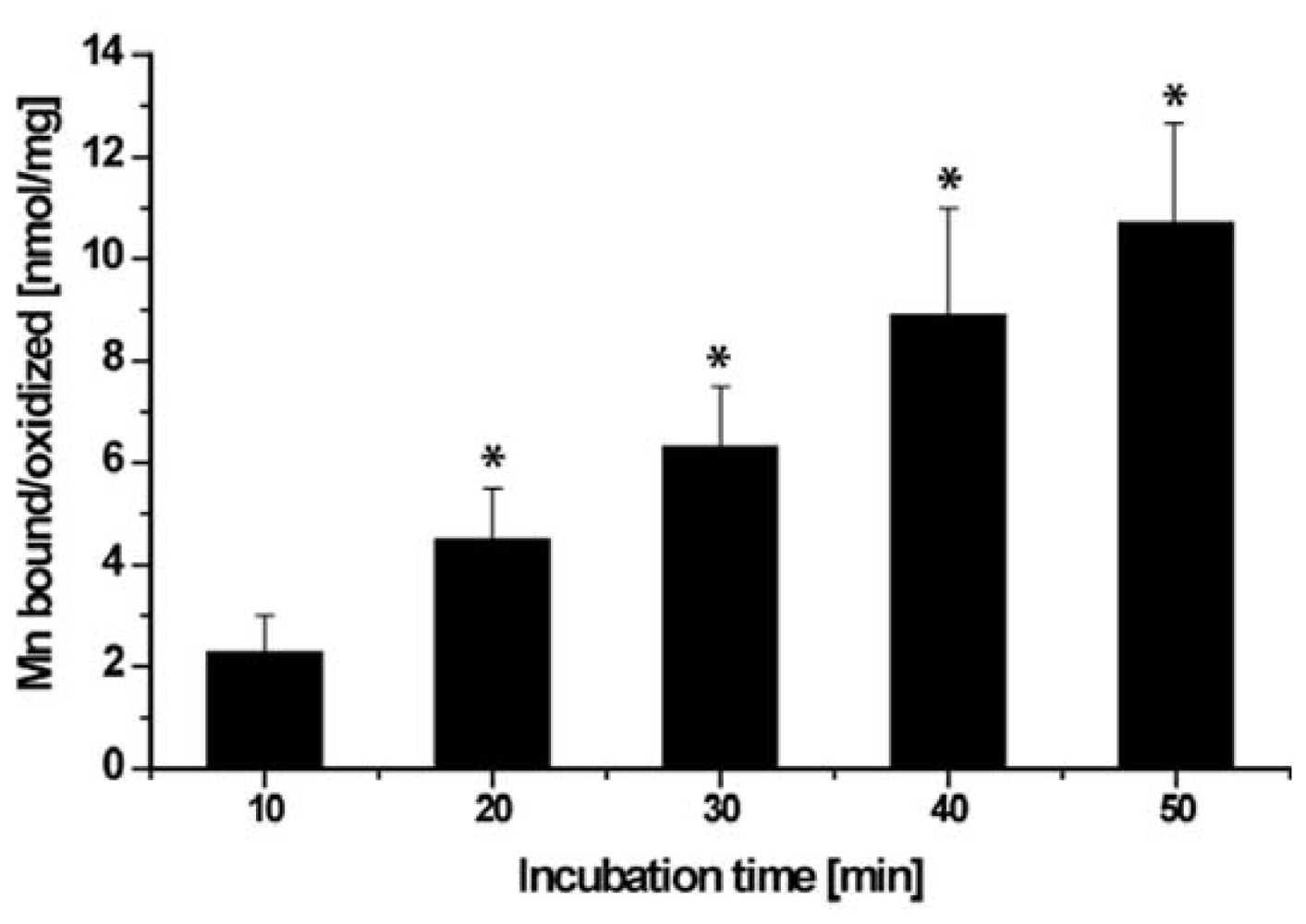
2.6. Binding/Oxidation of Manganese by the BAC-SubDo-03 Strain
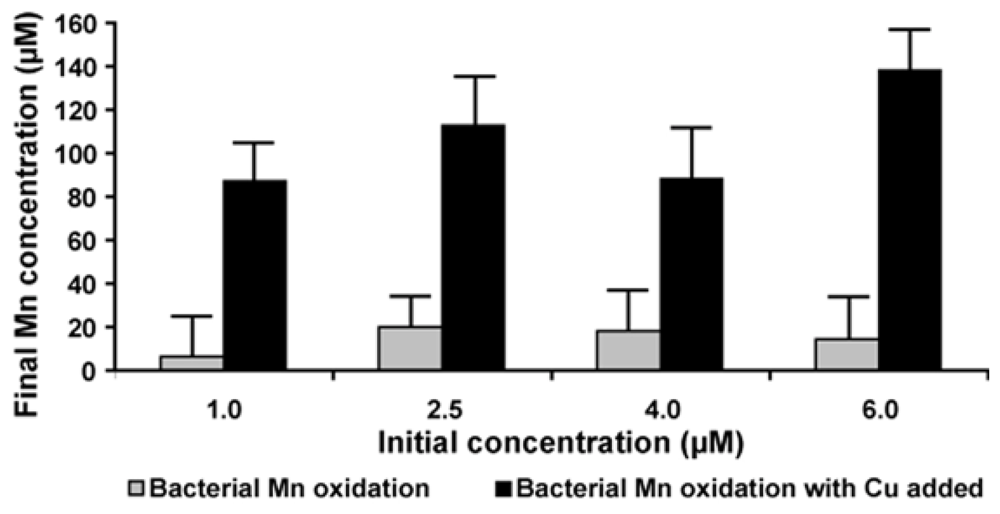
2.7. Effect of Copper on Manganese Oxidation
2.8. mnxG Gene in BAC-SubDo-3
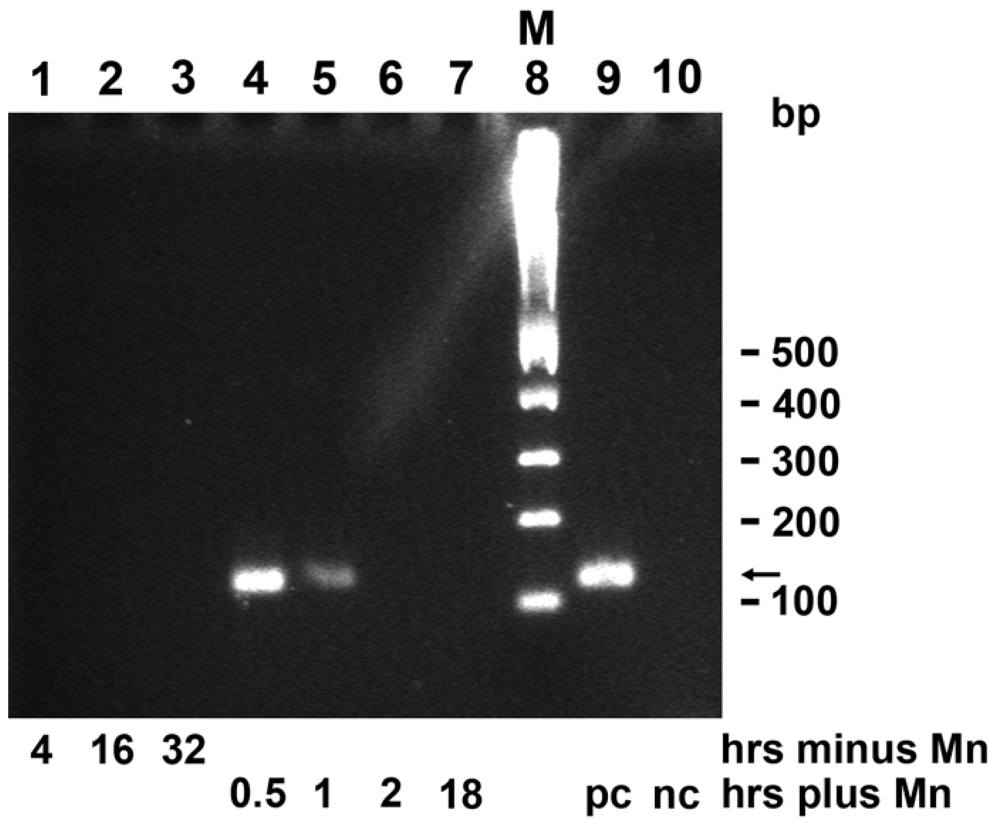
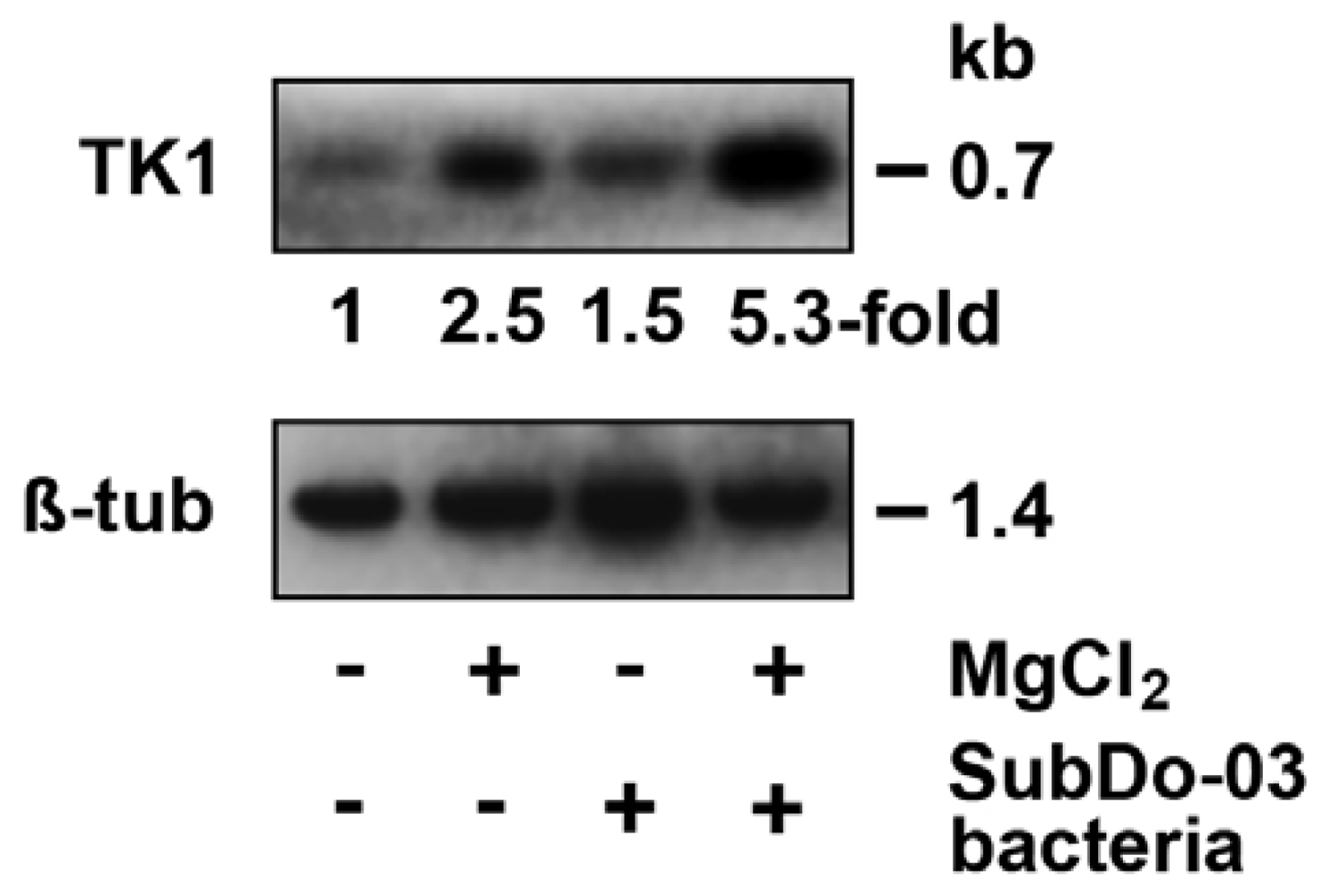
2.9. Induction of mnxG Gene after Incubation with MnCl2
2.10. Co-Incubation of S. domuncula Primmorphs with BAC-SubDo-03
2.11. Gene Expression Studies in Primmorphs
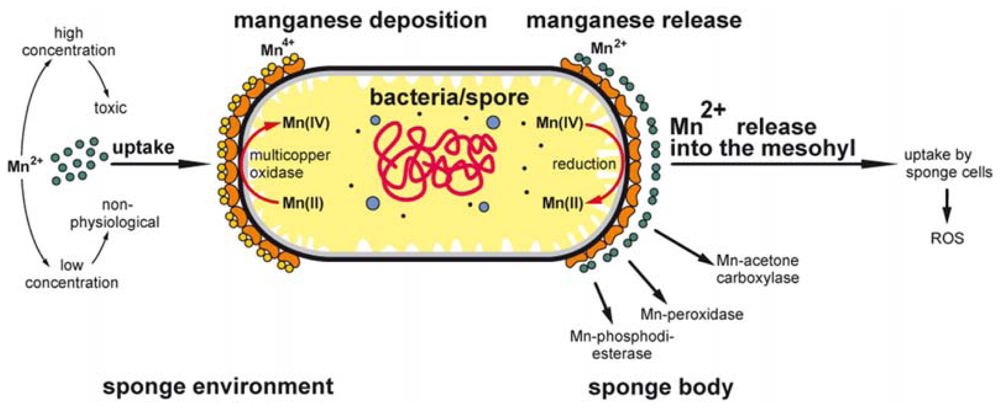
3. Discussion
4. Experimental Section
4.1. Materials
4.2. Isolation and Cultivation of Manganese Oxidizing Bacteria from S. domuncula
4.3. Isolation and Purification of Spores from the Bacterial Cultures
4.4. Detection of Mn(II)-Oxidizing Activity in Spores
4.5. Determination of Binding/Oxidation Capacity of BAC-SubDo-03 Strain for Manganese
4.6. Cloning and Sequencing of Bacterial 16S rDNA by PCR
4.7. Cloning and Sequencing of Multicopper Oxidase mnxG Gene by PCR
4.8. Expression Studies of mnxG-SubDo-03 Gene by RT-PCR
4.9. Data Analysis
4.10. Primmorphs and Co-Incubation with BAC-SubDo-03
4.11. Histological Analysis
4.12. RNA Extraction and Northern Blotting
4.13. Analytical Method
4.14. Statistical Analysis
Acknowledgements
References and Notes
- Glasby, GP. Manganese: predominant role of nodules and crusts. In Marine Geochemistry; 2nd revised, updated and extended ed; Schulz, HD, Zabel, M, Eds.; Springer-Verlag, Berlin: Germany, 2006; pp. 371–427. [Google Scholar]
- Ehrlich, HL. Geomicrobiology; Marcel Dekker: New York, NY, USA, 2002. [Google Scholar]
- Wang, XH; Müller, WEG. Contribution of biomineralization during growth of polymetallic nodules and ferromanganese crusts from the Pacific Ocean. Front Mater Sci China 2009, 3, 109–123. [Google Scholar]
- Wang, XH; Müller, WEG. Marine biominerals: perspectives and challenges for polymetallic nodules and crusts. Trends Biotechnol 2009, 27, 375–383. [Google Scholar]
- Wang, XH; Schröder, HC; Schloßmacher, U; Müller, WEG. Organized bacterial assemblies in manganese nodules: evidence for a role of S-layers in metal deposition. Geo-Mar Lett 2009, 29, 85–91. [Google Scholar]
- Wang, XH; Schröder, HC; Wiens, M; Schloßmacher, U; Müller, WEG. Manganese/polymetallic nodules: Micro-structural characterization of exolithobiontic- and endolithobiontic microbial biofilms by scanning electron microscopy. Micron 2009, 40, 350–358. [Google Scholar]
- Tebo, BM; Bargar, JR; Clement, BG; Dick, GJ; Murray, KJ; Parker, D; Verity, R; Webb, SM. Biogenic manganese oxides: properties and mechanisms of formation. Annu Rev Earth Planetary Sci 2004, 32, 287–328. [Google Scholar]
- Brouwers, G-J; de Vrind, JPM; Corstjens, PLAM; Cornelis, P; Baysse, C; de Vrind-de Jong, EW. cumA, a gene encoding a multicopper oxidase, is involved in Mn2+-oxidation in Pseudomonas putida GB-1. Appl Environ Microbiol 1999, 65, 1762–1768. [Google Scholar]
- Corstjens, PLAM; de Vrind, JPM; Goosen, T; de Vrind-de Jong, EW. Identification and molecular analysis of the Leptothrix discophora SS-1 mofA gene, a gene putatively encoding a manganese-oxidizing protein with copper domains. Geomicrobiol J 1997, 14, 91–108. [Google Scholar]
- van Waasbergen, LG; Hildebrand, M; Tebo, BM. Identification and characterization of a gene cluster involved in manganese oxidation by spores of the marine Bacillus sp. strain SG-1. J Bacteriol 1996, 178, 3517–3530. [Google Scholar]
- Dick, GJ; Lee, YE; Tebo, BM. Manganese(II)-oxidizing Bacillus spores in Guaymas Basin hydrothermal sediments and plumes. Appl Envir Microbiol 2006, 72, 3184–3190. [Google Scholar]
- Dick, GJ; Podell, S; Johnson, HA; Rivera-Espinoza, Y; Bernier-Latmani, R; McCarthy, JK; Torpey, JW; Clement, BG; Gaasterland, T; Tebo, BM. Genomic insights into Mn(II) oxidation by the marine Alphaproteobacterium Aurantimonas sp. strain SI85-9A1. Appl Envir Microbiol 2008, 74, 2646–2658. [Google Scholar]
- Dick, GJ; Torpey, JW; Beveridge, TJ; Tebo, BM. Direct Identification of a bacterial manganese(II) oxidase, the multicopper oxidase MnxG, from spores of several different marine Bacillus species. Appl Environ Microbiol 2008, 74, 1527–1534. [Google Scholar]
- Mayhew, LE; Swanner, ED; Martin, AP; Templeton, AS. Phylogenetic relationships and functional genes: distribution of a gene (mnxG) encoding a putative manganese-oxidizing enzyme in Bacillus species. Appl Environ Microbiol 2008, 74, 7265–7271. [Google Scholar]
- Ridge, JP; Lin, M; Larsen, EI; Fegan, M; McEwan, AG; Sly, LI. A multicopper oxidase is essential for manganese oxidation and laccase-like activity in Pedomicrobium sp. ACM 3067. Environ Microbiol 2007, 9, 944–953. [Google Scholar]
- Solomon, EI; Sundaram, UM; Machonkin, TE. Multicopper oxidases and oxygenases. Chem Rev 1996, 96, 2563–2605. [Google Scholar]
- Anderson, CR; Johnson, HA; Caputo, N; Davis, RE; Torpey, JW; Tebo, BM. Mn(II) oxidation is catalyzed by heme peroxidase in Aurantimonas manganoxydans strain SI85-9A1 and Erythrobacter sp. strain SD-21. Appl Environ Microbiol 2009, 75, 4130–4138. [Google Scholar]
- LePennec, G; Perović, S; Ammar, MSA; Grebenjuk, VA; Steffen, R; Müller, WEG. Cultivation of primmorphs from the marine sponge Suberites domuncula: Morphogenetic potential of silicon and iron. J Biotechnol 2003, 100, 93–108. [Google Scholar]
- Müller, WEG; Wiens, M; Batel, R; Steffen, R; Schröder, HC; Borojevic, R; Custodio, MR. Establishment of a primary cell culture from a sponge: Primmorphs from Suberites domuncula. Mar Ecol Progr Ser 1999, 178, 205–219. [Google Scholar]
- Bargar, JR; Tebo, BM; Villinski, JE. In situ characterization of Mn(II) oxidation by spores of the marine Bacillus sp. strain SG-1. Geochim Cosmochim Acta 2000, 64, 2775–2778. [Google Scholar]
- Rosson, RA; Nealson, KH. Manganese binding and oxidation by spores of a marine Bacillus. J Bacteriol 1982, 151, 1027–1034. [Google Scholar]
- Francis, CA; Tebo, BM. Enzymatic manganese(II) oxidation by metabolically dormant spores of diverse Bacillus species. Appl Environ Microbiol 2002, 68, 874–880. [Google Scholar]
- De Vrind, JPM; de Vrind-de Jong, EW; de Voogt, JWH; Westbroek, P; Boogerd, FC; Rosson, RA. Manganese oxidation by spores and spore coats of a marine bacillus species. Appl Environ Microbiol 1986, 52, 1096–1100. [Google Scholar]
- Müller, W; Zahn, RK; Beyer, R. Is “I-DNA” derived from nuclear DNA. Nature 1970, 227, 1211–1212. [Google Scholar]
- Mertz, W. The scientific and practical importance of trace elements. Phil Trans R Soc Lond B 1981, 294, 9–18. [Google Scholar]
- Böhm, M; Hentschel, U; Friedrich, A; Fieseler, L; Steffen, R; Gamulin, V; Müller, IM; Müller, WEG. Molecular response of the sponge Suberites domuncula to bacterial infection. Mar Biol 2001, 139, 1037–1045. [Google Scholar]
- Müller, WEG; Custódio, M; Wiens, M; Zilberberg, C; Châtel, A; Müller, IM; Schröder, HC. Effect of bacterial infection on stem cell pattern in Porifera. In Stem Cells in Marine Organisms; Rinkevich, B, Matranga, V, Eds.; Springer Press: Heidelberg, Germany, 2009; pp. 309–336. [Google Scholar]
- Segura-Pena, D; Lutz, S; Monnerjahn, C; Konrad, M; Lavie, A. Binding of ATP to TK1-like enzymes is associated with a conformational change in the quaternary structure. J Mol Biol 2007, 369, 129–141. [Google Scholar]
- Ferguson, RE; Carroll, HP; Harris, A; Maher, ER; Selby, PJ; Banks, RE. Housekeeping proteins: a preliminary study illustrating some limitations as useful references in protein expression studies. Proteomics 2005, 5, 566–571. [Google Scholar]
- Fabbri, D; Gabbianelli, G; Locatelli, C; Lubrano, D; Trombini, C; Vassura, I. Distribution of mercury and other heavy metals in core sediments of the Northern Adriatic Sea. Water Air Soil Pollut 2001, 129, 143–153. [Google Scholar]
- Howe, PD; Malcolm, HM; Dobson, S. Manganese and Its Compounds: Environmental Aspects. In Concise International Chemical Assessment Document 63; World Health Organization: Geneva, Switzerland, 2004. [Google Scholar]
- Langille, WM; Maclean, KS. Some essential nutrient elements in forest plants as related to species, plant part, season, and location. Plant Soil 1976, 45, 17–26. [Google Scholar]
- Hakala, M; Rantamäki, S; Puputti, E-M; Tyystjärvi, T; Tyystjärvi, E. Photoinhibition of manganese enzymes: insights into the mechanism of photosystem II photoinhibition. J Exp Bot 2006, 57, 1809–1816. [Google Scholar]
- Andreassen, OA; Ferrante, RJ; Dedeoglu, A; Albers, DW; Klivenyi, P; Carlson, EJ; Epstein, CJ; Beal, MF. Mice with a partial deficiency of manganese superoxide dismutase show increased vulnerability to the mitochondrial toxins malonate, 3-nitropropionic acid, and MPTP. Exptl Neurol 2001, 167, 189–195. [Google Scholar]
- Krasko, A; Schröder, HC; Batel, R; Grebenjuk, VA; Steffen, R; Müller, IM; Müller, WEG. Iron induces proliferation and morphogenesis in primmorphs from the marine sponge Suberites domuncula. DNA Cell Biol 2002, 21, 67–80. [Google Scholar]
- Wells, ML; Zorkin, NG; Lewis, AAG. The role of colloid chemistry in providing a source of iron to phytplankton. J Mar Res 1983, 41, 731–746. [Google Scholar]
- Pickell, L. Using flow injection analysis (FIA) to measure trace metal concentrations of iron and manganese in seawater. 2002. Available online: http://www.mbari.org/education/internship/02interns/02papers/lisa.pdf.
- Sung, JH; Kim, CY; Yang, SO; Khang, HS; Cheong, HK; Lee, JS; Song, CW; Park, JD; Han, JH; Chung, YH; Choi, BS; Kwon, IH; Cho, MH; Yu, IJ. Changes in blood manganese concentration and MRI T1 relaxation time during 180 days of stainless steel welding-fume exposure in Cynomolgus Monkeys. Inhal Toxicol 2007, 19, 47–55. [Google Scholar]
- Choi, JW; Kim, SK. Association between blood lead concentrations and body iron status in children. Arch Dis Child 2003, 88, 791–792. [Google Scholar]
- Boschwitz, H; Gofshtein-Gandman, L; Halvorson, HO; Keynan, A; Milner, Y. The possible involvement of trypsin-like enzymes in germination of spores of Bacillus cereus T and Bacillus subtilis 168. J Gen Microbiol 1991, 137, 1145–1153. [Google Scholar]
- Gilbert, PUPA; Abrecht, M; Frazer, BH. The organic-mineral interface in biominerals. Rev Mineral Geochem 2005, 59, 157–185. [Google Scholar]
- Vasantha, N; Freese, E. The role of manganese in growth and sporulation of Bacillus subtilis. J Gen Microbiol 1979, 112, 329–336. [Google Scholar]
- Sleytr, UB; Messner, P. Crystalline surface layers on bacteria. Annu Rev Microbiol 1983, 37, 311–339. [Google Scholar]
- Schultze-Lam, S; Beveridge, TJ. Physicochemical characteristics of the mineral-forming S-layer from the cyanobacterium Synechococcus strain GL24. Can J Microbiol 1994, 40, 216–223. [Google Scholar]
- Fortin, D; Ferris, FG; Beveridge, TJ. Surface-mediated mineral development by bacteria. Rev Mineral 1997, 35, 161–180. [Google Scholar]
- Bini, E; Dikshit, V; Dirksen, K; Drozda, M; Blum, P. Stability of mRNA in the hyperthermophilic archaeon Sulfolobus solfataricus. RNA 2002, 8, 1129–1136. [Google Scholar]
- Iafolla, MAJ; Dong, GQ; McMillen, DR. Increasing the efficiency of bacterial transcription simulations: When to exclude the genome without loss of accuracy. BMC Bioinformatics 2008, 9, 373. [Google Scholar]
- Jürgen, B; Schweder, T; Hecker, M. The stability of mRNA from the gsiB gene of Bacillus subtilis is dependent on the presence of a strong ribosome binding site. Mol Gen Genet 1998, 258, 538–545. [Google Scholar]
- Althoff, K; Schütt, C; Steffen, R; Batel, R; Müller, WEG. Evidence for a symbiosis between bacteria of the genus Rhodobacter and the marine sponge Halichondria panicea: Harbor also for putatively-toxic bacteria? Mar Biol 1998, 130, 529–536. [Google Scholar]
- Faulkner, DJ; Harper, MK; Haygood, MG; Salomon, CE; Schmidt, EW. Symbiotic Bacteria in Sponges: Sources of Bioactive Substances. In Drugs from the Sea; Fusetani, N, Ed.; Karger: Basel, Switzerland, 2000; pp. 107–119. [Google Scholar]
- Müller, WEG; Grebenjuk, VA; Le Pennec, G; Schröder, HC; Brümmer, F; Hentschel, U; Müller, IM; Breter, H-J. Sustainable production of bioactive compounds by sponges: Cell culture and gene cluster approach. Mar Biotechnol 2004, 6, 105–117. [Google Scholar]
- Weiss, B; Ebel, R; Elbrächter, M; Kirchner, M; Proksch, P. Defense metabolites from the marine sponge Verongia aerophoba. Biochem System Ecol 1996, 24, 1–7. [Google Scholar]
- Thakur, NL; Müller, WEG. Sponge-bacteria Association: A useful model to explore symbiosis in marine invertebrates. Symbiosis 2005, 39, 109–116. [Google Scholar]
- Kataoka, K; Sugiyama, R; Hirota, S; Inoue, M; Urata, K; Minagawa, Y; Seo, D; Sakurai, T. Four-electron reduction of dioxygen by a multicopper oxidase, CueO, and Roles of Asp112 and Glu506 located adjacent to the trinuclear copper center. J Biol Chem 2009, 284, 14405–14413. [Google Scholar]
- Ghiorse, WC. Biology of iron and manganese-depositing bacteria. Ann Rev Microbiol 1984, 38, 515–550. [Google Scholar]
- Rivera, ING; Lipp, EK; Gil, A; Choopun, N; Huq, A; Colwell, RR. Method of DNA extraction and application of multiplex polymerase chain reaction to detect toxigenic Vibrio cholerae O1 and O139 from aquatic ecosystems. Environ. Microbiol. 2003, 5, 599–606. [Google Scholar]
- Schaeffer, A; Fulton, D. A simplified method for staining spores. Science 1933, 77, 194–195. [Google Scholar]
- Francis, CA; Co, EM; Tebo, BM. Enzymatic manganese(II) oxidation by a marine aproteobacterium. Appl Environ Microbiol 2001, 67, 4024–4029. [Google Scholar]
- Krumbein, WE; Altmann, HJ. A new method for the detection and enumeration of manganese oxidizing and reducing microorganisms. Helgol Wiss Meeresunters 1973, 25, 347–356. [Google Scholar]
- Müller, WEG; Schuster, DK; Zahn, RK; Maidhof, A; Leyhausen, G; Falke, D; Koren, R; Umezawa, H. Properties and specificity of binding sites for the potential immunomodulator bestatin on the surface of mammalian cells. Int J Immunopharmacol 1982, 4, 393–400. [Google Scholar]
- Zheng, M; Liu, Q; Jin, N; Guo, J; Huang, X; Li, H; Zhu, W; Xiong, Y. A duplex PCR assay for simultaneous detection and differentiation of Capripoxvirus and Orf virus. Mol Cell Probes 2007, 21, 276–281. [Google Scholar]
- Lane, DL. 16S/23S rRNA sequencing. In Nucleic Acid Techniques in Bacterial Systematics; Stackebrandt, E, Goodfellow, M, Eds.; Wiley: New York, NY, USA, 1991; pp. 115–175. [Google Scholar]
- Müller, WEG; Kasueske, M; Wang, XH; Schröder, HC; Wang, Y; Pisignano, D; Wiens, M. Luciferase a light source for the silica-based optical waveguides (spicules) in the demosponge Suberites domuncula. Cell Mol Life Sci 2009, 66, 537–552. [Google Scholar]
- Dayhoff, MO; Schwartz, RM; Orcutt, BC. A model of evolutionary change in protein. In Atlas of Protein Sequence and Structure; Dayhoff, MO, Ed.; Nat Biomed Res Foundation: Washington, DC, USA, 1978; pp. 345–352. [Google Scholar]
- Nicholas, KB; Nicholas, HB, Jr. GeneDoc: a tool for editing and annotating multiple sequence alignments. Version 1.1.004. 1997. Distributed by the author; cris.com/~ketchup/genedoc.shtml; INTERNET.
- Wagner, C; Steffen, R; Koziol, C; Batel, R; Lacorn, M; Steinhart, H; Simat, T; Müller, WEG. Apoptosis in marine sponges: A biomarker for environmental stress (cadmium and bacteria). Mar Biol 1998, 131, 411–421. [Google Scholar]
- Huppert, M; Oliver, DJ; Sun, SH. Combined methenamine-silver nitrate and hematoxylin & eosin stain for fungi in tissues. J Clin Microbiol 1978, 8, 598–603. [Google Scholar]
- Wiens, M; Koziol, C; Hassanein, HMA; Batel, R; Müller, WEG. Expression of the chaperones 14-3-3 and HSP70 induced by PCB 118 (2,3′,4,4′,5-pentachlorobiphenyl) in the marine sponge Geodia cydonium. Mar Ecol Prog Ser 1998, 165, 247–257. [Google Scholar]
- Schröder, HC; Grebenjuk, VA; Binder, M; Skorokhod, A; Batel, R; Hassanein, H; Müller, WEG. Functional molecular biodiversity assessment of the immune status of two sponge populations (Suberites domuncula) on the molecular level. Mar Ecol 2004, 25, 93–108. [Google Scholar]
- Schütze, J; Skorokhod, A; Pahler, S; Müller, IM; Müller, WEG. Molecular evolution of metazoan extracellular matrix: cloning and expression of structural proteins from the demosponges Suberites domuncula andGeodia cydonium. J Mol Evol 2001, 53, 402–415. [Google Scholar]
- Compton, S; Jones, C. Mechanism of dye response and interference in the Bradford protein assay. Anal Biochem 1985, 151, 369–374. [Google Scholar]
- Sachs, L. Angewandte Statistik; Springer: Berlin, Germany, 1984. [Google Scholar]
- Sequences: The 16S rDNA fragment of the Bacillus-related strain (termed BAC-SubDo-3) that is associated with S. domuncula (16S-BAC-SubDo-3) has been deposited (EMBL/GenBank) under FN565385; in addition the fragment encoding the partial multicopper oxidase-like enzyme MnxG-SubDo-03 has been deposited as well (FN565386).


© 2011 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Wang, X.; Wiens, M.; Divekar, M.; Grebenjuk, V.A.; Schröder, H.C.; Batel, R.; Müller, W.E.G. Isolation and Characterization of a Mn(II)-Oxidizing Bacillus Strain from the Demosponge Suberites domuncula . Mar. Drugs 2011, 9, 1-28. https://doi.org/10.3390/md9010001
Wang X, Wiens M, Divekar M, Grebenjuk VA, Schröder HC, Batel R, Müller WEG. Isolation and Characterization of a Mn(II)-Oxidizing Bacillus Strain from the Demosponge Suberites domuncula . Marine Drugs. 2011; 9(1):1-28. https://doi.org/10.3390/md9010001
Chicago/Turabian StyleWang, Xiaohong, Matthias Wiens, Mugdha Divekar, Vladislav A. Grebenjuk, Heinz C. Schröder, Renato Batel, and Werner E. G. Müller. 2011. "Isolation and Characterization of a Mn(II)-Oxidizing Bacillus Strain from the Demosponge Suberites domuncula " Marine Drugs 9, no. 1: 1-28. https://doi.org/10.3390/md9010001





