Assessing Genetic Diversity after Mangrove Restoration in Brazil: Why Is It So Important?
Abstract
:1. Introduction
2. Materials and Methods
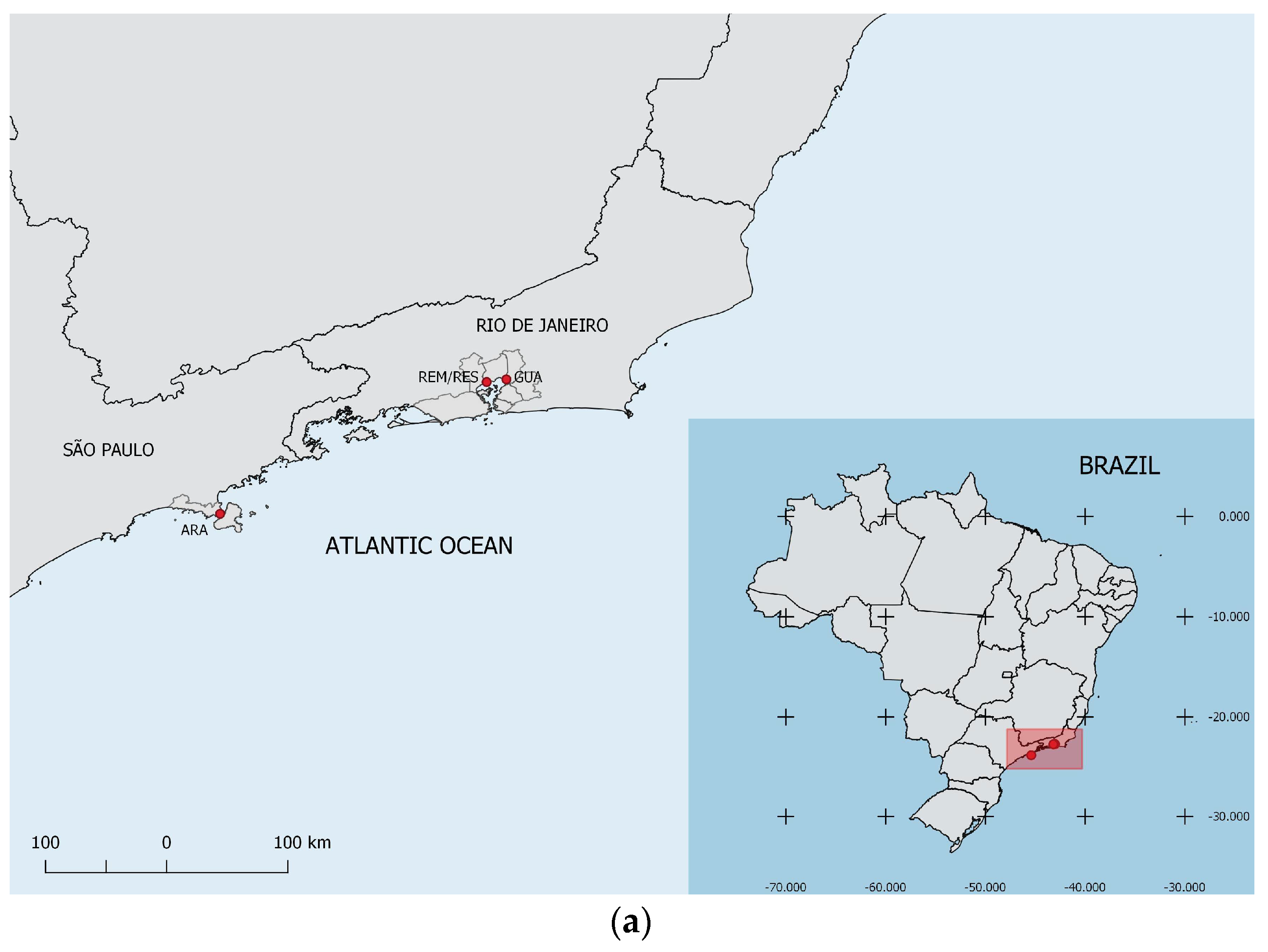
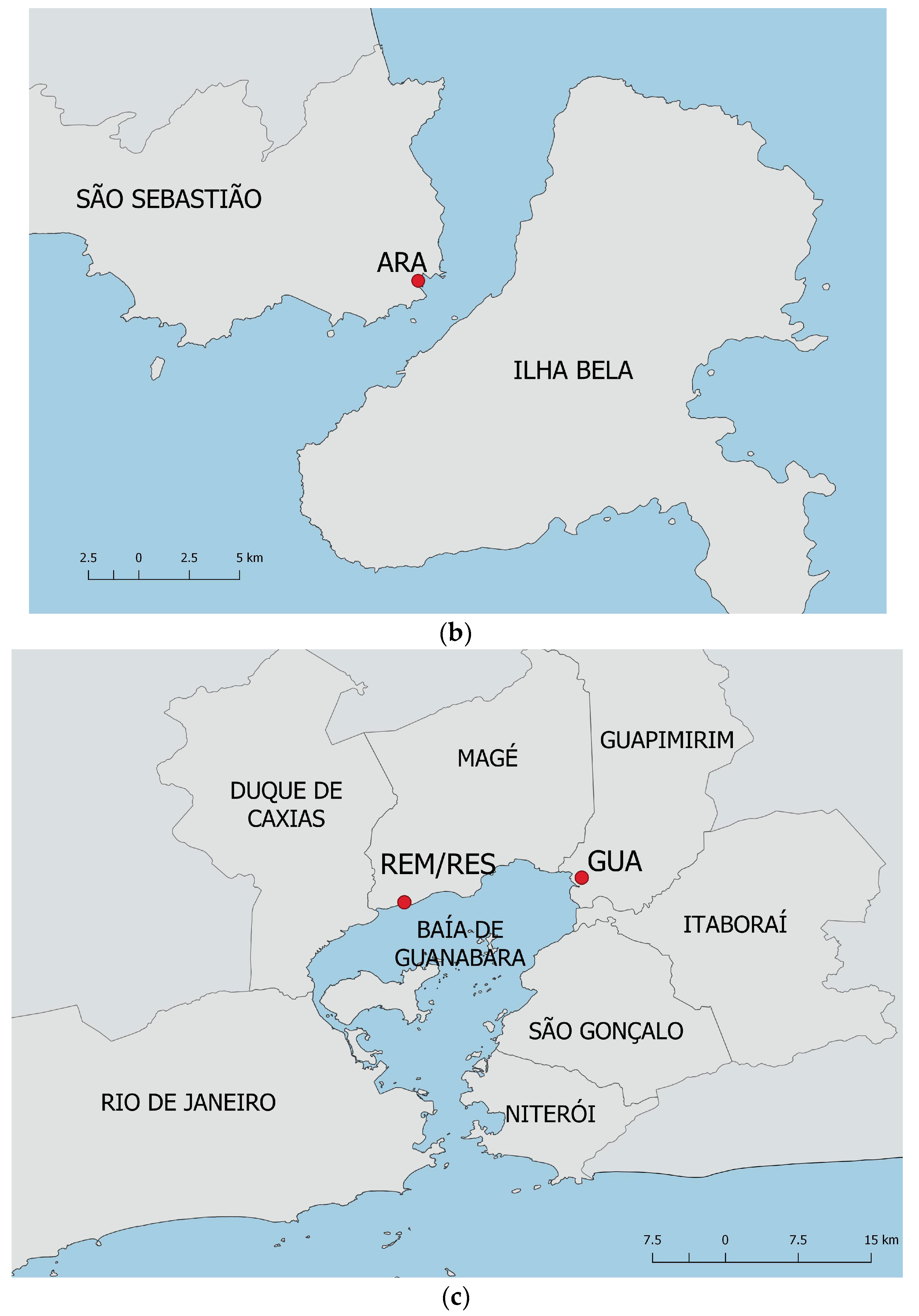
2.1. Studied Areas and Plant Material
2.2. DNA Isolation and PCR Amplification
2.3. Data Analysis
3. Results
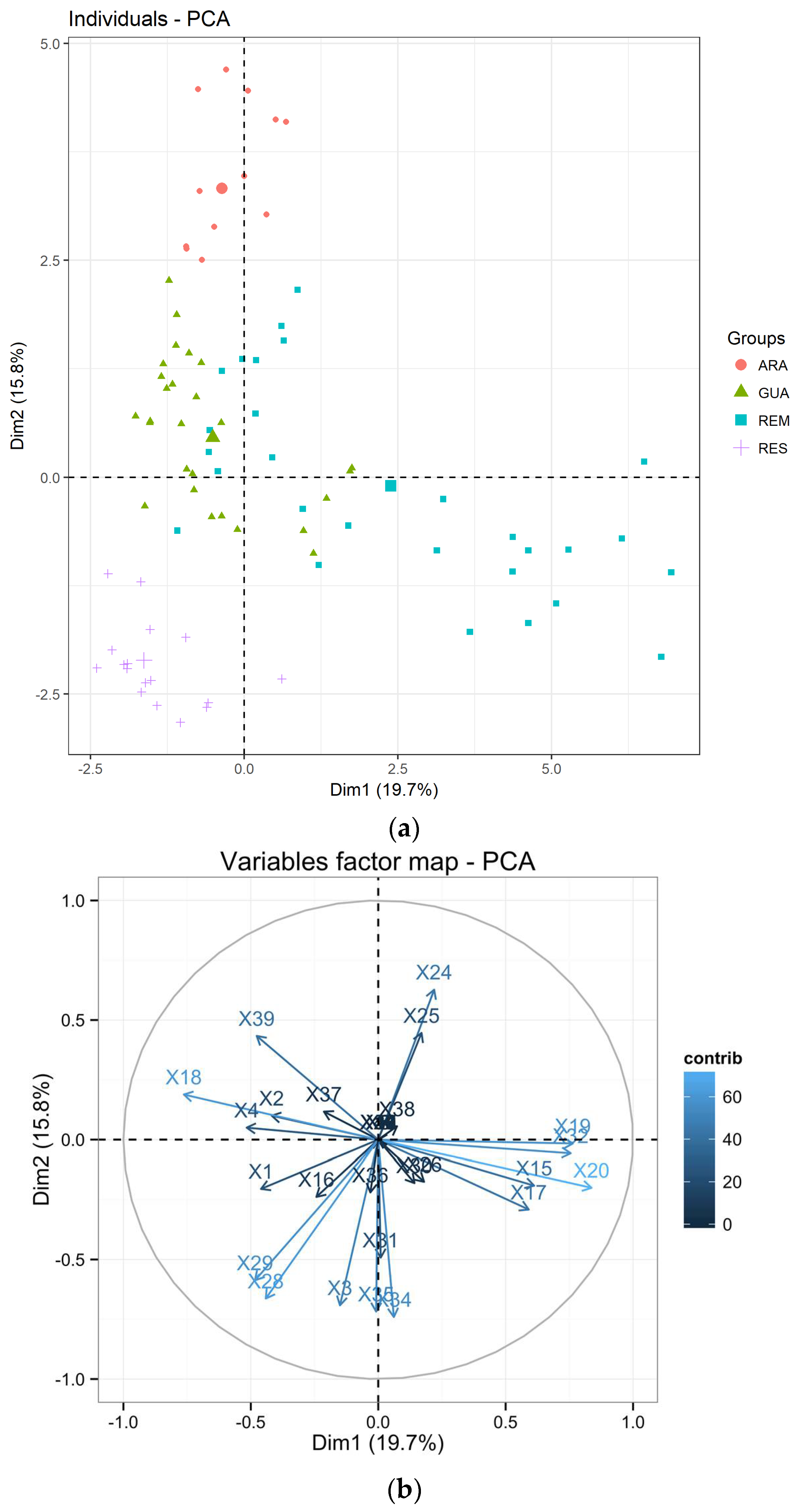
3.1. Genetic Diversity of Laguncularia racemosa
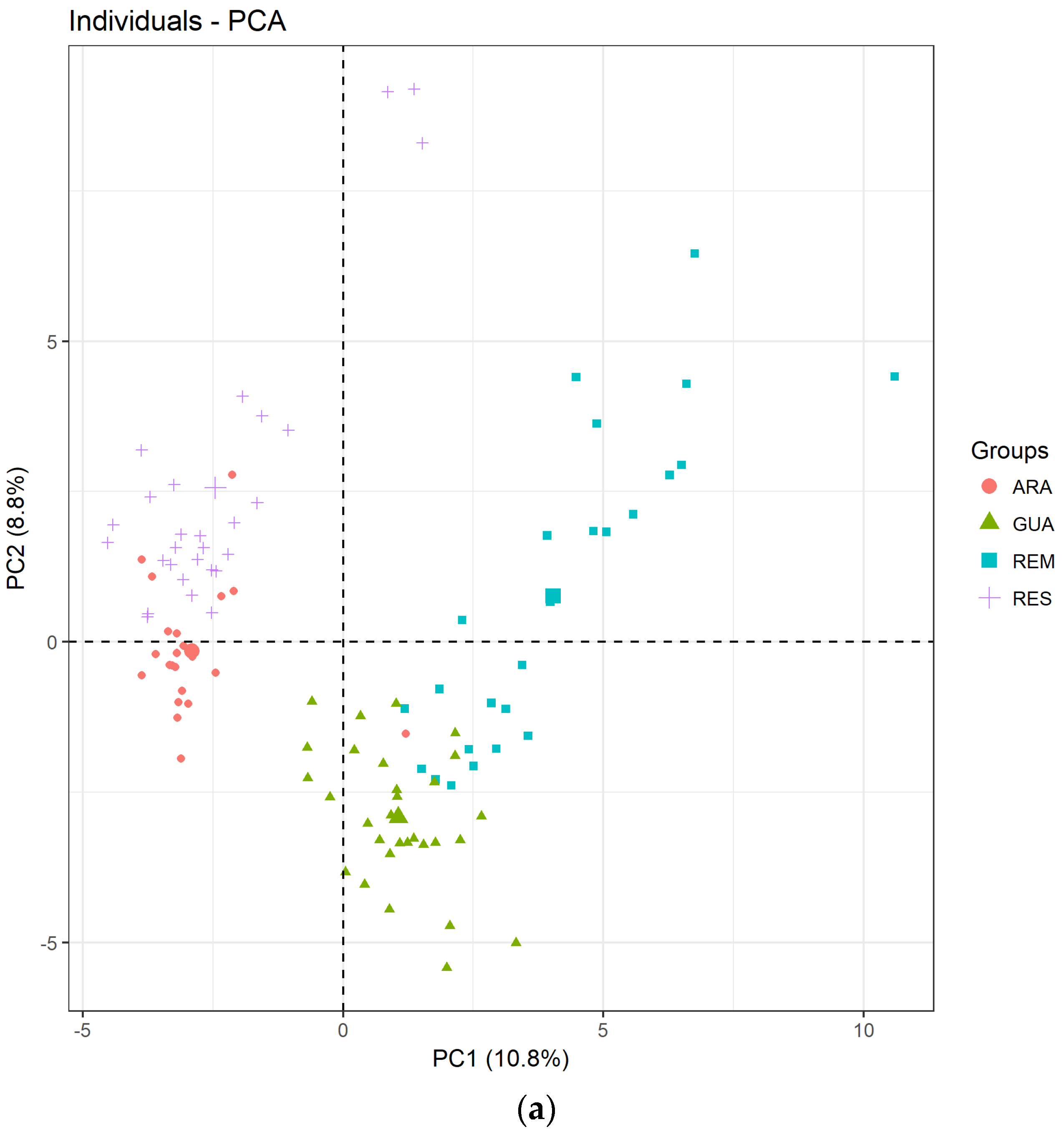
3.2. Genetic Diversity of Avicennia schaueriana
4. Discussion
4.1. Mangrove Restoration Practices Caused Bottleneck in Introduced Laguncularia racemosa
4.2. Avicennia schaueriana Has Similar Within-Population Genetic Diversity in Fragmented, Impacted, and Restored Areas
4.3. Pitfalls of Mangrove Restoration
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Schaeffer-Novelli, Y.; Cintron-Molero, G.; Soares, M.L.G. Chapter Nine Mangroves as indicators of sea level change in the muddy coasts of the world. In Proceedings in Marine Science; Elsevier: New York, NY, USA, 2002; Volume 4, pp. 245–262. ISBN 1568-2692. [Google Scholar]
- Tomlinson, P.B. The Botany of Mangroves; Cambridge University Press: Cambridge, UK, 1986. [Google Scholar]
- Erwin, K.L. Wetlands and global climate change: The role of wetland restoration in a changing world. Wetl. Ecol. Manag. 2009, 17, 71–84. [Google Scholar] [CrossRef]
- Duke, N.C.; Meynecke, J.-O.; Dittmann, S.; Ellison, A.M.; Anger, K.; Berger, U.; Cannicci, S.; Diele, K.; Ewel, K.C.; Field, C.D.; et al. A World Without Mangroves? Science 2007, 317, 41–42. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Alongi, D.M. Carbon Cycling and Storage in Mangrove Forests. Annu. Rev. Mar. Sci. 2014, 6, 195–219. [Google Scholar] [CrossRef] [PubMed]
- Donato, D.C.; Kauffman, J.B.; Murdiyarso, D.; Kurnianto, S.; Stidham, M.; Kanninen, M. Mangroves among the most carbon-rich forests in the tropics. Nat. Geosci. 2011, 4, 293–297. [Google Scholar] [CrossRef]
- Sánchez-Andrés, R.; Sánchez-Carrillo, S.; Alatorre, L.C.; Cirujano, S.; Álvarez-Cobelas, M. Litterfall dynamics and nutrient decomposition of arid mangroves in the Gulf of California: Their role sustaining ecosystem heterotrophy. Estuar. Coast. Shelf Sci. 2010, 89, 191–199. [Google Scholar] [CrossRef]
- Colmer, T.D.; Flowers, T.J. Flooding tolerance in halophytes. New Phytol. 2008, 179, 964–974. [Google Scholar] [CrossRef] [PubMed]
- Flowers, T.J.; Colmer, T.D. Salinity tolerance in halophytes. New Phytol. 2008, 179, 945–963. [Google Scholar] [CrossRef] [PubMed]
- Krauss, K.W.; Ball, M.C. On the halophytic nature of mangroves. Trees 2013, 27, 7–11. [Google Scholar] [CrossRef]
- Polidoro, B.A.; Carpenter, K.E.; Collins, L.; Duke, N.C.; Ellison, A.M.; Ellison, J.C.; Farnsworth, E.J.; Fernando, E.S.; Kathiresan, K.; Koedam, N.E.; et al. The Loss of Species: Mangrove Extinction Risk and Geographic Areas of Global Concern. PLoS ONE 2010, 5, e10095. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- FAO. The World’s Mangroves; FAO: Rome, Italy, 2007; ISBN 978-92-5-105856-5. [Google Scholar]
- Alongi, D.M. Present state and future of the world’s mangrove forests. Environ. Conserv. 2002, 29, 331–349. [Google Scholar] [CrossRef]
- Martinuzzi, S.; Gould, W.A.; Lugo, A.E.; Medina, E. Conversion and recovery of Puerto Rican mangroves: 200 years of change. For. Ecol. Manag. 2009, 257, 75–84. [Google Scholar] [CrossRef]
- Amador, E. Baía de Guanabara e Ecossistemas Periféricos: Homem e Natureza; Instituto de Geociências/UFRJ: Rio de Janeiro, Brazil, 1997. [Google Scholar]
- Fahrig, L. Effects of habitat fragmentation on biodiversity. Annu. Rev. Ecol. Evol. Syst. 2003, 34, 487–515. [Google Scholar] [CrossRef]
- Haddad, N.M.; Brudvig, L.A.; Clobert, J.; Davies, K.F.; Gonzalez, A.; Holt, R.D.; Lovejoy, T.E.; Sexton, J.O.; Austin, M.P.; Collins, C.D.; et al. Habitat fragmentation and its lasting impact on Earth’s ecosystems. Sci. Adv. 2015, 1, e1500052. [Google Scholar] [CrossRef] [PubMed]
- Kathiresan, K.; Bingham, B.L. Biology of mangroves and mangroves ecosystems. Adv. Mar. Biol. 2001, 40, 81–251. [Google Scholar]
- Friess, D.A.; Krauss, K.W.; Horstman, E.M.; Balke, T.; Bouma, T.J.; Galli, D.; Webb, E.L. Are all intertidal wetlands naturally created equal? Bottlenecks, thresholds and knowledge gaps to mangrove and saltmarsh ecosystems. Biol. Rev. 2012, 87, 346–366. [Google Scholar] [CrossRef] [PubMed]
- Gilman, E.L.; Ellison, J.; Duke, N.C.; Field, C. Threats to mangroves from climate change and adaptation options: A review. Aquat. Bot. 2008, 89, 237–250. [Google Scholar] [CrossRef]
- Young, A.; Boyle, T.; Brown, T. The population genetic consequences of habitat fragmentation for plants. Trends Ecol. Evol. 1996, 11, 413–418. [Google Scholar] [CrossRef]
- Frankham, R. Conservation genetics. Annu. Rev. Genet. 1995, 29, 305–327. [Google Scholar] [CrossRef] [PubMed]
- Hartl, D.L.; Clark, A.G. Principles of Population Genetics; Sinauer Associates: Sunderland, MA, USA, 1997. [Google Scholar]
- Frankham, R. Where are we in conservation genetics and where do we need to go? Conserv. Genet. 2010, 11, 661–663. [Google Scholar] [CrossRef]
- Triest, L. Molecular ecology and biogeography of mangrove trees towards conceptual insights on gene flow and barriers: A review. Aquat. Bot. 2008, 89, 138–154. [Google Scholar] [CrossRef]
- McKay, J.K.; Christian, C.E.; Harrison, S.; Rice, K.J. “How Local Is Local?” A review of practical and conceptual issues in the genetics of restoration. Restor. Ecol. 2005, 13, 432–440. [Google Scholar] [CrossRef]
- Grativol, C.; Lira-Medeiros, C.F.; Hemerly, A.S.; Ferreira, P.C.G. High efficiency and reliability of inter-simple sequence repeats (ISSR) markers for evaluation of genetic diversity in Brazilian cultivated Jatropha curcas L. accessions. Mol. Biol. Rep. 2011, 38, 4245–4256. [Google Scholar] [CrossRef] [PubMed]
- Kumar, A.; Mishra, P.; Singh, S.C.; Sundaresan, V. Efficiency of ISSR and RAPD markers in genetic divergence analysis and conservation management of Justicia adhatoda L., a medicinal plant. Plant Syst. Evol. 2014, 300, 1409–1420. [Google Scholar] [CrossRef]
- Huang, J.C.; Sun, M. Genetic diversity and relationships of sweetpotato and its wild relatives in Ipomoea series Batatas (Convolvulaceae) as revealed by inter-simple sequence repeat (ISSR) and restriction analysis of chloroplast DNA. Theor. Appl. Genet. 2000, 100, 1050–1060. [Google Scholar] [CrossRef]
- Bruschi, P.; Angeletti, C.; González, O.; Adele Signorini, M.; Bagnoli, F. Genetic and morphological variation of Rhizophora mangle (red mangrove) along the northern Pacific coast of Nicaragua. Nord. J. Bot. 2014, 32, 320–329. [Google Scholar] [CrossRef]
- Rosero-Galindo, C.; Gaitan-Solis, E.; Cardenas-Henao, H.; Tohme, J.; Toro-Perea, N. Polymorphic microsatellites in a mangrove species, Rhizophora mangle L. (Rhizophoraceae). Mol. Ecol. Notes 2002, 2, 281–283. [Google Scholar] [CrossRef]
- Núñez-Farfán, J.; Domínguez, C.A.; Eguiarte, L.E.; Cornejo, A.; Quijano, M.; Vargas, J.; Dirzo, R. Genetic divergence among Mexican populations of red mangrove (Rhizophora mangle). Evol. Ecol. Res. 2002, 4, 1049–1064. [Google Scholar]
- Dasgupta, N.; Nandy, P.; Sengupta, C.; Das, S. RAPD and ISSR marker mediated genetic polymorphism of two mangroves Bruguiera gymnorrhiza and Heritiera fomes from Indian Sundarbans in relation to their sustainability. Physiol. Mol. Biol. Plants 2015, 21, 375–384. [Google Scholar] [CrossRef] [PubMed]
- Ge, X.-J.; YU, Y.; Yuan, Y.-M.; Huang, H.-W.; Yan, C. Genetic diversity and geographic differentiation in endangered ammopiptanthus (Leguminosae) populations in desert regions of Northwest China as revealed by ISSR analysis. Ann. Bot. 2005, 95, 843–851. [Google Scholar] [CrossRef] [PubMed]
- Jian, S.-G.; Tang, T.; Zhong, Y.; Shi, S.-H. Conservation genetics of Heritiera littoralis (Sterculiaceae), a threatened mangrove in China, based on AFLP and ISSR markers. Biochem. Syst. Ecol. 2010, 38, 924–930. [Google Scholar] [CrossRef]
- Jian, S.; Tang, T.; Zhong, Y.; Shi, S. Variation in inter-simple sequence repeat (ISSR) in mangrove and non-mangrove populations of Heritiera littoralis (Sterculiaceae) from China and Australia. Aquat. Bot. 2004, 79, 75–86. [Google Scholar] [CrossRef]
- Jena, S.N.; Verma, S.; Nair, K.N.; Srivastava, A.K.; Misra, S.; Rana, T.S. Genetic diversity and population structure of the mangrove lime (Merope angulata) in India revealed by AFLP and ISSR markers. Aquat. Bot. 2015, 120, 260–267. [Google Scholar] [CrossRef]
- Ge, X.-J.; Sun, M. Population genetic structure of Ceriops tagal (Rhizophoraceae) in Thailand and China. Wetl. Ecol. Manag. 2001, 9, 213–219. [Google Scholar] [CrossRef]
- Ge, X.J.; Sun, M. Reproductive biology and genetic diversity of a cryptoviviparous mangrove Aegiceras corniculatum (Myrsinaceae) using allozyme and intersimple sequence repeat (ISSR) analysis. Mol. Ecol. 1999, 8, 2061–2069. [Google Scholar] [CrossRef] [PubMed]
- Schaeffer-Novelli, Y.; Cintrón-Molero, G.; Reis-Neto, A.S.; Abuchahla, G.M.O.; Neta, L.C.P.; Lira-Medeiros, C.F. The mangroves of Araçá Bay through time: An interdisciplinary approach for conservation of spatial diversity at large scale. Ocean Coast. Manag. 2018. [Google Scholar] [CrossRef]
- Lira-Medeiros, C.F.; Cardoso, M.; Fernandes, R.; Ferreira, P. Analysis of genetic diversity of two mangrove species with morphological alterations in a natural environment. Diversity 2015, 7, 105–117. [Google Scholar] [CrossRef]
- Holsinger, K.E.; Lewis, P.O.; Dey, D. A Bayesian approach to inferring population structure from domimant markers. EEB Artic. 2002, 1, 1–14. [Google Scholar] [CrossRef]
- Thioulouse, J.; Chessel, D.; Dole’dec, S.; Olivier, J.-M. ADE-4: A multivariate analysis and graphical display software. Stat. Comput. 1997, 7, 75–83. [Google Scholar] [CrossRef]
- Team, R.D.C. R: A Language and Environment for Statistical Computing; R Foundation for Statistical Computing: Vienna, Austria, 2008; ISBN 3-900051-07-0. [Google Scholar]
- Campos, L.C.T. Análise Das Populações de Mangue Branco (Laguncularia racemosa) e Mangue Preto (Avicennia schaueriana) do Estado do RJ Com Marcadores ISSR; Monograph; Faculdades Integradas Maria Thereza: Niterói, Brazil, 2011. [Google Scholar]
- Pil, M.W.; Boeger, M.R.T.; Muschner, V.C.; Pie, M.R.; Ostrensky, A.; Boeger, W.A. Postglacial north-south expansion of populations of Rhizophora mangle (Rhizophoraceae) along the Brazilian coast revealed by microsatellite analysis. Am. J. Bot. 2011, 98, 1031–1039. [Google Scholar] [CrossRef] [PubMed]
- Su, G.-H.; Huang, Y.-L.; Tan, F.-X.; Ni, X.-W.; Tang, T.; Shi, S.-H. Genetic variation in Lumnitzera racemosa, a mangrove species from the Indo-West Pacific. Aquat. Bot. 2006, 84, 341–346. [Google Scholar] [CrossRef]
- Reynolds, L.K.; McGlathery, K.J.; Waycott, M. Genetic diversity enhances restoration success by augmenting ecosystem services. PLoS ONE 2012, 7, e38397. [Google Scholar] [CrossRef] [PubMed]
- Williams, S.L. Reduced genetic diversity in eelgrass transplantations affects both population growth and individual fitness. Ecol. Appl. 2001, 11, 1472–1488. [Google Scholar] [CrossRef]
- Ellstrand, N.C.; Elam, D.R. Population genetic consequences of small population size: Implications for plant conservation. Annu. Rev. Ecol. Syst. 1993, 24, 217–242. [Google Scholar] [CrossRef]
- Mori, G.M.; Zucchi, M.I.; Sampaio, I.; Souza, A.P. Species distribution and introgressive hybridization of two Avicennia species from the Western Hemisphere unveiled by phylogeographic patterns. BMC Evol. Biol. 2015, 15, 61. [Google Scholar] [CrossRef] [PubMed]
- Ceron-Souza, I.; Toro-Perea, N.; Cardenas-Henao, H. Population genetic structure of neotropical mangrove species on the Colombian Pacific coast: Avicennia germinans (Avicenniaceae). Biotropica 2005, 37, 258–265. [Google Scholar] [CrossRef]
- Albrecht, M.; Kneeland, K.M.; Lindroth, E.; Foster, J.E. Genetic diversity and relatedness of the mangrove Rhizophora mangle L. (Rhizophoraceae) using amplified fragment polymorphism (AFLP) among locations in Florida, USA and the Caribbean. J. Coast. Conserv. 2013, 17, 483–491. [Google Scholar] [CrossRef]
- Dodd, R.S.; Afzal-Rafii, Z.; Kashani, N.; Budrick, J. Land barriers and open oceans: Effects on gene diversity and population structure in Avicennia germinans L. (Avicenniaceae). Mol. Ecol. 2002, 11, 1327–1338. [Google Scholar] [CrossRef] [PubMed]
- Ellison, A.M. Mangrove Restoration: Do We Know Enough? Restor. Ecol. 2000, 8, 219–229. [Google Scholar] [CrossRef]
- Lewis, R.R. Ecological engineering for successful management and restoration of mangrove forests. Ecol. Eng. 2005, 24, 403–418. [Google Scholar] [CrossRef]
- Hufford, K.M.; Mazer, S.J. Plant ecotypes: Genetic differentiation in the age of ecological restoration. Trends Ecol. Evol. 2003, 18, 147–155. [Google Scholar] [CrossRef]
| Primer | Primer Sequence | TA for L. racemosa | TA for A. schaueriana |
|---|---|---|---|
| 808 | 5′ [AG]8C 3′ | 46 °C | 46 °C |
| 809 | 5′ [AG]8G 3′ | 46 °C | 50 °C |
| 810 | 5′ [GA]8T 3′ | - | 52 °C |
| 811 | 5′ [GA]8C 3′ | 48 °C | 50 °C |
| 834 | 5′ [AG]8YT 3′ | 46 °C | 52 °C |
| 835 | 5′ [AG]8YC 3′ | - | 48 °C |
| 840 | 5′ [GA]8YT 3′ | 48 °C | 52 °C |
| 841 | 5′[GA]8YC 3′ | 46 °C | - |
| 842 | 5′ [GA]8YG 3′ | 48 °C | 54 °C |
| Populations | HS (SD) |
|---|---|
| RES | 0.108 (0.020) |
| REM | 0.239 (0.017) |
| GUA | 0.185 (0.021) |
| ARA | 0.157 (0.018) |
| Mean | 0.173 |
| Populations | HS (SD) |
|---|---|
| RES | 0.242 (0.015) |
| REM | 0.261 (0.012) |
| GUA | 0.238 (0.012) |
| ARA | 0.243 (0.014) |
| Mean | 0.246 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Granado, R.; Pinto Neta, L.C.; Nunes-Freitas, A.F.; Voloch, C.M.; Lira, C.F. Assessing Genetic Diversity after Mangrove Restoration in Brazil: Why Is It So Important? Diversity 2018, 10, 27. https://doi.org/10.3390/d10020027
Granado R, Pinto Neta LC, Nunes-Freitas AF, Voloch CM, Lira CF. Assessing Genetic Diversity after Mangrove Restoration in Brazil: Why Is It So Important? Diversity. 2018; 10(2):27. https://doi.org/10.3390/d10020027
Chicago/Turabian StyleGranado, Renan, Luiza C. Pinto Neta, André F. Nunes-Freitas, Carolina M. Voloch, and Catarina F. Lira. 2018. "Assessing Genetic Diversity after Mangrove Restoration in Brazil: Why Is It So Important?" Diversity 10, no. 2: 27. https://doi.org/10.3390/d10020027





