Meiotic and Mitotic Phenotypes Conferred by the blm1-1 Mutation in Saccharomyces cerevisiae and MSH4 Suppression of the Bleomycin Hypersusceptibility
Abstract
:Introduction
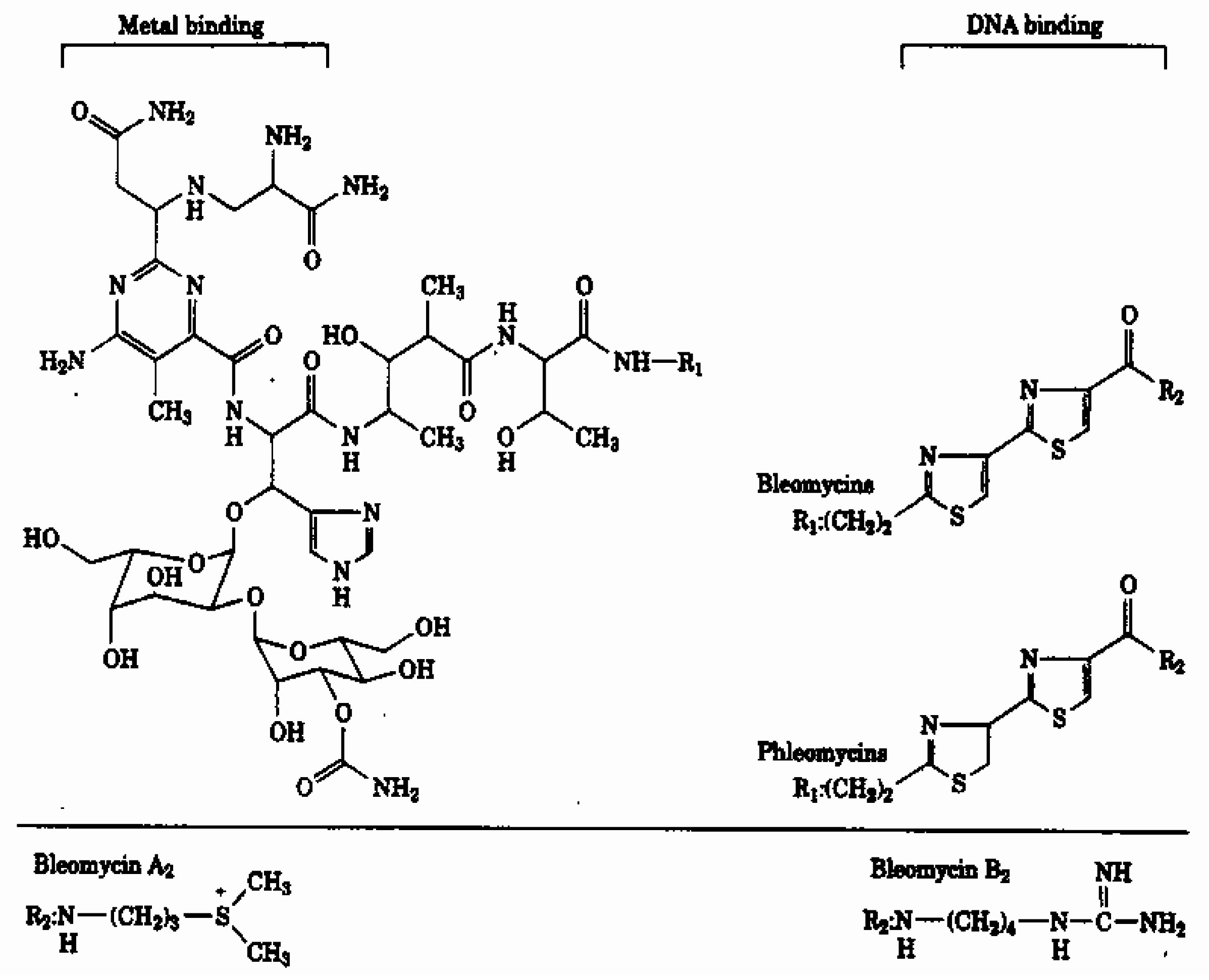
Materials and Methods
Strains and Plasmids
| Strain or Plasmid | Genotype or Selectable Markers | Source |
| Saccharomyces cerevisiae | ||
| CM1457-79A | MATα ade2-1 and/or ade2-40 trp1-1 and/or trp5b | This laboratory |
| CM1457-79B | MATa ade2-1 and/or ade2-40 ilv1-92 trp1-1 and/or trp5b | This laboratory |
| CM1457-79C | MATa ade2-1 and/or ade2-40 blm1-1 trp1-1 and/or trp5b leu2-3 and/or leu2-112 ura3-1 | This laboratory |
| CM1457-79D | MATα ade2-1 and/or ade2-40 blm1-1 ilv1-92 leu2-3 and/or leu2-112 trp1-1 and/or trp5b ura3-1 | This laboratory |
| PS593/6H | MATα his3Δ200 leu2-3 or leu2-112 msh4::LEU2 trp1-289 ura3-52 | Dr. Nancy Hollingsworth; Dr. Shirleen Roeder [15] |
| Plasmids | ||
| YCp50 | amp tet ARS1 CEN4 URA3 | Johnston and Davis, 1984 [16] |
| pPM118 | tet ARS1 CEN4 URA3 | This laboratory |
| pPM118-4 | tet ARS1 CEN4 URA3 | This laboratory |
Media and Growth Conditions
Sporulation and Dissection of Tetrads
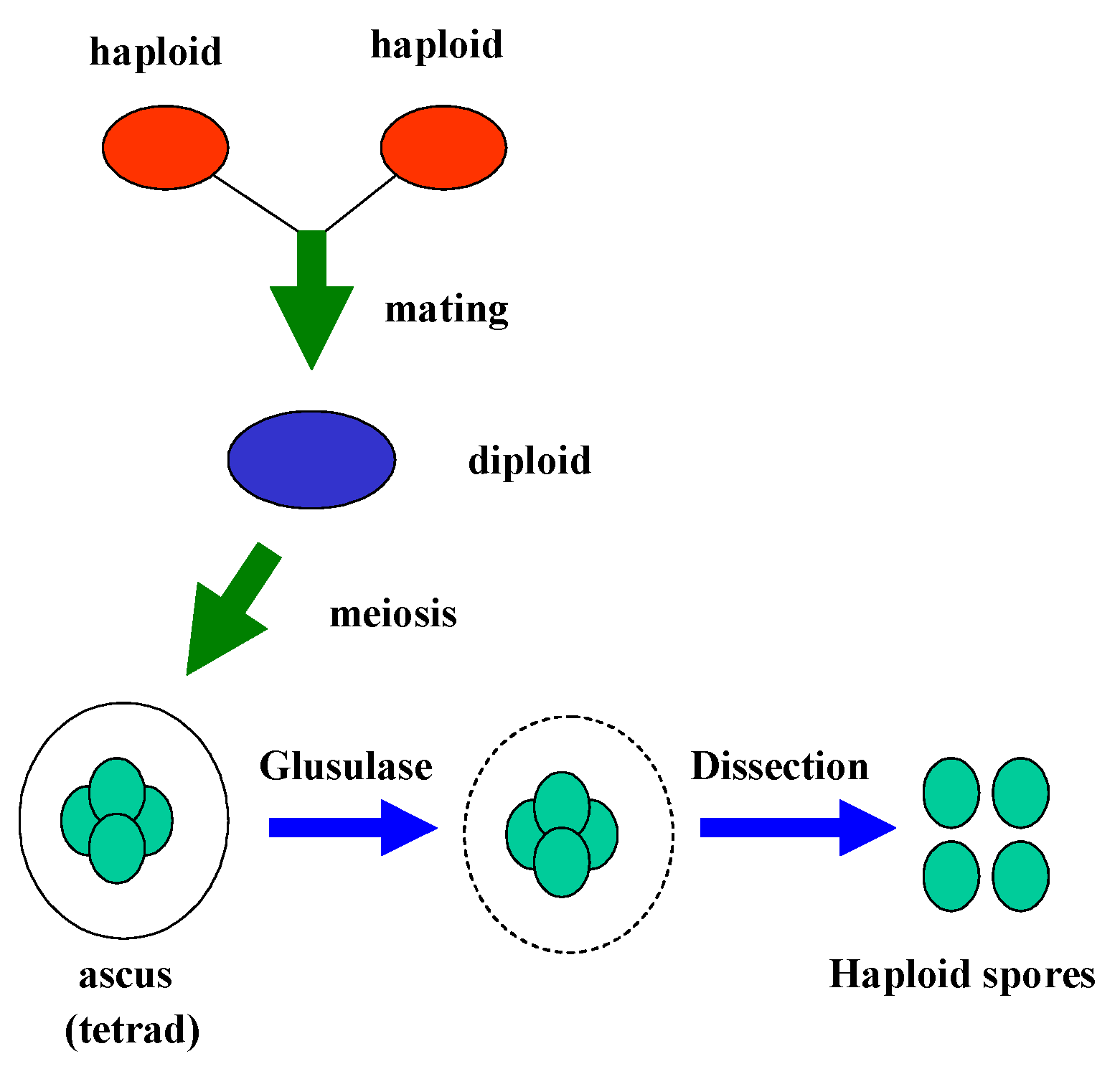
Zeocin Treatments
E. coli and Yeast Transformations
Results
Reduced Viability of Spores from Homozygous Mutant Strains
| Diploid Type | Total number of tetrads dissected | No viable spores | 1 viable spore | 2 viable spores | 3 viable spores | 4 viable spores |
| Homozygous normal | 16 | 0 | 1 | 0 | 6 | 9 |
| Heterozygous | 14 | 0 | 0 | 0 | 5 | 9 |
| Homozygous mutant | 31 | 0 | 7 | 20 | 4 | 0 |
| Diploid Type | Total number of tetrads dissected | Sporulation (%) | 1 viable spore (%) | 2 viable spores (%) | 3 viable spores (%) | 4 viable spores (%) |
| Homozygous normal | 16 | 70±5.81 | 6 | 0 | 38 | 56 |
| Heterozygous | 14 | 66.3±4.51 | 0 | 0 | 36 | 64 |
| Homozygous mutant | 31 | 52.1±4.11 | 23 | 65 | 13 | 0 |
Mitotic Phenotype of blm1-1 Mutant Strains: Reduced Growth Rate and Elevated Frequencies of Irregularly-shaped Cells
| Strains | Final Titers (cells/ml) | Number and Percent of Irregularly- shaped Cells 2 | Normal-sized Single Cells and Budded Cells 3 | Number and Percent of Enlarged Cells 4 | Total Number of Cells |
| Means of experiments one and two | |||||
| BLM1 (CM1457-79A) | 5.00 x 108 | 0% | (462) 84.6% | (84) 15.4% | 546 (100%) |
| Experiments three and four: means of both strains | |||||
| blm1-1 (CM1457-79C and CM1457-79D) | 3.60 x 108 | (28.5) 5.23% | (350) 64.1% | (167) 30.6% | 546 (100%) |
| Experiments five and six: means of both strains | |||||
| blm1-1 (CM1457-79C and CM1457-79D) | 3.78 x 108 | (29) 5.31% | (347) 63.6% | (170) 31.1% | 546 (100%) |
 .
. .
. .
.Isolation of Plasmids, Restriction Analyses, and DNA Sequence Analyses

Relief of Drug Hypersensitivity
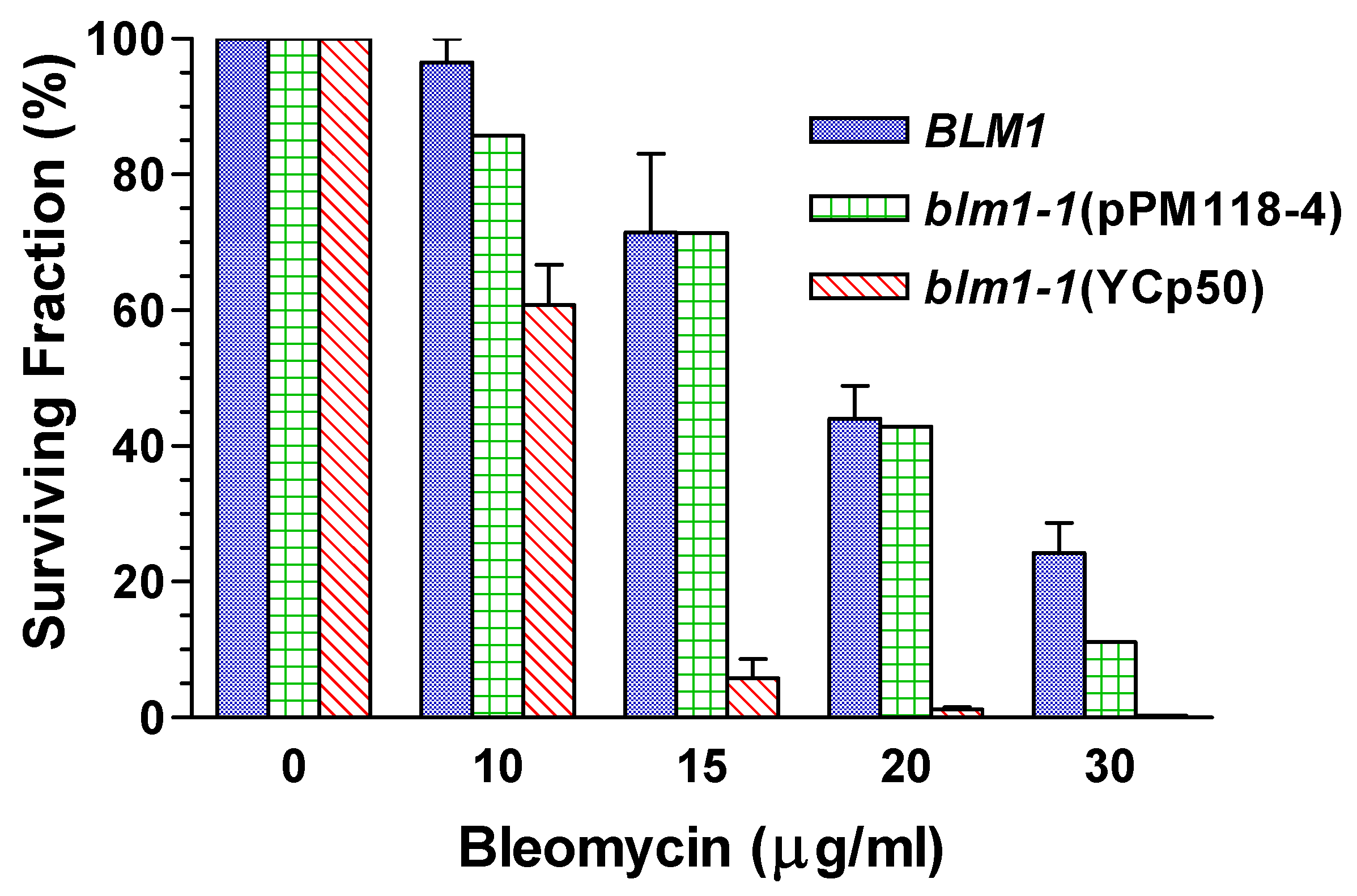
Genetic Mapping Showed that MSH4 is not the BLM1 Structural Gene
| Classes of tetrads | Number of tetrads | Number of spores | blm1-1, leu2 | BLM1, LEU2 | BLM1, leu2 | blm1-1, LEU2 |
| Parental Ditype (PD) | 5 | 20 | 10 | 10 | 0 | 0 |
| Non-Parental Ditype (NPD) | 6 | 24 | 0 | 0 | 12 | 12 |
| Tetratype (TT) | 3 | 12 | 3 | 3 | 3 | 3 |
| Total spores | 14 | 56 | 13 | 13 | 15 | 15 |
Discussion

Acknowledgements
References
- Moore, C. W. Internucleosomal cleavage and chromosomal degradation by bleomycin and phleomycin in yeast. Cancer Res. 1988, 48, 6837–6843. [Google Scholar] [PubMed]
- Beaudouin, R.; Lim, S. T.; Steide, J. A.; Powell, M.; McKoy, J.; Pramanik, A. J.; Johnson, E.; Moore, C. W.; Lipke, P. N. Bleomycin affects cell wall anchorage of mannoproteins in Saccharomyces cerevisiae. Antimicrob. Agents Chemother. 1993, 37, 1264–1269. [Google Scholar] [CrossRef] [PubMed]
- Burger, R. M. Cleavage of nucleic acids by bleomycin. Chemical Reviews 1998, 98, 1153–1170. [Google Scholar] [CrossRef] [PubMed]
- Petering, D. H.; Byrnes, R. W.; Antholine, W. E. The role of redox-active metals in the mechanism of action of bleomycin. Chem. Biol. Interact. 1990, 73, 133–182. [Google Scholar] [CrossRef] [PubMed]
- Steighner, R. J.; Povirk, L. F. Bleomycin-induced DNA lesions at mutational hot spots: implications for the mechanism of double-strand cleavage. Proc. Natl. Acad. Sci. U. S. A. 1990, 87, 8350–8354. [Google Scholar] [CrossRef] [PubMed]
- Moore, C. W. Responses of radiation-sensitive mutants of Saccharomyces cerevisiae to lethal effects of bleomycin. Mutat. Res. 1978, 51, 165–180. [Google Scholar] [CrossRef] [PubMed]
- Moore, C. W. Control of in vivo (cellular) phleomycin sensitivity by nuclear genotype, growth phase, and metal ions. Cancer Res. 1982, 42, 929–933. [Google Scholar] [PubMed]
- Moore, C. W. Ligase-deficient yeast cells exhibit defective DNA rejoining and enhanced gamma ray sensitivity. J. Bacteriol. 1982, 150, 1227–1233. [Google Scholar] [PubMed]
- Moore, C. W. Degradation of DNA and structure-activity relationship between bleomycins A2 and B2 in the absence of DNA repair. Biochemistry 1990, 29, 1342–1347. [Google Scholar] [CrossRef] [PubMed]
- Moore, C. W. Bleomycin. In Encyclopedia of Molecular Biology; Creighton, T., Ed.; John Wiley and Sons, Inc.: New York, 1999; pp. 292–297. [Google Scholar]
- Moore, C. W. Isolation and partial characterization of mutants of Saccharomyces cerevisiae altered in sensitivities to lethal effects of bleomycins. J. Antibiot. (Tokyo) 1980, 33, 1369–1375. [Google Scholar] [CrossRef] [Green Version]
- Moore, C. W. Further characterizations of bleomycin-sensitive (blm) mutants of Saccharomyces cerevisiae with implications for a radiomimetic model. J. Bacteriol. 1991, 173, 3605–3608. [Google Scholar] [PubMed]
- Ross-Macdonald, P.; Roeder, G. S. Mutation of a meiosis-specific MutS homolog decreases crossing over but not mismatch correction. Cell 1994, 79, 1069–1080. [Google Scholar]
- Eisen, J. A. A phylogenomic study of the MutS family of proteins. Nucleic Acids Res. 1998, 26, 4291–4300. [Google Scholar] [CrossRef] [PubMed]
- Novak, J. E.; Ross-Macdonald, P. B.; Roeder, G. S. The budding yeast Msh4 protein functions in chromosome synapsis and the regulation of crossover distribution. Genetics 2001, 158, 1013–1025. [Google Scholar] [PubMed]
- Johnston, M.; Davis, R. W. Sequences that regulate the divergent GAL1-GAL10 promoter in Saccharomyces cerevisiae. Mol. Cell Biol. 1984, 4, 1440–1448. [Google Scholar] [PubMed]
- Sambrook, J.; Fritsch, E. F.; Maniatis, T. Oligotide-Mediatede Mutagenesis by Selection Against Template Strands That Contain Uracil (Kunkel Method). In Molecular Cloning: a Laboratory Manual; Ford, N., Nolan, C., Ferguson, M., Eds.; Cold Spring Harbor Laboratory Press: Cold Spring Harbor, NY, 1989; pp. 15.74–15.79. [Google Scholar]
- Ito, H.; Fukuda, Y.; Murata, K.; Kimura, A. Transformation of intact yeast cells treated with alkali cations. J. Bacteriol. 1983, 153, 163–168. [Google Scholar] [PubMed]
- Rose, A. H. Isolations of Genes by Complementation in Yeast. In Methods in Enzymology; Academic Press: New York, 2002; pp. 481–504. [Google Scholar]
- Rose, M. D.; Broach, J. R. Cloning Genes by Complementation in Yeast. In Methods in Enzymology; Academic Press: New York, 1991; pp. 195–230. [Google Scholar]
- Moore, C. W. Further characterizations of bleomycin-sensitive (blm) mutants of Saccharomyces cerevisiae with implications for a radiomimetic model. J. Bacteriol. 1991, 173, 3605–3608. [Google Scholar] [PubMed]
- Pramanik, A.; Martinez, M.; McKoy, J. F.; Ospina, R.; Daly, J.; Moore, C. W. A mutant approach and molecular strategy to study fungal cell walls. Cell. Mol. Biol. (Noisy. -le-grand) 1995, 41 Suppl 1, S73–S81. [Google Scholar]
- Altschul, S. F.; Gish, W.; Miller, W.; Myers, E. W.; Lipman, D. J. Basic local alignment search tool. J. Mol. Biol. 1990, 215, 403–410. [Google Scholar] [CrossRef] [PubMed]
- Kaiser, C.; Michaelis, S.; Mitchell, A. Meiotic Mapping. In Methods in Yeast Genetics: a Cold Spring Harbor Laboratory Course Manual; Cold Spring Harbor Laboratory Press, 1994; pp. 25–35. [Google Scholar]
- Roeder, G. S. Meiotic chromosomes: it takes two to tango. Genes Dev. 1997, 11, 2600–2621. [Google Scholar]
© 2003 by MDPI (http://www.mdpi.org).
Share and Cite
Anyatonwu, G.; Garcia, E.; Pramanik, A.; Powell, M.; Moore, C.W. Meiotic and Mitotic Phenotypes Conferred by the blm1-1 Mutation in Saccharomyces cerevisiae and MSH4 Suppression of the Bleomycin Hypersusceptibility. Int. J. Mol. Sci. 2003, 4, 1-12. https://doi.org/10.3390/i4010001
Anyatonwu G, Garcia E, Pramanik A, Powell M, Moore CW. Meiotic and Mitotic Phenotypes Conferred by the blm1-1 Mutation in Saccharomyces cerevisiae and MSH4 Suppression of the Bleomycin Hypersusceptibility. International Journal of Molecular Sciences. 2003; 4(1):1-12. https://doi.org/10.3390/i4010001
Chicago/Turabian StyleAnyatonwu, Georgia, Ediberto Garcia, Ajay Pramanik, Marie Powell, and Carol Wood Moore. 2003. "Meiotic and Mitotic Phenotypes Conferred by the blm1-1 Mutation in Saccharomyces cerevisiae and MSH4 Suppression of the Bleomycin Hypersusceptibility" International Journal of Molecular Sciences 4, no. 1: 1-12. https://doi.org/10.3390/i4010001




