Antibody Fragments as Probe in Biosensor Development
Abstract
:1. Antibodies in sensor applications
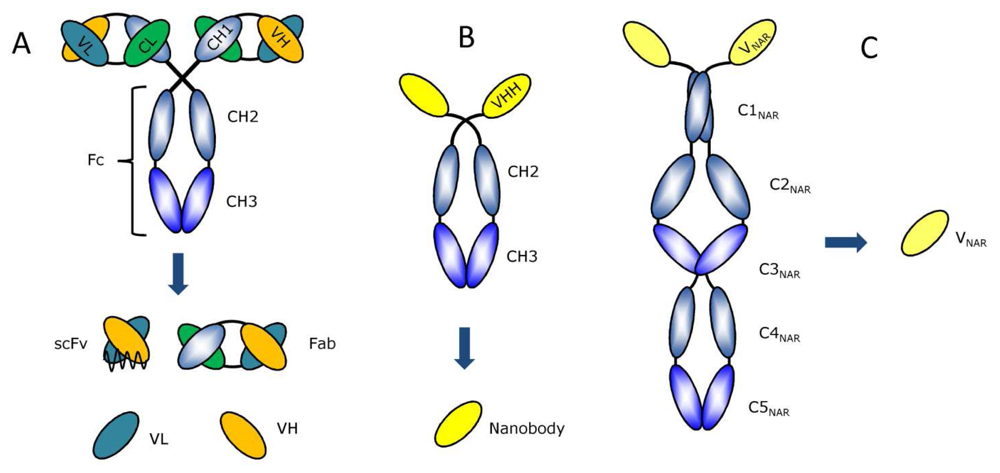
2. Available antibody fragment formats
3. Generation of antibody fragments libraries
4. Selection of antigen-specific antibody fragments
5. Affinity engineering
6. Stability engineering
7. Immobilization of antibody fragments
8. Examples of antibody fragments in biosensor development
9. Concluding remarks
Acknowledgments
References and Notes
- Holthoff, E.L.; Bright, F.V. Molecularly imprinted xerogels as platforms for sensing. Acc. Chem. Res. 2007, 40, 756–767. [Google Scholar]
- Wink, T.; van Zuilen, S.J.; Bult, A; van Bennkom, W.P. Self-assembled monolayers for biosensors. Analyst 1997, 122, 43R–50R. [Google Scholar]
- Holt, L.J.; Enever, C.; de Wildt, R.M.; Tomlinson, I.M. The use of recombinant antibodies in proteomics. Curr. Opin. Biotechnol. 2000, 11, 445–449. [Google Scholar]
- Briand, E.; Salmain, M.; Compere, C.; Pradier, C.M. Immobilization of Protein A on SAMs for the elaboration of immunosensors. Colloids Surf. B. Biointerfaces 2006, 53, 215–224. [Google Scholar]
- Pavlickova, P.; Schneider, E.M.; Hug, H. Advances in recombinant antibody microarrays. Clin. Chim. Acta 2004, 343, 17–35. [Google Scholar]
- Nelson, P.N.; Reynolds, G.M.; Waldron, E.E.; Ward, E.; Giannopoulos, K.; Murray, P.G. Monoclonal antibodies. Mol. Pathol. 2000, 53, 111–117. [Google Scholar]
- Ohara, R.; Knappik, A.; Shimada, K.; Frisch, C.; Ylera, F.; Koga, H. Antibodies for proteomic research: comparison of traditional immunization with recombinant antibody technology. Proteomics 2006, 6, 2638–2646. [Google Scholar]
- Hoogenboom, H.R. Selecting and screening recombinant antibody libraries. Nat. Biotechnol. 2005, 23, 1105–1116. [Google Scholar]
- Haab, B.B.; Dunham, M.J.; Brown, P.O. Protein microarrays for highly parallel detection and quantitation of specific proteins and antibodies in complex solutions. Genome Biol. 2001, 2, research0004.1–research0004.13. [Google Scholar]
- Jung, Y.; Jeong, J.Y.; Chung, B.H. Recent advances in immobilization methods of antibodies on solid supports. Analyst 2008, 133, 697–701. [Google Scholar]
- Lu, B.; Smyth, M.R.; O'Kennedy, R. Oriented immobilization of antibodies and its applications in immunoassays and immunosensors. Analyst 1996, 121, 29R–32R. [Google Scholar]
- Honegger, A. Engineering antibodies for stability and efficient folding. Handb. Exp. Pharmacol. 2008, 47–68. [Google Scholar]
- Hust, M.; Jostock, T.; Menzel, C.; Voedisch, B.; Mohr, A.; Brenneis, M.; Kirsch, M.I.; Meier, D.; Dubel, S. Single chain Fab (scFab) fragment. BMC Biotechnol. 2007, 7, 14. [Google Scholar]
- Holt, L.J.; Herring, C.; Jespers, L.S.; Woolven, B.P.; Tomlinson, I.M. Domain antibodies: proteins for therapy. Trends Biotechnol. 2003, 21, 484–490. [Google Scholar]
- Hamers-Casterman, C.; Atarhouch, T.; Muyldermans, S.; Robinson, G.; Hamers, C.; Songa, E.B.; Bendahman, N.; Hamers, R. Naturally occurring antibodies devoid of light chains. Nature 1993, 363, 446–448. [Google Scholar]
- Nguyen, V.K.; Hamers, R.; Wyns, L.; Muyldermans, S. Loss of splice consensus signal is responsible for the removal of the entire C(H)1 domain of the functional camel IGG2A heavy-chain antibodies. Mol. Immunol. 1999, 36, 515–524. [Google Scholar]
- Muyldermans, S. Single domain camel antibodies: current status. J. Biotechnol. 2001, 74, 277–302. [Google Scholar]
- Saerens, D.; Stijlemans, B.; Baral, T.N.; Nguyen Thi, G.T.; Wernery, U.; Magez, S.; De Baetselier, P.; Muyldermans, S.; Conrath, K. Parallel selection of multiple anti-infectome Nanobodies without access to purified antigens. J. Immunol. Methods 2008, 329, 138–150. [Google Scholar]
- Muyldermans, S.; Atarhouch, T.; Saldanha, J.; Barbosa, J.A.; Hamers, R. Sequence and structure of VH domain from naturally occurring camel heavy chain immunoglobulins lacking light chains. Protein Eng. 1994, 7, 1129–1135. [Google Scholar]
- Muyldermans, S.; Lauwereys, M. Unique single-domain antigen binding fragments derived from naturally occurring camel heavy-chain antibodies. J. Mol. Recognit. 1999, 12, 131–140. [Google Scholar]
- Dooley, H.; Flajnik, M.F.; Porter, A.J. Selection and characterization of naturally occurring single-domain (IgNAR) antibody fragments from immunized sharks by phage display. Mol. Immunol. 2003, 40, 25–33. [Google Scholar]
- Dumoulin, M.; Conrath, K.; Van Meirhaeghe, A.; Meersman, F.; Heremans, K.; Frenken, L.G.; Muyldermans, S.; Wyns, L.; Matagne, A. Single-domain antibody fragments with high conformational stability. Protein Sci. 2002, 11, 500–515. [Google Scholar]
- Fellouse, F.A.; Esaki, K.; Birtalan, S.; Raptis, D.; Cancasci, V.J.; Koide, A.; Jhurani, P.; Vasser, M.; Wiesmann, C.; Kossiakoff, A.A.; Koide, S.; Sidhu, S.S. High-throughput generation of synthetic antibodies from highly functional minimalist phage-displayed libraries. J. Mol. Biol. 2007, 373, 924–940. [Google Scholar]
- Silacci, M.; Brack, S.; Schirru, G.; Marlind, J.; Ettorre, A.; Merlo, A.; Viti, F.; Neri, D. Design, construction, and characterization of a large synthetic human antibody phage display library. Proteomics 2005, 5, 2340–2350. [Google Scholar]
- Desiderio, A.; Franconi, R.; Lopez, M.; Villani, M.E.; Viti, F.; Chiaraluce, R.; Consalvi, V.; Neri, D.; Benvenuto, E. A semi-synthetic repertoire of intrinsically stable antibody fragments derived from a single-framework scaffold. J. Mol. Biol. 2001, 310, 603–615. [Google Scholar]
- Liu, J.L.; Anderson, G.P.; Goldman, E.R. Isolation of anti-toxin single domain antibodies from a semi-synthetic spiny dogfish shark display library. BMC Biotechnol. 2007, 7, 78. [Google Scholar]
- Fellouse, F.A.; Li, B.; Compaan, D.M.; Peden, A.A.; Hymowitz, S.G.; Sidhu, S.S. Molecular recognition by a binary code. J. Mol. Biol. 2005, 348, 1153–1162. [Google Scholar]
- Mondon, P.; Souyris, N.; Douchy, L.; Crozet, F.; Bouayadi, K.; Kharrat, H. Method for generation of human hyperdiversified antibody fragment library. J. Biotechnol. 2007, 2, 76–82. [Google Scholar]
- Qin, W.; Zhao, A.; Han, Y.; Wen, W.; Li, Y.; Chen, G.; Zhang, Z.; Wang, H. A novel technique for efficient construction of large scFv libraries. Mol. Biotechnol. 2007, 37, 201–205. [Google Scholar]
- Verheesen, P.; Roussis, A.; de Haard, H.J.; Groot, A.J.; Stam, J.C.; den Dunnen, J.T.; Frants, R.R.; Verkleij, A.J.; Theo Verrips, C.; van der Maarel, S.M. Reliable and controllable antibody fragment selections from Camelid non-immune libraries for target validation. Biochim. Biophys. Acta 2006, 1764, 1307–1319. [Google Scholar]
- Conrath, K.E.; Lauwereys, M.; Galleni, M.; Matagne, A.; Frere, J.M.; Kinne, J.; Wyns, L.; Muyldermans, S. Beta-lactamase inhibitors derived from single-domain antibody fragments elicited in the camelidae. Antimicrob. Agents Chemother. 2001, 45, 2807–2812. [Google Scholar]
- Dolk, E.; van der Vaart, M.; Lutje Hulsik, D.; Vriend, G.; de Haard, H.; Spinelli, S.; Cambillau, C.; Frenken, L.; Verrips, T. Isolation of llama antibody fragments for prevention of dandruff by phage display in shampoo. Appl. Environ. Microbiol. 2005, 71, 442–450. [Google Scholar]
- Saerens, D.; Kinne, J.; Bosmans, E.; Wernery, U.; Muyldermans, S.; Conrath, K. Single domain antibodies derived from dromedary lymph node and peripheral blood lymphocytes sensing conformational variants of prostate-specific antigen. J. Biol. Chem. 2004, 279, 51965–51972. [Google Scholar]
- Azzazy, H.M.; Highsmith, W.E., Jr. Phage display technology: clinical applications and recent innovations. Clin. Biochem. 2002, 35, 425–445. [Google Scholar]
- Liu, B.; Huang, L.; Sihlbom, C.; Burlingame, A.; Marks, J.D. Towards proteome-wide production of monoclonal antibody by phage display. J. Mol. Biol. 2002, 315, 1063–1073. [Google Scholar]
- Smith, G.P.; Petrenko, V.A. Phage Display. Chem. Rev. 1997, 97, 391–410. [Google Scholar]
- He, M.; Taussig, M.J. Rapid discovery of protein interactions by cell-free protein technologies. Biochem. Soc. Trans. 2007, 35, 962–965. [Google Scholar]
- Schaffitzel, C.; Hanes, J.; Jermutus, L.; Pluckthun, A. Ribosome display: an in vitro method for selection and evolution of antibodies from libraries. J. Immunol. Methods 1999, 231, 119–135. [Google Scholar]
- Wang, M.; He, M. The rapid discovery of engineered antibodies. IDrugs 2007, 10, 562–565. [Google Scholar]
- Zahnd, C.; Amstutz, P.; Pluckthun, A. Ribosome display: selecting and evolving proteins in vitro that specifically bind to a target. Nat. Methods 2007, 4, 269–279. [Google Scholar]
- Kopsidas, G.; Carman, R.K.; Stutt, E.L.; Raicevic, A.; Roberts, A.S.; Siomos, M.A.; Dobric, N.; Pontes-Braz, L.; Coia, G. RNA mutagenesis yields highly diverse mRNA libraries for in vitro protein evolution. BMC Biotechnol. 2007, 7, 18. [Google Scholar]
- Adamson, C.S.; Yao, Y.; Vasiljevic, S.; Sy, M.S.; Ren, J.; Jones, I.M. Novel single chain antibodies to the prion protein identified by phage display. Virology 2007, 358, 166–177. [Google Scholar]
- Oh, M.Y.; Joo, H.Y.; Hur, B.U.; Jeong, Y.H.; Cha, S.H. Enhancing phage display of antibody fragments using gIII-amber suppression. Gene 2007, 386, 81–89. [Google Scholar]
- Steinhauer, C.; Wingren, C.; Hager, A.C.; Borrebaeck, C.A. Single framework recombinant antibody fragments designed for protein chip applications. Biotechniques 2002, Suppl, 38–45. [Google Scholar]
- Jespers, L.; Schon, O.; Famm, K.; Winter, G. Aggregation-resistant domain antibodies selected on phage by heat denaturation. Nat. Biotechnol. 2004, 22, 1161–1165. [Google Scholar]
- Yan, X.H.; Xu, Z.R. Production of human single-chain variable fragment (scFv) antibody specific for digoxin by ribosome display. Indian J. Biochem. Biophys. 2005, 42, 350–357. [Google Scholar]
- Backmann, N.; Zahnd, C.; Huber, F.; Bietsch, A.; Pluckthun, A.; Lang, H.P.; Guntherodt, H.J.; Hegner, M.; Gerber, C. A label-free immunosensor array using single-chain antibody fragments. Proc. Natl. Acad. Sci. U S A 2005, 102, 14587–14592. [Google Scholar]
- Harrison, J.S.; Gill, A.; Hoare, M. Stability of a single-chain Fv antibody fragment when exposed to a high shear environment combined with air-liquid interfaces. Biotechnol. Bioeng. 1998, 59, 517–519. [Google Scholar]
- Arndt, K.M.; Muller, K.M.; Pluckthun, A. Helix-stabilized Fv (hsFv) antibody fragments: substituting the constant domains of a Fab fragment for a heterodimeric coiled-coil domain. J. Mol. Biol. 2001, 312, 221–228. [Google Scholar]
- Webber, K.O.; Reiter, Y.; Brinkmann, U.; Kreitman, R.; Pastan, I. Preparation and characterization of a disulfide-stabilized Fv fragment of the anti-Tac antibody: comparison with its single-chain analog. Mol. Immunol. 1995, 32, 249–258. [Google Scholar]
- Young, N.M.; MacKenzie, C.R.; Narang, S.A.; Oomen, R.P.; Baenziger, J.E. Thermal stabilization of a single-chain Fv antibody fragment by introduction of a disulphide bond. FEBS Lett. 1995, 377, 135–139. [Google Scholar]
- Quintero-Hernandez, V.; Juarez-Gonzalez, V.R.; Ortiz-Leon, M.; Sanchez, R.; Possani, L.D.; Becerril, B. The change of the scFv into the Fab format improves the stability and in vivo toxin neutralization capacity of recombinant antibodies. Mol. Immunol. 2007, 44, 1307–1315. [Google Scholar]
- Monsellier, E.; Bedouelle, H. Quantitative measurement of protein stability from unfolding equilibria monitored with the fluorescence maximum wavelength. Protein Eng. Des. Sel. 2005, 18, 445–456. [Google Scholar]
- Jung, S.; Pluckthun, A. Improving in vivo folding and stability of a single-chain Fv antibody fragment by loop grafting. Protein Eng. 1997, 10, 959–966. [Google Scholar]
- Krauss, J.; Arndt, M.A.; Martin, A.C.; Liu, H.; Rybak, S.M. Specificity grafting of human antibody frameworks selected from a phage display library: generation of a highly stable humanized anti-CD22 single-chain Fv fragment. Protein Eng. 2003, 16, 753–759. [Google Scholar]
- Saerens, D.; Pellis, M.; Loris, R.; Pardon, E.; Dumoulin, M.; Matagne, A.; Wyns, L.; Muyldermans, S.; Conrath, K. Identification of a Universal VHH Framework to Graft Non-canonical Antigen-binding Loops of Camel Single-domain Antibodies. J. Mol. Biol. 2005, 352, 597–607. [Google Scholar]
- Saerens, D.; Conrath, K.; Govaert, J.; Muyldermans, S. Disulfide bond introduction for general stabilization of immunoglobulin heavy-chain variable domains. J. Mol. Biol. 2008, 377, 478–488. [Google Scholar]
- Hagihara, Y.; Mine, S.; Uegaki, K. Stabilization of an immunoglobulin fold domain by an engineered disulfide bond at the buried hydrophobic region. J. Biol. Chem. 2007, 282, 36489–36495. [Google Scholar]
- Harmsen, M.M.; van Solt, C.B.; van Zijderveld-van Bemmel, A.M.; Niewold, T.A.; van Zijderveld, F.G. Selection and optimization of proteolytically stable llama single-domain antibody fragments for oral immunotherapy. Appl. Microbiol. Biotechnol. 2006, 72, 544–551. [Google Scholar]
- Jung, S.; Honegger, A.; Pluckthun, A. Selection for improved protein stability by phage display. J. Mol. Biol. 1999, 294, 163–180. [Google Scholar]
- van der Linden, R.H.; de Geus, B.; Frenken, G.J.; Peters, H.; Verrips, C.T. Improved production and function of llama heavy chain antibody fragments by molecular evolution. J. Biotechnol. 2000, 80, 261–270. [Google Scholar]
- Saerens, D.; Frederix, F.; Reekmans, G.; Conrath, K.; Jans, K.; Brys, L.; Huang, L.; Bosmans, E.; Maes, G.; Borghs, G.; Muyldermans, S. Engineering Camel Single-Domain Antibodies and Immobilization Chemistry for Human Prostate-Specific Antigen Sensing. Anal. Chem. 2005, 77, 7547–7555. [Google Scholar]
- Cao, T.; Wang, A.; Liang, X.; Tang, H.; Auner, G.W.; Salley, S.O.; Ng, K.Y. Investigation of spacer length effect on immobilized Escherichia coli pili-antibody molecular recognition by AFM. Biotechnol. Bioeng. 2007, 98, 1109–1122. [Google Scholar]
- Huang, L.; Reekmans, G.; Saerens, D.; Friedt, J.M.; Frederix, F.; Francis, L.; Muyldermans, S.; Campitelli, A.; Hoof, C.V. Prostate-specific antigen immunosensing based on mixed self-assembled monolayers, camel antibodies and colloidal gold enhanced sandwich assays. Biosens. Bioelectron. 2005, 21, 483–490. [Google Scholar]
- Norde, W. Adsorption of proteins from solution at the solid-liquid interface. Adv. Colloid. Interface Sci. 1986, 25, 267–340. [Google Scholar]
- Oh, B.K.; Kim, Y.K.; Lee, W.; Bae, Y.M.; Lee, W.H.; Choi, J.W. Immunosensor for detection of Legionella pneumophila using surface plasmon resonance. Biosens. Bioelectron. 2003, 18, 605–611. [Google Scholar]
- Boltovets, P.M.; Boyko, V.R.; Kostikov, I.Y.; Dyachenko, N.S.; Snopok, B.A.; Shirshov, Y.M. Simple method for plant virus detection: effect of antibody immobilization technique. J. Virol. Methods 2002, 105, 141–146. [Google Scholar]
- Bonroy, K.; Frederix, F.; Reekmans, G.; Dewolf, E.; De Palma, R.; Borghs, G.; Declerck, P.; Goddeeris, B. Comparison of random and oriented immobilisation of antibody fragments on mixed self-assembled monolayers. J. Immunol. Methods 2006, 312, 167–81. [Google Scholar]
- Ionescu, R.E.; Jaffrezic-Renault, N.; Bouffier, L.; Gondran, C.; Cosnier, S.; Pinacho, D.G.; Marco, M.P.; Sanchez-Baeza, F.J.; Healy, T.; Martelet, C. Impedimetric immunosensor for the specific label free detection of ciprofloxacin antibiotic. Biosens. Bioelectron. 2007, 23, 549–555. [Google Scholar]
- Sun, Y.; Bai, Y.; Song, D.; Li, X.; Wang, L.; Zhang, H. 2007. Design and performances of immunoassay based on SPR biosensor with magnetic microbeads. Biosens. Bioelectron. 2007, 23, 473–478. [Google Scholar]
- Lee, W.; Oh, B.K.; Lee, W.H.; Choi, J.W. Immobilization of antibody fragment for immunosensor application based on surface plasmon resonance. Colloids. Surf. B Biointerfaces 2005, 40, 143–148. [Google Scholar]
- Torrance, L.; Ziegler, A.; Pittman, H.; Paterson, M.; Toth, R.; Eggleston, I. Oriented immobilisation of engineered single-chain antibodies to develop biosensors for virus detection. J. Virol. Methods 2006, 134, 164–170. [Google Scholar]
- Lee, K.G.; Pillai, S.R.; Singh, S.R.; Willing, G.A. The investigation of Protein A and Salmonella antibody adsorption onto biosensor surfaces by atomic force microscopy. Biotechnol. Bioeng. 2007, 99(4), 949–959. [Google Scholar]
- Shen, Z.; Yan, H.; Parl, F.F.; Mernaugh, R.L.; Zeng, X. Recombinant antibody piezoimmunosensors for the detection of cytochrome P450 1B1. Anal. Chem. 2007, 79, 1283–1289. [Google Scholar]
- Cho, I.H.; Paek, E.H.; Lee, H.; Kang, J.Y.; Kim, T.S.; Paek, S.H. Site-directed biotinylation of antibodies for controlled immobilization on solid surfaces. Anal. Biochem. 2007, 365, 14–23. [Google Scholar]
- Scholler, N.; Garvik, B.; Quarles, T.; Jiang, S.; Urban, N. Method for generation of in vivo biotinylated recombinant antibodies by yeast mating. J. Immunol. Methods 2006, 317, 132–143. [Google Scholar]
- Pleschberger, M.; Saerens, D.; Weigert, S.; Sleytr, U.B.; Muyldermans, S.; Sara, M.; Egelseer, E.M. An S-layer heavy chain camel antibody fusion protein for generation of a nanopatterned sensing layer to detect the prostate-specific antigen by surface plasmon resonance technology. Bioconjug. Chem. 2004, 15, 664–671. [Google Scholar]
- Nanduri, V.; Bhunia, A.K.; Tu, S.I.; Paoli, G.C.; Brewster, J.D. SPR biosensor for the detection of L. monocytogenes using phage-displayed antibody. Biosens. Bioelectron. 2007, 23, 248–252. [Google Scholar]
- Gilchuk, P.V.; Volkov, G.L. Immobilization of mouse single-chain antibodies for affinity chromatography using the cellulose-binding protein. Ukr Biokhim Zh. 2006, 78, 160–163. [Google Scholar]
- Shen, Z.; Mernaugh, R.L.; Yan, H.; Yu, L.; Zhang, Y.; Zeng, X. Engineered recombinant single-chain fragment variable antibody for immunosensors. Anal. Chem. 2005, 77, 6834–6842. [Google Scholar]
- Steinhauer, C.; Wingren, C.; Khan, F.; He, M.; Taussig, M.J.; Borrebaeck, C.A. Improved affinity coupling for antibody microarrays: engineering of double-(His)6-tagged single framework recombinant antibody fragments. Proteomics 2006, 6, 4227–4234. [Google Scholar]
- Watanabe, H.; Tsumoto, K.; Taguchi, S.; Yamashita, K.; Doi, Y.; Nishimiya, Y.; Kondo, H.; Umetsu, M.; Kumagai, I. A human antibody fragment with high affinity for biodegradable polymer film. Bioconjug. Chem. 2007, 18, 645–651. [Google Scholar]
- Hattori, T.; Umetsu, M.; Nakanishi, T.; Tsumoto, K.; Ohara, S.; Abe, H.; Naito, M.; Asano, R.; Adschiri, T.; Kumagai, I. Grafting of material-binding function into antibodies Functionalization by peptide grafting. Biochem. Biophys. Res. Commun. 2008, 365, 751–757. [Google Scholar]
- Mechaly, A.; Zahavy, E.; Fisher, M. Development and implementation of a scFv antibody for specific detection of B. anthracis spores. Appl. Environ. Microbiol. 2007, 74(3), 818–822. [Google Scholar]
- Hu, X.; O'Hara, L.; White, S.; Magner, E.; Kane, M.; Wall, J.G. Optimisation of production of a domoic acid-binding scFv antibody fragment in Escherichia coli using molecular chaperones and functional immobilisation on a mesoporous silicate support. Protein. Expr. Purif. 2007, 52, 194–201. [Google Scholar]
- Deng, X.K.; Nesbit, L.A.; Morrow, K.J., Jr. Recombinant single-chain variable fragment antibodies directed against Clostridium difficile toxin B produced by use of an optimized phage display system. Clin. Diagn. Lab. Immunol. 2003, 10, 587–595. [Google Scholar]
- Carlsson, A.; Wingren, C.; Ingvarsson, J.; Ellmark, P.; Baldertorp, B.; Ferno, M.; Olsson, H.; Borrebaeck, C.A. Serum proteome profiling of metastatic breast cancer using recombinant antibody microarrays. Eur. J. Cancer. 2007, 44(3), 472–480. [Google Scholar]
- Ymeti, A.; Subramaniam, V.; Beumer, T.A.; Kanger, J.S. An ultrasensitive young interferometer handheld sensor for rapid virus detection. Expert Rev. Med. Devices 2007, 4, 447–454. [Google Scholar]
| Immobilization method | Advantages | Disadvantages |
|---|---|---|
| Adsorption | Minimal manipulation | Random orientation |
| No Ab modification | Ab denaturation | |
| Mostly high immobilization level | Non-specific protein binding | |
| Leakage of Ab from surface | ||
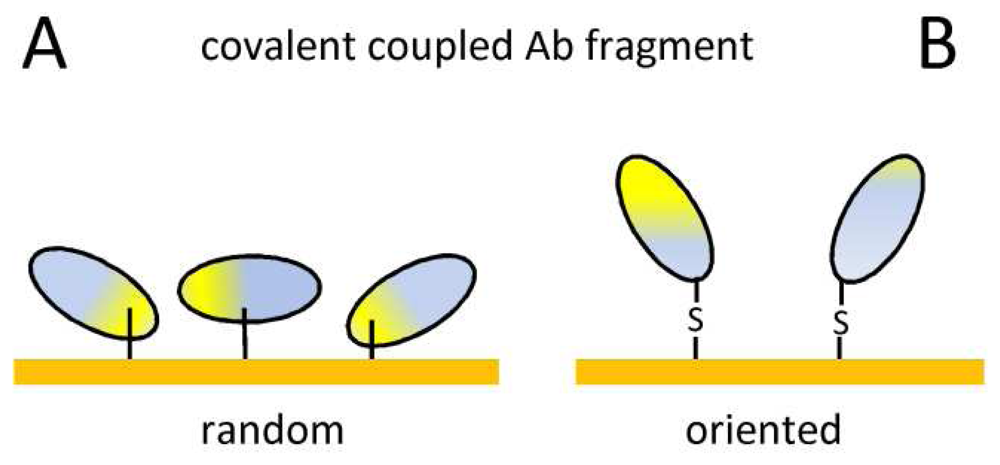
| Covalent coupling | Stable immobilization | Random orientation |
| Commercially available surfaces | Ab modification, possible denaturation | |
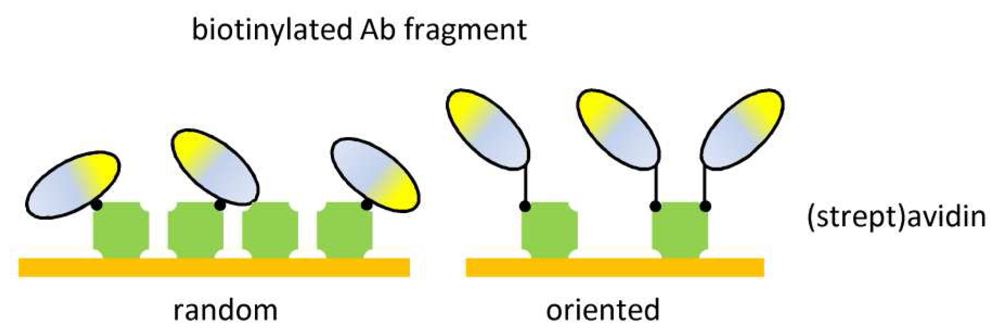
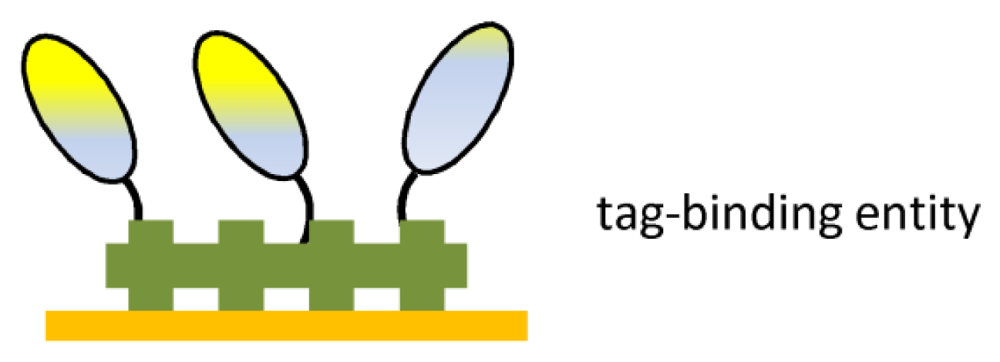
| Ab fragment tag | Oriented immobilization | Surface stability |
| Mild incubation | ||
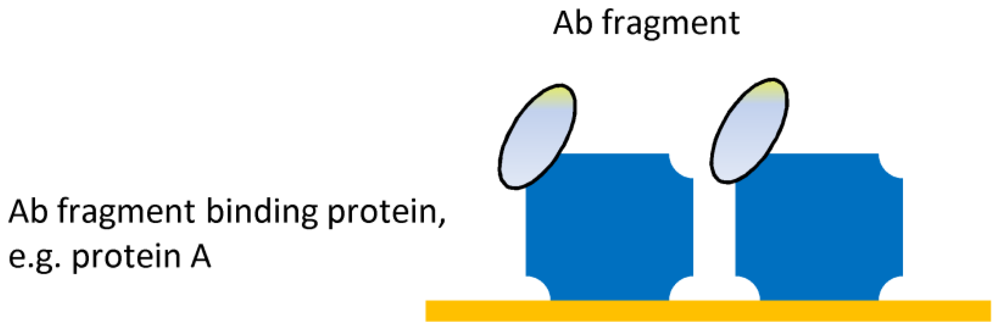
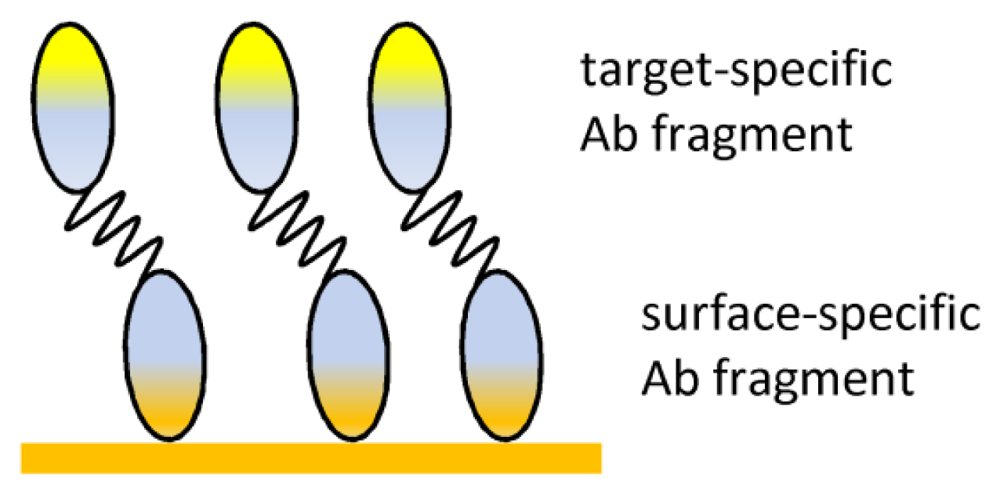
| Ab-binding proteins | Oriented immobilization | Surface stability |
| No Ab modification | ||
| Mild incubation | ||
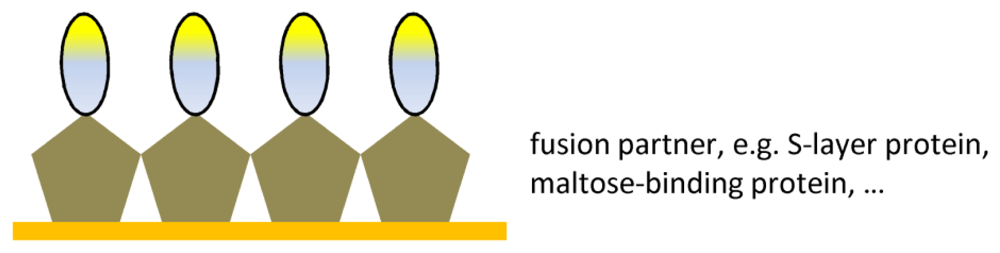
| Ab fragment fusions | Oriented immobilization | Compatibility between Ab fragment and |
| Surface stability | fusion partner |
© 2008 by the authors; licensee Molecular Diversity Preservation International, Basel, Switzerland. This article is an open-access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Saerens, D.; Huang, L.; Bonroy, K.; Muyldermans, S. Antibody Fragments as Probe in Biosensor Development. Sensors 2008, 8, 4669-4686. https://doi.org/10.3390/s8084669
Saerens D, Huang L, Bonroy K, Muyldermans S. Antibody Fragments as Probe in Biosensor Development. Sensors. 2008; 8(8):4669-4686. https://doi.org/10.3390/s8084669
Chicago/Turabian StyleSaerens, Dirk, Lieven Huang, Kristien Bonroy, and Serge Muyldermans. 2008. "Antibody Fragments as Probe in Biosensor Development" Sensors 8, no. 8: 4669-4686. https://doi.org/10.3390/s8084669
APA StyleSaerens, D., Huang, L., Bonroy, K., & Muyldermans, S. (2008). Antibody Fragments as Probe in Biosensor Development. Sensors, 8(8), 4669-4686. https://doi.org/10.3390/s8084669




