A Centrifugation-based Method for Preparation of Gold Nanoparticles and its Application in Biodetection
Abstract
:1. Introduction
2. Results and Discussion
Conclusion
3. Experimental Section
3.1 Reagents and instrumentation
3.2 Synthesis of 13-nm AuNPs
3.3 Evaluation of salt-resistance of AuNPs
3.4 Synthesis of DNA-AuNPs conjugates
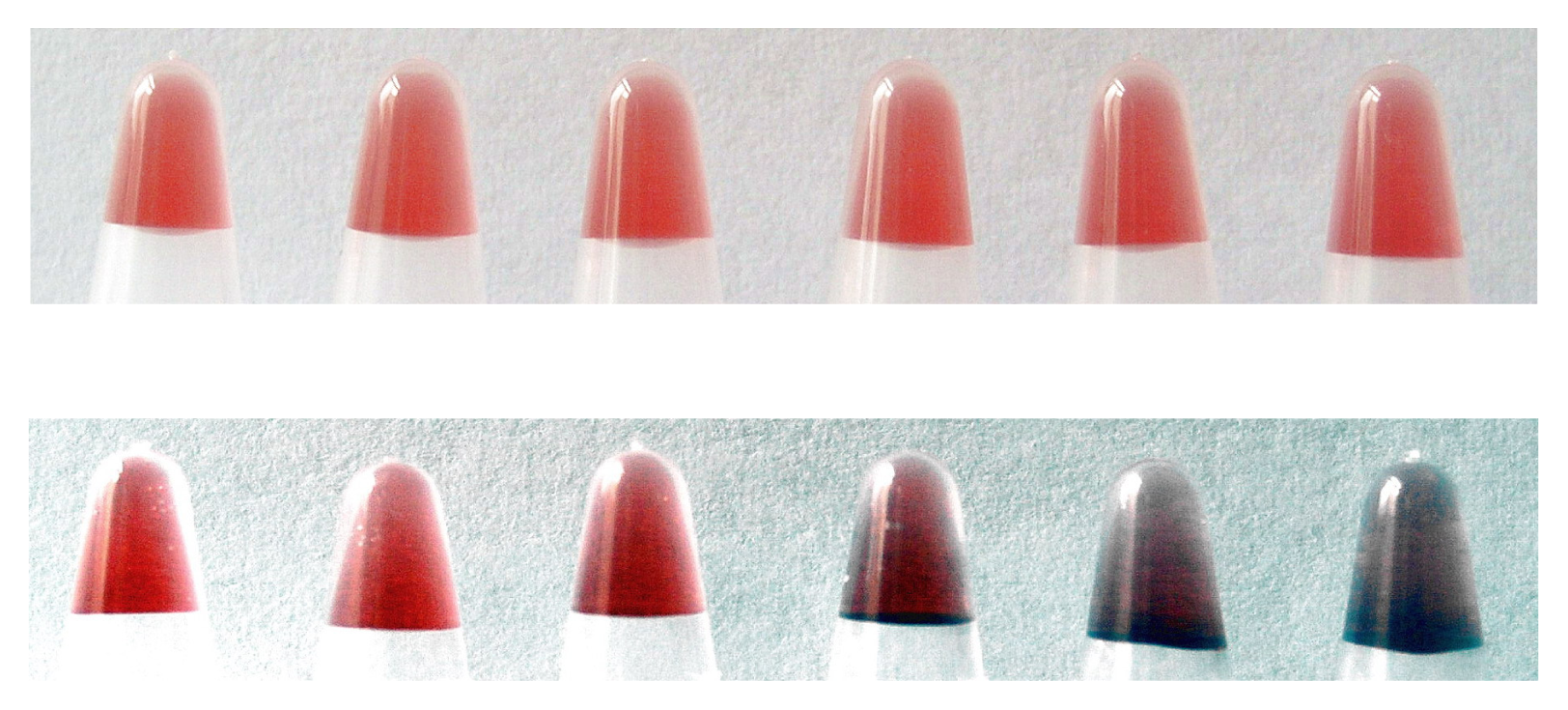
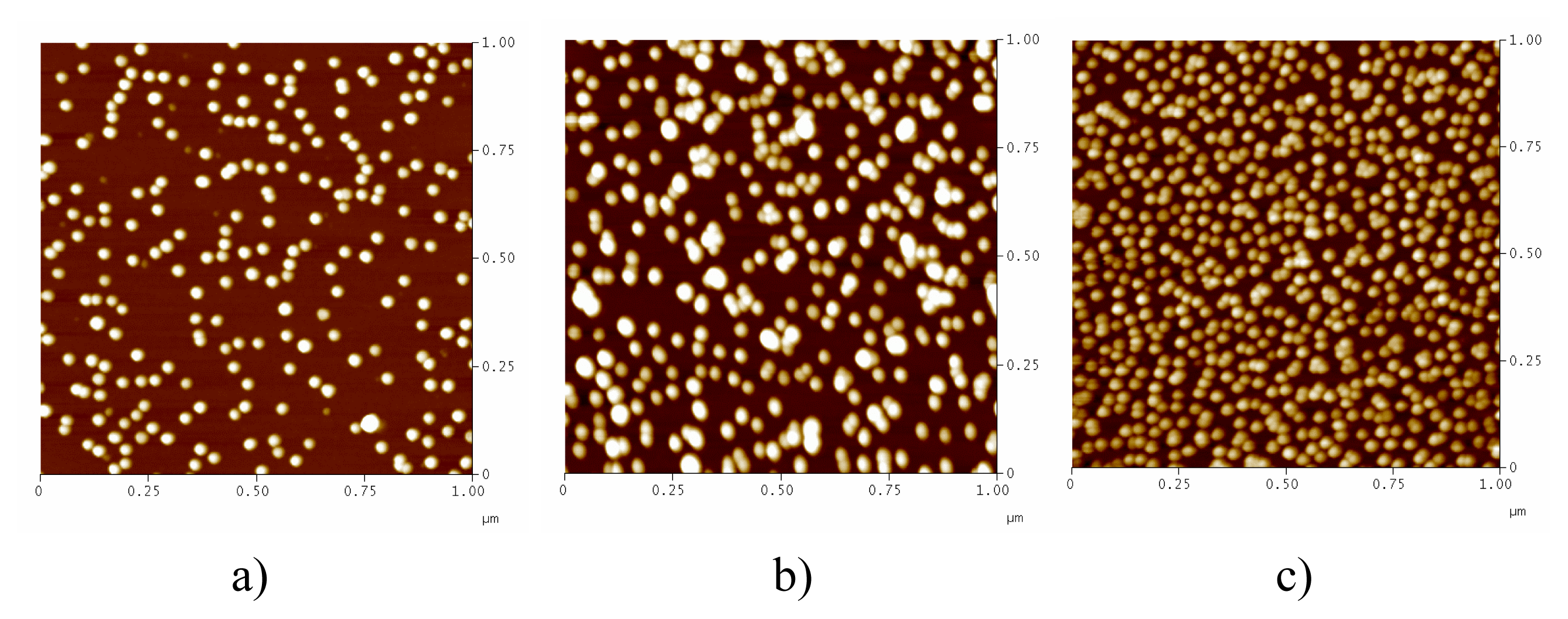
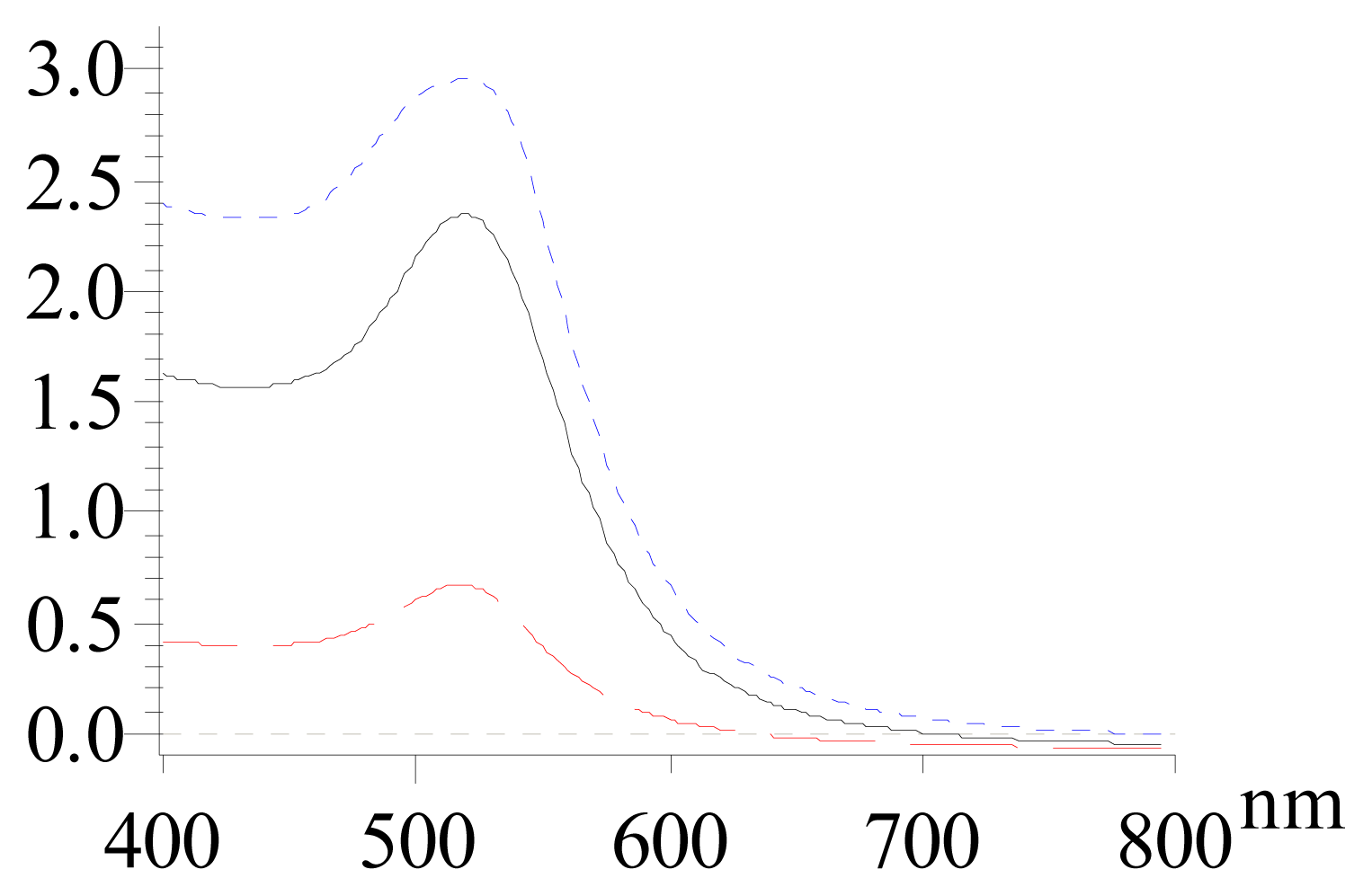

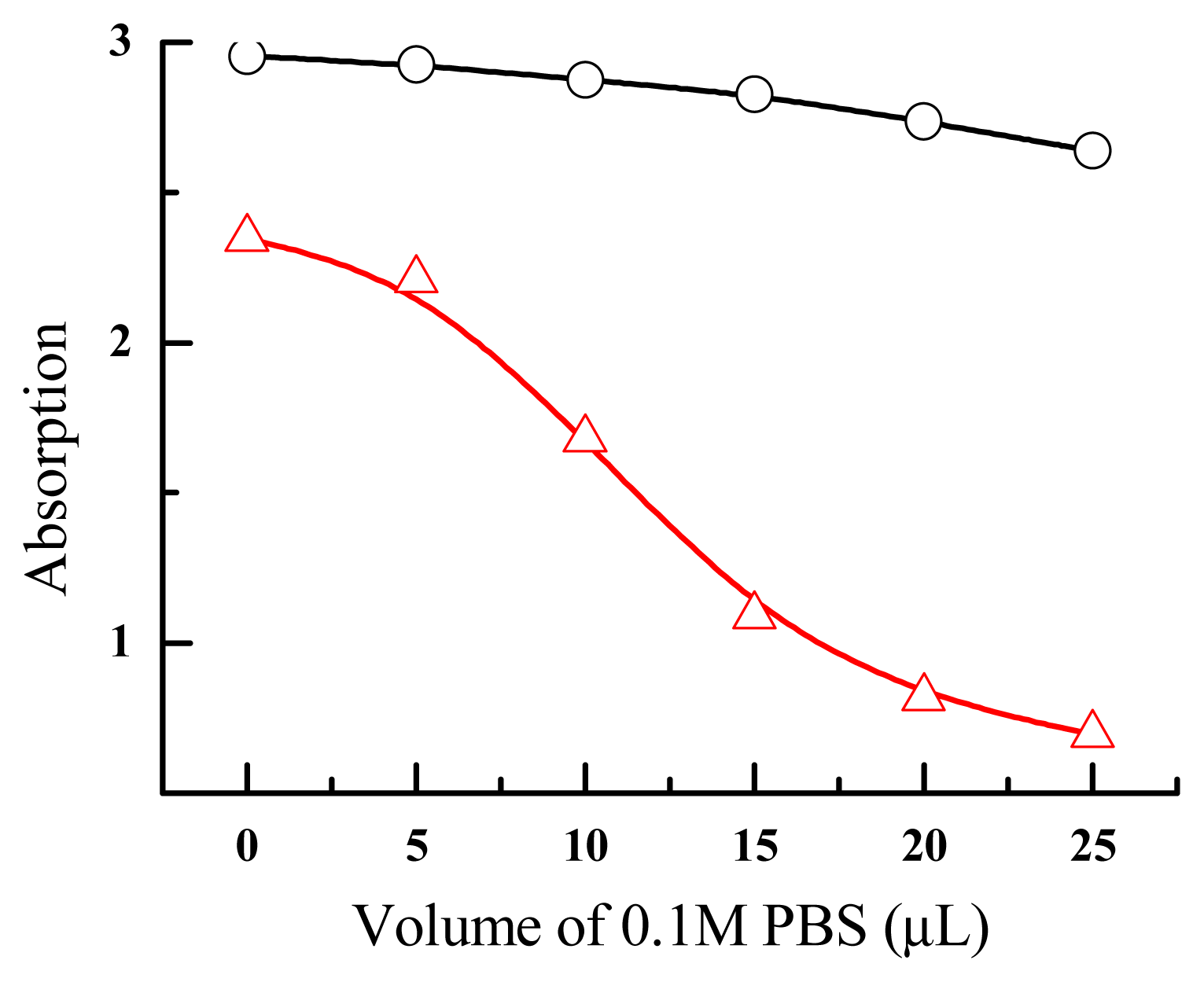
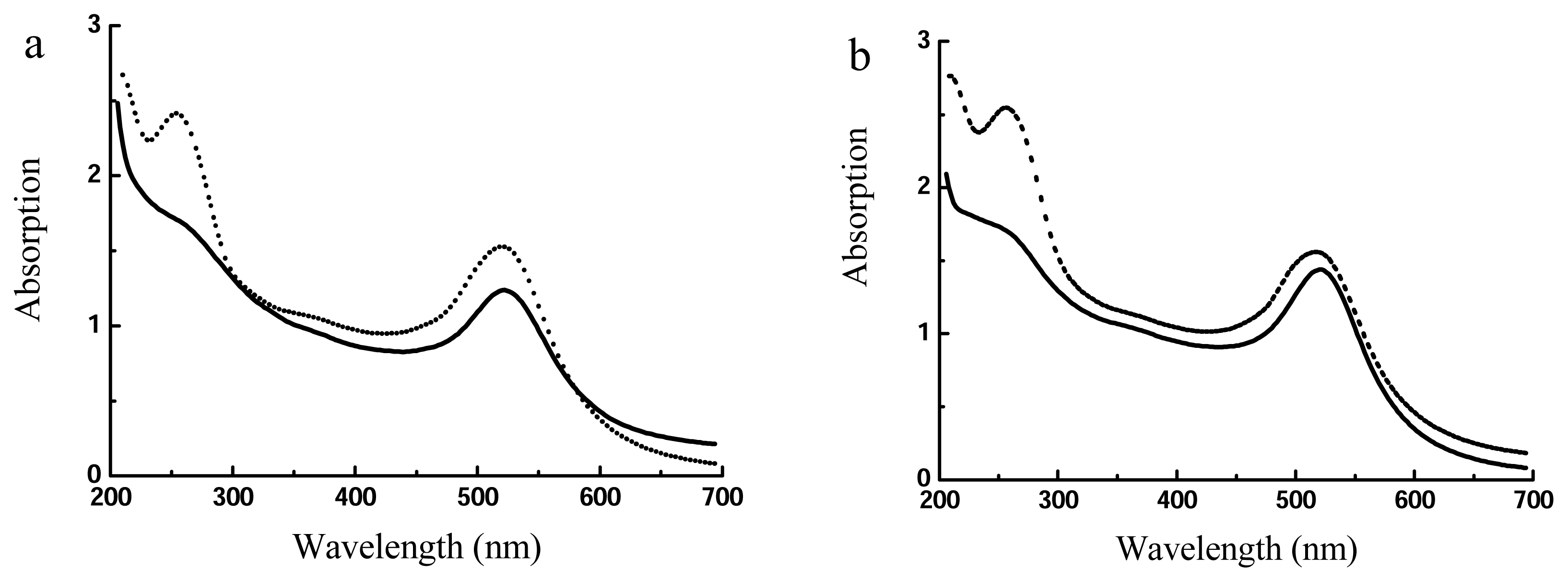
| Size (nm) | Absorption (520 nm) | Salt-resistance of AuNPs | Salt-resistance of DNA-AuNPs cojugates | Storage stability of DNA-AuNPs cojugates | |
|---|---|---|---|---|---|
| Method I (direct reduction) | 14 ± 5 | 2.346 | 9 mM | 1.5 M | < 7 days |
| Method II (centrifuging) | 12.5 ± 2.3 | 2.954 | 44 mM | 2.6 M | > 30 days |
Acknowledgements
References and Notes
- Hacker, G.W.; Gu, J. Gold and Silver Staining: Techniques in Molecular Morphology; CRC Press: Boca Raton, 2002. [Google Scholar]
- Mirkin, C.A.; Letsinger, R.L.; Mucic, R.C.; Storhoff, J.J. A DNA-based method for rationally assembling nanoparticles into macroscopic materials. Nature 1996, 382, 607–609. [Google Scholar]
- Alivisatos, A.P.; Johnsson, K.P.; Peng, X.G.; Wilson, T.E.; Loweth, C.J.; Bruchez, M.P.; Schultz, P.G. Organization of 'nanocrystal molecules' using DNA. Nature 1996, 382, 609–611. [Google Scholar]
- Taton, T.A.; Mirkin, C.A.; Letsinger, R.L. Scanometric DNA Array Detection with Nanoparticle Probes. Science 2000, 289, 1757–1760. [Google Scholar]
- Cao, Y.W.C.; Jin, R.C.; Mirkin, C.A. Nanoparticles with Raman Spectroscopic Fingerprints for DNA and RNA Detection. Science 2002, 297, 1536–1540. [Google Scholar]
- Rosi, N.L.; Mirkin, C.A. Nanostructures in biodiagnostics. Chem. Rev 2005, 105, 1547–1562. [Google Scholar]
- Nam, J.M.; Thaxon, C.S.; Mirkin, C.A. Nanoparticle-based bio-bar codes for the ultrasensitive detection of proteins. Science 2003, 301, 1884–1886. [Google Scholar]
- Oh, B.K.; Nam, J.M.; Lee, S.W.; Mirkin, C.A. A fluorophore-based bio-barcode amplification assay for proteins. Small 2006, 2, 103–108. [Google Scholar]
- Li, H.; Rothberg, L. Colorimetric detection of DNA sequences based on electrostatic interactions with unmodified gold nanoparticles. Proc. Natl. Acad. Sci., U.S.A 2004, 101, 14036–14039. [Google Scholar]
- Li, H.; Rothberg, L.J. Label-free colorimetric detection of specific sequences in genomic DNA amplified by the polymerase chain reaction. J. Am. Chem. Soc 2004, 126, 10958–10961. [Google Scholar]
- Li, H.; Huang, J.; Lv, J.; An, H.; Zhang, X.; Zhang, Z.; Fan, C.; Hu, J. Nanoparticle PCR: Nanogold-assisted PCR with enhanced specificity. Angew. Chem. Int. Ed 2005, 44, 5100–5103. [Google Scholar]
- Elghanian, R.; Storhoff, J.J.; Mucic, R.C.; Letsinger, R.L.; Mirkin, C.A. DNA-Directed Synthesis of Binary Nanoparticle Network Materials. Science 1997, 277, 1078–1081. [Google Scholar]
- Grabar, K.C.; Freeman, R.G.; Hommer, M.B.; Natan, M.J. Preparation and Characterization of Au Colloid Monolayers. Anal. Chem 1995, 67, 735–743. [Google Scholar]
- Rosi, N.L.; Giljohann, D.A.; Thaxton, C.S.; Lytton-Jean, A.K.R.; Han, M.S.; Mirkin, C.A. Oligonucleotide-Modified Gold Nanoparticles for Intracellular Gene Regulation. Science 2006, 312, 1027–1030. [Google Scholar]
© 2007 by MDPI Reproduction is permitted for noncommercial purposes.
Share and Cite
Liang, Z.; Zhang, J.; Wang, L.; Song, S.; Fan, C.; Li, G. A Centrifugation-based Method for Preparation of Gold Nanoparticles and its Application in Biodetection. Int. J. Mol. Sci. 2007, 8, 526-532. https://doi.org/10.3390/i8060526
Liang Z, Zhang J, Wang L, Song S, Fan C, Li G. A Centrifugation-based Method for Preparation of Gold Nanoparticles and its Application in Biodetection. International Journal of Molecular Sciences. 2007; 8(6):526-532. https://doi.org/10.3390/i8060526
Chicago/Turabian StyleLiang, Zhiqiang, Juan Zhang, Lihua Wang, Shiping Song, Chunhai Fan, and Genxi Li. 2007. "A Centrifugation-based Method for Preparation of Gold Nanoparticles and its Application in Biodetection" International Journal of Molecular Sciences 8, no. 6: 526-532. https://doi.org/10.3390/i8060526




