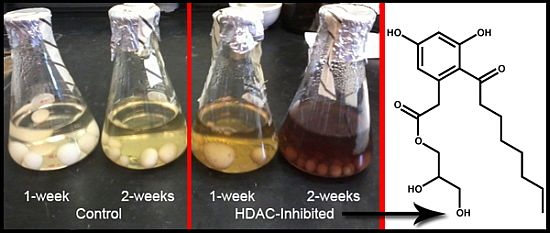
Epigenetic Tailoring for the Production of Anti-Infective Cytosporones from the Marine Fungus Leucostoma persoonii
Abstract
:1. Introduction

2. Results and Discussion
2.1. Fungal Isolation and Identification
2.2. Malaria Screening
2.3. Antibiotic Screening
2.4. Isolation of Active Compounds
2.5. Bioactivity of Cytosporones B, C and E
| Compound | MRSA (µM) | A549 (µM) | |||
|---|---|---|---|---|---|
| MIC | MBC90 | MBEC90 | IC50 | IC90 | |
| Cytosporone B (1) | 78 | 93 | 110 | 170 | 190 |
| Cytosporone C (2) | >358 | NT a | NT | 690 | 840 |
| Cytosporone E (3) | 72 | 45 | 39 | 280 | 440 |
2.6. Epigenetics Studies
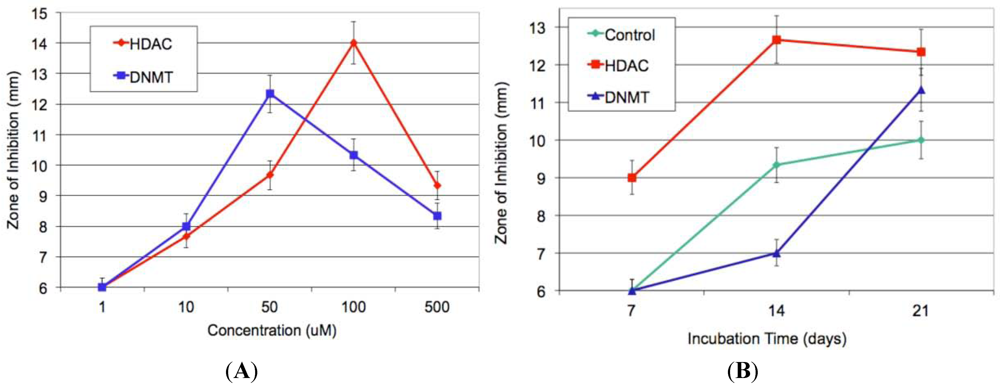
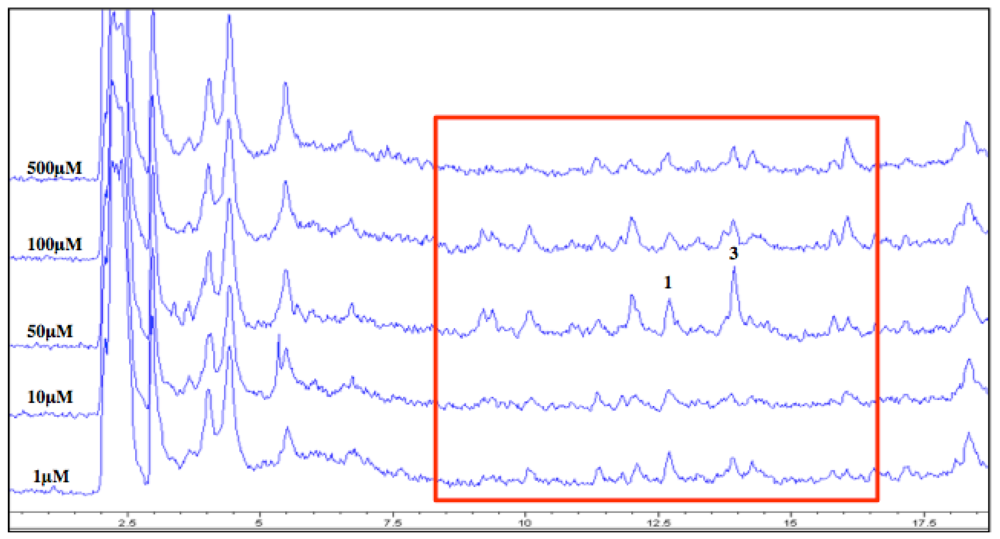
2.7. Increase in Production and Biological Evaluation of Compounds
| Compound | Control (mg) | HDAC Inhibited (mg) | Increase (%) |
|---|---|---|---|
| Cytosporone B (1) | 2.4 | 8.7 | 330 |
| Cytosporone C (2) | 0.7 | 3.9 | 510 |
| Cytosporone E (3) | 3.6 | 32.1 | 820 |
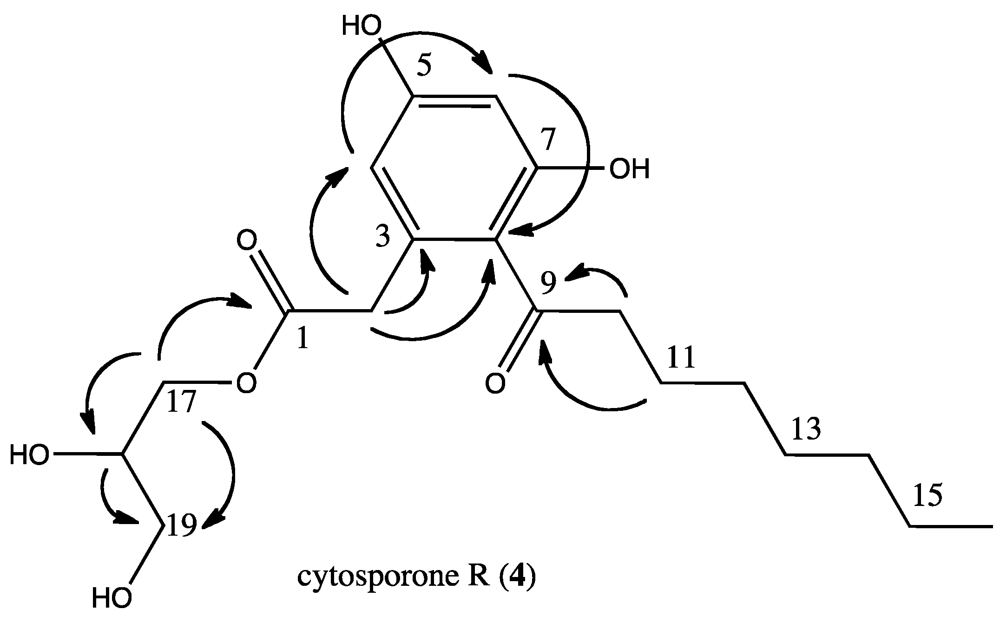
2.8. Chemical Analysis of Additional Isolate
| Cytosporone R | Cytosporone B | |||||
|---|---|---|---|---|---|---|
| Position | δC, mult b | δH (mult, J in Hz) | HMBC | Position | δC | δH |
| 1 | 173.5, C | 2, 17 | 1 | 171.6 | ||
| 2 | 40.5, CH2 | 3.61, s | 4 | 2 | 40.1 | 3.67 |
| 3 | 137.1, C | 2 | 3 | 137.0 | ||
| 4 | 112.0, CH | 6.22, d (2.2) | 2, 6 | 4 | 102.5c | 6.37 |
| 5 | 161.6, C | 4, 6 | 5 | 160.7 | ||
| 6 | 102.8, CH | 6.27, d (2.2) | 4 | 6 | 111.8c | 6.31 |
| 7 | 160.0, C | 6 | 7 | 159.7 | ||
| 8 | 121.1, C | 4, 6 | 8 | 120.8 | ||
| 9 | 209.1, C | 10, 11 | 9 | 206.3 | ||
| 10 | 45.2, CH2 | 2.87, m | 11, 12, 13 | 10 | 44.4 | 2.90 |
| 11 | 25.6, CH2 | 1.60, br, m | 10, 12, 13 | 11 | 25.0 | 1.63 |
| 12 | 30.5, CH2 | 1.25–1.36, br, m | 10, 11, 13, 14 | 12 | 30.0 | 1.26–1.32 |
| 13 | 30.3, CH2 | 1.25–1.36, br, m | 11, 12, 14, 15 | 13 | 29.9 | 1.26–1.32 |
| 14 | 32.9, CH2 | 1.25–1.36, br, m | 12, 13, 15, 16 | 14 | 32.5 | 1.26–1.32 |
| 15 | 23.7, CH2 | 1.25–1.36, br, m | 13, 14, 16 | 15 | 23.3 | 1.26–1.32 |
| 16 | 14.4, CH3 | 0.90, t | 14, 15 | 16 | 14.3 | 0.88 |
| 17 | 66.8, CH2 | 4.16, dd (11.3, 5.4) | 18, 19 | 17 | 61.0 | 4.08 |
| 4.08, dd (11.3, 5.4) | 18, 19 | |||||
| 18 | 71.1, CH | 3.84, quin (5.4) | 17, 19 | 18 | 14.5 | |
| 19 | 64.1, CH2 | 3.54, m | 17, 18 | |||
3. Experimental Section
3.1. General Experimental Procedures
3.2. Biological Material
3.3. DNA Extraction
3.4. PCR Parameters
3.5. Gel Step and Isolation of DNA
3.6. Fungus Identification
3.7. Initial Culture and Screening
3.8. Malaria Assay
3.9. Methicillin-Resistant Staphylococcus Aureus Strain
3.10. Disk Diffusion Assay
3.11. Natural Product Extraction and Isolation
3.12. Microtiter MIC and MBC Determination Assays
3.13. Biofilm Assay
3.14. Cytotoxicity Assay
3.15. Epigenetics Studies
3.6. Cytosporone R (4)
4. Conclusions
Acknowledgments
References
- Mumby, P.J.; Edwards, A.J.; Arias-González, J.E.; Lindeman, K.C.; Blackwell, P.G.; Gall, A.; Gorczynska, M.I.; Harborne, A.R.; Pescod, C.L.; Renken, H.; et al. Mangroves enhance the biomass of coral reef fish communities in the Caribbean. Nature 2004, 427, 533–536. [Google Scholar]
- Liu, D.; Li, X.M.; Meng, L.; Li, C.S.; Gao, S.S.; Shang, Z.; Proksch, P.; Huang, C.G.; Wang, B.G. Asperolides A–C, tetranorlabdane diterpenoids from the marine alga-derived endophytic fungus Aspergillus wentii EN-48. J. Nat. Prod. 2011, 74, 1787–1791. [Google Scholar] [CrossRef]
- Cao, S.; Clardy, J. New naphthoquinones and a new delta-lactone produced by endophytic fungi from Costa Rica. Tetrahedron Lett. 2011, 52, 2206–2208. [Google Scholar]
- Huang, H.; Feng, X.; Xiao, Z.; Liu, L.; Li, H.; Ma, L.; Lu, Y.; Ju, J.; She, Z.; Lin, Y. Azaphilones and p-terphenyls from the mangrove endophytic fungus Penicillium chermesinum (ZH4-E2) isolated from the South China Sea. J. Nat. Prod. 2011, 74, 997–1002. [Google Scholar]
- Crawford, J.M.; Clardy, J. Bacterial symbionts and natural products. Chem. Commun. 2011, 47, 7559–7566. [Google Scholar]
- Cai, S.; Sun, S.; Zhou, H.; Kong, X.; Zhu, T.; Li, D.; Gu, Q. Prenylated polyhydroxy-p-terphenyls from Aspergillus taichungensis ZHN-7-07. J. Nat. Prod. 2011, 74, 1106–1110. [Google Scholar] [CrossRef]
- Cao, S.; Ross, L.; Tamayo, G.; Clardy, J. Asterogynins: Secondary metabolites from a costa rican endophytic fungus. Org. Lett. 2010, 12, 4661–4663. [Google Scholar]
- Lu, Z.; Zhu, H.; Fu, P.; Wang, Y.; Zhang, Z.; Lin, H.; Liu, P.; Zhuang, Y.; Hong, K.; Zhu, W. Cytotoxic polyphenols from the marine-derived fungus Penicillium expansum. J. Nat. Prod. 2010, 73, 911–914. [Google Scholar]
- Wen, L.; Cai, X.; Xu, F.; She, Z.; Chan, W.L.; Vrijmoed, L.L.P.; Jones, E.B.G.; Lin, Y. Three metabolites from the mangrove endophytic fungus Sporothrix sp. (#4335) from the South China Sea. J. Org. Chem. 2009, 74, 1093–1098. [Google Scholar]
- Vervoort, H.C.; Draskovic, M.; Crews, P. Azonazine, a novel dipeptide from a hawaiian marine sediment-derived fungus, Aspergillus insulicola. Org. Lett. 2011, 13, 410–413. [Google Scholar]
- Wang, X.; Filho, J.G.S.; Hoover, A.R.; King, J.B.; Ellis, T.K.; Powell, D.R.; Cichewicz, R.H. Chemical epigenetics alters the secondary metabolite composition of guttate excreted by an Atlantic-Forest-derived Penicillium citreonigrum. J. Nat. Prod. 2010, 73, 942–948. [Google Scholar] [CrossRef]
- Scherlach, K.; Hertweck, C. Triggering cryptic natural product biosynthesis in microorganisms. Org. Biomol. Chem. 2009, 7, 1753–1760. [Google Scholar]
- Schwab, E.K.; Bok, J.W.; Tribus, M.; Galehr, J.; Graessle, S.; Keller, N.P. Histone deacetylase activity regulates chemical diversity in Aspergillus. Eukaryot. Cell 2007, 6, 1656–1664. [Google Scholar] [CrossRef]
- Brady, S.F.; Wagenaar, M.M.; Singh, M.P.; Janso, J.E.; Clardy, J. The cytosporones, new octaketide antibiotics isolated from an endophytic fungus. Org. Lett. 2000, 2, 4043–4046. [Google Scholar]
- Paranagama, P.A.; Wijeratne, E.M.; Gunatilaka, A.A. Uncovering biosynthetic potential of plant-associated fungi: Effect of culture conditions on metabolite production by Paraphaeosphaeria quadriseptata and Chaetomium chiversii. J. Nat. Prod. 2007, 70, 1939–1945. [Google Scholar] [CrossRef]
- Xu, J.; Kjer, J.; Sendker, J.; Wray, V.; Guan, H.; Edrada, R.; Müller, W.E.; Bayer, M.; Lin, W.; Wu, J.; Proksch, P. Cytosporones, coumarins, and an alkaloid from the endophytic fungus Pestalotiopsis sp. isolated from the Chinese mangrove plant Rhizophoramucronata. Bioorg. Med. Chem. 2009, 17, 7362–7367. [Google Scholar]
- Abrei, L.M.; Phipps, R.K.; Pfenning, L.H.; Gotfredsen, C.H.; Takahashi, J.A.; Larsen, T.O. ChemInform abstract: Cytosporones O (I), P (IIa) and Q (IIb) from an endophytic Cytospora sp. Tetrahedron Lett. 2010, 41, 1803–1805. [Google Scholar]
- Trager, W.; Jenson, J.B. Cultivation of malarial parasites. Nature 1978, 273, 621–622. [Google Scholar]
- Zhan, Y.; Du, X.; Chen, H.; Liu, J.; Zhao, B.; Huang, D.; Li, G.; Xu, Q.; Zhang, M.; Weimer, B.C.; et al. Cytosporone B is an agonist for nuclear orphan receptor Nur77. Nat. Chem. Biol. 2008, 4, 548–556. [Google Scholar] [CrossRef]
- Borneman, J.; Hartin, R.J. PCR primers that amplify fungal rRNA genes from environmental samples. Appl. Environ. Microbiol. 2000, 66, 4356–4360. [Google Scholar]
- Lebar, M.D.; Hahn, K.N.; Mutka, T.; Maignan, P.; van Olphen, A.; Kyle, D.E.; McClintock, J.B.; Amsler, C.D.; Baker, B.J. CNS and antimalarial activity of synthetic meridianin and psammopemmin analogs. Bioorg. Med. Chem. 2011, 19, 5756–5762. [Google Scholar]
- Burda, W.N.; Fields, K.B.; Gill, J.B.; Burt, R.; Shepherd, M.; Zhang, X.P.; Shaw, L.N. Neutral metallated and meso-substituted porphyrins as antimicrobial agents against gram-positive pathogens. Eur.J. Clin. Microbiol. Infect. Dis. 2012, 31, 327–335. [Google Scholar]
- Beenken, K.E.; Blevins, J.S.; Smeltzer, M.S. Mutation of sarA in Staphylococcus aureus limits biofilm formation. Infect. Immun. 2003, 71, 4206–4211. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Beau, J.; Mahid, N.; Burda, W.N.; Harrington, L.; Shaw, L.N.; Mutka, T.; Kyle, D.E.; Barisic, B.; Van Olphen, A.; Baker, B.J. Epigenetic Tailoring for the Production of Anti-Infective Cytosporones from the Marine Fungus Leucostoma persoonii. Mar. Drugs 2012, 10, 762-774. https://doi.org/10.3390/md10040762
Beau J, Mahid N, Burda WN, Harrington L, Shaw LN, Mutka T, Kyle DE, Barisic B, Van Olphen A, Baker BJ. Epigenetic Tailoring for the Production of Anti-Infective Cytosporones from the Marine Fungus Leucostoma persoonii. Marine Drugs. 2012; 10(4):762-774. https://doi.org/10.3390/md10040762
Chicago/Turabian StyleBeau, Jeremy, Nida Mahid, Whittney N. Burda, Lacey Harrington, Lindsey N. Shaw, Tina Mutka, Dennis E. Kyle, Betty Barisic, Alberto Van Olphen, and Bill J. Baker. 2012. "Epigenetic Tailoring for the Production of Anti-Infective Cytosporones from the Marine Fungus Leucostoma persoonii" Marine Drugs 10, no. 4: 762-774. https://doi.org/10.3390/md10040762
APA StyleBeau, J., Mahid, N., Burda, W. N., Harrington, L., Shaw, L. N., Mutka, T., Kyle, D. E., Barisic, B., Van Olphen, A., & Baker, B. J. (2012). Epigenetic Tailoring for the Production of Anti-Infective Cytosporones from the Marine Fungus Leucostoma persoonii. Marine Drugs, 10(4), 762-774. https://doi.org/10.3390/md10040762