New Peptides Isolated from Lyngbya Species: A Review
Abstract
:1. Introduction
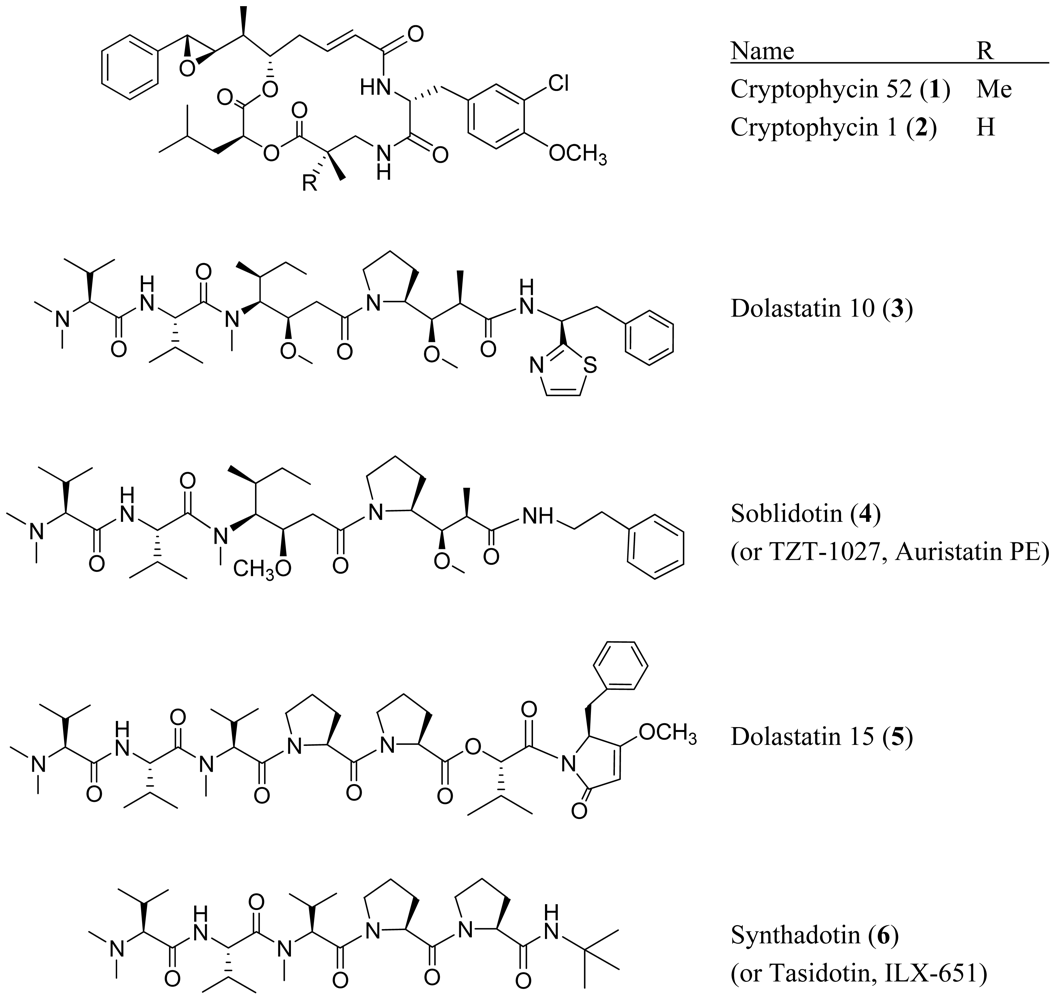
2. Acyclic Peptides
 | |||||
|---|---|---|---|---|---|
| Name | R1 | R2 | R3 | R4 | Acyl |
| Dragonamide A (7) | i-Pr | i-Pr | i-Pr | PhCH2- |  |
| Carmabin A (8) | Bn | Me | Me | 4-MeO-PhCH2- |  |
| Dragomabin (9) | Bn | Me | Me | 4-MeO-PhCH2- | 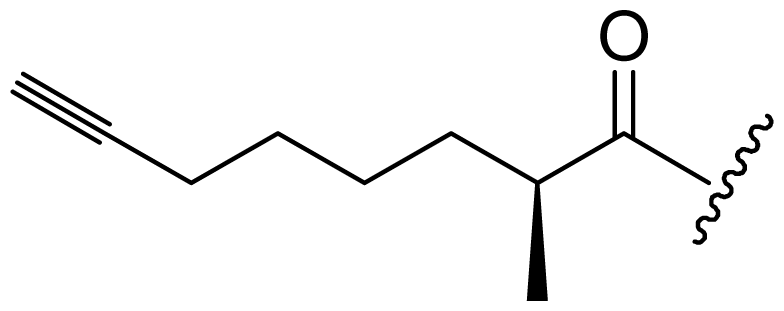 |
| Dragonamide B (10) | i-Pr | i-Pr | i-Pr | i-Pr |  |
| Dragonamide C (11) | i-Pr | i-Pr | i-Pr | i-Pr | 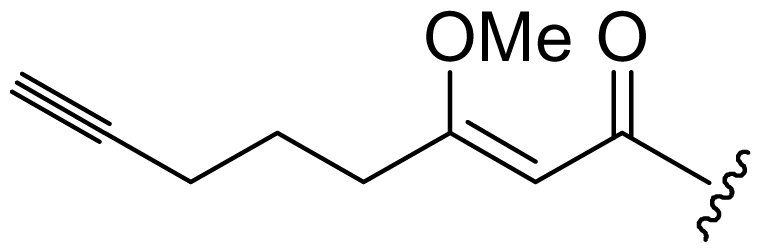 |
| Dragonamide D (12) | i-Pr | i-Pr | i-Pr | i-Pr | 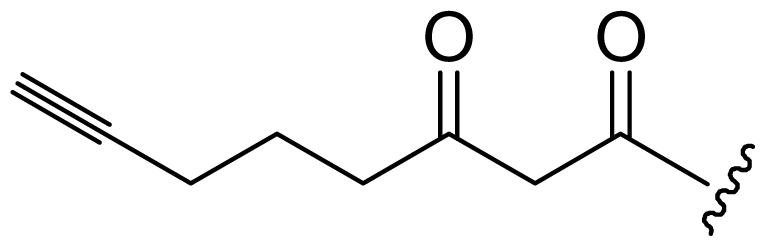 |
| Dragonamide E (13) | i-Pr | i-Pr | i-Pr | PhCH2- |  |
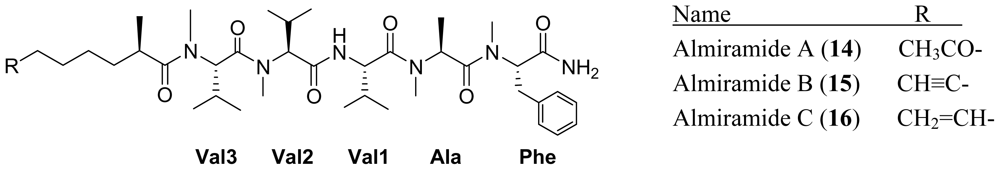
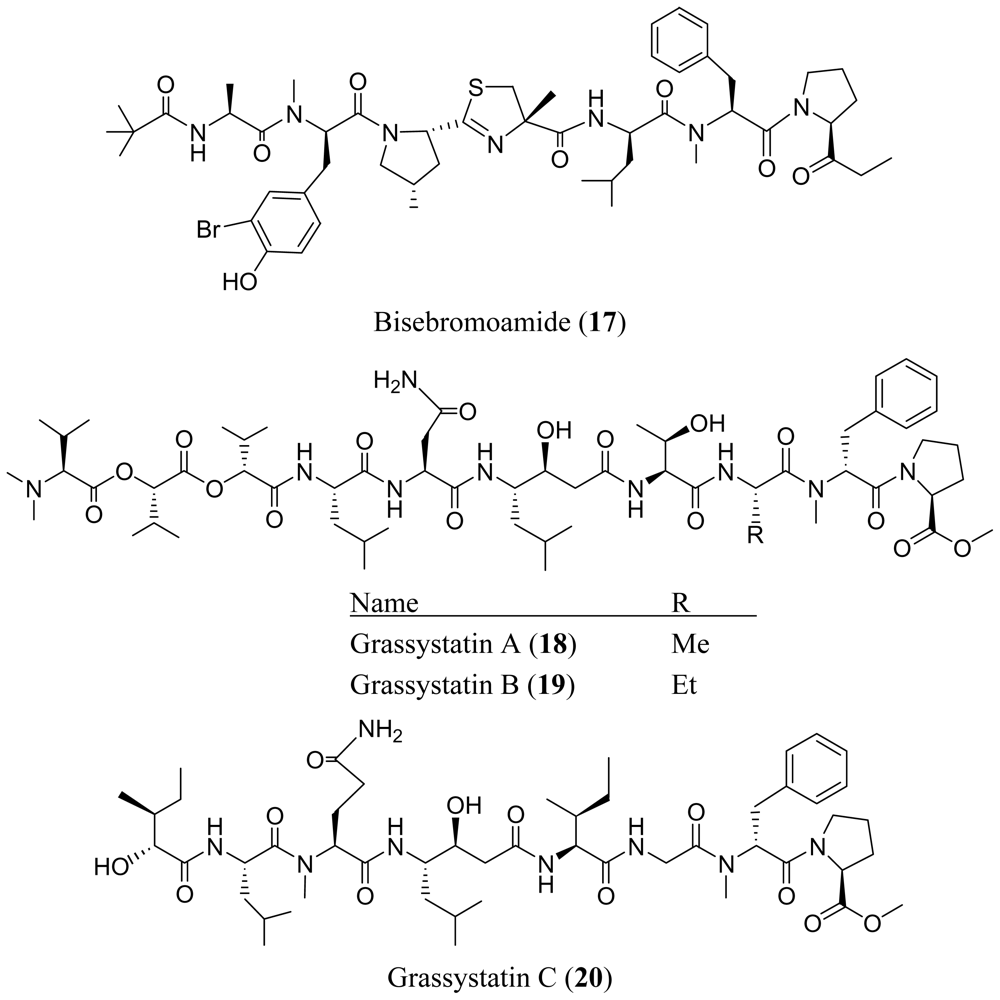
3. Cyclic Peptides
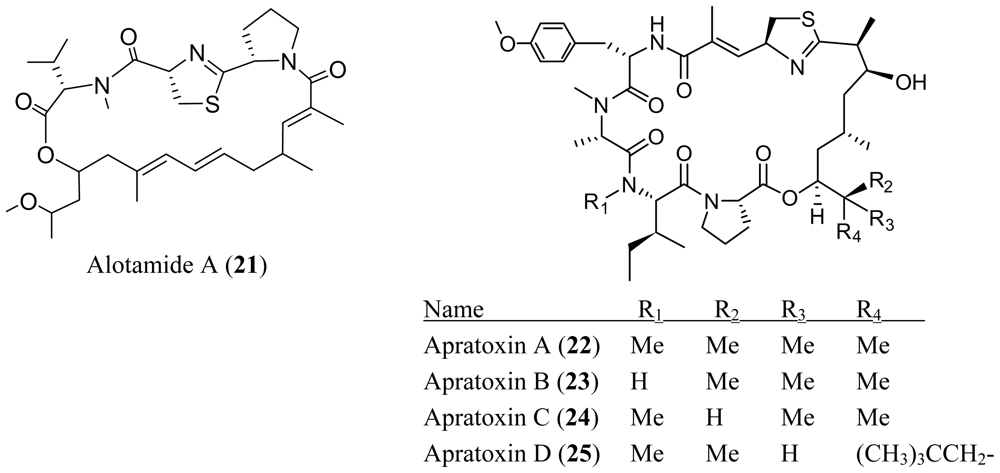
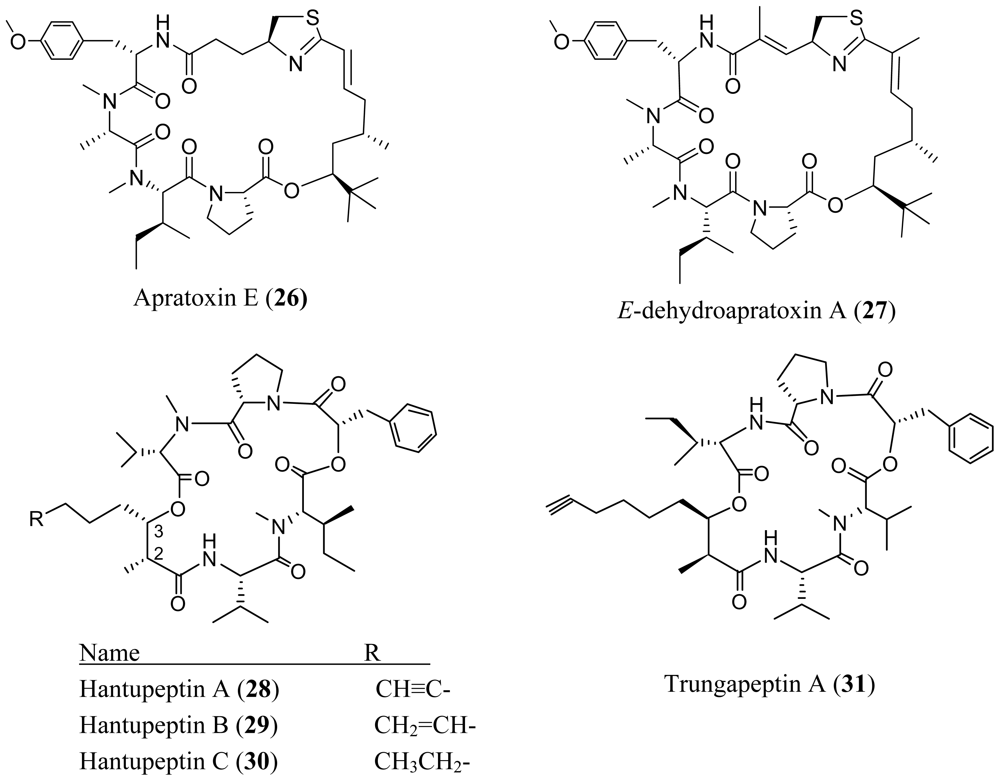
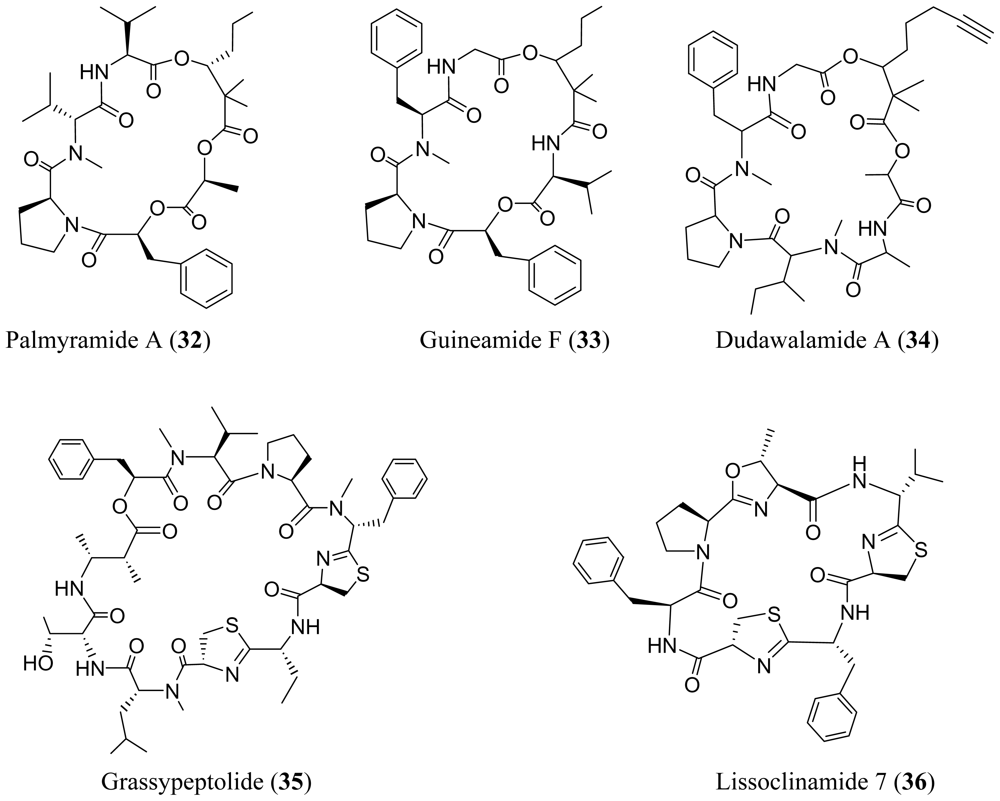
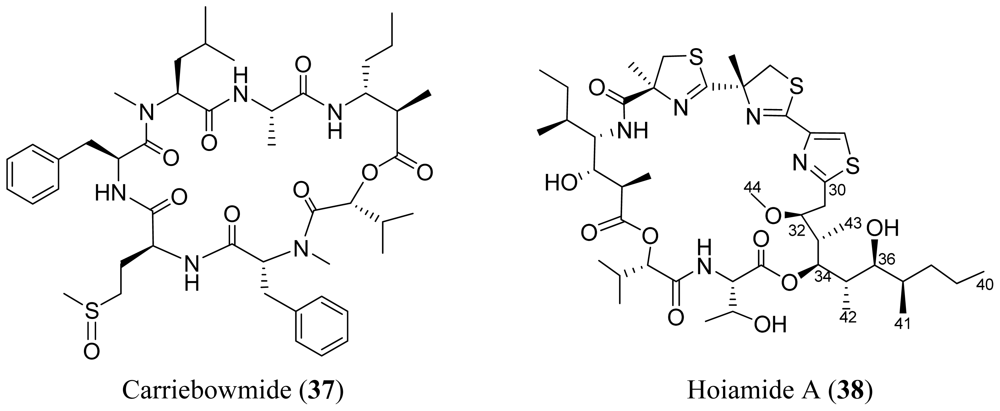
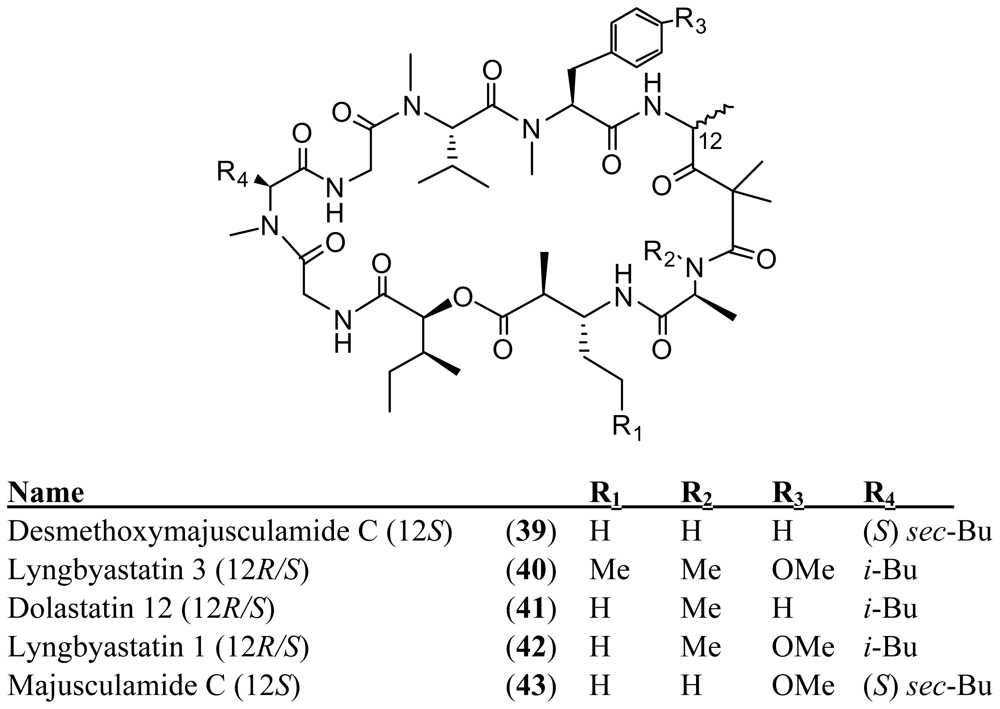
4. Cyclic Peptides with One or More Amide or Ester Branches
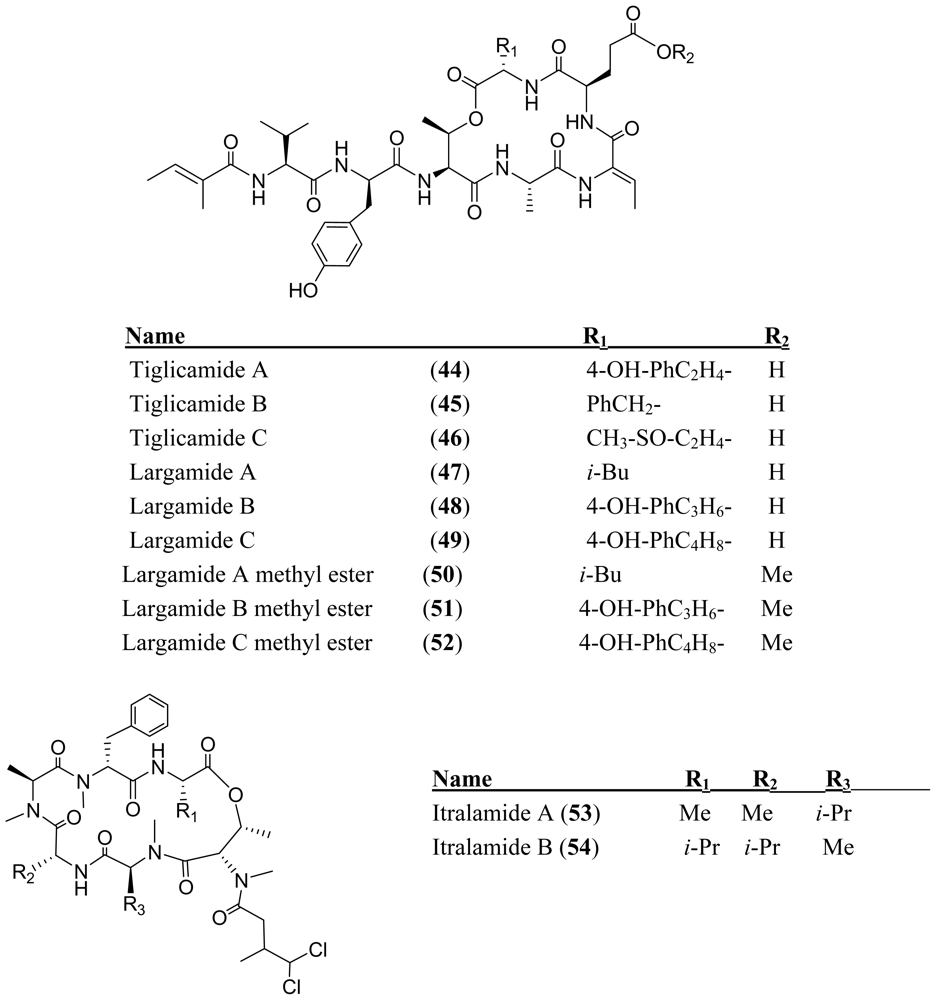
Acknowledgements
References and Notes
- Stanley, SM. Earth System History, 2nd ed; WH Freeman & Co: New York, NY, USA, 2004; p. 263. [Google Scholar]
- Tan, LT. Bioactive natural products from marine cyanobacteria for drug discovery. Phytochemistry 2007, 68, 954–979. [Google Scholar]
- Gerwick, WH; Roberts, MA; Proteau, PJ; Chen, JL. Screening cultured marine microalgae for anticancer-type activity. J. Appl. Phycol 1994, 6, 143–149. [Google Scholar]
- Gerwick, WH; Tan, LT; Siachitta, N. Nitrogen-containing metabolites from marine cyanobacteria. In The Alkaloids; Academic Press: San Diego, CA, USA, 2001; pp. 75–184. [Google Scholar]
- Namikoshi, M; Rinehart, KL. Bioactive compounds produced by cyanobacteria. J. Ind. Microbial. Biotechnol 1996, 17, 373–384. [Google Scholar]
- Mayer, MS; Gustafson, KR. Antitumor and cytotoxic compounds. Int. J. Cancer 2003, 105, 291–299. [Google Scholar]
- Trimurtulu, G; Ohtani, I; Patterson, GML; Moore, RE; Corbett, TH; Valeriote, FA; Dechik, L. Total structures of cryptophycins, potent antitumor depsipeptides from the blue-green alga Nostoc sp. strain GSV 224. J. Am. Chem. Soc 1994, 116, 4729–4737. [Google Scholar]
- D’Agostino, G; del Campo, J; Mellado, B; Izquierdo, MA; Minarik, T; Cirri, L; Marini, L; Perez-Gracia, JL; Scambia, G. A multicenter phase II study of the cryptophycin analog LY355703 in patients with platinum-resistant ovarian cancer. Int. J. Gynecol. Cancer 2006, 16, 71–76. [Google Scholar]
- Simmons, TL; Andrianasolo, E; McPhail, K; Flatt, P; Gerwick, WH. Marine natural products as anticancer drugs. Mol. Cancer Ther 2005, 4, 333–342. [Google Scholar]
- Shnyder, SD; Cooper, PA; Millington, NJ; Pettit, GR; Bibby, MC. Auristatin PYE, a novel synthetic derivative of dolastatin 10, is highly effective in human colon tumour models. Int. J. Oncol 2007, 31, 353–360. [Google Scholar]
- Ebbinghaus, S; Hersh, E; Cunningham, CC; O’Day, S; McDermott, D; Stephenson, J; Richards, DA; Eckardt, J; Haider, OL; Hammond, LA. Phase II study of synthadotin (SYN-D; ILX651) administered daily for 5 consecutive days once every 3 weeks in patients with inoperable locally advanced or metastatic melanoma. Proceedings of the 2004 American Society of Clinical Oncology Annual Meeting, New Orleans, LA, USA, June 2004. Abstr. #7530.
- Welker, M; von Döhren, H. Cyanobacterial peptides-Nature’s own combinatorial biosynthesis. FEMS Microiol. Rev 2006, 30, 530–563. [Google Scholar]
- Speziale, BJ; Dyck, LA. Lyngbya infestations: comparative taxonomy of Lyngbya wollei comb. nov. (Cyanobacteria). J. Phycol 1992, 28, 693–706. [Google Scholar]
- Jiménez, JI; Scheuer, PJ. New lipopeptides from the Caribbean cyanobacterium Lyngbya majuscula. J. Nat. Prod 2001, 64, 200–203. [Google Scholar]
- McPhail, KL; Correa, J; Linington, RG; Gonzalez, J; Ortega-Barría, E; Capson, TL; Gerwick, WH. Antimalarial linear lipopeptides from a Panamanian strain of the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod 2007, 70, 984–988. [Google Scholar]
- Gunasekera, SP; Ross, C; Paul, VJ; Matthew, S; Luesch, H. Dragonamides C and D, linear lipopeptides from the marine cyanobacterium brown Lyngbya polychroa. J. Nat. Prod 2008, 71, 887–890. [Google Scholar]
- Balunas, MJ; Linington, RG; Tidgewell, K; Fenner, AM; Ureña, LD; Togna, GD; Kyle, DE; Gerwick, WH. Dragonamide E, a modified linear lipopeptide from Lyngbya majuscula with antileishmanial activity. J. Nat. Prod 2010, 73, 60–66. [Google Scholar]
- Sanchez, LM; Lopez, D; Vesely, BA; Della Togna, G; Gerwick, WH; Kyle, DE; Linington, RG. Almiramides A–C: discovery and development of a new class of leishmaniasis lead compounds. J. Med. Chem 2010. [Epub ahead of print]. [Google Scholar]
- Teruya, T; Sasaki, H; Fukazawa, H; Suenaga, K. Bisebromoamide, a potent cytotoxic peptide from the marine cyanobacterium Lyngbya sp.: isolation, stereostructure, and biological activity. Org. Lett 2009, 11, 5062–5065. [Google Scholar]
- Kwan, JC; Eksioglu, EA; Liu, C; Paul, VJ; Luesch, H. Grassystatins A–C from marine cyanobacteria, potent cathepsin E inhibitors that reduce antigen presentation. J. Med. Chem 2009, 52, 5732–5747. [Google Scholar]
- Soria-Mercado, IE; Pereira, A; Cao, Z; Murray, TF; Gerwick, WH. Alotamide A, a novel neuropharmacological agent from the marine cyanobacterium Lyngbya bouillonii. Org. Lett 2009, 11, 4704–4707. [Google Scholar]
- Gutiérrez, M; Suyama, TL; Engene, N; Wingerd, JS; Matainaho, T; Gerwick, WH. Apratoxin D, a potent cytotoxic cyclodepsipeptide from Papua New Guinea collections of the marine cyanobacteria Lyngbya majuscula and Lyngbya sordida. J. Nat. Prod 2008, 71, 1099–1103. [Google Scholar]
- Simmons, TL; Coates, RC; Clark, BR; Engene, N; Gonzalez, D; Esquenazi, E; Dorrestein, PC; Gerwick, WH. Biosynthetic origin of natural products isolated from marine microorganism-invertebrate assemblages. Proc. Natl. Acad. Sci. USA 2008, 105, 4587–4594. [Google Scholar]
- Matthew, S; Schupp, PJ; Luesch, H. Apratoxin E, a cytotoxic peptolide from a Guamanian collection of the marine cyanobacterium Lyngbya bouillonii. J. Nat. Prod 2008, 71, 1113–1116. [Google Scholar]
- Tripathi, A; Puddick, J; Prinsep, MR; Lee, PPF; Tan, LT. Hantupeptin A, a cytotoxic cyclic depsipeptide from a Singapore collection of Lyngbya majuscula. J. Nat. Prod 2009, 72, 29–32. [Google Scholar]
- Tripathi, A; Puddick, J; Prinsep, MR; Lee, PPF; Tan, LT. Hantupeptins B and C, cytotoxic cyclodepsipeptides from the marine cyanobacterium Lyngbya majuscula. Phytochemistry 2009, 72, 29–32. [Google Scholar]
- Bunyajetpong, S; Yoshida, WY; Sitachitta, N; Kaya, K. Trungapeptins A–C, cyclodepsipeptides from the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod 2006, 69, 1539–1542. [Google Scholar]
- Taniguchi, M; Nunnery, JK; Engene, N; Esquenazi, E; Byrum, T; Dorrestein, PC; Gerwick, WH. Palmyramide A, a cyclic depsipeptide from a Palmyra Atoll collection of the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod 2010, 73, 393–398. [Google Scholar]
- Tan, LT; Sitachitta, N; Gerwick, WH. The guineamides, novel cyclic depsipeptides from a Papua New Guinea collection of the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod 2003, 66, 764–771. [Google Scholar]
- Liu, WT; Ng, J; Meluzzi, D; Bandeira, N; Gutierrez, M; Simmons, TL; Schultz, AW; Linington, RG; Moore, BS; Gerwick, WH; Pevzner, PA; Dorrestein, PC. Interpretation of tandem mass spectra obtained from cyclic nonribosomal peptides. Anal. Chem 2009, 81, 4200–4209. [Google Scholar]
- Malloy, K; Engene, N; Pedler, B; Clark, BR; Gerwick, WH. Isolation, structure elucidation, and SAR perspectives of cyanobacterial cyclic depsipeptides containing the unique Dhoya (3-hydroxy-2,2-dimethyl-7-octynoic acid) fragment and its derivatives. 42nd Western Regional Meeting of the American Chemical Society, Las Vegas, NV, USA, September 2008.
- Kwan, JC; Rocca, JR; Abboud, KA; Paul, VJ; Luesch, H. Total structure determination of grassypeptolide, a new marine cytotoxin. Org. Lett 2008, 10, 789–792. [Google Scholar]
- Hawkins, CJ; Lavin, MF; Marshall, KA; van den Brenk, AL; Watters, DJ. Structure-activity relationships of the lissoclinamides: cytotoxic cyclic peptides from the ascidian Lissoclinum patella. J. Med. Chem 1990, 33, 1634–1638. [Google Scholar]
- Wipf, P; Fritch, PC; Geib, SJ; Sefler, AM. Conformational studies and structure–activity analysis of lissoclinamide 7 and related cyclopeptide alkaloids. J. Am. Chem. Soc 1998, 120, 4105–4112. [Google Scholar]
- Gunasekera, SP; Ritson-Williams, R; Paul, VJ. Carriebowmide, a new cyclodepsipeptide from the marine cyanobacterium Lyngbya polychroa. J. Nat. Prod 2008, 71, 2060–2063. [Google Scholar]
- Pereira, A; Cao, Z; Murray, TF; Gerwick, WH. Hoiamide A, a sodium channel activator of unusual architecture from a consortium of two Pupa New guinea cyanobactiera. Chem. Biol 2009, 16, 893–906. [Google Scholar]
- Simmons, TL; Nogle, LM; Media, J; Valeriote, FA; Mooberry, SL; Gerwick, WH. Desmethoxymajusculamide C, a cyanobacterial depsipeptide with potent cytotoxicity in both cyclic and ring-opened forms. J. Nat. Prod 2009, 72, 1011–1016. [Google Scholar]
- Matthew, S; Paul, VJ; Luesch, H. Tiglicamides A–C, cyclodepsipeptides from the marine cyanobacterium Lyngbya confervoides. Phytochemistry 2009, 70, 2058–2063. [Google Scholar]
- Matthew, S; Paul, VJ; Luesch, H. Largamides A–C, tiglic acid-containing cyclodepsipeptides with elastase-inhibitory activity from the marine cyanobacterium Lyngbya confervoides. Planta Med 2009, 75, 528–533. [Google Scholar]
- Kleinkauf, H; Von Döhren, H. A nonribosomal system of peptide biosynthesis. Eur. J. Biochem 1996, 236, 335–351. [Google Scholar]
- Jiménez, JI; Vansach, T; Yoshida, WY; Sakamoto, B; Pörzgen, P; Horgen, FD. Halogenated fatty acid amides and cyclic depsipeptides from an eastern Caribbean collection of the cyanobacterium Lyngbya majuscula. J. Nat. Prod 2009, 72, 1573–1578. [Google Scholar]
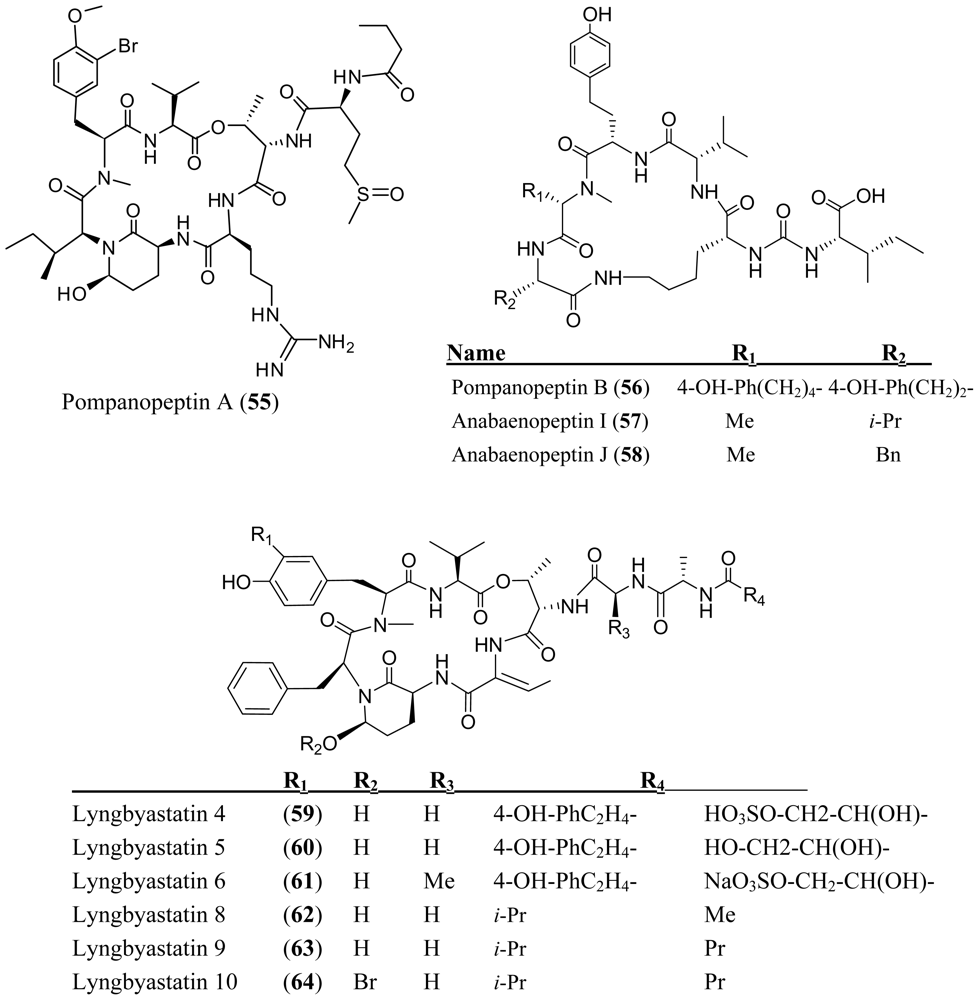
- Matthew, S; Ross, C; Paul, VJ; Luesch, H. Pompanopeptins A and B, new cyclic peptides from the marine cyanobacterium Lyngbya confervoides. Tetrahedron 2008, 64, 4081–4089. [Google Scholar]
- Murakami, M; Suzuki, S; Itou, Y; Kodani, S; Ishida, K. New anabaenopeptins, potent carboxypeptidase-A inhibitors from the cyanobacterium Aphanizomenon flos-aquae. J. Nat. Prod 2000, 63, 1280–1282. [Google Scholar]
- Matthew, S; Ross, C; Rocca, JR; Paul, VJ; Luesch, H. Lyngbyastatin 4, a dolastatin 13 analog with elastase and chymotrypsin inhibitory activity from the marine cyanobacterium Lyngbya confervoides. J. Nat. Prod 2007, 70, 124–127. [Google Scholar]
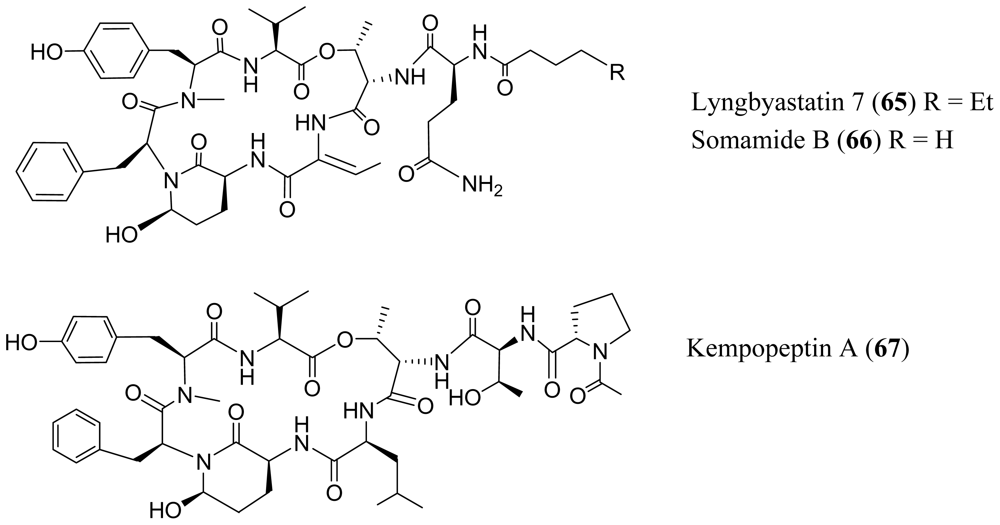
- Taori, K; Matthew, S; Rocca, JR; Paul, VJ; Luesch, H. Lyngbyastatins 5–7, potent elastase inhibitors from Floridian marine cyanobacteria, Lyngbya spp. J. Nat. Prod 2007, 70, 1593–1600. [Google Scholar]
- Kwan, JC; Taori, K; Paul, VJ; Luesch, H. Lyngbyastatins 8–10, elastase inhibitors with cyclic depsipeptide scaffolds isolated from the marine cyanobacterium Lyngbya semiplena. Mar. Drugs 2009, 7, 528–538. [Google Scholar]
- Nogle, LM; Williamson, RT; Gerwick, WH. Somamides A and B, two new depsipeptide analogs of dolastatin 13 from a Fijian cyanobacterial assemblage of Lyngbya majuscula and Schizothrix species. J. Nat. Prod 2001, 64, 716–719. [Google Scholar]
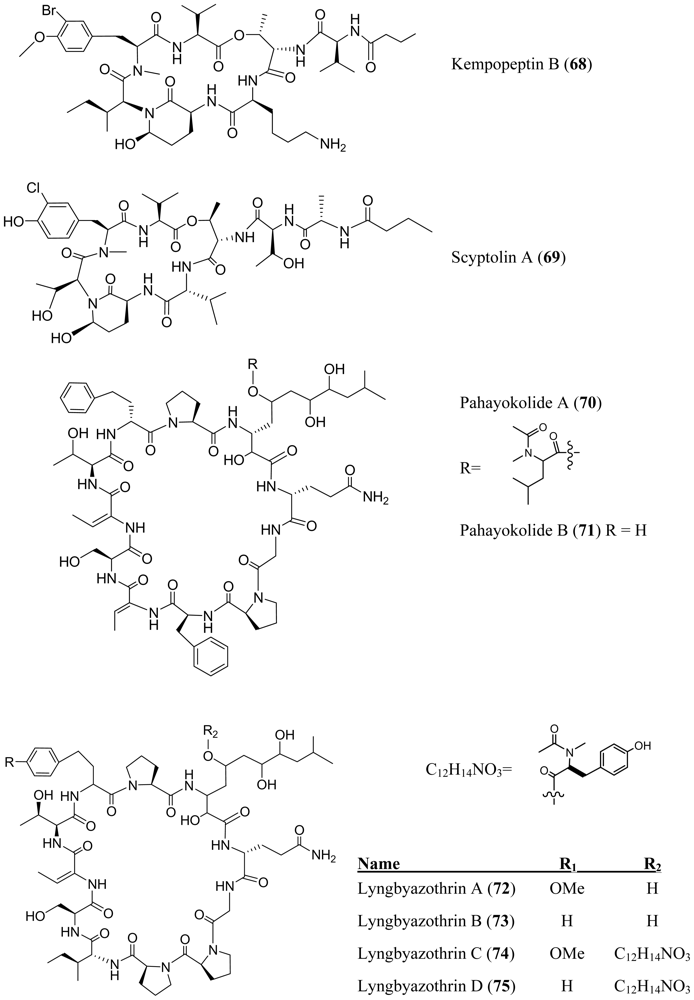
- Taori, K; Paul, VJ; Luesch, H. Kempopeptins A and B, serine protease inhibitors with different selectivity profiles from a marine cyanobacterium, Lyngbya sp. J. Nat. Prod 2008, 71, 1625–1629. [Google Scholar]
- Matern, U; Schleberger, C; Jelakovic, S; Weckesser, J; Schulz, EG. Binding structure of elastase inhibitor scyptolin A. Chem. Biol 2003, 10, 997–1001. [Google Scholar]
- Matern, U; Oberer, L; Falchetto, RA; Erhard, M; König, WA; Herdman, M; Weckesser, J. Scyptolin A and B, cyclic depsipeptides from axenic cultures of Scytonema hofmanni PCC 7110. Phytochemistry 2001, 58, 1087–1095. [Google Scholar]
- Lee, AY; Smitka, TA; Bonjouklian, R; Clardy, J. Atomic structure of the trypsin-A90720A complex: a unified approach to structure and function. Chem. Biol 1994, 1, 113–117. [Google Scholar]
- An, T; Krishnaswamy, T; Kumar, S; Wang, M; Liu, L; Lay, J, Jr; Liyanage, R; Berry, J; Gantar, M; Marks, V; Gawley, RE; Rein, KS. Structures of pahayokolides A and B, two cyclic peptides from a Lyngbya sp. J. Nat. Prod 2007, 70, 730–735. [Google Scholar]
- Zainuddin, NE; Jansen, R; Nimtz, M; Wray, V; Preisitsch, M; Lalk, M; Mundt, S. Lyngbyazothrins A–D, antimicrobial cyclic undecapeptides from the cultured cyanobacterium Lyngbya sp. J. Nat. Prod 2009, 72, 1373–1378. [Google Scholar]
- Zainuddin, NE; Jansen, R; Nimtz, M; Wray, V; Preisitsch, M; Lalk, M; Mundt, S. Corrigendum to: Lyngbyazothrins A–D, antimicrobial cyclic undecapeptides from the cultured cyanobacterium Lyngbya sp. J. Nat. Prod 2009, 72, 2080, The correct absolute configuration of the Gln residue is R as showed in the discussion. [Google Scholar]
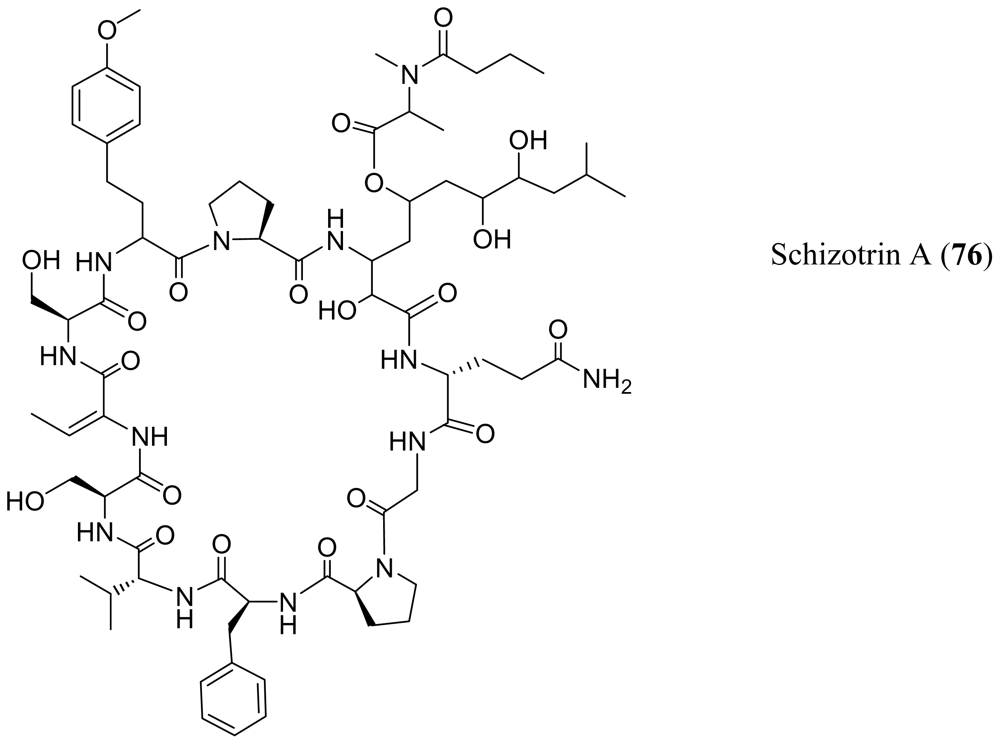
- Pergament, I; Carmeli, S. Schizotrin A, a novel antimicrobial cyclic peptide from a cyanobacterium. Tetrahedron Lett 1994, 35, 8473–8476. [Google Scholar]
- Mehner, C; Mueller, D; Krick, A; Kehraus, S; Löeser, R; Güetschow, M; Maier, A; Fiebig, HF; Brun, R; Köenig, GM. A novel β-amino acid in cytotoxic peptides from the cyanobacterium Tychonema sp. Eur. J. Org. Chem 2008, 10, 1732–1739. [Google Scholar]
- Berry, JP; Gantar, M; Gawley, RE; Wang, M; Rein, KS. Pharmacology and toxicology of pahayokolide A, a bioactive metabolite from a freshwater species of Lyngbya isolated from the Florida Everglades. Comp. Biochem. Physiol., Part C: Toxicol. Pharmacol 2004, 139, 231–238. [Google Scholar]
- Magarvey, NA; Beck, ZQ; Golakoti, T; Ding, Y; Huber, U; Hemscheidt, TK; Abelson, D; Moore, RE; Sherman, DH. Biosynthetic characterization and chemoenzymatic assembly of the cryptophycins. Potent anticancer agents from cyanobionts. ACS Chem. Biol 2006, 1, 766–779. [Google Scholar]
- Watts, RE; Tse, ML; Khosla, C. Substrate tolerance of module 6 of the epothilone synthetase. Biochemistry 2007, 46, 3385–3393. [Google Scholar]
- Challis, GL; Ravel, J; Townsend, CA. Predictive, structure-based model of amino acid recognition by nonribosomal peptide synthetase adenylation domains. Chem. Biol 2000, 7, 211–224. [Google Scholar]
- Chang, Z; Flatt, P; Gerwick, WH; Nguyen, VA; Willis, CL; Sherman, DH. The barbamide biosynthetic gene cluster: a novel marine cyanobacterial system of mixed polyketide synthase (PKS)-non-ribosomal peptide synthetase (NRPS) origin involving an unusual trichloroleucyl starter unit. Gene 2002, 296, 235–247. [Google Scholar]
- Chang, Z; Sitachitta, N; Rossi, JV; Roberts, MA; Flatt, PM; Jia, J; Sherman, DH; Gerwick, WH. Biosynthetic pathway and gene cluster analysis of curacin A, an antitubulin natural product from the tropical marine cyanobacterium Lyngbya majuscula. J. Nat. Prod 2004, 67, 1356–1367. [Google Scholar]
- Edwards, DJ; Gerwick, WH. Lyngbyatoxin biosynthesis: sequence of biosynthetic gene cluster and identification of a novel aromatic prenyltransferase. J. Am. Chem. Soc 2004, 126, 11432–11433. [Google Scholar]
- Edwards, DJ; Marquez, BL; Nogle, LM; McPhail, K; Goeger, DE; Roberts, MA; Gerwick, WH. Structure and biosynthesis of the jamaicamides, new mixed polyketide-peptide neurotoxins from the marine cyanobacterium Lyngbya majuscula. Chem. Biol 2004, 11, 817–833. [Google Scholar]
- Ramaswamy, AV; Sorrels, CM; Gerwick, WH. Cloning and biochemical characterization of the hectochlorin biosynthetic gene cluster from the marine cyanobacterium Lyngbya majuscula. J. Nat. Prod 2007, 70, 1977–1986. [Google Scholar]
| Name | Elastase | Chymotrypsin | Trypsin | Reference |
|---|---|---|---|---|
| Tiglicamide A (44) | 2.14 | >50 | >50 | [38] |
| Tiglicamide B (45) | 6.99 | >50 | >50 | [38] |
| Tiglicamide C (46) | 7.28 | >50 | >50 | [38] |
| Largamide A (47) | 1.41 | >50 | >50 | [39] |
| Largamide B (48) | 0.53 | >50 | >50 | [39] |
| Largamide C (49) | 1.15 | >50 | >50 | [39] |
| Pompanopeptin A (55) | 2.4 | [42] | ||
| Lyngbyastatin 4 (59) | 0.0139 or 0.03 | 4.3 or 0.3 | >30 | [44,45] |
| Lyngbyastatin 5 (60) | 0.0032 | 2.8 | >30 | [45] |
| Lyngbyastatin 6 (61) | 0.0033 | 2.5 | >30 | [45] |
| Lyngbyastatin 8 (62) | 0.0083 or 0.047 | 2.5 | >30 | [45,46] |
| Lyngbyastatin 9 (63) | 0.123 | / | / | [46] |
| Lyngbyastatin 10 (64) | 0.210 | / | / | [46] |
| Lyngbyastatin 7 (65) | 0.120 | / | / | [46] |
| Somamide B (66) | 0.0095 | 4.2 | >30 | [47] |
| Kempopeptin A (67) | 0.32 | 2.6 | >67 | [48] |
| Kempopeptin B (68) | >67 | >67 | 8.4 | [48] |
| Scyptolin A (69) | 2.8 | / | >446 | [50] |
© 2008 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Liu, L.; Rein, K.S. New Peptides Isolated from Lyngbya Species: A Review. Mar. Drugs 2010, 8, 1817-1837. https://doi.org/10.3390/md8061817
Liu L, Rein KS. New Peptides Isolated from Lyngbya Species: A Review. Marine Drugs. 2010; 8(6):1817-1837. https://doi.org/10.3390/md8061817
Chicago/Turabian StyleLiu, Li, and Kathleen S. Rein. 2010. "New Peptides Isolated from Lyngbya Species: A Review" Marine Drugs 8, no. 6: 1817-1837. https://doi.org/10.3390/md8061817




