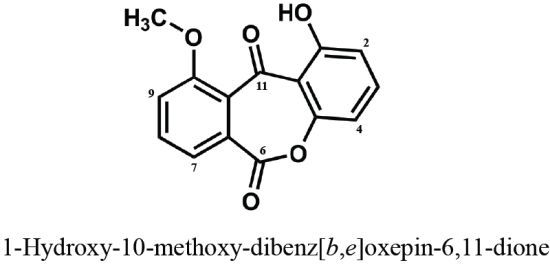
A New Dibenz[b,e]oxepine Derivative, 1-Hydroxy-10-methoxy-dibenz[b,e]oxepin-6,11-dione, from a Marine-Derived Fungus, Beauveria bassiana TPU942
Abstract
:1. Introduction
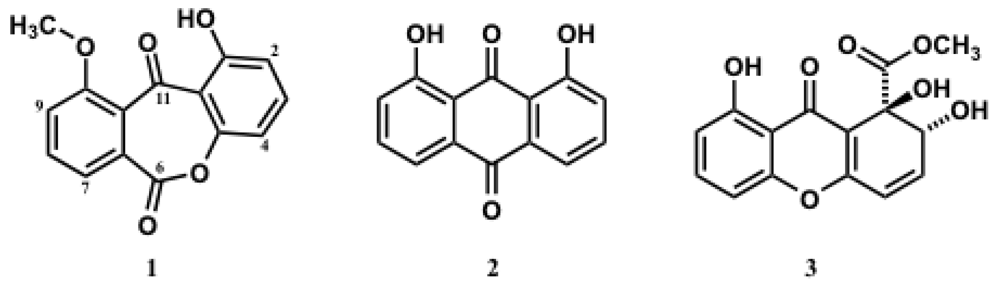
2. Results and Discussion
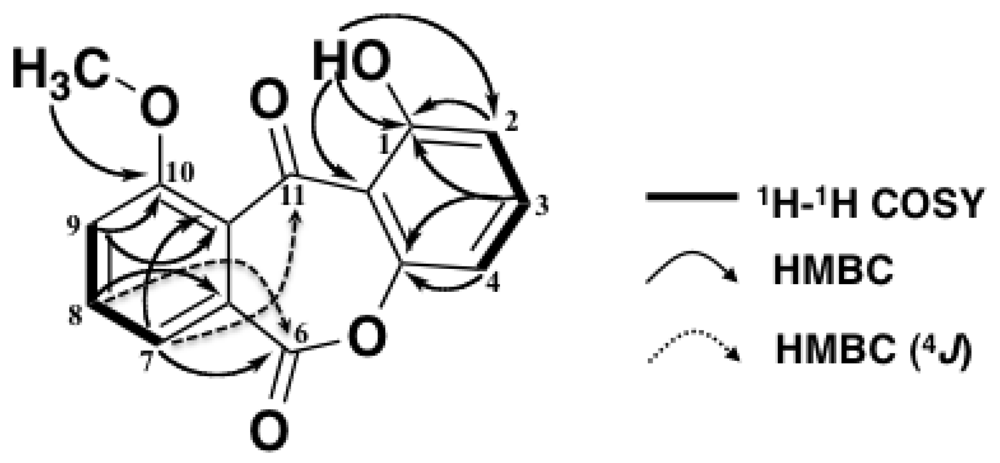
| No. | δC | δH (J in Hz) | HMBC |
|---|---|---|---|
| 1 | 161.8 | - | 1, 2, 11a |
| 2 | 111.0 | 6.82 dd (8.3, 1.0) | 1, 4 |
| 3 | 137.1 | 7.61 t (8.5) | 1, 4a |
| 4 | 106.9 | 6.94 dd (8.3, 1.0) | 2, 4a |
| 4a | 155.8 | - | - |
| 5 | |||
| 6 | 156.1 | - | - |
| 6a | 133.7 | - | - |
| 7 | 119.5 | 7.56 dd (8.3, 1.0) | 6, 9, 10a |
| 8 | 135.0 | 7.77 dd (8.5, 7.1) | 6, 6a |
| 9 | 122.7 | 7.33 dd (7.3, 1.0) | 10, 10a |
| 10 | 169.5 | - | - |
| 10a | 117.5 | - | - |
| 11 | 181.0 | - | - |
| 11a | 109.0 | - | - |
| 1-OH | - | 12.2 s | 1, 2, 11a |
| 10-OCH3 | 53.1 | 4.03 s | 10 |

| Compound | IC50 (μM) | Inhibition zone (mm) | ||||||||||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Cytotoxicity | Antimicrobial activity (10 μg/disk) | |||||||||||||||||||
| HCT-15 | Jurkat | S. aureus | E. coli | C. albicans | M. hiemalis | |||||||||||||||
| 1 | > | 30 | > | 30 | – a | – a | – a | – a | ||||||||||||
| 2 | > | 30 | > | 30 | – a | – a | 7 | – a | ||||||||||||
| 3 | 10.7 | 2.3 | – a | – a | 7 | – a | ||||||||||||||
3. Experimental Section
3.1. General
3.2. Materials
3.3. Fermentation and Isolation
3.4. Antimicrobial Assay
3.5. Cytotoxicity Assay
4. Conclusions
Acknowledgments
- Samples Availability: Not available.
References
- Faulkner, D.J. Marine natural products. Nat. Prod. Rep. 2002, 19, 1–48. [Google Scholar]
- Blunt, J.W.; Copp, B.R.; Munro, M.H.G.; Northcote, P.T.; Prinsep, M.R. Marine natural products. Nat. Prod. Rep. 2012, 29, 144–222. [Google Scholar] [CrossRef]
- Robert, A.H. Marine natural products. Annu. Rep. Prog. Chem. 2012, 108, 131–146. [Google Scholar]
- Mostafa, E.R.; Rainer, E. Secondary metabolites of fungi from marine habitats. Nat. Prod. Rep. 2011, 28, 290–344. [Google Scholar]
- Namikoshi, M. Biologically Active Natural Products from Marine Fungi. In Biomaterials from Aquatic and Terrestrial Organism; Fingerman, M., Nagabhushanam, R., Eds.; Science Publishers: Enfield, NH, USA, 2006; pp. 285–359. [Google Scholar]
- Fujimoto, H.; Inagaki, M.; Satoh, Y.; Yoshida, E.; Yamazaki, M. Monoamine oxidase-inhibitory components from an ascomycete, Coniochaeta tetraspora. Chem. Phram. Bull. 1996, 44, 1090–1092. [Google Scholar] [CrossRef]
- Carvalhoa, M.R.; de Almeida Barbosab, L.C.; de Queiroz, J.H.; Howarthc, O.W. Novel lactones from Aspergillus versicolor. Tetrahedron Lett. 2001, 42, 809–811. [Google Scholar] [CrossRef]
- Rosario, M.A.; Rodrigues, F.E.; Moitinho, M.L.; Santos, L.S. Biologically active polyketides produced by Penicillium janthinellum isolated as an endophytic fungus from fruits of Melia azedarach. J. Braz. Chem. Soc. 2005, 16, 280–283. [Google Scholar] [CrossRef]
- Huang, H.; Li, Q.; Feng, X.; Chen, B.; Wang, J.; Liu, L.; She, Z.; Lin, Y. Structural elucidation and NMR assignments of four aromatic lactones from amangrove endophytic fungus (No. GX4-1B). Magn. Reson. Chem. 2010, 48, 496–499. [Google Scholar]
- Muhlemann, H. Anthrachinons and anthrachinon glycosides. Pharm. Acta Helv. 1952, 27, 9–10. [Google Scholar]
- Wijeratne, E.M.; Turbyville, T.J.; Fritz, A.; Whitesell, L.; Gunatilaka, A.A. A new dihydroxanthenone from a plant-associated strain of the fungus Chaetomium globosum demonstrates anticancer activity. Bioorg. Med. Chem. 2006, 14, 7917–7923. [Google Scholar] [CrossRef]
- Mosmann, T. Rapid colorimetric assay for cellular growth and survival: Application to proliferation and cytotoxicity assays. J. Immunol. Methods 1983, 65, 55–63. [Google Scholar] [CrossRef]
© 2012 by the authors; licensee MDPI, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Yamazaki, H.; Rotinsulu, H.; Kaneko, T.; Murakami, K.; Fujiwara, H.; Ukai, K.; Namikoshi, M. A New Dibenz[b,e]oxepine Derivative, 1-Hydroxy-10-methoxy-dibenz[b,e]oxepin-6,11-dione, from a Marine-Derived Fungus, Beauveria bassiana TPU942. Mar. Drugs 2012, 10, 2691-2697. https://doi.org/10.3390/md10122691
Yamazaki H, Rotinsulu H, Kaneko T, Murakami K, Fujiwara H, Ukai K, Namikoshi M. A New Dibenz[b,e]oxepine Derivative, 1-Hydroxy-10-methoxy-dibenz[b,e]oxepin-6,11-dione, from a Marine-Derived Fungus, Beauveria bassiana TPU942. Marine Drugs. 2012; 10(12):2691-2697. https://doi.org/10.3390/md10122691
Chicago/Turabian StyleYamazaki, Hiroyuki, Henki Rotinsulu, Tsuyoshi Kaneko, Kazuki Murakami, Hiromu Fujiwara, Kazuyo Ukai, and Michio Namikoshi. 2012. "A New Dibenz[b,e]oxepine Derivative, 1-Hydroxy-10-methoxy-dibenz[b,e]oxepin-6,11-dione, from a Marine-Derived Fungus, Beauveria bassiana TPU942" Marine Drugs 10, no. 12: 2691-2697. https://doi.org/10.3390/md10122691
APA StyleYamazaki, H., Rotinsulu, H., Kaneko, T., Murakami, K., Fujiwara, H., Ukai, K., & Namikoshi, M. (2012). A New Dibenz[b,e]oxepine Derivative, 1-Hydroxy-10-methoxy-dibenz[b,e]oxepin-6,11-dione, from a Marine-Derived Fungus, Beauveria bassiana TPU942. Marine Drugs, 10(12), 2691-2697. https://doi.org/10.3390/md10122691