Marine Myxobacteria as a Source of Antibiotics—Comparison of Physiology, Polyketide-Type Genes and Antibiotic Production of Three New Isolates of Enhygromyxa salina
Abstract
:1. Introduction
2. Results and Discussion
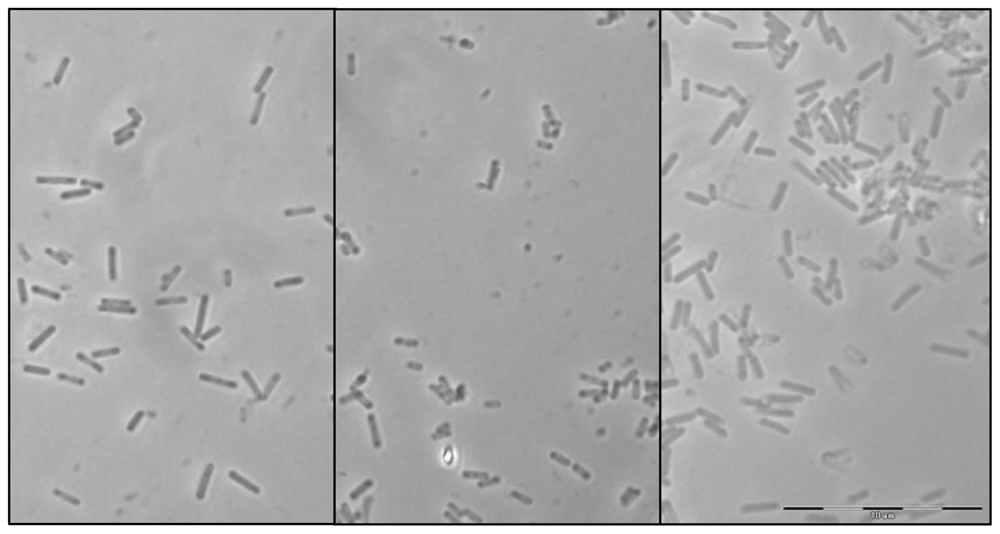
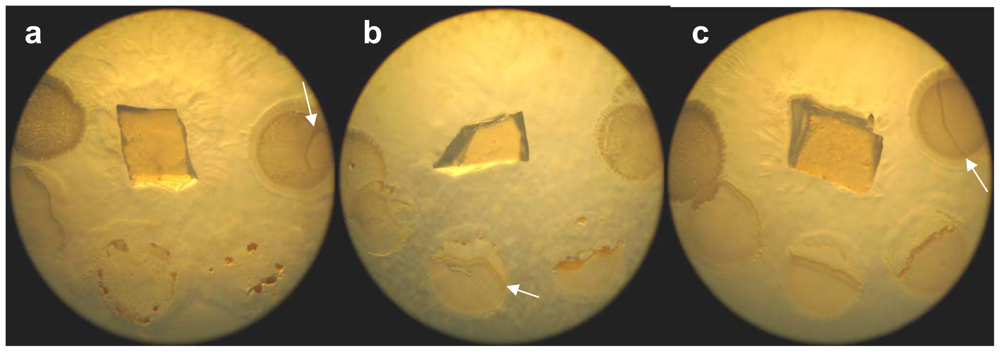
2.1. Morphological characterization
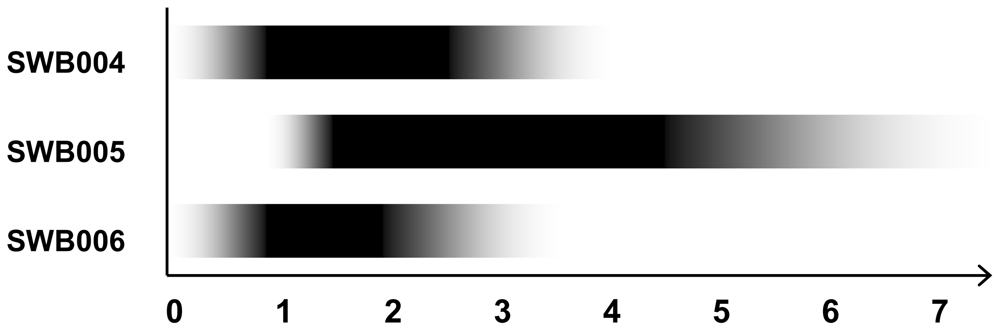
2.2. Physiological Characterization
2.3. Chemotaxonomy
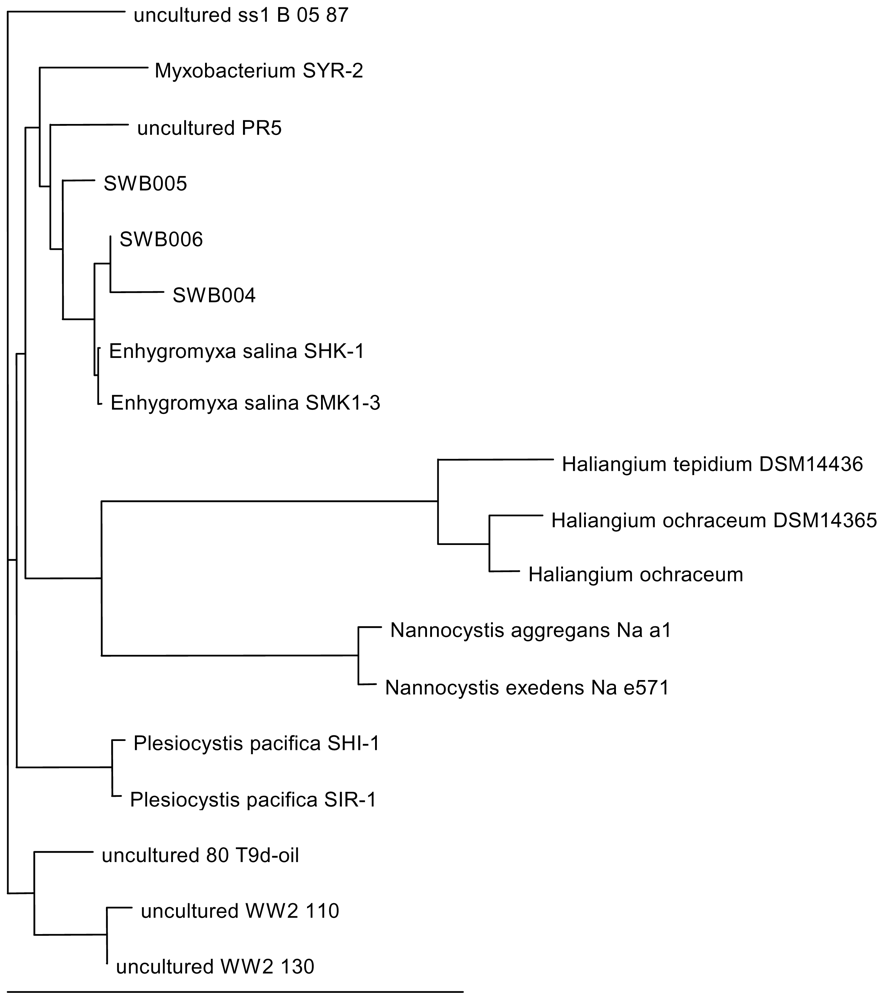
2.4. DNA analysis
2.5. The ability to produce secondary metabolites with antibacterial activity
3. Experimental Section
3.1. Sample collection, isolation and bacterial strains
3.2. Morphological and physiological characterization
3.3. Chemotaxonomy
3.4. DNA analysis
3.5. Agar diffusion assay
Acknowledgements
- Samples Availability: Available from the authors.
References
- Reichenbach, H; Dworkin, M. Balows, A, Trüper, HG, Dworkin, M, Harder, W, Schleifer, KH, Eds.; The myxobacteria. In The Prokaryotes, 2nd ed; Springer-Verlag: New York, NY, USA, 1992; pp. 3416–3487. [Google Scholar]
- Li, YZ; Hu, W; Zhang, YQ; Qiu, ZJ; Zhang, Y; Wu, BH. A simple method to isolate salt-tolerant myxobacteria from marine samples. J. Microbiol. Methods 2002, 50, 205–209. [Google Scholar]
- Reichenbach, H. Dworkin, M, Kaiser, D, Eds.; Biology of myxobacteria: Ecology and taxonomy. In Myxobacteria II; American Society for Microbiology: Washington, DC, USA, 1993; pp. 13–62. [Google Scholar]
- Fudou, R; Jojima, Y; Iizuka, T; Yamanaka, S. Haliangium ochraceum gen. nov., sp.nov. and Haliangium tepidum sp. nov.: Novel moderately halophilic myxobacteria isolated from coastal saline environments. J. Gen. Appl. Microbiol 2002, 48, 109–115. [Google Scholar]
- Iizuka, T; Jojima, Y; Fudou, R; Hiraishi, A; Ahn, J-W; Yamanaka, S. Plesiocystis pacifica gen. nov., sp. nov., a marine myxobacterium that contains dihydrogenated menaquinone, isolated from the Pacific coasts of Japan. Int. J. Syst. Evol. Microbiol 2003, 53, 189–195. [Google Scholar]
- Iizuka, T; Jojima, Y; Fudou, R; Tokura, M; Hiraishi, A; Yamanaka, S. Enhygromyxa salina gen. nov., sp. nov., a slightly halophilic myxobacterium isolated from the coastal areas of Japan. Syst. Appl. Microbiol 2003, 26, 189–196. [Google Scholar]
- Schneiker, S; Perlova, O; Kaiser, O; Gerth, K; Alici, A; Altmeyer, MO; Bartels, D; Bekel, T; Beyer, S; Bode, E; et al. Complete genome sequenece of the myxobacterium Sorangium cellulosum. Nat. Biotechnol 2007, 25, 1281–1289. [Google Scholar]
- Bode, HB; Müller, R. Analysis of myxobacterial secondary metabolism goes molecular. J. Ind. Microbiol. Biotechnol 2006, 33, 577–588. [Google Scholar]
- Reichenbach, H. Myxobacteria, producers of novel bioactive substances. J. Ind. Microbiol. Biotechnol 2001, 27, 149–156. [Google Scholar]
- Irschik, H; Schummer, D; Höfle, G; Reichenbach, H; Steinmetz, H; Jansen, R. Etnangien, a macrolide-polyene antibiotic from Sorangium cellulosum that inhibits nucleic acid polymerase. J. Nat. Prod 2007, 70, 1060–1063. [Google Scholar]
- Menche, D; Arikan, F; Perlova, O; Horstmann, N; Ahlbrecht, W; Wenzel, SC; Jansen, R; Irschik, H; Müller, R. Stereochemical determination and complex biosynthetic assembly of etnangien, a highly potent RNA polymerase inhibitor from the myxobacterium Sorangium cellulosum. J. Am. Chem. Soc 2008, 130, 14234–14243. [Google Scholar]
- Erol, O; Schäberle, TF; Schmitz, A; Rachid, S; Gurgui, C; El Omari, M; Lohr, F; Kehraus, S; Piel, J; Müller, R; König, GM. Biosynthesis of the myxobacterial antibiotic corallopyronin A. Chembiochem 2010, 11, 1253–1265. [Google Scholar]
- Irschik, H; Jansen, R; Höfle, G; Gerth, K; Reichenbach, H. The corallopyronins, new inhibitors of bacterial RNA synthesis from myxobacteria. J. Antibiot 1985, 38, 145–152. [Google Scholar]
- Wenzel, SC; Mülller, R. The biosynthetic potential of myxobacteria and their impact in drug discovery. Curr. Opin. Drug. Discov. Devel 2009, 12, 220–230. [Google Scholar]
- Thomas, ES; Gomez, HL; Li, RK; Chung, HC; Fein, LE; Chan, VF; Jassem, J; Pivot, XB; Klimovsky, JV; de Mendoza, FH; et al. Ixabepilone plus capecitabine for metastatic brest cancer progression after anthracycline and taxane treatment. J. Clin. Oncol 2007, 25, 5210–5217. [Google Scholar]
- MacLeod, RA. The question of the existence of specific marine bacteria. Bacteriol. Rev 1965, 29, 9–24. [Google Scholar]
- Kushner, DJ. Kushner, DJ, Ed.; Life in high salt and solute concentrations: halophilic bacteria. In Microbial Life in Extreme Environments; Academic: New York, NY, USA, 1978; pp. 318–346. [Google Scholar]
- Irschik, H; Augustiniak, H; Gerth, K; Höfle, G; Reichenbach, H. The ripostatins, novel inhibitors of eubacterial RNA polymerase isolated from myxobacteria. J. Antibiot 1995, 48, 787–792. [Google Scholar]
- Komaki, H; Fudou, R; Iizuka, T; Nakajima, D; Okazaki, K; Shibata, D; Ojika, M; Harayama, S. PCR detection of type I polyketide synthase genes in myxobacteria. Appl. Environ. Microbiol 2008, 74, 5571–5574. [Google Scholar]
- Fudou, R; Iizuka, T; Yamanaka, S. Haliangicin, a novel antifungal metabolite produced by a marine myxobacterium. J. Antibiot 2001, 54, 149–152. [Google Scholar]
- Gontang, EA; Gaudêncio, SP; Fenical, W; Jensen, PR. Sequence-based analysis of secondary-metabolite biosynthesis in marine actinobacteria. Appl. Environ. Microbiol 2010, 76, 2487–2499. [Google Scholar]
- Altschul, SF; Gish, W; Miller, W; Myers, EW; Lipman, DJ. Basic local alignment search tool. J. Mol. Biol 1990, 215, 403–410. [Google Scholar]
- Larkin, MA; Blackshields, G; Brown, NP; Chenna, R; McGettigan, PA; McWilliam, H; Valentin, F; Wallace, IM; Wilm, A; Lopez, R; Thompson, JD; Gibson, TJ; Higgins, DG. ClustalW and ClustalX version 2. Bioinformatics 2007, 23, 2947–2948. [Google Scholar]
- Mesbah, M; Premachandran, U; Whitman, WB. Precise measurement of the G + C content of deoxyribonucleic acid by high performance liquid chromathography. Int. J. Syst. Bacteriol 1989, 39, 159–167. [Google Scholar]
| SWB005 | SWB006 | SWB004 | DSM 15201 | |
|---|---|---|---|---|
| Colony color (on VY1/4) | colorless | colorless | colorless | colorless |
| Rod cell #: | ||||
| Diameter (μm) | ca. 0.35 | ca. 0.35 | ca. 0.35 | 0.5–0.8 * |
| Length (μm) | 1.2–1.8 | 1.3–2.6 | 1.2–2.6 | 1.5–7.0 * |
| Myxospore: | ||||
| Shape | spherical | spherical | spherical | spherical * |
| Diameter (μm) | 0.3–0.5 | 0.3–0.6 | 0.3–0.6 | 0.5–0.7 * |
| Growth at: | ||||
| 19 °C | yes | yes | yes | yes * |
| 30 °C | yes | yes | yes | yes |
| 37 °C | no | no | no | no * |
| Growth salinity (%NaCl): | ||||
| Range | 1.0–7.0 | 0.1–3.5 | 0.1–4.0 | 0.1–4.0 * |
| Optimum | 1.5–4.5 | 1.0–2.0 | 1.0–2.5 | 1.0–2.0 * |
| Cation requirement | Ca2+ or Mg2+ or K+ | Ca2+ and Mg2+ | Ca2+ or Mg2+ or K+ | Ca2+ or Mg2+ or K+ |
| Oxidase | yes | yes | yes | yes |
| Catalase | yes | yes | yes | yes |
| Hydrolysis of: | ||||
| Alginate | no growth | no growth | no growth | no |
| Cellulose | no | no | no | no |
| Chitin | no | no | no | no |
| Starch | no | no | no | no |
| Tween 20 | no | no | no | no growth |
| Yeast cells (autoclaved) | weak | yes | yes | yes |
| E. coli cells (alive) | yes | yes | yes | yes |
| G + C content (mol%) | 63.0 | 67.3 | 63.7 | 65–67 * |
| Preferred artificial seawater | ASW | SWS | SWS | SWS |
| Sequence | Identity | Strain and (putative) function | Accession number |
|---|---|---|---|
| SWB004 #1 | 512/690 (74%) | Frankia alni str. ACN14A, putative 6-methylsalicylic acid synthase | CT573213.2 |
| SWB004 #2 | 512/690 (74%) | Frankia alni str. ACN14A, putative 6-methylsalicylic acid synthase | CT573213.2 |
| SWB004 #3 | 512/690 (74%) | Frankia alni str. ACN14A, putative 6-methylsalicylic acid synthase | CT573213.2 |
| SWB004 #4 | 519/714 (72%) | Polyangium cellulosum So0157-2-KS1 beta-ketoacyl synthase gene | DQ359910.1 |
| SWB004 #5 | 512/714 (71%) | Polyangium cellulosum So0157-2-KS1 beta-ketoacyl synthase gene | DQ359910.1 |
| SWB004 #6 | 512/690 (74%) | Frankia alni str. ACN14A, putative 6-methylsalicylic acid synthase | CT573213.2 |
| SWB005 #1 | 567/638 (88%) | Enhygromyxa sp. SYM-1 gene for polyketide synthase | AB376534.1 |
| SWB005 #2 | 530/712 (74%) | Streptomyces abikoensis gene for PKS | AB430936.1 |
| SWB005 #3 | 541/706 (76%) | Polyangium cellulosum So ce26-KS7 beta-ketoacyl synthase gene | DQ359879.1 |
| SWB005 #4 | 530/712 (74%) | Streptomyces abikoensis gene for PKS | AB430936.1 |
| SWB006 #1 | 367/499 (73%) | Streptomyces sp. NRRL 11266 tetronomycin gene cluster | AB193609.1 |
| SWB006 #2 | 449/607 (73%) | Streptomyces sp. ID05-A0343, gene for PKS | AB431909.1 |
| SWB006 #3 | 350/474 (73%) | Actinoplanes sp. ID05-A0405, gene for PKS | AB432011.1 |
| SWB006 #4 | 511/694 (73%) | Streptomyces sp. ID05-A0343, gene for PKS | AB431909.1 |
| SWB006 #5 | 288/412 (69%) | Streptomyces noursei ATCC 11455, nystatin biosynthetic gene cluster | AF263912.1 |
| Strain\Extract | Dichloromethane | Ethylacetate | Acetone | Methanol |
|---|---|---|---|---|
| MRSA LT1334 | 13 | 14 | 11 | 8 |
| MRSA LT1338 | 10 | 10 | 8 | 8 |
| MRSE LT1324 | - | - | m.i. | 11 |
| CNS I-10925 | - | - | - | m.i. |
| Escherichia coli I-11276b | - | - | - | - |
| Escherichia coli O-19592 | - | - | - | - |
| Klebsiella pneumoniae I-10910 | - | - | - | - |
| Pseudomonas aeruginosa 4991 | - | - | - | - |
| Pseudomonas aeruginosa I-10968 | - | - | - | - |
| Citrobacter freundii I-11090 | - | - | - | - |
| Staphylococcus aureus I-11574 | - | - | m.i. | 9 |
| Staphylococcus aureus 5185 | - | - | m.i. | 9 |
| Stenotrophomonas maltophilia O-16451 | - | - | - | - |
| Stenotrophomonas maltophilia I-10717 | - | - | - | m.i. |
| Enterococcus I-11305b | - | - | - | 9 |
| Enterococcus I-11054 | - | - | m.i. | 9 |
| Staphylococcus simulans 22 | - | - | m.i. | 21 |
| Micrococcus luteus ATCC 4658 | - | - | 10 | 18 |
| Staphylococcus epidermidis 25 | - | - | - | 13 |
| Staphylococcus aureus SG511 | - | - | 9 | 14 |
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Schäberle, T.F.; Goralski, E.; Neu, E.; Erol, Ö.; Hölzl, G.; Dörmann, P.; Bierbaum, G.; König, G.M. Marine Myxobacteria as a Source of Antibiotics—Comparison of Physiology, Polyketide-Type Genes and Antibiotic Production of Three New Isolates of Enhygromyxa salina. Mar. Drugs 2010, 8, 2466-2479. https://doi.org/10.3390/md8092466
Schäberle TF, Goralski E, Neu E, Erol Ö, Hölzl G, Dörmann P, Bierbaum G, König GM. Marine Myxobacteria as a Source of Antibiotics—Comparison of Physiology, Polyketide-Type Genes and Antibiotic Production of Three New Isolates of Enhygromyxa salina. Marine Drugs. 2010; 8(9):2466-2479. https://doi.org/10.3390/md8092466
Chicago/Turabian StyleSchäberle, Till F., Emilie Goralski, Edith Neu, Özlem Erol, Georg Hölzl, Peter Dörmann, Gabriele Bierbaum, and Gabriele M. König. 2010. "Marine Myxobacteria as a Source of Antibiotics—Comparison of Physiology, Polyketide-Type Genes and Antibiotic Production of Three New Isolates of Enhygromyxa salina" Marine Drugs 8, no. 9: 2466-2479. https://doi.org/10.3390/md8092466





