Crop Classification in Satellite Images through Probabilistic Segmentation Based on Multiple Sources †
Abstract
:1. Introduction
2. Previous Works
3. The Proposal
3.1. Training Stage
3.2. Segmentation Stage through Multiples Sources and Probabilistic Approach
- For a given image, we compute the feature spaces provided by the mapping , .
- Then, the likelihood is assigned according to the following equationsuch that , , . The likelihood is obtained by normalizing with respect to the classes.
- Here we propose a robust GMMF approach that generalizes the proposal in [28] including more feature spaces and with weight functions in both the data and regularization terms. Equations (10)–(12) indicate the modifications.where is an uncertainty measure of the information source f at pixel r, for example, the measure of entropy (5) used in [28]; is the regularization parameter, controls the relative importance of the likelihood for different sources. When is very large, the contribution of the likelihood for all sources tends to be the same and when it is close to zero, the functional in (10) tends to select the likelihood corresponding to the lowest uncertainty, i.e., this tends to the solution proposed in [28] when using the entropy as uncertainty measure. The weight function allows to control the edges between classes, here we usesuch that if the sites very probably belong to the same class and otherwise. The solution of the optimization problem (10) yields the following Gauss-Seidel scheme:which is similar to the Gauss-Seidel scheme (2), but now the term , Equation (14), is a mixture term that allows us to combine or fuse different likelihoods. In addition, the above formula also includes function weights to control the edges between classes. Note that, the first term in the numerator of the Equation (15) is a convex linear combination of likelihoods derived from different sources. Feature spaces with lower uncertainty have a greater and therefore they have a greater contribution to the data term in Equation (10).
3.3. Validation
4. Experiments and Discussion
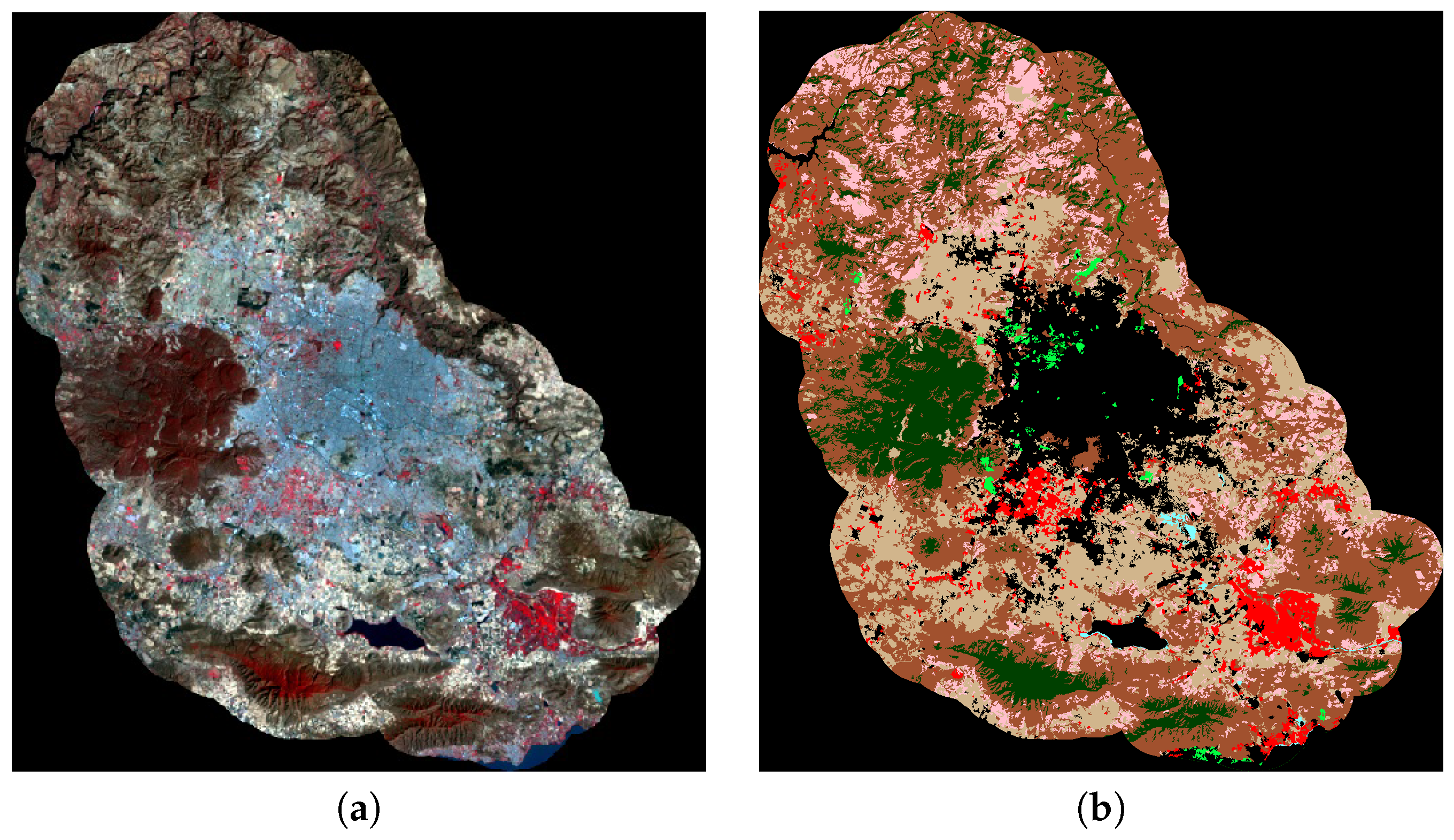
4.1. Study Area
4.2. Data Sources
4.3. Studied Feature Spaces
- Space 2: it contains the first three principal components from the PCA applied on 10 vegetation indices, see Table 3. Such indices are based on mathematical operations on spectral bands and they allow to enhance the information related to vegetation. In order to compute the indices we calculate the reflectance values, , corresponding to the acquired images, using the algorithms in [48], see also the procedures given in [49,50,51,52]. The included indices appear in Table 3. Symbols , , and denote the reflectance values for the red, blue, green and infrared bands respectively. We followed the recommendation given in [35], and set to compute the SAVI and SARVI expressions. For SARVI we considered as authors in [34].
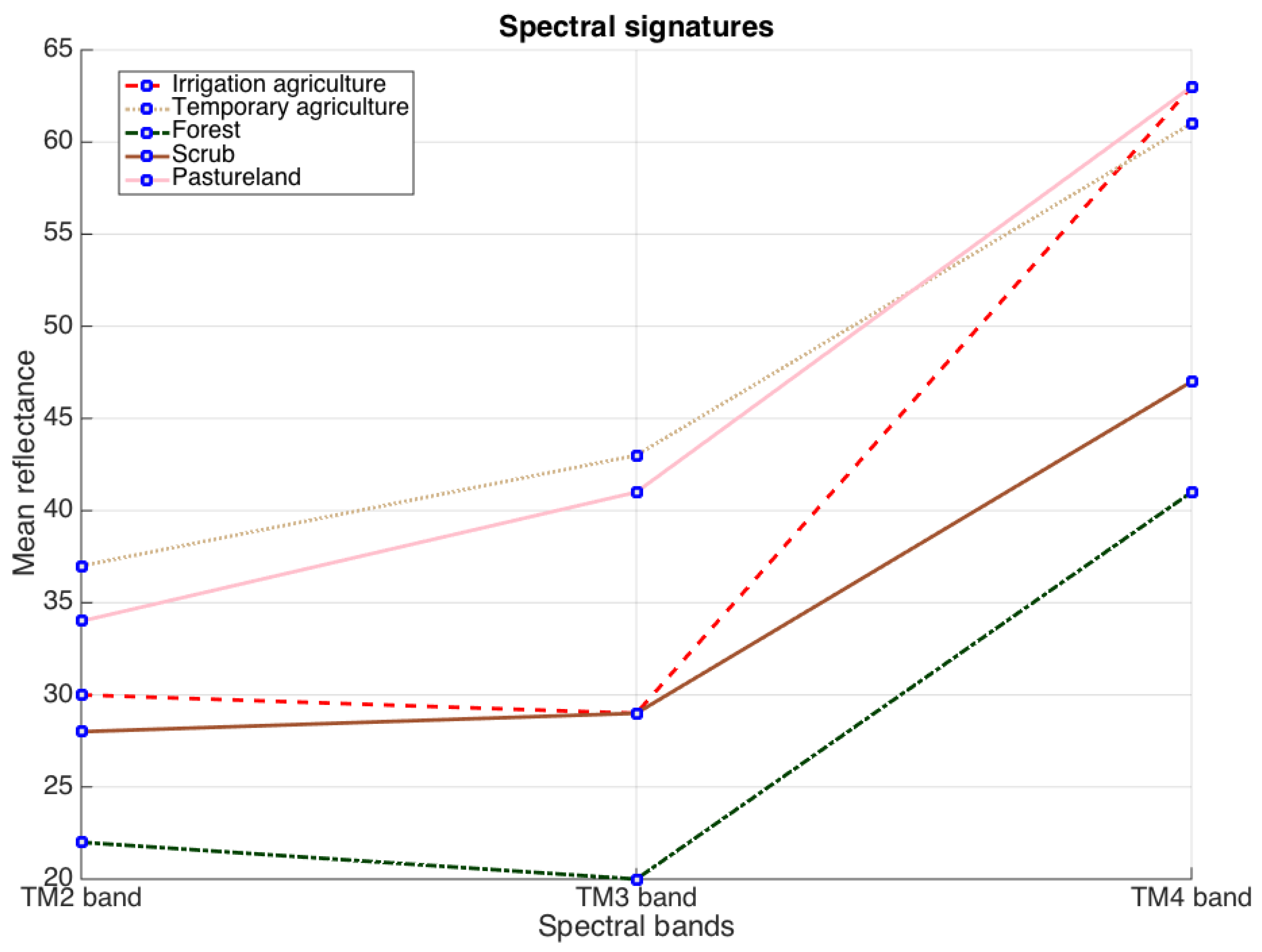
- Space 3: this space contains three principal components from PCA applied on the spectral bands TM1, TM2, TM3, TM4, TM5 and TM7, see Table 2. Although TM2, TM3 and TM4 bands more accurately describes information related to vegetation [30], in this investigation PCA on all six mentioned bands is applied in order to include information from other spectral regions.
4.4. Comparison Measures
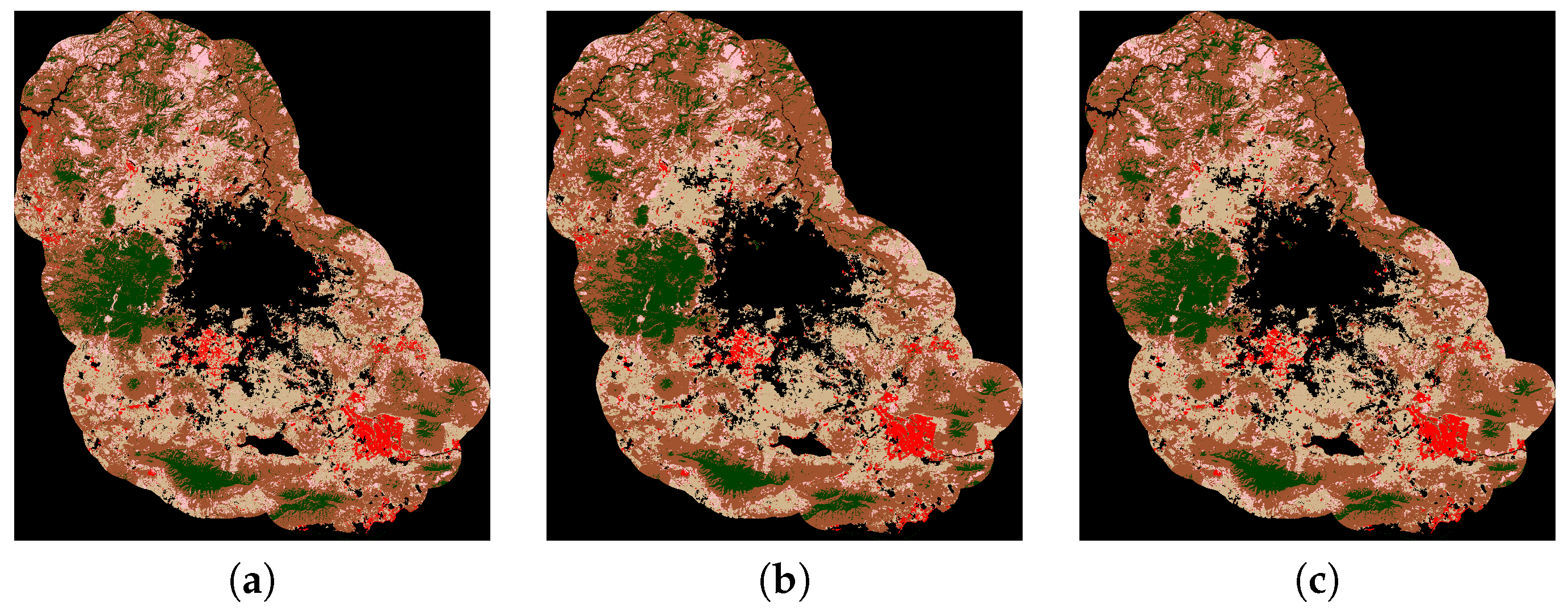
4.5. Results and Discussion
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Jain, A.; Dubes, R.C. Algorithm for Clustering Data; Prentice Hall: Upper Saddle River, NJ, USA, 1988. [Google Scholar]
- Li, S.Z. Markov Random Field Modeling in Image Analysis; Springer: Tokyo, Japan, 2001. [Google Scholar]
- Geman, S.; Geman, D. Stochastic relaxation, Gibbs distributions and bayesian restoration of images. IEEE Trans. Pattern Anal. Mach. Intell. 1984, 6, 721–741. [Google Scholar] [CrossRef] [PubMed]
- Marroquin, J.L.; Rivera, M.; Nakamura, M. Gauss-Markov measure field models for low-level vision. IEEE Trans. Pattern Anal. Mach. Intell. 2001, 23, 337–348. [Google Scholar] [CrossRef]
- Cedeño, O.D.; Rivera, M.; Mayorga, P.P. Computing the Alpha-Channel with Probabilistic Segmentation for Image Colorization. In Proceedings of the IEEE 11th International Conference on Computer Vision (ICCV 2007), Rio de Janeiro, Brazil, 14–21 October 2007; pp. 1–7. [Google Scholar]
- Dalmau, O.; Rivera, M. A General Bayesian Markov Random Field Model for Probabilistic Image Segmentation. In Combinatorial Image Analysis: Lecture Notes in Computer Science; Wiederhold, P., Barneva, R., Eds.; Springer: Berlin/Heidelberg, Germany, 2009; Volume 5852, pp. 149–161. [Google Scholar]
- Dalmau, O.; Rivera, M. Beta-Measure for Probabilistic Segmentation. In Advances in Artificial Intelligence Part I, Proceedings of the 9th Mexican International Conference on Artificial Intelligence (MICAI 2010), Pachuca, Mexico, 8–13 November 2010; Springer: Berlin/Heidelberg, Germany, 2010; pp. 312–324. [Google Scholar]
- Marroquin, J.L.; Arce, E.; Botello, S. Hidden Markov measure field models for image segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2003, 25, 1380–1387. [Google Scholar] [CrossRef]
- Rivera, M.; Ocegueda, O.; Marroquin, J.L. Entropy controlled gauss-markov random measure field models for early vision. In Proceedings of the Variational, Geometric, and Level Set Methods in Computer Vision (VLSM), Beijing, China, 16 October 2005; Volume 3572, pp. 137–148. [Google Scholar]
- Marroquin, J.L.; Vemuri, B.C.; Botello, S.; Calderon, E.; Fernandez-Bouzas, A. An accurate and efficient Bayesian method for automatic segmentation of brain MRI. IEEE Trans. Med. Imaging 2002, 5, 934–945. [Google Scholar] [CrossRef] [PubMed]
- Gijbels, I.; Lambert, A.; Qiu, P. Edge-preserving image denoising and estimation of discontinuous surfaces. IEEE Trans. Pattern Anal. Mach. Intell. 2006, 28, 1075–1087. [Google Scholar] [CrossRef] [PubMed]
- Ocegueda, O.; Shah, S.K.; Kakadiaris, I.A. Which parts of the face give out your identity? In Proceedings of the 2011 IEEE Conference on Computer Vision and Pattern Recognition (CVPR), Colorado Springs, CO, USA, 20–25 June 2011; pp. 641–648. [Google Scholar]
- Ocegueda, O.; Fang, T.; Shah, S.K.; Kakadiaris, I.A. 3D face discriminant analysis using Gauss-Markov posterior marginals. IEEE Trans. Pattern Anal. Mach. Intell. 2013, 35, 728–739. [Google Scholar] [CrossRef] [PubMed]
- Bogoliubova, A.; Tymkow, P. Land cover changes and dynamics of yuntolovsky reserve. Electron. J. Pol. Agric. Univ. 2014, 17. Available online: http://www.ejpau.media.pl/volume17/issue3/art-03.html (accessed on 13 June 2017).
- Ustuner, M.; Sanli, F.; Abdikan, S.; Esetlili, M.; Kurucu, Y. Crop type classification using vegetation indices of RapidEye imagery. In Proceedings of the International Archives of the Photogrammetry, Remote Sensing and Spatial Information Sciences, Istanbul, Turkey, 29 September–2 October 2014; pp. 195–198. [Google Scholar]
- Tucker, C.J. Detection of Red and photographic infrared linear combinations for monitoring vegetation. Remote Sens. Environ. 1979, 8, 127–150. [Google Scholar] [CrossRef]
- Gitelson, A.A.; Merzlyak, M.N. Detection of Red Edge Position and Chlorophyll Content by Reflectance Measurements Near 700 nm. J. Plant Physiol. 1996, 148, 501–508. [Google Scholar] [CrossRef]
- Barnes, E.M.; Clarke, T.R. Coincident detection of crop water stress, nitrogen status and canopy density using ground based multispectral data. In Proceedings of the Fifth International Conference on Precision Agriculture, Bloomington, MN, USA, 16–19 July 2000. [Google Scholar]
- Vapnik, V. The Nature of Statistical Learning Theory; Springer: Berlin, Germany, 2013. [Google Scholar]
- Vikesh, K.; Vinod, K.; Kamal, J. Development of Spectral Signature and Classification of Sugarcane using ASTER DATA. Int. J. Comput. Sci. Commun. 2010, 1, 245–251. [Google Scholar]
- Yang, C.; Everit, J.H.; Murden, D. Evaluating high resolution {SPOT} 5 satellite imagery for crop identification. Comput. Electron. Agric. 2011, 75, 347–354. [Google Scholar] [CrossRef]
- Kruse, F.A.; Lefkoff, A.B.; Boardman, J.B.; Heidebrecht, K.B.; Shapiro, A.T.; Barloon, P.J.; Goetz, A.F.H. The Spectral Image Processing System (SIPS)—Interactive Visualization and Analysis of Imaging Spectrometer Data. Remote Sens. Environ. 1993, 44, 145–163. [Google Scholar] [CrossRef]
- Nawaz, A.; Iqbal, Z.; Ullah, S. Performance analysis of supervised image classification techniques for the classification of multispectral satellite imagery. In Proceedings of the ICASE 2015—4th IEEE International Conference on Aerospace Science and Engineering, Islamabad, Pakistan, 2–4 September 2015. [Google Scholar]
- Dash, M.; Liu, H. Feature selection for classification. Intell. Data Anal. 1997, 1, 131–156. [Google Scholar] [CrossRef]
- Luis Carlos Molina, A.L.B.; Nebot, A. Feature selection algorithms: A survey and experimental evaluation. In Proceedings of the IEEE International Conference on Data Mining (ICDM), Maebashi City, Japan, 9–12 December 2002. [Google Scholar]
- Peng, H.; Long, F.; Ding, C. Feature selection based on mutual information criteria of max-dependency, max-relevance, and min-redundancy. IEEE Trans. Pattern Anal. Mach. Intell. 2005, 27, 1226–1238. [Google Scholar] [CrossRef] [PubMed]
- Pena-Barragan, J.M.; Ngugi, M.K.; Plant, R.E.; Six, J. Object-based crop identification using multiple vegetation indices, textural features and crop phenology. Remote Sens. Environ. 2011, 115, 1301–1316. [Google Scholar] [CrossRef]
- Oliva, F.E.; Dalmau, O.S.; Alarcón, T.E. Classification of Different Vegetation Types Combining Two Information Sources Through a Probabilistic Segmentation Approach. In Proceedings of the Mexican International Conference on Artificial Intelligence, Cuernavaca, Mexico, 25–31 October 2015; Volume 9414, pp. 382–392. [Google Scholar]
- Oliva, F.E.; Dalmau, O.S.; Alarcón, T.E. A Supervised Segmentation Algorithm for Crop Classification Based on Histograms Using Satellite Images. In Proceedings of the Mexican International Conference on Artificial Intelligence, Tuxtla Gutiérrez, Mexico, 16–22 November 2014; Volume 8856, pp. 327–335. [Google Scholar]
- Sabins, F.F. Remote Sensing: Principles and Applications; Waveland Press Inc.: Long Grove, IL, USA, 2007. [Google Scholar]
- Jordan, C. Derivation of leaf area index from quality of light on the forest floor. Ecology 1969, 50, 663–666. [Google Scholar] [CrossRef]
- Gitelson, A.A.; Merzlyak, M.N. Relationships between leaf chlorophyll content and spectral reflectance and algorithms for non-destructive chlorophyll assessment in higher plant leaves. J. Plant Physiol. 2003, 160, 271–282. [Google Scholar] [CrossRef] [PubMed]
- Huete, A.R.; Liu, H.; Batchily, K.; Van Leeuwen, W. A comparison of vegetation indices over a global set of TM images for EOS-MODIS. Remote Sens. Environ. 1997, 59, 440–451. [Google Scholar] [CrossRef]
- Kaufman, Y.; Tanre, D. Atmospherically resistant vegetation index (ARVI) for EOS-MODIS. Geosci. Remote Sens. 1992, 30, 261–270. [Google Scholar] [CrossRef]
- Jordan, C. A soil-adjusted vegetation index (SAVI). Remote Sens. Environ. 1988, 25, 295–309. [Google Scholar]
- Broge, N.H.; Leblanc, E. Comparing prediction power and stability of broadband and hyperspectral vegetation indices for estimation of green leaf area index and canopy chlorophyll density. Remote Sens. Environ. 2001, 76, 156–172. [Google Scholar] [CrossRef]
- Gitelson, A.A. Wide dynamic range vegetation index for remote quantification of biophysical characteristics of vegetation. J. Plant Physiol. 2004, 161, 165–173. [Google Scholar] [CrossRef] [PubMed]
- Shannon, C.E. A Mathematical Theory of Communication. Bell Syst. Tech. J. 1948, 27, 379–423. [Google Scholar] [CrossRef]
- Marroquin, J.L.; Velasco, F.A.; Rivera, M.; Nakamura, M. Gauss-Markov Measure Field Models for Low-Level Vision. IEEE Trans. Pattern Anal. Mach. Intell. 2001, 23, 337–348. [Google Scholar] [CrossRef]
- Rivera, M.; Dalmau, O.S. Variational Viewpoint of the Quadratic Markov Measure Field Models: Theory and Algorithms. IEEE Trans. Image Process. 2011, 21, 1246–1257. [Google Scholar] [CrossRef] [PubMed]
- Cohen, J. A coefficient of agreement for nominal scales. Educ. Psychol. Meas. 1960, 20, 37–46. [Google Scholar] [CrossRef]
- Kohavi, R.; Provost, F. Guest Editors’ Introduction: On Applied Research in Machine Learning. Mach. Learn. 1998, 30, 127–132. [Google Scholar]
- Sokolova, M.; Lapalme, G. A systematic analysis of performance measures for classification tasks. Inf. Process. Manag. 2009, 45, 427–437. [Google Scholar] [CrossRef]
- Pulido, H.G.; Bautista, A.M.; Guevara, R.M. Jalisco Territorio y Problemas de Desarrollo; Instituto de Información Estadística y Geográfica: Zapopan, Mexico, 2013.
- Northrop, A.; Team, L.S. Ideas-Lansat Products Description Document, Technical Report; Telespazio VEGA UK Ltd.: Luton, UK, 2015. [Google Scholar]
- Visualization Viewer Site. Available online: http://glovis.usgs.gov/ (accessed on 13 June 2017).
- Tomasi, C.; Manduchi, R. Bilateral Filtering for Gray and Color Images. In Proceedings of the Sixth International Conference on Computer Vision, Bombay, India, 7 January 1998; pp. 839–846. [Google Scholar]
- Chen, D.; Huan, J.; Jackson, T.J. Vegetation water content estimation for corn and soybeans using spectral indices derived from MODIS near- and short-wave infrared bands. Remote Sens. Environ. 2005, 98, 225–236. [Google Scholar] [CrossRef]
- Jiang, H.; Feng, M.; Zhu, Y.; Lu, N.; Huang, J.; Xiao, T. An Automated Method for Extracting Rivers and Lakes from Landsat Imagery. Remote Sens. 2014, 6, 5067–5089. [Google Scholar] [CrossRef]
- Rokni, K.; Ahmad, A.; Selamat, A.; Hazini, S. Water Feature Extraction and Change Detection Using Multitemporal Landsat Imagery. Remote Sens. 2014, 6, 4173–4189. [Google Scholar] [CrossRef]
- Yashon, O.; Tateishi, R. A water index for rapid mapping of shoreline changes of five East African Rift Valley lakes: an empirical analysis using Landsat TM and ETM+ data. Int. J. Remote Sens. 2006, 27, 3153–3181. [Google Scholar]
- Ji, L.; Zhang, L.; Wylie, B. Analysis of Dynamic Thresholds for the Normalized Difference Water Index. Photogramm. Eng. Remote Sens. 2009, 75, 1307–1317. [Google Scholar] [CrossRef]
- Viera, A.; Garrett, J. Understanding interobserver agreement: The kappa statistic. Fam. Med. 2005, 37, 360–363. [Google Scholar] [PubMed]
- Marroquin, J.L.; Botello, S.; Calderon, F.; Vemuri, B.C. The MPM-MAP Algorithm for image segmentation. Pattern Recognit. 2000, 1, 303–308. [Google Scholar]
- MultiSpec Software. Available online: https://engineering.purdue.edu/~biehl/MultiSpec/ (accessed on 13 June 2017).
- Su, B.; Noguchi, N. Agricultural land use information extraction in Miyajimanuma wetland area based on remote sensing imagery. Environ. Control Biol. 2012, 50, 277–287. [Google Scholar] [CrossRef]
- Omkar, S.N.; Senthilnath, J.; Mudigere, D.; Kumar, M.M. Crop classification using biologically-inspired techniques with high resolution satellite image. J. Indian Soc. Remote Sens. 2008, 36, 175–182. [Google Scholar] [CrossRef]
- Karakahya, H.; Yazgan, B.; Ersoy, O.K. A Spectral-Spatial Classification Algorithm for Multispectral Remote Sensing Data. In Proceedings of the Artificial Neural Networks and Neural Information Processing, Istanbul, Turkey, 26–29 June 2003; pp. 1011–1017. [Google Scholar]
- Kettig, R.L.; Landgrebe, D.A. Computer Classification of Remotely Sensed Multispectral Image Data by Extraction and Classification of Homogeneous Objects. IEEE Trans. Geosci. Electron. 1976, 14, 19–26. [Google Scholar] [CrossRef]
- Landgrebe, D. The Development of a Spectral-Spatial Classifier for Earth Observational Data. Pattern Recognit. 1980, 12, 165–175. [Google Scholar] [CrossRef]
- Wolfowitz, J. The Minimum Distance Method. Ann. Math. Statist. 1957, 28, 75–88. [Google Scholar] [CrossRef]
- Hazewinkel, M. Maximum likelihood method. In Encyclopedia of Mathematics; Springer: Berlin, Germany, 2001. [Google Scholar]
- Mika, S.; Ratsch, G.; Weston, J.; Scholkopf, B.; Mullers, K.R. Fisher discriminant analysis with kernels. In Proceedings of the Neural Networks for Signal Processing IX, Madison, WI, USA, 25 August 1999; pp. 41–48. [Google Scholar]



| Class | Vegetation Name |
|---|---|
| C1 | Irrigation agriculture |
| C2 | Temporary agriculture |
| C3 | Forest |
| C4 | Scrub |
| C5 | Pastureland |
| TM Bands | Wavelength () | Features |
|---|---|---|
| TM1 | 0.45–0.52 | B (Blue) |
| TM2 | 0.52–0.60 | G (Green) |
| TM3 | 0.63–0.69 | R (Red) |
| TM4 | 0.76–0.90 | near infrared |
| TM5 | 1.55–1.75 | mid-infrared |
| TM6 | 10.4–12.50 | thermal infrared |
| TM7 | 2.08–2.35 | mid-infrared |
| Name VI | Formula | References |
|---|---|---|
| MSR | [31] | |
| CI | [32] | |
| midrule NDVI | [15] | |
| GNDVI | [16] | |
| EVI | [33] | |
| SARVI | [34] | |
| RDVI | [31] | |
| SAVI | [35] | |
| MSAVI | [36] | |
| WDRVI | [37] |
| Cohen’s Kappa | Interpretation |
|---|---|
| <0 | Poor agreement |
| 0.00–0.20 | Slight agreement |
| 0.21–0.40 | Fair agreement |
| 0.41–0.60 | Moderate agreement |
| 0.61–0.80 | Substantial agreement |
| 0.81–1.00 | Almost perfect agreement |
| N. Combination | Feature Space Combination |
|---|---|
| 1 | Space 1 |
| 2 | Space 2 |
| 3 | Space 3 |
| 4 | Space 1 + Space 2 |
| 5 | Space 1 + Space 3 |
| 6 | Space 2 + Space 3 |
| 7 | Space 1 + Space 2 + Space 3 |
| Experiment | Precision | Overall Accuracy | Cohen’s Kappa | ||||
|---|---|---|---|---|---|---|---|
| C1 | C2 | C3 | C4 | C5 | |||
| 1 | 0.86 | 0.83 | 0.89 | 0.85 | 0.63 | 0.8331 | 0.7520 |
| 2 | 0.84 | 0.81 | 0.89 | 0.87 | 0.62 | 0.8324 | 0.7528 |
| 3 | 0.83 | 0.82 | 0.91 | 0.89 | 0.65 | 0.8506 | 0.7801 |
| Experiment | Precision | Overall Accuracy | Cohen’s Kappa | ||||
|---|---|---|---|---|---|---|---|
| C1 | C2 | C3 | C4 | C5 | |||
| 4 | 0.86 | 0.83 | 0.92 | 0.87 | 0.67 | 0.8472 | 0.7726 |
| 5 | 0.89 | 0.84 | 0.94 | 0.88 | 0.70 | 0.8624 | 0.7955 |
| 6 | 0.85 | 0.83 | 0.91 | 0.89 | 0.64 | 0.8485 | 0.7770 |
| 7 | 0.87 | 0.84 | 0.93 | 0.88 | 0.67 | 0.8588 | 0.7904 |
| Experiment | Precision | Overall Accuracy | Cohen’s Kappa | ||||
|---|---|---|---|---|---|---|---|
| C1 | C2 | C3 | C4 | C5 | |||
| 8 | 0.88 | 0.83 | 0.92 | 0.87 | 0.67 | 0.8497 | 0.7767 |
| 9 | 0.90 | 0.84 | 0.94 | 0.89 | 0.71 | 0.8649 | 0.7993 |
| 10 | 0.88 | 0.82 | 0.91 | 0.89 | 0.66 | 0.8517 | 0.7812 |
| 11 | 0.90 | 0.83 | 0.93 | 0.89 | 0.69 | 0.8600 | 0.7923 |
| Method | Feature Space | C1 | C2 | C3 | C4 | C5 | Overall Accuracy | Kappa |
|---|---|---|---|---|---|---|---|---|
| MED [61] | Landsat-5 TM | 0.82 | 0.73 | 0.54 | 0.80 | 0.29 | 0.6244 | 0.4781 |
| ML [62] | Landsat-5 TM | 0.65 | 0.73 | 0.72 | 0.76 | 0.40 | 0.6928 | 0.5558 |
| FLL [63] | Landsat-5 TM | 0.88 | 0.74 | 0.71 | 0.75 | 0.46 | 0.7105 | 0.5743 |
| ESS [60] | Landsat-5 TM | 0.76 | 0.70 | 0.80 | 0.74 | 0.56 | 0.7257 | 0.5811 |
| SVM linear | Landsat-5 TM | 0.77 | 0.74 | 0.96 | 0.73 | 0.19 | 0.7498 | 0.6071 |
| SVM rbf | Landsat-5 TM | 0.79 | 0.83 | 0.90 | 0.90 | 0.66 | 0.8536 | 0.7859 |
| MICAI 2014 [29] | Space 1 | 0.86 | 0.83 | 0.89 | 0.85 | 0.63 | 0.8331 | 0.7520 |
| MICAI | Space 3 | 0.83 | 0.82 | 0.91 | 0.89 | 0.65 | 0.8506 | 0.7801 |
| MICAI 2015 [28] | Spaces 1 & 2 | 0.86 | 0.83 | 0.92 | 0.87 | 0.67 | 0.8472 | 0.7726 |
| MICAI | Spaces 1 & 3 | 0.89 | 0.84 | 0.94 | 0.88 | 0.70 | 0.8624 | 0.7955 |
| Proposal | Spaces 1 & 3 | 0.90 | 0.84 | 0.94 | 0.89 | 0.71 | 0.8649 | 0.7993 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dalmau, O.S.; Alarcón, T.E.; Oliva, F.E. Crop Classification in Satellite Images through Probabilistic Segmentation Based on Multiple Sources. Sensors 2017, 17, 1373. https://doi.org/10.3390/s17061373
Dalmau OS, Alarcón TE, Oliva FE. Crop Classification in Satellite Images through Probabilistic Segmentation Based on Multiple Sources. Sensors. 2017; 17(6):1373. https://doi.org/10.3390/s17061373
Chicago/Turabian StyleDalmau, Oscar S., Teresa E. Alarcón, and Francisco E. Oliva. 2017. "Crop Classification in Satellite Images through Probabilistic Segmentation Based on Multiple Sources" Sensors 17, no. 6: 1373. https://doi.org/10.3390/s17061373





