A Fast, Efficient Domain Adaptation Technique for Cross-Domain Electroencephalography(EEG)-Based Emotion Recognition
Abstract
:1. Introduction
- In contrast to the conventional domain adaptation method, we combine the marginal and conditional distribution adaptation strategies in the subspace, meaning that the new feature is effective and robust for substantial distribution discrepancies;
- Although the classifier in our proposed method needs to be updated along with the newly obtained EEG data, the computational efficiency is much better than standard domain adaptation, and it indicates the capability for the online implementation of the proposed algorithm when the number of training samples and test samples is controlled within certain range;
- Our work is intended to provide evaluation and analysis of several state-of-the-art domain adaptation techniques in the field of pattern recognition for researchers aiming to apply these to non-stationary EEG signal classification.
2. Experiment Design
3. Proposed Method
3.1. Feature Extraction and Normalization
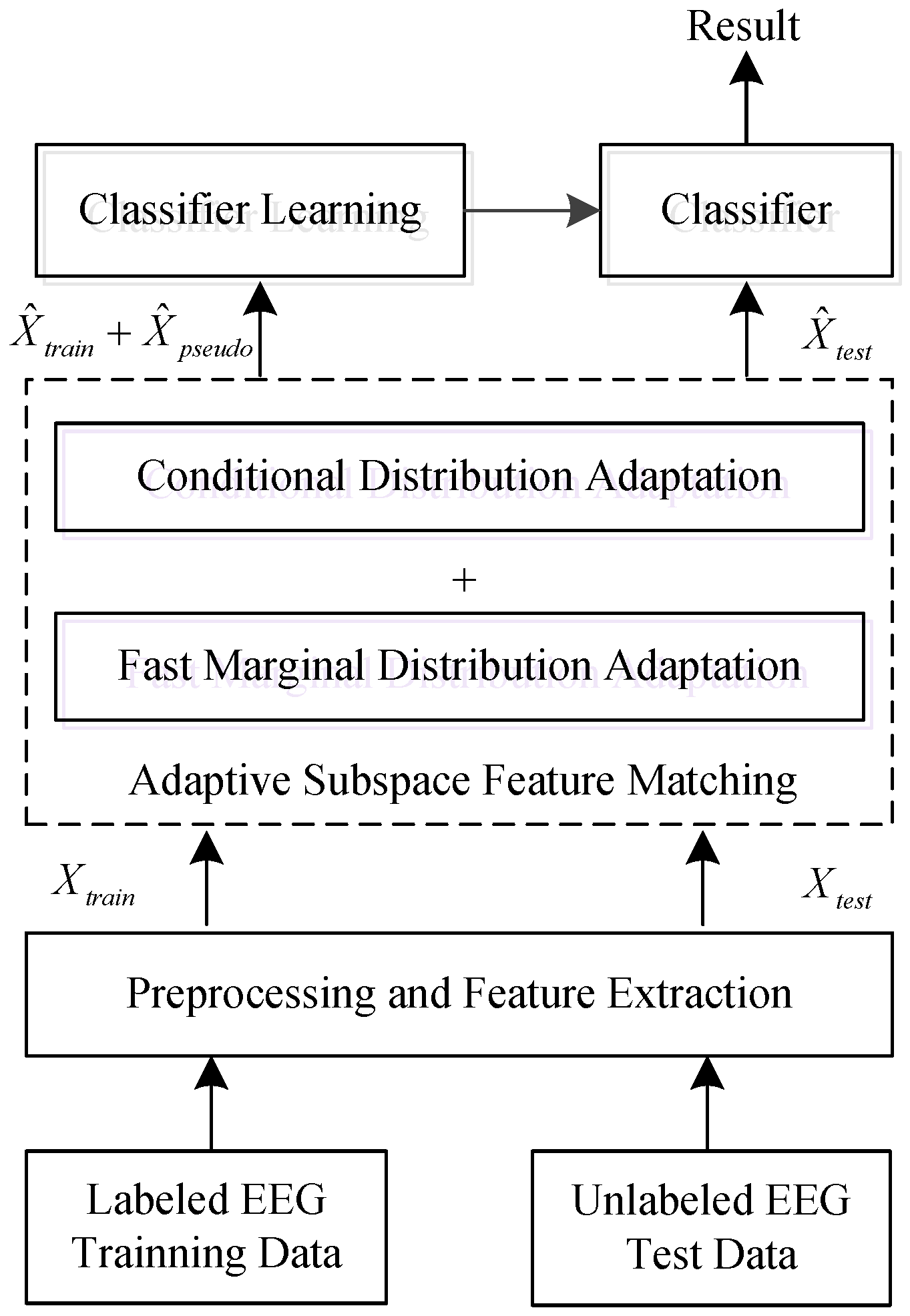
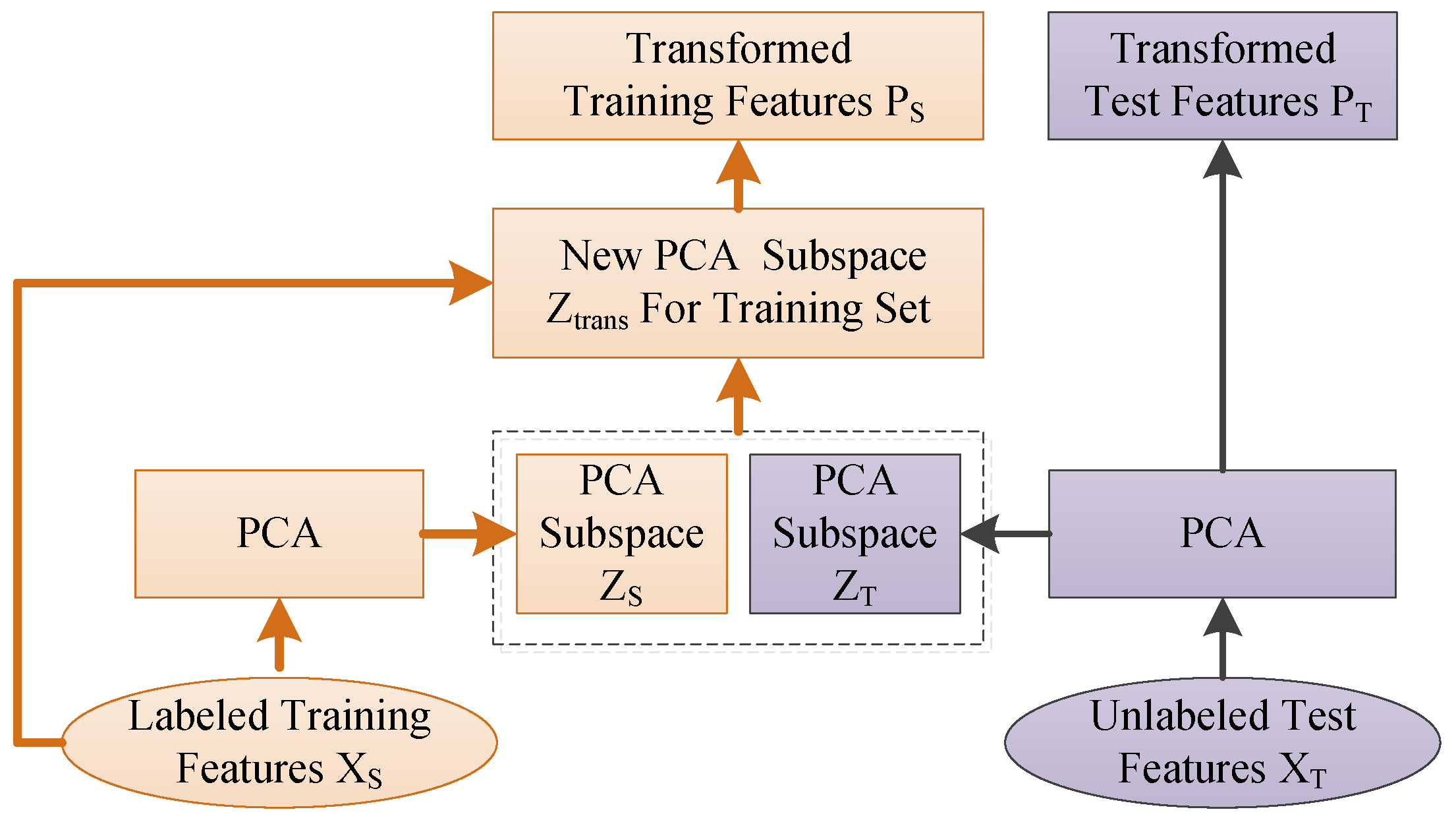
3.2. Adaptive Subspace Feature Matching
3.2.1. Fast Marginal Distribution Adaptation
3.2.2. Conditional Distribution Adaptation
| Algorithm 1 ASFM: adaptive subspace feature matching and classification. |
| Require: Source data , Target data , Source labels , Threshold , Subspace dimension g, Maximum Iterations: maxepoch Ensure: are the predicted labels for Target data .
|
4. Experiment Results
4.1. Off-Line Evaluation
- Auto-encoder (AE) [28];
- Transfer sparse coding (TSC) [27];
- Transfer component analysis (TCA) [24].
- Transfer joint matching (TJM) [26];
- Subspace alignment auto-encoder (SAAE) [34];
- Subspace feature matching (SFM) without adaptive strategy;
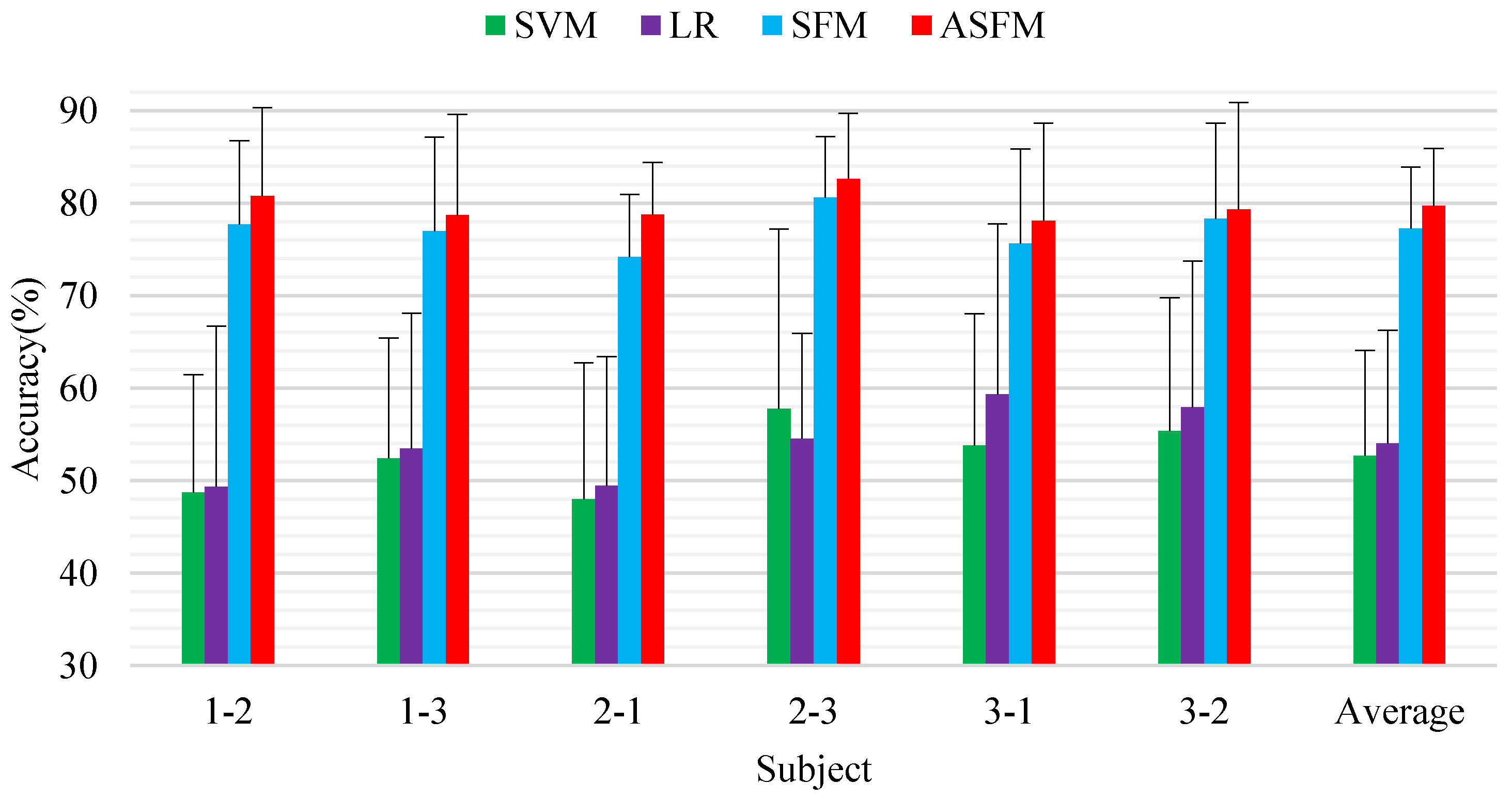
4.1.1. Subject-To-Subject Experimental Results
4.1.2. Session-to-Session Experimental Results
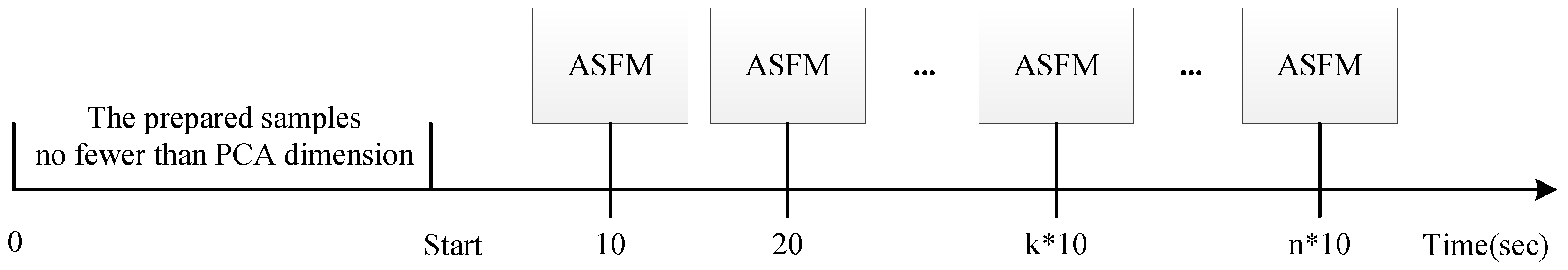
4.2. On-Line Evaluation
5. Discussion
5.1. Comparison with State-of-the-Art Methods
5.2. Computation Time Analysis
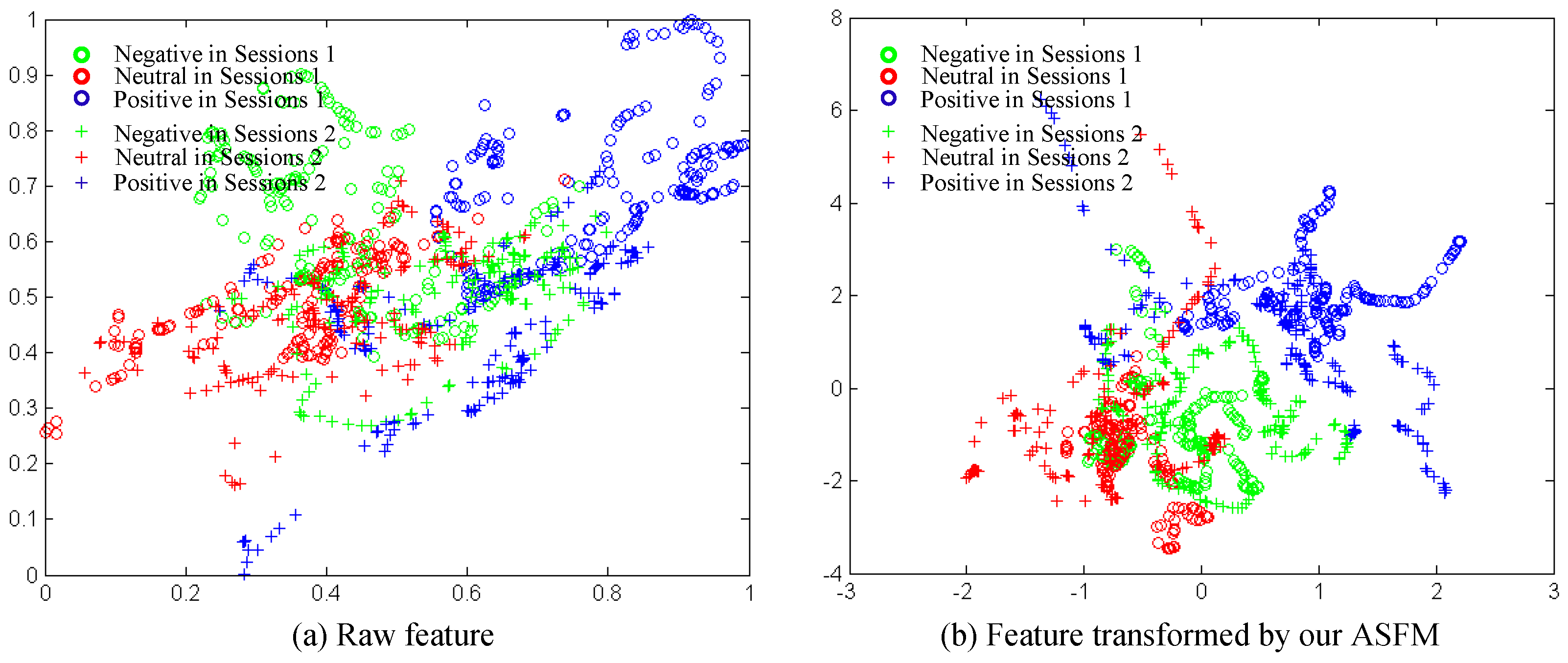
5.3. Effectiveness Analysis
5.4. Online Trajectory Analysis
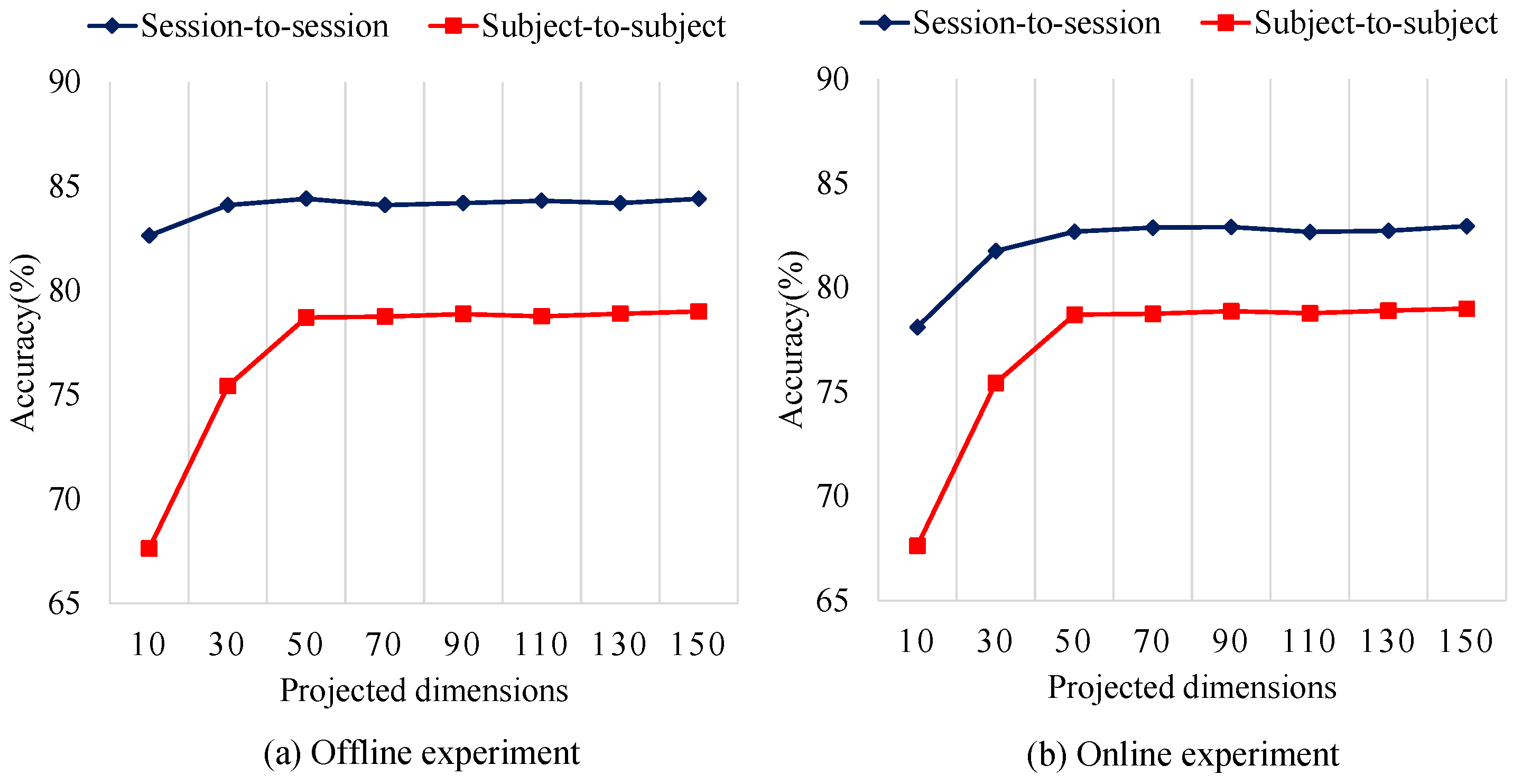
5.5. Parameter Sensitivity
5.6. The limitations of ASFM
- (1)
- As some works [39,40,41] have mentioned, although there are many successful applications, the power of transfer learning still has its limit, which is named “negative transfer”. Negative transfer refers to the phenomenon that, instead of improving performance, transfer learning from source domain degrades the performance on the target domain. In general, transfer learning methods treat knowledge from every source domain as a valuable contribution to the task on the target domain. However, when knowledge is transferred from highly irrelevant sources, “negative transfer” will happen. In fact, given multiple source domains, in the worst case, the majority of the source domains could be irrelevant to the target domain. In reality, ASFM is not effective on DEAP [32] which is a frequently-used database for EEG-based emotion estimation. This is probably because there are too many irrelevant subsets between each other in this dataset. So far, it is still difficult to estimate the relevance between two subjects without labeled data in the target subject. Therefore, unsupervised transfer learning (or domain adaptation) research for EEG-based emotion estimation on DEAP database is a greater challenge, and an important research aspect in our future work.
- (2)
- The complexity of our method is still limited by the number of training samples and test samples. In order to use our method for online implementation, the number of training samples and test samples should be paid attention, for example by enlarging the sampling interval of the EEG features when necessary.
6. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Deshmukh, S.; Patwardhan, M.; Mahajan, A. Survey on real-time facial expression recognition techniques. IET Biom. 2016, 5, 155–163. [Google Scholar]
- Yan, J.; Zheng, W.; Xu, Q. Sparse Kernel Reduced-Rank Regression for Bimodal Emotion Recognition From Facial Expression and Speech. IEEE Trans. Multimed. 2016, 18, 1319–1329. [Google Scholar] [CrossRef]
- Agrafioti, F.; Hatzinakos, D.; Anderson, A.K. ECG Pattern Analysis for Emotion Detection. IEEE Trans. Affect. Comput. 2014, 5, 227–237. [Google Scholar] [CrossRef]
- Gruebler, A.; Suzuki, K. Design of a Wearable Device for Reading Positive Expressions from Facial EMG Signals. IEEE Trans. Affect. Comput. 2011, 3, 102–115. [Google Scholar] [CrossRef]
- Liu, N.; Chiang, C.; Hsu, H. Improving driver alertness through music selection using a mobile EEG to detect brainwaves. Sensors 2013, 13, 8199–8221. [Google Scholar] [CrossRef] [PubMed]
- Sauvet, F.; Bougard, C.; Coroenne, M. In flight automatic detection of vigilance states using a single EEG channel. IEEE Trans. Biomed. Eng. 2014, 61, 2840–2847. [Google Scholar] [CrossRef] [PubMed]
- Muhl, C.; Jeunet, C.; Lotte, F. EEG-based workload estimation across affective contexts. Front. Neurosci. 2014, 8, 114. [Google Scholar] [PubMed]
- Chung, M.; Cheung, W.; Scherer, R.; Rao, R.P. A hierarchical architecture or adaptive brain-computer interfacing. In Proceedings of the International Joint Conference on Artificial Intelligence, Barcelona, Spain, 16–22 July 2011; pp. 1647–1652. [Google Scholar]
- Zander, T.; Jatzev, S. Context-aware brain–computer interfaces: Exploring the information space of user, technical system and environment. J. Neural Eng. 2012, 9, 16003–16012. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Chen, M.; Zhao, S. ReliefF-Based EEG Sensor Selection Methods for Emotion Recognition. Sensors 2016, 16, 1558. [Google Scholar] [CrossRef] [PubMed]
- Sh, L.; Jiao, Y.; Lu, B. Differential entropy feature for EEG-based vigilance estimation. In Proceedings of the 35th Annual International Conference of the IEEE Engineering in Medicine and Biology Society (EMBC), Osaka, Japan, 3–7 July 2013; pp. 6627–6630. [Google Scholar]
- Duan, R.; Zhu, J.; Lu, B. Differential entropy feature for EEG-based emotion classification. In Proceedings of the 6th International IEEE/EMBS Conference on Neural Engineering (NER), San Diego, CA, USA, 6–8 November 2013; pp. 81–84. [Google Scholar]
- Zheng, W.L.; Lu, B.L. Investigating critical frequency bands and channels for EEG-based emotion recognition with deep neural networks. IEEE Trans. Auton. Ment. Dev. 2015, 7, 162–175. [Google Scholar] [CrossRef]
- Morioka, H.; Kanemura, A. Learning a common dictionary for subject-transfer decoding with resting calibration. NeuroImage 2015, 111, 167–178. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Kambara, H.; Koike, Y.; Sugiyama, M. Application of covariate shift adaptation techniques in brain-computer interfaces. IEEE Trans. Bio-Med. Eng. 2010, 57, 1318–1324. [Google Scholar]
- Buttfield, A.; Ferrez, P.W.; Millán, J.d.R. Towards a robust BCI: Error potentials and online learning. IEEE Trans. Neural Syst. Rehabil. Eng. 2006, 14, 164–168. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.; Miyapuram, K.P.; Bapi, R.S. Detection of cognitive states from fMRI data using machine learning techniques. In Proceedings of the International Joint Conference on Artificial Intelligence, Hyderabad, India, 6–12 January 2007; pp. 587–592. [Google Scholar]
- Jenke, R.; Peer, A.; Buss, M. Feature extraction and selection for emotion recognition from EEG. IEEE Trans. Affect. Comput. 2014, 5, 327–339. [Google Scholar] [CrossRef]
- Abraham, A.; Pedregosa, F. Machine learning for neuroimaging with scikit-learn. Front. Neuroinform. 2014, 8, 14. [Google Scholar] [CrossRef] [PubMed]
- Luo, J.; Feng, Z.; Zhang, J.; Lu, N. Dynamic frequency feature selection based approach for classification of motor imageries. Comput. Biol. Med. 2016, 75, 45–53. [Google Scholar] [CrossRef] [PubMed]
- Zheng, W.L.; Zhang, Y.Q.; Zhu, J.; Lu, B.L. Transfer components between subjects for EEG-based emotion recognition. In Proceedings of the International Conference on Affective Computing and Intelligent Interaction, Xi’an, China, 21–24 September 2015; pp. 917–922. [Google Scholar]
- Zheng, W.L.; Lu, B.L. Personalizing EEG-based Affective Models with Transfer Learning. In Proceedings of the 25th International Joint Conference on Artificial Intelligence, New York, NY, USA, 9–15 July 2016; pp. 2732–3738. [Google Scholar]
- Jayaram, V.; Alamgir, M.; Altun, Y. Transfer Learning in Brain-Computer Interfaces. IEEE Comput. Intell. Mag. 2016, 11, 20–31. [Google Scholar] [CrossRef]
- Pan, S.J.; Tsang, I.W.; Kwok, J.T.; Yang, Q. Domain Adaptation via Transfer Component Analysis. IEEE Trans. Neural Netw. 2011, 22, 199–210. [Google Scholar] [CrossRef] [PubMed]
- Gretton, A.; Borgwardt, K.M.; Rasch, M. A kernel method for the two-sample-problem. In Advances in Neural Information Processing Systems; The MIT Press: Cambridge, MA, USA, 2006; pp. 513–520. [Google Scholar]
- Long, M.; Wang, J.; Ding, G. Transfer Joint Matching for Unsupervised Domain Adaptation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Columbus, OH, USA, 23–28 June 2014; pp. 1410–1417. [Google Scholar]
- Mingsheng, L.; Guiguang, D.; Jianmin, W. Transfer Sparse Coding for Robust Image Representation. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Portland, OR, USA, 23–28 June 2013; Volume 9, pp. 407–414. [Google Scholar]
- Glorot, X.; Bordes, A.; Bengio, Y. Domain Adaptation for Large-Scale Sentiment Classification: A Deep Learning Approach. In Proceedings of the International Conference on Machine Learning, Bellevue, WA, USA, 28 June–2 July 2011; pp. 513–520. [Google Scholar]
- Deng, J.; Zhang, Z.; Eyben, F.; Schuller, B. Autoencoder-based Unsupervised Domain Adaptation for Speech Emotion Recognition. IEEE Signal Process. Lett. 2014, 21, 1068–1072. [Google Scholar]
- Kan, M.; Shan, S.; Chen, X. Bi-Shifting Auto-Encoder for Unsupervised Domain Adaptation. In Proceedings of the IEEE International Conference on Computer Vision, Santiago, Chile, 7–13 December 2015; pp. 3846–3854. [Google Scholar]
- Yin, Z.; Wang, Y.; Liu, L.; Zhang, W.; Zhang, J. Cross-subject EEG feature selection for emotion recognition using transfer recursive feature elimination. Front. Neurorobot. 2017. [Google Scholar] [CrossRef] [PubMed]
- Koelstra, S.; Muhl, C.; Soleymani, M.; Lee, J.S.; Yazdani, A.; Ebrahimi, T.; Pun, T.; Nijholt, A.; Patras, I. DEAP: A Database for Emotion Analysis Using Physiological Signals. IEEE Trans. Affect. Comput. 2012, 3, 18–31. [Google Scholar] [CrossRef]
- Yin, Z.; Zhang, J. Cross-session classification of mental workload levels using EEG and an adaptive deep learning model. Biomed. Signal Process. Control 2017, 33, 30–47. [Google Scholar] [CrossRef]
- Chai, X.; Wang, Q.; Zhao, Y. Unsupervised domain adaptation techniques based on auto-encoder for non-stationary EEG-based emotion recognition. Comput. Biol. Med. 2016, 79, 205–214. [Google Scholar] [CrossRef] [PubMed]
- Fernando, B.; Habrard, A.; Sebban, M. Unsupervised Visual Domain Adaptation Using Subspace Alignment. In Proceedings of the International Conference on Computer Vision, Sydney, Australia, 1–8 December 2013; pp. 2960–2967. [Google Scholar]
- Chang, C.; Lin, C. Libsvm: A library for support vector machines. ACM Trans. Intell. Syst. Technol. 2011, 2, 27. [Google Scholar] [CrossRef]
- Fan, R.; Chang, K.; Hsieh, C.; Wang, X.; Lin, C. LIBLINEAR: A library for large linear classification. J. Mach. Learn. Res. 2008, 9, 1871–1874. [Google Scholar]
- Zheng, W.L.; Zhu, J.Y.; Lu, B. Identifying Stable Patterns over Time for Emotion Recognition from EEG. arXiv, 2016; arXiv:1601.02197. [Google Scholar]
- Pan, S.J.; Yang, Q. A Survey on Transfer Learning. IEEE Trans. Knowl. Data Eng. 2010, 22, 1345–1359. [Google Scholar] [CrossRef]
- Weiss, K.; Khoshgoftaar, T.; Wang, D. A survey of transfer learning. J. Big Data 2016, 3, 9. [Google Scholar] [CrossRef]
- Ge, L.; Gao, J.; Ngo, H.; Li, K.; Zhang, A. On Handling Negative Transfer and Imbalanced Distributions in Multiple Source Transfer Learning. Stat. Anal. Data Min. 2014, 7, 254–271. [Google Scholar] [CrossRef]



| Method | Parameter Details |
|---|---|
| SVM | Linear kernel; the parameter C was set by searching ; |
| LR | L2-regularized logistic regression; We set the parameter C by searching ; |
| AE | Structure with 2 hidden layers; The number of hidden neurons was set by searching ; |
| TSC | Number of basis vectors was 150; Sparsity regularization was 0.1; Number of iterations per TSC was set as 15; |
| TCA | Subspace bases was set as 80; Regularizer was set by searching ; |
| TJM | Subspace bases was set as 80; The number of iterations for TJM to converge is T = 3. Regularizer was set by searching ; |
| ASFM | The dimension of the PCA subspace is set to all; Threshold is set to 0.45; the number of iterations is set to 1. |
| Task Group No. | Experimental Session 1 | Experimental Session 2 | Experimental Session 3 | Average |
|---|---|---|---|---|
| SVM | 60.31/9.53 | 53.37/12.97 | 58.20/12.40 | 57.29/7.42 |
| LR | 59.84/10.12 | 54.32/14.42 | 58.12/12.58 | 57.43/7.95 |
| AE [28] | 65.64/10.44 | 55.82/11.52 | 63.67/12.31 | 61.71/8.09 |
| TSC [27] | 73.33/7.61 | 72.16/9.08 | 67.03/9.79 | 70.84/6.44 |
| TCA [24] | 75.91/11.52 | 74.19/12.26 | 75.87/6.87 | 75.32/11.07 |
| TJM [26] | 77.62/10.90 | 74.30/12.06 | 76.79/5.69 | 76.24/10.38 |
| SAAE * [34] | 80.22/8.00 | 74.68/12.76 | 78.73/12.96 | 77.88/7.33 |
| SFM | 80.34/6.13 | 74.81/10.51 | 77.74/10.15 | 77.63/5.84 |
| ASFM |
| Task Group No. | Average | ||||||
|---|---|---|---|---|---|---|---|
| SVM | 67.63/12.73 | 64.21/13.00 | 67.62/14.73 | 70.37/19.48 | 72.69/14.21 | 66.64/14.38 | 68.19/11.41 |
| LR | 60.64/17.38 | 60.42/14.61 | 58.82/13.97 | 65.30/11.37 | 63.89/18.44 | 63.85/15.79 | 62.15/9.51 |
| GELM * [38] | 72.55/10.29 | 67.22/10.42 | 75.86/7.71 | 76.62/15.34 | 76.28/11.47 | 78.17/13.41 | 74.45/8.20 |
| AE [28] | 76.66/8.92 | 75.30/10.83 | 77.47/11.54 | 77.20/15.35 | 77.02/12.81 | 78.21/13.15 | 76.98/9.52 |
| TSC [27] | 79.85/12.12 | 80.71/10.70 | 82.07/8.08 | 80.24/10.32 | 79.92/7.71 | 77.99/11.36 | 80.13/8.52 |
| TCA [24] | 81.56/11.52 | 79.35/12.26 | 81.56/6.87 | 82.83/11.07 | 80.84/8.00 | 77.97/13.90 | 80.68/7.70 |
| TJM [26] | 82.43/10.90 | 80.89/12.06 | 82.36/5.69 | 84.04/10.38 | 80.57/9.97 | 79.09/11.69 | 81.56/7.47 |
| SAAE * [34] | 80.04/13.03 | 84.31/7.21 | 83.09/9.98 | 80.20/9.99 | 78.77/10.49 | 81.81/7.56 | |
| SFM | 83.09/10.12 | 81.61/10.42 | 84.11/6.23 | 84.21/8.19 | 84.62/9.59 | 82.16/11.06 | 83.30/7.12 |
| ASFM | 84.34/10.18 |
| Method | Parameter Details |
|---|---|
| SVM | Linear kernel; the parameter C was set by searching ; |
| LR | L2-regularized logistic regression; we set the parameter C by searching ; |
| SFM | The dimension of the PCA subspace is set to all. |
| ASFM | The dimension of the PCA subspace is set to all; Threshold is set to 0.41; The number of iterations is set to 1. |
| SVM | LR | AE | TSC | TCA | TJM | SAAE * | SFM | ASFM | |
|---|---|---|---|---|---|---|---|---|---|
| Training time (s) | 0.20 | 0.29 | 90.14 | 246.39 | 58.08 | 164.92 | 121.81 | 0.39 | 1.39 |
| Subspace Dimension | 10 | 30 | 50 | 70 | 90 | 110 | 130 | 150 | 250 | 325 |
|---|---|---|---|---|---|---|---|---|---|---|
| Training time (s) | 0.19 | 0.26 | 0.30 | 0.37 | 0.45 | 0.50 | 0.58 | 0.63 | 1.01 | 1.39 |
| 0.1 | 0.2 | 0.3 | 0.4 | 0.45 | 0.5 | 0.6 | 0.7 | 0.8 | 0.9 | |
|---|---|---|---|---|---|---|---|---|---|---|
| Classification accuracy (%) | 77.45 | 77.48 | 78.23 | 79.49 | 80.46 | 79.73 | 78.31 | 77.95 | 77.82 | 77.71 |
| Number of Iterations | 0 | 1 | 2 | 3 | 4 | 5 | 6 | 7 |
|---|---|---|---|---|---|---|---|---|
| Classification accuracy (%) | 77.63 | 80.46 | 80.69 | 80.90 | 81.05 | 81.08 | 81.09 | 81.09 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Chai, X.; Wang, Q.; Zhao, Y.; Li, Y.; Liu, D.; Liu, X.; Bai, O. A Fast, Efficient Domain Adaptation Technique for Cross-Domain Electroencephalography(EEG)-Based Emotion Recognition. Sensors 2017, 17, 1014. https://doi.org/10.3390/s17051014
Chai X, Wang Q, Zhao Y, Li Y, Liu D, Liu X, Bai O. A Fast, Efficient Domain Adaptation Technique for Cross-Domain Electroencephalography(EEG)-Based Emotion Recognition. Sensors. 2017; 17(5):1014. https://doi.org/10.3390/s17051014
Chicago/Turabian StyleChai, Xin, Qisong Wang, Yongping Zhao, Yongqiang Li, Dan Liu, Xin Liu, and Ou Bai. 2017. "A Fast, Efficient Domain Adaptation Technique for Cross-Domain Electroencephalography(EEG)-Based Emotion Recognition" Sensors 17, no. 5: 1014. https://doi.org/10.3390/s17051014






