Lobophytones O–T, New Biscembranoids and Cembranoid from Soft Coral Lobophytum pauciflorum
Abstract
:1. Introduction
2. Results and Discussion
3. Experimental Section
3.1. General Procedures
3.2. Animal Material
3.3. Extraction and Isolation
3.3.1. Lobophytone O (1)
3.3.2. Lobophytone P (2)
3.3.3. Lobophytone Q (3)
3.3.4. Lobophytone R (4)
3.3.5. Lobophytone S (5)
3.3.6. Lobophytone T (6)
3.4. Assay for Inhibition of Nitric Oxide (NO) Production
3.5. Antimicrobial and Antifungal Assays
Supplementary Material
marinedrugs-08-02837-s001.pdfAcknowledgements
References
- Su, J; Long, K; Peng, T; He, C; Clardy, J. The structure of methyl isosartortuoate, a novel tetracyclic tetraterpenoid from the soft coral Sarcophyton tortuosum. J. Am. Chem. Soc. 1986, 108, 177–178. [Google Scholar]
- Kusumi, T; Igari, M; Ishitsuka, MO; Ichikawa, A; Itezono, Y; Nakayama, N; Kakisawa, H. A novel chlorinated biscembranoid from the marine soft coral Sarcophyton glaucum. J. Org. Chem. 1990, 55, 6286–6289. [Google Scholar]
- Leone, PA; Bowden, BF; Carroll, AR; Coll, JC; Meehan, GV. Studies of Australian soft corals, XLIX: A new biscembranoid and its probable biosynthetic precursors from the soft coral Sarcophyton tortuosum. J. Nat. Prod. 1993, 56, 521–526. [Google Scholar]
- Zeng, L; Lan, W; Su, J; Zhang, G; Feng, X; Liang, Y; Yang, X. Two new cytotoxic tetracyclic tetraterpenoids from the soft coralSarcophyton tortuosum. J. Nat. Prod. 2004, 67, 1915–1918. [Google Scholar]
- Iwagawa, T; Hashimoto, K; Okamura, H; Kurawaki, J; Nakatani, M; Hou, DX; Fujii, M; Doe, M; Morimoto, Y; Takemura, K. Biscembranes from the soft coral Sarcophyton glaucum. J. Nat. Prod. 2006, 69, 1130–1133. [Google Scholar]
- Jia, R; Guo, Y; Chen, P; Yang, Y; Mollo, E; Gavagnin, M; Cimino, G. Biscembranoids and their probable biogenetic precursor from the Hainan soft coral Sarcophyton tortuosum. J. Nat. Prod. 2007, 70, 1158–1166. [Google Scholar]
- Bishara, A; Rudi, A; Benayahu, Y; Kashman, Y. Three biscembranoids and their monomeric counterpart cembranoid, a biogenetic Diels–Alder precursor, from the soft coral Sarcophyton elegans. J. Nat. Prod. 2007, 70, 1951–1954. [Google Scholar]
- Iwagawa, T; Hashimoto, K; Yokogawa, Y; Okamura, H; Nakatani, M; Doe, M; Morimoto, Y; Takemura, K. Cytotoxic biscembranes from the soft coral Sarcophyton glaucum. J. Nat. Prod. 2009, 72, 946–949. [Google Scholar]
- Yan, P; Lv, Y; Ofwegen, L; Proksch, P; Lin, W. Lobophytones A–G, new isobiscembranoids from the soft coral Lobophytum pauciflorum. Org. Lett. 2010, 2, 2484–2487. [Google Scholar]
- Li, E; Jiang, L; Guo, L; Zhang, H; Che, Y. Pestalachlorides A–C, antifungal metabolites from the plant endophytic fungus Pestalotiopsis adusta. Bioorg. Med. Chem. 2008, 16, 7894–7899. [Google Scholar]

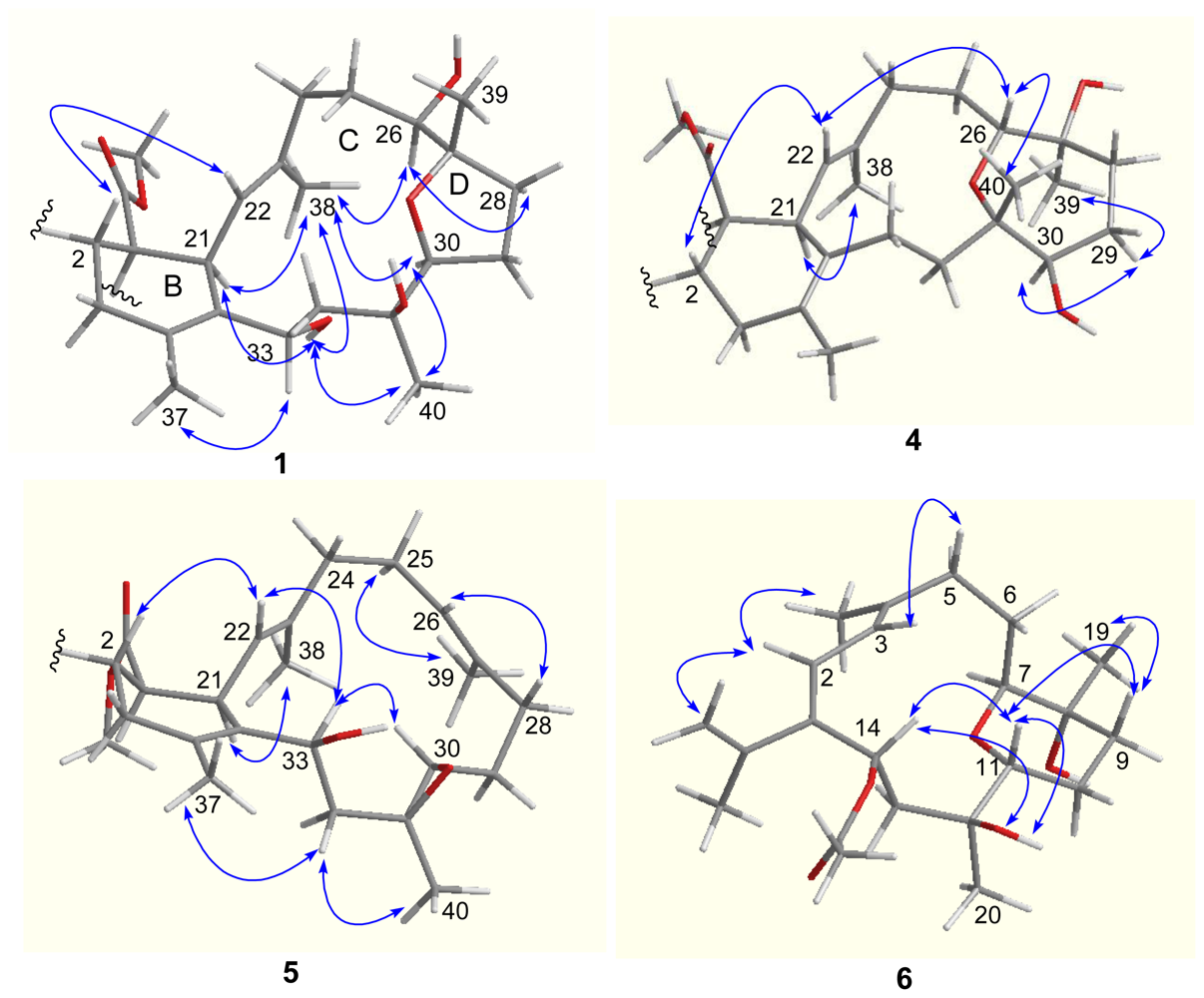
| Position | 1 | 2 | 3 | 4 | 5 |
|---|---|---|---|---|---|
| 1 | 49.4 (Cq) | 49.4 (Cq) | 49.3 (Cq) | 48.5 (Cq) | 49.9 (Cq) |
| 2 | 43.9 (CH) | 43.8 (CH) | 43.9 (CH) | 45.9 (CH) | 44.0 (CH) |
| 3 | 213.8 (Cq) | 213.8 (Cq) | 213.8 (Cq) | 212.1 (Cq) | 213.9 (Cq) |
| 4 | 54.1 (CH2) | 54.0 (CH2) | 54.1 (CH2) | 50.4 (CH2) | 53.9 (CH2) |
| 5 | 27.3 (CH) | 27.3 (CH) | 27.3 (CH) | 26.9 (CH) | 27.3 (CH) |
| 6 | 38.0 (CH2) | 37.9 (CH2) | 38.0 (CH2) | 37.0 (CH2) | 38.0 (CH2) |
| 7 | 26.0 (CH2) | 26.0 (CH2) | 26.0 (CH2) | 24.4 (CH2) | 26.0 (CH2) |
| 8 | 34.1 (CH2) | 34.0 (CH2) | 34.1 (CH2) | 32.7 (CH2) | 34.1 (CH2) |
| 9 | 47.6 (CH) | 47.5 (CH) | 47.7 (CH) | 46.4 (CH) | 47.8 (CH) |
| 10 | 213.3 (Cq) | 213.3 (Cq) | 213.3 (Cq) | 213.6 (Cq) | 213.1 (Cq) |
| 11 | 30.9 (CH2) | 31.0 (CH2) | 30.9 (CH2) | 34.9 (CH2) | 30.9 (CH2) |
| 12 | 51.8 (CH) | 51.8 (CH) | 51.7 (CH) | 51.7 (CH) | 51.3 (CH) |
| 13 | 210.0 (Cq) | 209.9 (Cq) | 209.9 (Cq) | 211.5 (Cq) | 210.3 (Cq) |
| 14 | 45.9 (CH2) | 45.8 (CH2) | 45.9 (CH2) | 46.9 (CH2) | 46.2 (CH2) |
| 15 | 28.7 (CH) | 28.7 (CH) | 28.7 (CH) | 28.8 (CH) | 28.9 (CH) |
| 16 | 21.5 (CH3) | 21.5 (CH3) | 21.5 (CH3) | 21.2 (CH3) | 21.5 (CH3) |
| 17 | 17.6 (CH3) | 17.6 (CH3) | 17.6 (CH3) | 18.3 (CH3) | 17.7 (CH3) |
| 18 | 17.6 (CH3) | 17.6 (CH3) | 17.6 (CH3) | 17.1 (CH3) | 17.7 (CH3) |
| 19 | 22.9 (CH3) | 22.9 (CH3) | 22.9 (CH3) | 22.1 (CH3) | 22.8 (CH3) |
| 20 | 174.8 (Cq) | 174.7 (Cq) | 174.7 (Cq) | 173.8 (Cq) | 174.6 (Cq) |
| 21 | 43.7 (CH) | 43.5 (CH) | 43.7 (CH) | 50.2 (CH) | 43.2 (CH) |
| 22 | 128.6 (CH) | 129.4 (CH) | 130.0 (CH) | 124.6 (CH) | 124.6 (CH) |
| 23 | 137.8 (Cq) | 136.9 (Cq) | 136.2 (Cq) | 135.9 (Cq) | 133.7 (Cq) |
| 24 | 33.5 (CH2) | 32.9 (CH2) | 34.0 (CH2) | 36.7 (CH2) | 37.3 (CH2) |
| 25 | 33.1 (CH2) | 31.1 (CH2) | 34.8 (CH2) | 24.4 (CH2) | 23.8 (CH2) |
| 26 | 70.4 (CH) | 73.5 (CH) | 65.1 (CH) | 77.8 (CH) | 125.1 (CH) |
| 27 | 85.1 (Cq) | 83.2 (Cq) | 84.8 (Cq) | 74.7 (Cq) | 132.8 (Cq) |
| 28 | 35.9 (CH2) | 36.1 (CH2) | 36.8 (CH2) | 39.7 (CH2) | 35.6 (CH2) |
| 29 | 28.1 (CH2) | 28.0 (CH2) | 28.0 (CH2) | 27.6 (CH2) | 25.1 (CH2) |
| 30 | 89.2 (CH) | 89.6 (CH) | 90.1 (CH) | 80.0 (CH) | 59.8 (CH) |
| 31 | 73.6 (Cq) | 73.3 (Cq) | 73.2 (Cq) | 80.5 (Cq) | 59.7 (Cq) |
| 32 | 40.8 (CH2) | 41.5 (CH2) | 40.7 (CH2) | 41.5 (CH2) | 39.6 (CH2) |
| 33 | 64.0 (CH) | 64.1 (CH) | 63.9 (CH) | 24.4 (CH2) | 62.6 (CH) |
| 34 | 135.6 (Cq) | 135.2 (Cq) | 135.2 (Cq) | 132.8 (Cq) | 133.1 (Cq) |
| 35 | 123.2 (Cq) | 123.2 (Cq) | 123.6 (Cq) | 123.3 (Cq) | 127.6 (Cq) |
| 36 | 32.5 (CH2) | 32.6 (CH2) | 32.5 (CH2) | 32.1 (CH2) | 32.6 (CH2) |
| 37 | 18.4 (CH3) | 18.4 (CH3) | 18.3 (CH3) | 20.1 (CH3) | 18.7 (CH3) |
| 38 | 20.6 (CH3) | 20.3 (CH3) | 20.2 (CH3) | 17.9 (CH3) | 17.6 (CH3) |
| 39 | 20.3 (CH3) | 20.8 (CH3) | 20.3 (CH3) | 25.4 (CH3) | 17.3 (CH3) |
| 40 | 23.5 (CH3) | 23.4 (CH3) | 23.5 (CH3) | 15.5 (CH3) | 20.3 (CH3) |
| OMe | 51.2 (CH3) | 51.1 (CH3) | 51.2 (CH3) | 51.0 (CH3) | 51.2 (CH3) |
| Ac | 171.3 (Cq) | ||||
| 21.4 (CH3) |
| Position | 1 | 2 | 3 | 4 | 5 |
|---|---|---|---|---|---|
| 2 | 3.47 t (8.3) | 3.45 m | 3.47 t (8.8) | 3.49 m | 3.59 t (8.1) |
| 4 | 3.09 dd (10.3, 19.6) | 3.09 dd (10.3, 19.5) | 3.09 dd (10.3, 20.0) | 2.61 dd (6.9, 17.8) | 3.06 dd (10.3, 19.6) |
| 2.40 d (19.6) | 2.40 d (19.5) | 2.40 d (20.0) | 2.32 m | 2.44 d (19.6) | |
| 5 | 1.68 m | 1.69 m | 1.67 m | 1.85 m | 1.63 m |
| 6 | 1.01 m | 1.01 m | 0.97 m | 1.03 m | 0.96 m |
| 1.07 m | 1.06 m | 0.87 m | 1.00 m | ||
| 7 | 1.10 m | 1.12 m | 1.08 m | 0.95 m | 0.94 m |
| 1.01 m | 1.00 m | 1.00 m | 1.07 m | ||
| 8 | 1.43 m | 1.49 m | 1.43 m | 1.40 m | 1.38 m |
| 1.36 m | 1.35 m | 1.35 m | 1.40 m | ||
| 9 | 2.33 m | 2.33 m | 2.32 m | 2.38 m | 2.30 m |
| 11 | 2.80 dd (10.3, 16.6) | 2.80 m | 2.80 dd (10.5, 16.9) | 2.73 dd (2.7, 16.9) | 1.88 d (20.0) |
| 1.84 d (16.6) | 1.84 m | 1.83 d (16.9) | 2.15 dd (2.4, 16.9) | 2.82 dd (10.5, 20.0) | |
| 12 | 2.88 m | 2.87 m | 2.89 m | 2.89 m | 2.89 brd (10.5) |
| 14 | 2.86 d (19.3) | 2.88 d (19.3) | 2.86 d (19.6) | 2.96 m | 2.94 d (19.6) |
| 2.72 d (19.3) | 2.74 d (19.3) | 2.72 d (19.6) | 2.90 m | 2.69 d (19.6) | |
| 15 | 2.33 m | 2.34 m | 2.32 m | 2.07 m | 2.22 m |
| 16 | 0.95 d (6.7) | 0.96 d (6.6) | 0.95 d (6.9) | 0.91 d (6.9) | 0.92 d (6.6) |
| 17 | 0.63 d (6.7) | 0.63 d (6.6) | 0.63 d (6.9) | 0.71 d (6.9) | 0.61 d (6.6) |
| 18 | 1.05 d (6.9) | 1.05 d (6.6) | 1.05 d (7.0) | 1.07 d (7.1) | 1.04 d (6.9) |
| 19 | 0.81 d (6.9) | 0.81 d (6.6) | 0.81 d (6.9) | 0.81 d (6.9) | 0.81 d (6.6) |
| 21 | 3.38 d (10.5) | 3.47 d (10.3) | 3.38 d (10.3) | 3.23 m | 3.20 d (11.0) |
| 22 | 4.78 d (10.5) | 4.85 d (10.3) | 4.87 d (10.3) | 4.94 d (11.5) | 4.70 d (11.0) |
| 24 | 2.02 m | 2.13 m | 2.18 m | 2.38 m | 2.07 m |
| 1.91 m | 1.51 m | 1.96 m | 1.87 m | 1.92 m | |
| 25 | 1.65 m | 1.80 m | 2.08 m | 1.82 m | 2.22 m |
| 1.36 m | 1.60 m | 1.66 m | 1.63 m | 2.00 m | |
| 26 | 3.01 brd (10.5) | 4.56 d (10.8) | 3.55 d (11.7) | 3.39 m | 4.72 br |
| 28 | 2.25 m | 1.71 m | 2.32 m | 1.58 m | 2.10 m |
| 1.38 m | 1.45 m | 1.62 m | 1.42 m | 1.91 m | |
| 29 | 1.67 m | 1.71 m | 1.75 m | 1.85 m | 1.76 m |
| 1.36 m | 1.43 m | 1.40 m | 1.31 m | 1.47 m | |
| 30 | 3.77 dd (5.3, 10.5) | 3.80 dd (5.1, 10.5) | 3.86 dd (5.4, 11.5) | 2.91 m | 2.30 m |
| 32 | 1.65 m | 1.71 m | 1.62 m | 2.07 m | 1.96 dd (10.0, 15.4) |
| 1.09 m | 1.13 m | 1.09 m | 1.04 m | 1.70 dd (2.0, 15.4) | |
| 33 | 4.80 m | 4.82 m | 4.82 brt | 2.10 m | 4.26 dd (2.0, 10.0) |
| 1.83 m | |||||
| 36 | 2.18 dd (8.3, 19.3) | 2.21 dd (8.8, 19.3) | 2.21 m | 2.47 m | 2.12 d (8.1) |
| 2.06 dd (8.8, 19.3) | 2.03 m | 2.05 m | 1.98 dd (5.4, 18.3) | 2.12 d (8.1) | |
| 37 | 1.66 s | 1.67 s | 1.67 s | 1.54 s | 1.63 s |
| 38 | 1.65 s | 1.64 s | 1.67 s | 1.69 s | 1.53 s |
| 39 | 0.99 s | 1.09 s | 1.19 s | 0.98 s | 1.52 s |
| 40 | 0.97 s | 0.99 s | 0.99 s | 0.92 s | 1.06 s |
| OMe | 3.42 s | 3.39 s | 3.42 s | 3.39 s | 3.40 s |
| Ac | 2.06 s | ||||
| OH-26 | 4.53 br | ||||
| OH-27 | 4.18 s | ||||
| OH-30 | 4.49 br | ||||
| OH-31 | 3.98 s | 4.14 s | 4.21 s | ||
| OH-33 | 4.40 br | 4.45 br | 4.46 br | 4.63 br |
© 2010 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Yan, P.; Deng, Z.; Ofwegen, L.v.; Proksch, P.; Lin, W. Lobophytones O–T, New Biscembranoids and Cembranoid from Soft Coral Lobophytum pauciflorum. Mar. Drugs 2010, 8, 2837-2848. https://doi.org/10.3390/md8112848
Yan P, Deng Z, Ofwegen Lv, Proksch P, Lin W. Lobophytones O–T, New Biscembranoids and Cembranoid from Soft Coral Lobophytum pauciflorum. Marine Drugs. 2010; 8(11):2837-2848. https://doi.org/10.3390/md8112848
Chicago/Turabian StyleYan, Pengcheng, Zhiwei Deng, Leen van Ofwegen, Peter Proksch, and Wenhan Lin. 2010. "Lobophytones O–T, New Biscembranoids and Cembranoid from Soft Coral Lobophytum pauciflorum" Marine Drugs 8, no. 11: 2837-2848. https://doi.org/10.3390/md8112848







