Biodiversity of Actinobacteria from the South Pacific and the Assessment of Streptomyces Chemical Diversity with Metabolic Profiling
Abstract
:1. Introduction
2. Materials and Methods
2.1. Marine Sampling
2.2. Sponge Sample Processing
2.3. Isolation of Actinobacteria
2.3.1. Isolation Media
2.3.2. Isolation Methods
2.4. Molecular Identification and Phylogenetic Analysis
2.5. Detection of PKS and NRPS Genes
2.6. Antibacterial Activity Screening
2.7. LC-HRMS Analysis
3. Results and Discussion
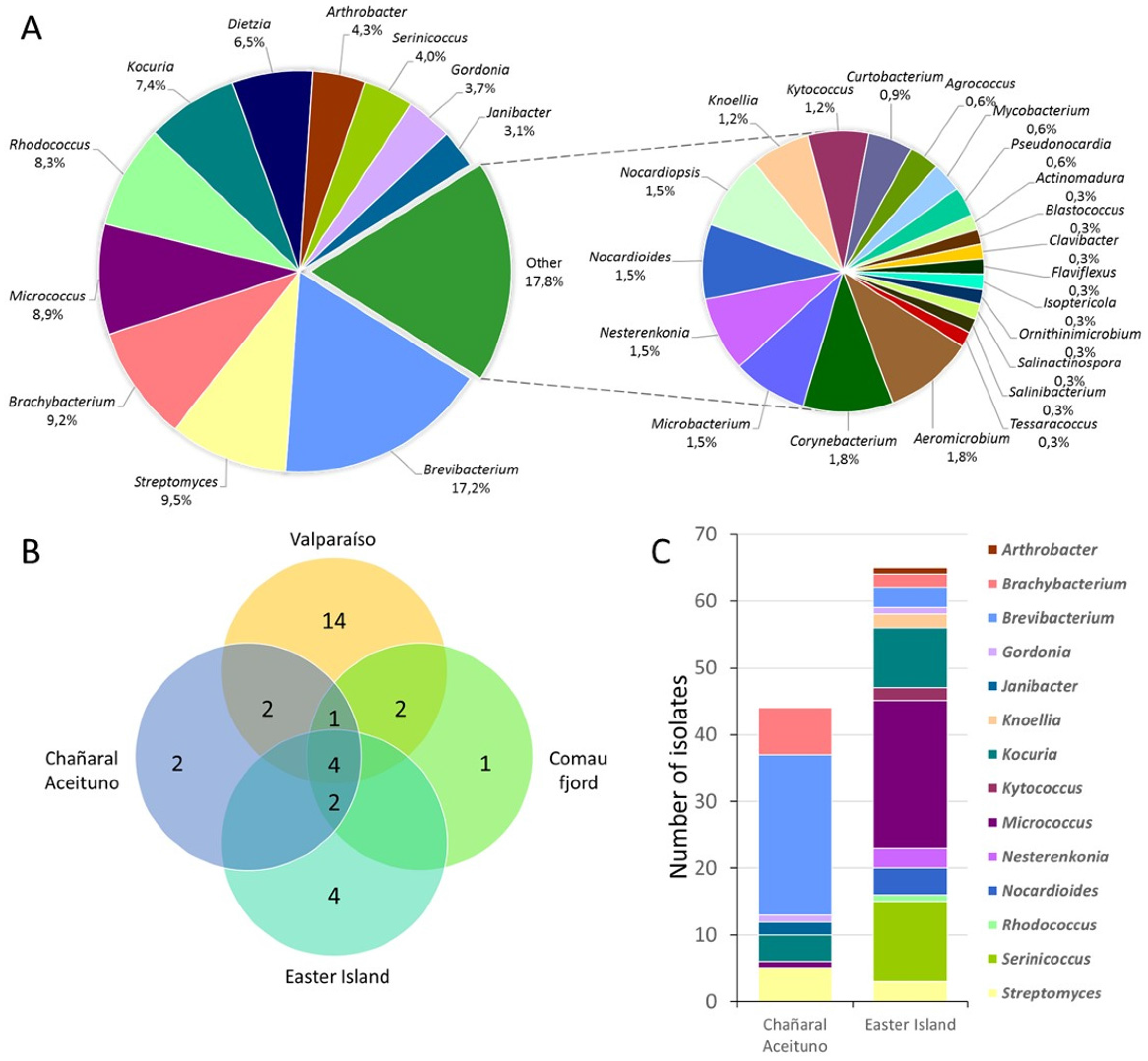
3.1. Biodiversity of Marine Actinobacteria
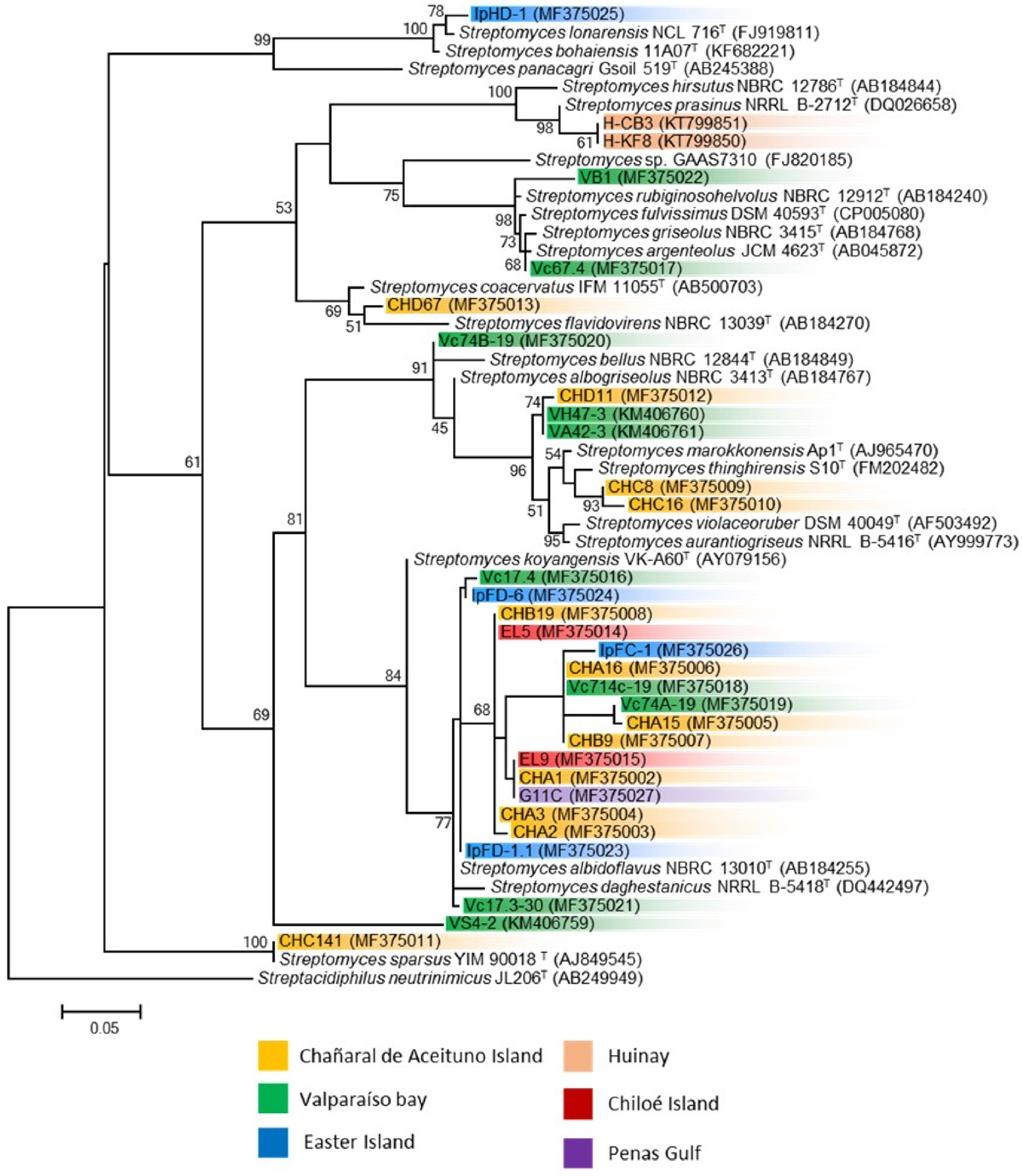
3.2. Phylogenetic Analysis of Marine Streptomyces and Presence of Biosynthetic Genes
3.3. Antimicrobial Potential of Marine Streptomyces
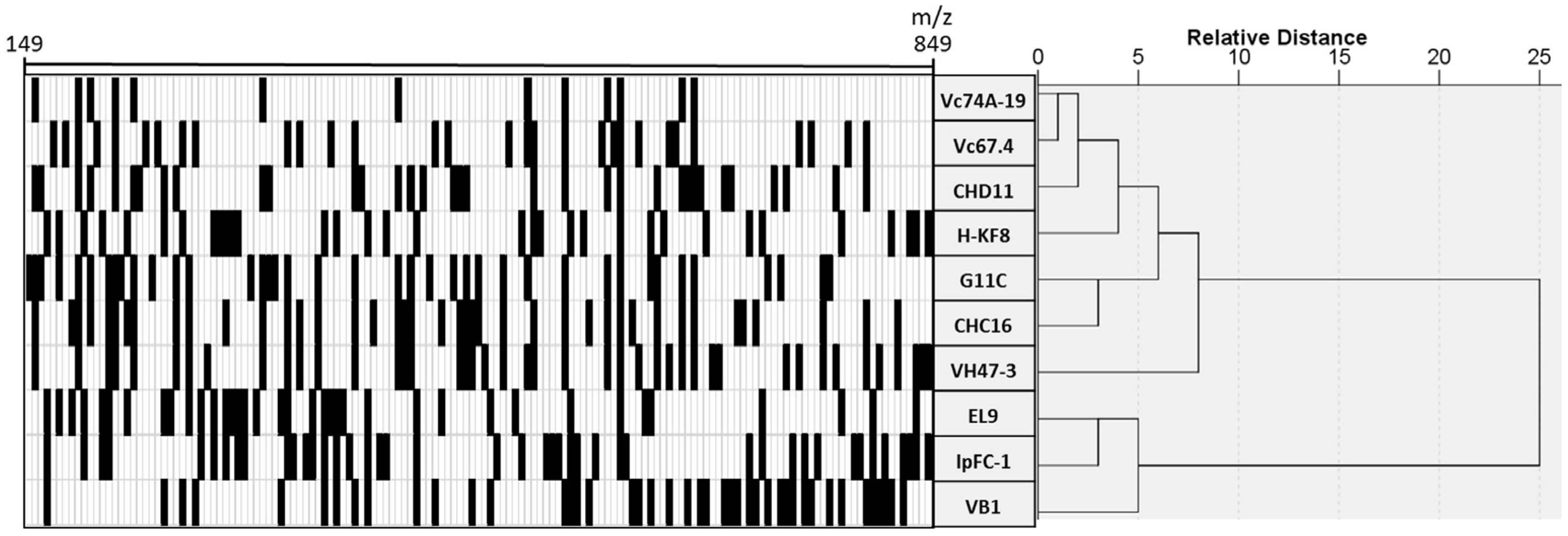
3.4. Chemical Profiling of Selected Marine Streptomyces
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- World Health Organization (WHO). Antimicrobial Resistance; World Health Organization: Geneva, Switzerland, 2014; Volume 61.
- Genilloud, O. The re-emerging role of microbial natural products in antibiotic discovery. Antonie Van Leeuwenhoek 2014, 106, 173–188. [Google Scholar] [CrossRef] [PubMed]
- Newman, D.J.; Cragg, G.M. Natural products as sources of new drugs from 1981 to 2014. J. Nat. Prod. 2016, 79, 629–661. [Google Scholar] [CrossRef] [PubMed]
- Joint, I.; Mühling, M.; Querellou, J. Culturing marine bacteria—An essential prerequisite for biodiscovery: Minireview. Microb. Biotechnol. 2010, 3, 564–575. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Williams, P.G. Panning for chemical gold: Marine bacteria as a source of new therapeutics. Trends Biotechnol. 2009, 27, 45–52. [Google Scholar] [CrossRef] [PubMed]
- Mincer, T.J.; Jensen, P.R.; Kauffman, C.A.; Fenical, W. Widespread and persistent populations of a major new marine actinomycete taxon in ocean sediments. Society 2002, 68, 5005–5011. [Google Scholar] [CrossRef]
- Prieto, A.; Villarreal, L.; Forschner, S.; Bull, A.; Stach, J.; Smith, D.; Rowley, D.; Jensen, P. Targeted search for actinomycetes from near-shore and deep sea marine sediments. FEMS 2014, 84, 510–518. [Google Scholar] [CrossRef]
- Bull, A.T.; Stach, J.E.; Ward, A.C.; Goodfellow, M. Marine actinobacteria: Perspectives, challenges, future directions. Antonie Van Leeuwenhoek 2005, 87, 65–79. [Google Scholar] [CrossRef] [PubMed]
- Undabarrena, A.; Ugalde, J.A.; Seeger, M.; Cámara, B. Genomic data mining of the marine actinobacteria Streptomyces sp. H-KF8 unveils insights into multi-stress related genes and metabolic pathways involved in antimicrobial synthesis. PeerJ 2017, 5, e2912. [Google Scholar] [CrossRef] [PubMed]
- Dalisay, D.S.; Williams, D.E.; Wang, X.L.; Centko, R.; Chen, J.; Raymond, J. Marine sediment-derived Streptomyces bacteria from British Columbia, Canada are a promising microbiota resource for the discovery of antimicrobial natural products. PLoS ONE 2013, 8, 1–14. [Google Scholar] [CrossRef] [PubMed]
- Donia, M.; Hamann, M.T. Marine natural products and their potential applications as anti-infective agents. Lancet Infect. Dis. 2003, 3, 338–348. [Google Scholar] [CrossRef]
- Duncan, K.; Haltli, B.; Gill, K.A.; Kerr, R.G. Bioprospecting from marine sediments of New Brunswick, Canada: Exploring the relationship between total bacterial diversity and actinobacteria diversity. Mar. Drugs 2014, 12, 899–925. [Google Scholar] [CrossRef] [PubMed]
- Penesyan, A.; Kjelleberg, S.; Egan, S. Development of novel drugs from marine surface associated microorganisms. Mar. Drugs 2010, 8, 438–459. [Google Scholar] [CrossRef] [PubMed]
- Vicente, J.; Stewart, A.; Song, B.; Hill, R.T.; Wright, J.L. Biodiversity of actinomycetes associated with caribbean sponges and their potential for natural product discovery. Mar. Biotechnol. 2013, 15, 413–424. [Google Scholar] [CrossRef] [PubMed]
- Graça, A.P.; Bondoso, J.; Gaspar, H.; Xavier, J.R.; Monteiro, M.C.; De La Cruz, M.; Oves-Costales, D.; Vicente, F.; Lage, O.M. Antimicrobial activity of heterotrophic bacterial communities from the marine sponge Erylus discophorus (Astrophorida, Geodiidae). PLoS ONE 2013, 8. [Google Scholar] [CrossRef]
- Montalvo, N.F.; Mohamed, N.M.; Enticknap, J.J.; Hill, R.T. Novel actinobacteria from marine sponges. Antonie Van Leeuwenhoek 2005, 87, 29–36. [Google Scholar] [CrossRef] [PubMed]
- Thomas, T.R.A.; Kavlekar, D.P.; LokaBharathi, P.A. Marine drugs from sponge-microbe association—A review. Mar. Drugs 2010, 8, 1417–1468. [Google Scholar] [CrossRef] [PubMed]
- Khan, S.T.; Musarrat, J.; Alkhedhairy, A.A.; Kazuo, S. Diversity of bacteria and polyketide synthase associated with marine sponge Haliclona sp. Ann. Microbiol. 2014, 64, 199–207. [Google Scholar] [CrossRef]
- Blunt, J.W.; Copp, B.R.; Munro, M.H.G.; Northcote, P.T.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2016, 27, 165–237. [Google Scholar] [CrossRef] [PubMed]
- Blunt, J.W.; Copp, B.R.; Munro, M.H.G.; Northcote, P.T.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2015, 22, 15–61. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Graça, A.P.; Viana, F.; Bondoso, J.; Correia, M.I.; Gomes, L.; Humanes, M.; Reis, A.; Xavier, J.R.; Gaspar, H.; Lage, O.M. The antimicrobial activity of heterotrophic bacteria isolated from the marine sponge Erylus deficiens (Astrophorida, Geodiidae). Front. Microbiol. 2015, 6, 1–14. [Google Scholar] [CrossRef]
- Selvin, J.; Joseph, S.; Asha, K.R.T.; Manjusha, W.A.; Sangeetha, V.S.; Jayaseema, D.M.; Antony, M.C.; Denslin Vinitha, A.J. Antibacterial potential of antagonistic Streptomyces sp. isolated from marine sponge Dendrilla nigra. FEMS Microbiol. Ecol. 2004, 50, 117–122. [Google Scholar] [CrossRef] [PubMed]
- Thakur, N.L.; Muller, W.E.G. Biotechnological potential of marine sponges. Curr. Sci. 2004, 86, 1506–1512. [Google Scholar] [CrossRef]
- Undabarrena, A.; Beltrametti, F.; Claverias, F.P.; Gonzalez, M.; Moore, E.R.B.; Seeger, M.; Camara, B. Exploring the diversity and antimicrobial potential of marine actinobacteria from the comau fjord in Northern Patagonia, Chile. Front. Microbiol. 2016, 7, 1–16. [Google Scholar] [CrossRef] [PubMed]
- Claverías, F.P.; Undabarrena, A.; González, M.; Seeger, M.; Cámara, B. Culturable diversity and antimicrobial activity of actinobacteria from marine sediments in Valparaíso bay, Chile. Front. Microbiol. 2015, 6, 1–11. [Google Scholar] [CrossRef] [PubMed]
- Undabarrena, A.; Ugalde, J.A.; Castro-Nallar, E.; Seeger, M.; Cámara, B. Genome sequence of Streptomyces sp. H-KF8, a marine Actinobacterium isolated from a northern chilean patagonian fjord. Genome Announc. Am. Soc. Microbiol. 2017, 5, 8–9. [Google Scholar] [CrossRef] [PubMed]
- Gregersen, T. Rapid method for distinction of gram-negative from gram-positive Bacteria. Eur. J. Appl. Microbiol. Biotechnol. 1978, 5, 123–127. [Google Scholar] [CrossRef]
- Stach, J.E.M.; Maldonado, L.A.; Ward, A.C.; Goodfellow, M.; Bull, A.T. New primers for the class Actinobacteria: Application to marine and terrestrial environments. Environ. Microbiol. 2003, 5, 828–841. [Google Scholar] [CrossRef] [PubMed]
- Lane, D. 16S/23S rRNA Sequencing. In Nucleic Acid Techniques in Bacterial Systematics; Stackebrandt, E., Goodfellow, M., Eds.; Wiley: Hoboken, NJ, USA; New York, NY, USA, 1991; pp. 115–175. [Google Scholar]
- Guindon, S.; Dufayard, J.F.; Lefort, V.; Anisimova, M.; Hordijk, W.; Gascuel, O. New algorithms and methods to estimate maximum-likelihood phylogenies: Assessing the performance of PhyML 3.0. Syst. Biol. 2010, 59, 307–321. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783. [Google Scholar] [CrossRef] [PubMed]
- Posada, D. jModelTest: Phylogenetic model averaging. Mol. Biol. Evol. 2008, 25, 1253–1256. [Google Scholar] [CrossRef] [PubMed]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular evolutionary genetics analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Gontang, E.A.; Gaude, S.P.; Fenical, W.; Jensen, P.R. Sequence-based analysis of secondary-metabolite biosynthesis in marine actinobacteria. Appl. Environ. Microbiol. 2010, 76, 2487–2499. [Google Scholar] [CrossRef] [PubMed]
- Ayuso, A.; Clark, D.; González, I.; Salazar, O.; Anderson, A.; Genilloud, O. A novel actinomycete strain de-replication approach based on the diversity of polyketide synthase and nonribosomal peptide synthetase biosynthetic pathways. Appl. Microbiol. Biotechnol. 2005, 67, 795–806. [Google Scholar] [CrossRef] [PubMed]
- Ayuso-Sacido, A.; Genilloud, O. New PCR primers for the screening of NRPS and PKS-I systems in actinomycetes: Detection and distribution of these biosynthetic gene sequences in major taxonomic groups. Microb. Ecol. 2005, 49, 10–24. [Google Scholar] [CrossRef] [PubMed]
- Haber, M.; Ilan, M. Diversity and antibacterial activity of bacteria cultured from Mediterranean Axinella spp. sponges. J. Appl. Microbiol. 2013, 116, 519–532. [Google Scholar] [CrossRef] [PubMed]
- Lyman, J.; Fleming, R. Composition of seawater. J. Mar. Res. 1940, 3, 134–146. [Google Scholar]
- De la Cruz, M.; González, I.; Parish, C.A.; Onishi, R.; Tormo, J.R.; Martín, J.; Peláez, F.; Zink, D.; El Aouad, N.; Reyes, F.; et al. Production of ramoplanin and ramoplanin analogs by actinomycetes. Front. Microbiol. 2017. [Google Scholar] [CrossRef] [PubMed]
- Chambers, M.; Maclean, B.; Burke, R.; Amodei, D.; Ruderman, D.; Neumann, S.; Gatto, L.; Fischer, B.; Agus, D.; MacCoss, M.; et al. A cross-platform toolkit for mass spectrometry and proteomics. Nat. Biotechnol. 2012. [Google Scholar] [CrossRef] [PubMed]
- Pluskal, T.; Castillo, S.; Villar-briones, A.; Ore, M. MZmine 2: Modular framework for processing, visualizing, and analyzing mass spectrometry-based molecular profile data. BMC Bioinform. 2010, 11, 395. [Google Scholar] [CrossRef] [PubMed]
- Forner, D.; Berrué, F.; Correa, H.; Duncan, K.; Kerr, R.G. Chemical dereplication of marine actinomycetes by liquid chromatography-high resolution mass spectrometry profiling and statistical analysis. Anal. Chim. Acta 2013, 805, 70–79. [Google Scholar] [CrossRef] [PubMed]
- Ward, J. Hierarchical grouping to optimize an objective function. J. Am. Stat. Assoc. 1963, 58, 236–244. [Google Scholar] [CrossRef]
- Gagnon, K.; Chadwell, C.D.; Norabuena, E. Measuring the onset of locking in the Peru-Chile trench with GPS and acoustic measurements. Nature 2005, 434, 205–208. [Google Scholar] [CrossRef] [PubMed]
- Baltz, R.H. Marcel Faber Roundtable: Is our antibiotic pipeline unproductive because of starvation, constipation or lack of inspiration? J. Ind. Microbiol. Biotechnol. 2006, 33, 507–513. [Google Scholar] [CrossRef] [PubMed]
- Lazzarini, A.; Cavaletti, L.; Toppo, G.; Marinelli, F. Rare genera of actinomycetes as potential producers of new antibiotics. Antonie Van Leeuwenhoek 2000, 78, 399–405. [Google Scholar] [CrossRef] [PubMed]
- Jose, P.A.; Jebakumar, S.R.D. The evolving role of natural products in drug discovery. Nat. Rev. Drug Discov. 2013, 4, 206–220. [Google Scholar] [CrossRef]
- Fenical, W. Chemical studies of marine bacteria: Developing a new resource. Chem. Rev. 1993, 93, 1673–1683. [Google Scholar] [CrossRef]
- Fenical, W.; Jensen, P.R. Developing a new resource for drug discovery: Marine actinomycete bacteria. Nat. Chem. Biol. 2006, 2, 666–673. [Google Scholar] [CrossRef] [PubMed]
- Magarvey, N.A.; Keller, J.M.; Bernan, V.; Dworkin, M.; Sherman, D.H.; Magarvey, N.A.; Keller, J.M.; Bernan, V.; Dworkin, M.; Sherman, D.H. Isolation and characterization of novel marine-derived actinomycete taxa rich in bioactive metabolites. Appl. Environ. Microbiol. 2004, 70, 7520–7529. [Google Scholar] [CrossRef] [PubMed]
- Jensen, P.R.; Mincer, T.J.; Williams, P.G.; Fenical, W. Marine actinomycete diversity and natural product discovery. Antonie Van Leeuwenhoek 2005, 87, 43–48. [Google Scholar] [CrossRef] [PubMed]
- Bredholdt, H.; Galatenko, O.A.; Engelhardt, K.; Fjærvik, E.; Terekhova, L.P.; Zotchev, S.B. Rare actinomycete bacteria from the shallow water sediments of the Trondheim fjord, Norway: Isolation, diversity and biological activity. Environ. Microbiol. 2007, 9, 2756–2764. [Google Scholar] [CrossRef] [PubMed]
- Gontang, E.A.; Fenical, W.; Jensen, P.R. Phylogenetic diversity of gram-positive bacteria cultured from marine sediments. Appl. Environ. Microbiol. 2007, 73, 3272–3282. [Google Scholar] [CrossRef] [PubMed]
- León, J.; Liza, L.; Soto, I. Actinomycetes bioactivos de sedimento marino de la costa central del Perú. Rev. Peru. Biol. 2007, 14, 259–270. [Google Scholar] [CrossRef]
- Maldonado, L.A.; Stach, J.E.M.; Ward, A.C.; Bull, A.T.; Goodfellow, M. Characterisation of micromonosporae from aquatic environments using molecular taxonomic methods. Antonie van Leeuwenhoek 2008, 94, 289–298. [Google Scholar] [CrossRef] [PubMed]
- Yuan, M.; Yu, Y.; Li, H.R.; Dong, N.; Zhang, X.H. Phylogenetic diversity and biological activity of actinobacteria isolated from the Chukchi Shelf marine sediments in the Arctic Ocean. Mar. Drugs 2014, 12, 1281–1297. [Google Scholar] [CrossRef] [PubMed]
- Hentschel, U.; Schmid, M.; Wagner, M.; Fieseler, L.; Gernert, C.; Hacker, J. Isolation and phylogenetic analysis of bacteria with antimicrobial activities from the Mediterranean sponges Aplysina aerophoba and Aplysina cavernicola. FEMS Microbiol. Ecol. 2001, 35, 305–312. [Google Scholar] [CrossRef] [PubMed]
- Abdelmohsen, U.R.; Pimentel-Elardo, S.M.; Hanora, A.; Radwan, M.; Abou-El-Ela, S.H.; Ahmed, S.; Hentschel, U. Isolation, phylogenetic analysis and anti-infective activity screening of marine sponge-associated actinomycetes. Mar. Drugs 2010, 8, 399–412. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kim, T.K.; Garson, M.J.; Fuerst, J.A. Marine actinomycetes related to the “Salinospora” group from the Great Barrier Reef sponge Pseudoceratina clavata. Environ. Microbiol. 2005, 7. [Google Scholar] [CrossRef] [PubMed]
- Li, Z.Y.; Liu, Y. Marine sponge Craniella austrialiensis-associated bacterial diversity revelation based on 16S rDNA library and biologically active Actinomycetes screening, phylogenetic analysis. Lett. Appl. Microbiol. 2006, 43, 410–416. [Google Scholar] [CrossRef] [PubMed]
- Jiang, S.; Sun, W.; Chen, M.; Dai, S.; Zhang, L.; Liu, Y.; Lee, K.J.; Li, X. Diversity of culturable actinobacteria isolated from marine sponge Haliclona sp. Antonie Van Leeuwenhoek 2007, 92, 405–416. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Lee, Y.K.; Zhang, W.; Lee, H.K. Culturable actinobacteria from the marine sponge Hymeniacidon perleve: Isolation and phylogenetic diversity by 16S rRNA gene-RFLP analysis. Antonie Van Leeuwenhoek 2006, 90, 159–169. [Google Scholar] [CrossRef] [PubMed]
- Radjasa, O.K.; Sabdono, A.; Junaidi, J.; Zocchi, E. Richness of secondary metabolites-producing marine bacteria associated with sponge Haliclona sp. Int. J. Pharmacol. 2007, 3, 275–279. [Google Scholar]
- Abdelmohsen, U.R.; Yang, C.; Horn, H.; Hajjar, D.; Ravasi, T.; Hentschel, U. Actinomycetes from red sea sponges: Sources for chemical and phylogenetic diversity. Mar. Drugs 2014, 12, 2771–2789. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Schneemann, I.; Nagel, K.; Kajahn, I.; Labes, A.; Wiese, J.; Imhoff, J.F. Comprehensive investigation of marine actinobacteria associated with the sponge Halichondria panicea. Appl. Environ. Microbiol. 2010, 76, 3702–3714. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Matobole, R.M.; van Zyl, L.J.; Parker-Nance, S.; Davies-Coleman, M.T.; Trindade, M. Antibacterial activities of bacteria isolated from the marine sponges Isodictya compressa and Higginsia bidentifera collected from Algoa Bay, South Africa. Mar. Drugs 2017, 15, 47. [Google Scholar] [CrossRef] [PubMed]
- Sarmiento-vizcaíno, A.; Braña, A.F.; González, V.; Nava, H.; Molina, A.; Llera, E.; Fiedler, H.; Rico, J.M. Atmospheric dispersal of bioactive Streptomyces albidoflavus strains among terrestrial and marine environments. Microb. Ecol. 2016, 71, 375–386. [Google Scholar] [CrossRef] [PubMed]
- Miller, S.I. Antibiotic resistance and regulation of the gram-negative bacterial outer membrane barrier by host innate immune molecules. MBio 2016, 7, e01541-16. [Google Scholar] [CrossRef] [PubMed]
- Delcour, A.H. Outer membrane permeability and antibiotic resistance. Biochim Biophys. Acta 2009, 1794, 808–816. [Google Scholar] [CrossRef] [PubMed]
- Vandermolen, K.M.; Raja, H.A.; El-Elimat, T.; Oberlies, N.H. Evaluation of culture media for the production of secondary metabolites in a natural products screening program. AMB Express 2013, 3, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Bode, H.B.; Bethe, B.; Höfs, R.; Zeeck, A. Big effects from small changes: Possible ways to explore nature’s chemical diversity. ChemBioChem 2002, 3, 619–627. [Google Scholar] [CrossRef]
- Sánchez, S.; Chávez, A.; Forero, A.; García-Huante, Y.; Romero, A.; Sánchez, M.; Rocha, D.; Sánchez, B.; Avalos, M.; Guzmán-Trampe, S.; et al. Carbon source regulation of antibiotic production. J. Antibiot. 2010, 63, 442–459. [Google Scholar] [CrossRef] [PubMed]
- Patin, N.V.; Duncan, K.R.; Dorrestein, P.C.; Jensen, P.R. Competitive strategies differentiate closely related species of marine actinobacteria. ISME J. 2015, 1–13. [Google Scholar] [CrossRef] [PubMed]
- Kiranmayi, M.U.; Sudhakar, P.; Sreenivasulu, K.; Vijayalakshmi, M. Optimization of culturing conditions for improved production of bioactive metabolites by Pseudonocardia sp. VUK-10. Mycobiology 2011, 39, 174–181. [Google Scholar] [CrossRef] [PubMed]
- Huang, R.; Yi, X.; Zhou, Y.; Su, X.; Peng, Y.; Gao, C. An Update on 2,5-Diketopiperazines from Marine Organisms. Mar. Drugs 2014, 12, 6213–6235. [Google Scholar] [CrossRef] [PubMed]
- Komiyama, T.; Matsuzawa, Y.; Oki, T.; Inui, T.; Takahashi, Y.; Naganawa, H.; Takeuchi, T.; Umezawa, H. Baumycins, new antitumor antibiotics related to daunomycin. J. Antibiot. 1977, 30, 619–621. [Google Scholar] [CrossRef]
- Xu, Z.; Schenk, A.; Hertweck, C. Molecular analysis of the benastatin biosynthetic pathway and genetic engineering of altered fatty acid-polyketide hybrids. J. Am. Chem. Soc. 2007, 129, 6022–6030. [Google Scholar] [CrossRef] [PubMed]
- Takada, K.; Ninomiya, A.; Naruse, M.; Sun, Y.; Miyazaki, M.; Nogi, Y.; Okada, S.; Matsunaga, S. Surugamides A–E, Cyclic Octapeptides with Four D-Amino Acid Residues, from a Marine Streptomyces sp.: LC–MS-Aided Inspection of Partial Hydrolysates for the Distinction of d- and l-Amino Acid Residues in the Sequence. J. Org. Chem. 2013, 50, 3–7. [Google Scholar]
- Wang, X.; Shaaban, K.; Elshahawi, S.; Ponomareva, L.; Sunkara, M.; Copley, G.; Hower, J.; Morris, A.; Kharel, M.; Thorson, J. Mullinamides A and B, new cyclopeptides produced by the Ruth Mullins coal mine fire isolate Streptomyces sp. RM-27-46. J. Antibiot. 2016, 23, 1780–1789. [Google Scholar] [CrossRef] [PubMed]
- Macintyre, L.; Zhang, T.; Viegelmann, C.; Martinez, I.J.; Cheng, C.; Dowdells, C.; Abdelmohsen, U.R.; Gernert, C.; Hentschel, U.; Edrada-Ebel, R.A. Metabolomic tools for secondary metabolite discovery from marine microbial symbionts. Mar. Drugs 2014, 12, 3416–3448. [Google Scholar] [CrossRef] [PubMed] [Green Version]
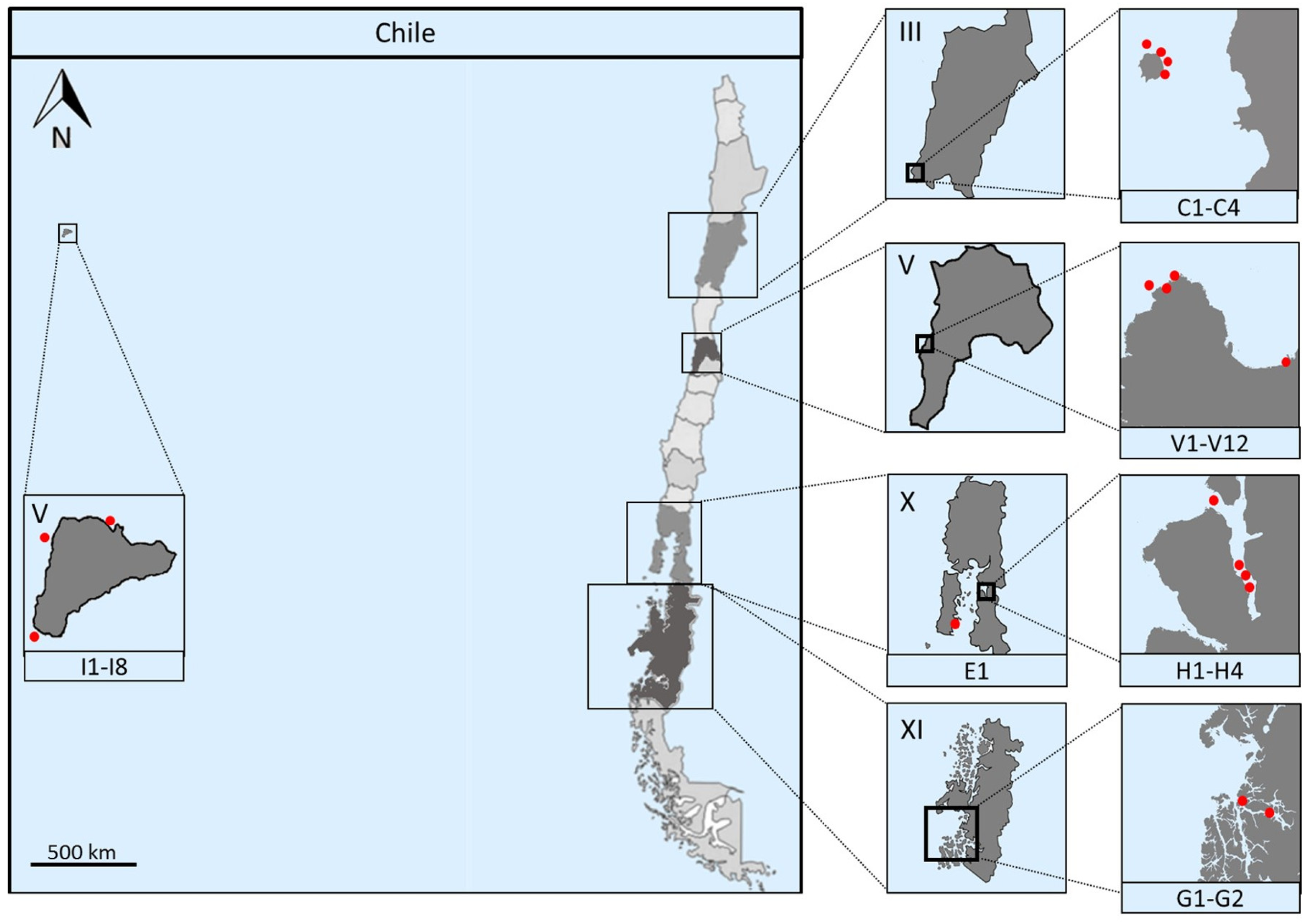
| Sample | Sampling Site | Isolation Method | Isolation Media | Temperature |
|---|---|---|---|---|
| C1 | Chañaral de Aceituno Island | Direct serially diluted (10ࢤ4) Stamping technique capillary technique | MA ISP2 prepared with ASW actinomycete isolation agar sea urchin agar marine sediment agar | 20 °C |
| C2 | ||||
| C3 | MA ISP2 prepared with ASW actinomycete isolation agar M3 sponge agar | |||
| C4 | ||||
| V1 | Valparaíso | Direct | Actinomycete isolation agar actinomycete isolation agar prepared with ASW | 4 °C 20 °C 30 °C |
| V2 | ||||
| V3 | ||||
| V4 | Direct | ISP2 prepared with ASW | 30 °C | |
| V5 | Direct heat treatment | Sea urchin agar marine sediment agar | 20 °C 30 °C | |
| V6 | ||||
| I1 | Easter Island | Direct serially diluted (10−4) heat treatment serially diluted (10−4) with heat treatment | MA actinomycete isolation agar M1 prepared with ASW marine sediment agar | 20 °C |
| I2 | MA actinomycete isolation agar M1 prepared with ASW marine sediment agar sponge agar | |||
| I3 | ||||
| I4 | MA actinomycete isolation agar M1 prepared with ASW marine sediment agar | |||
| I5 | ||||
| I6 | MA actinomycete isolation agar M1 prepared with ASW marine sediment agar sponge agar | |||
| I7 | ||||
| I8 | ||||
| E1 | Chiloé Island | Direct heat treatment | Sea urchin agar marine sediment agar | 20 °C 30 °C |
| G1 | Penas Gulf | Direct heat treatment | MA actinomycete isolation agar ISP2 prepared with ASW | 4 °C 20 °C |
| G2 |
| Strain | Closest Type Strain (Accession Number) (% Identity) | Sample Information | Biosynthetic Genes | Reference | |||
|---|---|---|---|---|---|---|---|
| Sampling Site | Sample Type | PKS I | PKS II | NRPS | |||
| CHA1 | S. albidoflavus NBRC 13010T (AB184255) (99.51) | Chañaral de Aceituno Island | Marine sediment | - | - | + | This study |
| CHA2 | S. albidoflavus NBRC 13010T (AB184255) (99.23) | - | - | + | This study | ||
| CHA3 | S. albidoflavus NBRC 13010T (AB184255) (99.65) | - | + | + | This study | ||
| CHA15 | S. albidoflavus NBRC 13010T (AB184255) (99.40) | - | + | - | This study | ||
| CHA16 | S. albidoflavus NBRC 13010T (AB184255) (99.62) | - | + | + | This study | ||
| CHB9 | S. albidoflavus NBRC 13010T (AB184255) (99.65) | - | - | + | This study | ||
| CHB19 | S. albidoflavus NBRC 13010T (AB184255) (99.44) | - | - | + | This study | ||
| CHC8 | S. thinghirensis S10T (FM202482) (99.59) | Marine sponge | - | - | + | This study | |
| CHC16 | S. thinghirensis S10T (FM202482) (99.52) | - | - | + | This study | ||
| CHC141 | S. sparsus YIM 90018T (AJ849545) (100) | + | + | + | This study | ||
| CHD11 | S. aurantiogriseus NRRL B-5416T (AY999773) (99.38) | + | + | + | This study | ||
| CHD67 | S. coacervatus AS-0823T (AB500703) (99.59) | + | - | + | This study | ||
| VA42-3 | S. aurantiogriseus NRRL B-5416T (AY999773) (99.37) | Valparaíso | Marine sediment | - | - | + | [25] |
| VH47-3 | S. aurantiogriseus NRRL B-5416T (AY999773) (99.37) | - | - | + | [25] | ||
| VS4-2 | S. fabae T66T (KM229360) (98.32) | - | + | - | [25] | ||
| Vc17.3-30 | S. albidoflavus NBRC 13010T (AB184255) (99.93) | - | - | - | This study | ||
| Vc17.4 | S. exfoliatus NBRC 13475T (AB184868) (99.79) | - | - | + | This study | ||
| Vc67-4 | S. argenteolus AS 4.1693T (D44272) (99.93) | + | + | + | This study | ||
| Vc714c-19 | S. albidoflavus NBRC 13010T (AB184255) (99.59) | - | - | + | This study | ||
| Vc74A-19 | S. albidoflavus NBRC 13010T (AB184255) (99.45) | - | - | + | This study | ||
| Vc74B-19 | S. albogriseolus NBRC 3413T (AB184315) (99.72) | - | - | - | This study | ||
| VB1 | S. pratensis ch24T (JQ824046) (99.86) | - | - | - | This study | ||
| IpFC-1 | S. albidoflavus NBRC 13010T (AB184255) (99.50) | Easter Island | Marine sponge | + | - | + | This study |
| IpFD-1.1 | S. albidoflavus NBRC 13010T (AB184255) (99.93) | + | - | + | This study | ||
| IpFD-6 | S. albidoflavus NBRC 13010T (AB184255) (99.79) | + | + | + | This study | ||
| IpHD-1 | S. lonarensis NCL 716T (FJ919811) (99.73) | Marine sediment | + | - | + | This study | |
| EL5 | S. albidoflavus NBRC 13010T (AB184255) (99.65) | Chiloé Island | Sea Urchin | - | - | - | This study |
| EL9 | S. albidoflavus NBRC 13010T (AB184255) (99.71) | - | + | - | This study | ||
| H-CB3 | S. prasinus NRRL B-2712T (DQ026658) (99.86) | Huinay | Marine sediment | - | - | + | [24] |
| H-KF8 | S. prasinus NRRL B-2712T (DQ026658) (99.93) | - | + | + | [24] | ||
| G11C | S. albidoflavus NBRC 13010T (AB184255) (99.64) | Penas Gulf | Marine sediment | - | - | + | This study |
| Strains | STAU | LIMO | PSAU | SAEN | ESCO | |||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| TSA | MA | ISP1 | ISP2 | TSA | MA | ISP1 | ISP2 | TSA | MA | ISP1 | ISP2 | TSA | MA | ISP1 | ISP2 | TSA | MA | ISP1 | ISP2 | |
| CHA1 | - | +/-- | +/- | + | - | +/- | - | - | - | - | - | - | - | - | - | - | + | - | - | - |
| CHA2 | -- | +/-- | - | + | - | +/- | - | + | - | - | - | - | - | - | - | - | + | - | - | - |
| CHA3 | -- | -- | - | + | - | - | - | + | - | - | - | - | - | - | - | - | ++ | - | - | - |
| CHA15 | -- | +/-- | +/- | + | - | +/- | - | +/- | - | - | - | - | - | - | - | - | + | - | - | - |
| CHA16 | -- | +/-- | +/- | + | - | +/- | +/- | +/- | - | - | - | - | - | - | - | - | + | - | - | - |
| CHB9 | -- | +/-- | +/- | + | - | +/- | +/- | - | - | - | - | - | - | - | - | - | + | - | - | - |
| CHB19 | -- | +/-- | +/- | + | - | +/- | +/- | + | - | - | - | - | - | - | - | - | + | - | - | - |
| CHC8 | + | +++ | +++ | + | - | + | - | - | - | + | - | - | - | ++ | - | - | + | +++ | +++ | - |
| CHC16 | +++ | + | +++ | + | - | - | - | - | +/- | - | - | - | + | + | - | - | ++ | + | +++ | - |
| CHC141 | -- | -- | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| CHD11 | -- | -- | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| VA42-3 | -- | -- | - | - | - | - | - | - | - | - | - | - | - | - | - | +/- | - | - | - | - |
| VH47-3 | -- | -- | - | - | - | - | - | + | - | - | - | - | - | - | - | +/- | - | - | - | - |
| VS4-2 | -- | +/-- | - | +/- | - | + | - | +/- | - | - | - | - | - | - | - | +/- | - | - | - | +/- |
| Vc17.3-30 | -- | +/-- | - | + | - | +/- | - | - | - | - | - | - | - | - | - | - | +/- | - | - | +/- |
| Vc17.4 | +/-- | +/-- | - | - | - | +/- | - | - | - | - | - | - | - | - | - | - | - | - | - | - |
| Vc67-4 | +/-- | -- | +/- | +++ | - | - | +/- | +++ | - | - | - | +/- | +/- | - | - | +/- | - | - | - | +/- |
| Vc714c-19 | -- | +/- | - | + | - | +/- | - | + | - | - | - | - | - | - | - | - | - | - | - | - |
| Vc74A-19 | +/-- | - | - | - | - | - | - | +++ | - | - | - | - | - | - | - | - | +/- | - | - | - |
| Vc74B-19 | -- | - | - | + | - | - | - | +/- | - | - | - | - | - | - | - | - | - | - | - | - |
| VB1 | +++ | ++ | + | +++ | +++ | +++ | ++ | +++ | ++ | - | - | - | - | +/- | - | - | +++ | - | +/- | - |
| IpFC-1 | +/-- | + | +/- | + | - | - | - | +/- | - | - | - | - | - | - | - | - | +++ | - | - | - |
| IpFD-1.1 | -- | + | +/- | + | - | +/- | - | - | - | - | - | - | - | - | - | - | +++ | - | - | - |
| IpFD-6 | +/-- | + | +/- | +/- | - | - | +/- | +/- | - | - | - | - | - | - | - | - | ++ | - | - | - |
| IpHD-1 | -- | - | - | ++ | - | +/- | - | +/- | - | - | - | - | - | - | - | - | ++ | - | - | - |
| EL5 | + | +/- | +/- | + | +/- | +/- | +/- | +/- | +/- | - | - | +/- | - | - | - | - | +/- | - | - | +/- |
| EL9 | + | +/- | +/- | + | - | - | +/- | +/- | - | - | - | +/- | +/- | - | - | - | + | - | - | - |
| H-CB3 | +++ | +++ | +++ | +++ | +/- | +++ | +++ | +/- | - | - | - | +/- | - | - | +++ | - | ++ | ++ | +++ | + |
| H-KF8 | +++ | +++ | +++ | +++ | +/- | +++ | +++ | +/- | - | - | - | +/- | - | - | +++ | - | + | ++ | +++ | + |
| G11C | -- | +/- | +/- | + | - | +/- | - | +/- | +++ | - | - | - | - | - | - | - | +++ | - | ++ | - |
| Streptomyces Strain | Model Bacteria | ||||
|---|---|---|---|---|---|
| STAU | LIMO | PSAU | SAEN | ESCO | |
| CHC16 | + | - | - | - | - |
| CHC8 | - | - | - | - | - |
| CHA3 | - | - | - | - | - |
| CHD11 | + | - | - | - | - |
| EL9 | + | - | - | - | - |
| H-CB3 | - | - | - | - | - |
| H-KF8 | + | - | - | - | - |
| VH47-3 | + | + | - | - | - |
| Vc67-4 | + | + | - | - | - |
| Vc74A-19 | + | - | - | - | - |
| VB1 | + | + | - | - | - |
| IpFC-1 | + | - | - | - | - |
| IpHD-1 | - | - | - | - | - |
| G11C | + | - | - | - | - |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cumsille, A.; Undabarrena, A.; González, V.; Claverías, F.; Rojas, C.; Cámara, B. Biodiversity of Actinobacteria from the South Pacific and the Assessment of Streptomyces Chemical Diversity with Metabolic Profiling. Mar. Drugs 2017, 15, 286. https://doi.org/10.3390/md15090286
Cumsille A, Undabarrena A, González V, Claverías F, Rojas C, Cámara B. Biodiversity of Actinobacteria from the South Pacific and the Assessment of Streptomyces Chemical Diversity with Metabolic Profiling. Marine Drugs. 2017; 15(9):286. https://doi.org/10.3390/md15090286
Chicago/Turabian StyleCumsille, Andrés, Agustina Undabarrena, Valentina González, Fernanda Claverías, Claudia Rojas, and Beatriz Cámara. 2017. "Biodiversity of Actinobacteria from the South Pacific and the Assessment of Streptomyces Chemical Diversity with Metabolic Profiling" Marine Drugs 15, no. 9: 286. https://doi.org/10.3390/md15090286





