DDX5 RNA Helicases: Emerging Roles in Viral Infection
Abstract
:1. Introduction
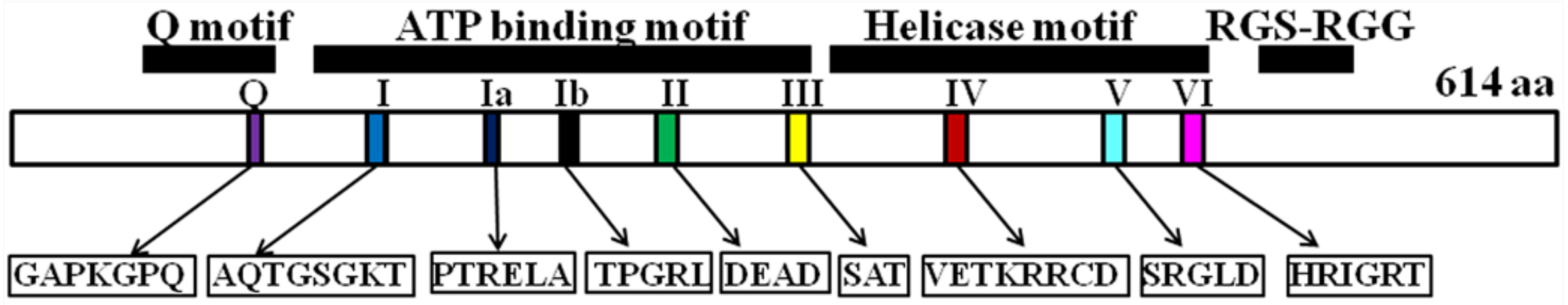
2. Modular Domain Structure of DDX5
3. DDX5 Is a Host Factor Influencing Viral Infection
3.1. DDX5 Interacts with Severe Acute Respiratory Syndrome Coronavirus Helicase
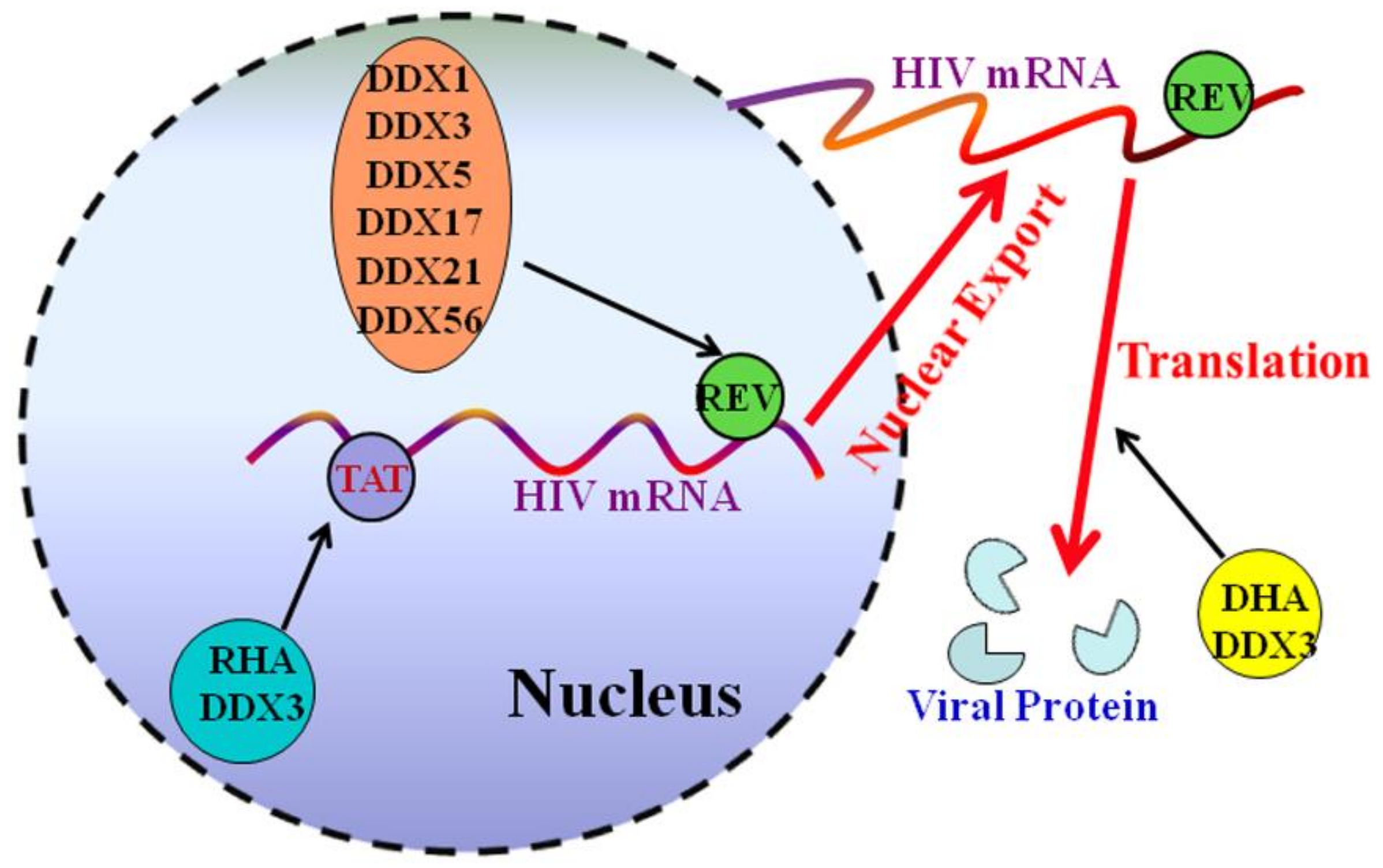
3.2. DDX5 Facilitates Human Immunodeficiency Virus 1 mRNA Export
3.3. DDX5 Is Involved in the Hepatitis C Virus (HCV) Life Cycle
3.4. DDX5 Acts as a Positive Regulator of Japanese Encephalitis Virus Replication
3.5. DDX5 Positively Regulates the Replication of Porcine Reproductive and Respiratory Syndrome Virus
3.6. DDX5 Enhances the Polymerase Activity of Influenza Virus
3.7. DDX5 Interacts with EBNA2 of Epstein-Barr Virus
3.8. DDX5 Restricts Myxoma Virus Replication and HBV Transcription
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lane, D.P.; Hoeffler, W.K. SV40 large T shares an antigenic determinant with a cellular protein of molecular weight 68,000. Nature 1980, 288, 167–170. [Google Scholar] [CrossRef] [PubMed]
- Hirling, H.; Scheffner, M.; Restle, T.; Stahl, H. RNA helicase activity associated with the human p68 protein. Nature 1989, 339, 562–564. [Google Scholar] [CrossRef] [PubMed]
- Iggo, R.; Picksley, S.; Southgate, J.; McPheat, J.; Lane, D.P. Identification of a putative RNA helicase in E. coli. Nucleic Acids Res. 1990, 18, 5413–5417. [Google Scholar] [CrossRef] [PubMed]
- Iggo, R.D.; Jamieson, D.J.; MacNeill, S.A.; Southgate, J.; McPheat, J.; Lane, D.P. 68 RNA helicase: Identification of a nucleolar form and cloning of related genes containing a conserved intron in yeasts. Mol. Cell Biol. 1991, 11, 1326–1333. [Google Scholar] [CrossRef] [PubMed]
- Watanabe, M.; Yanagisawa, J.; Kitagawa, H.; Takeyama, K.; Ogawa, S.; Arao, Y.; Suzawa, M.; Kobayashi, Y.; Yano, T.; Yoshikawa, H.; et al. A subfamily of RNA-binding DEAD-box proteins acts as an estrogen receptor α coactivator through the N-terminal activation domain (AF-1) with an RNA coactivator, SRA. EMBO J. 2001, 20, 1341–1352. [Google Scholar] [CrossRef] [PubMed]
- Liu, Z.R. p68 RNA helicase is an essential human splicing factor that acts at the U1 snRNA-5′ splice site duplex. Mol. Cell Biol. 2002, 22, 5443–5450. [Google Scholar] [CrossRef] [PubMed]
- Wilson, B.J.; Bates, G.J.; Nicol, S.M.; Gregory, D.J.; Perkins, N.D.; Fuller-Pace, F.V. The p68 and p72 DEAD-box RNA helicases interact with HDAC1 and repress transcription in a promoter-specific manner. BMC Mol. Biol. 2004, 5, 11. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Lin, C.; Yang, L.; Yang, J.J.; Huang, Y.; Liu, Z.R. ATPase/helicase activities of p68 RNA helicase are required for pre-mRNA splicing but not for assembly of the spliceosome. Mol. Cell Biol. 2005, 25, 7484–7493. [Google Scholar] [CrossRef] [PubMed]
- Jalal, C.; Uhlmann-Schiffler, H.; Stahl, H. Redundant role of DEAD-box proteins p68 (DDX5) and p72/p82 (Ddx17) in ribosome biogenesis and cell proliferation. Nucleic Acids Res. 2007, 35, 3590–3601. [Google Scholar] [CrossRef] [PubMed]
- Zonta, E.; Bittencourt, D.; Samaan, S.; Germann, S.; Dutertre, M.; Auboeuf, D. The RNA helicase DDX5/p68 is a key factor promoting c-fos expression at different levels from transcription to mRNA export. Nucleic Acids Res. 2013, 41, 554–564. [Google Scholar] [CrossRef] [PubMed]
- Dardenne, E.; Espinoza, M.P.; Fattet, L.; Germann, S.; Lambert, M.P.; Neil, H.; Zonta, E.; Mortada, H.; Gratadou, L.; Deygas, M.; et al. RNA helicases DDX5 and DDX17 dynamically orchestrate transcription, miRNA, and splicing programs in cell differentiation. Cell Rep. 2014, 7, 1900–1913. [Google Scholar] [CrossRef] [PubMed]
- Saporita, A.J.; Chang, H.C.; Winkeler, C.L.; Apicelli, A.J.; Kladney, R.D.; Wang, J.; Townsend, R.R.; Michel, L.S.; Weber, J.D. RNA helicase DDX5 is a p53-independent target of ARF that participates in ribosome biogenesis. Cancer Res. 2011, 71, 6708–6717. [Google Scholar] [CrossRef] [PubMed]
- Jost, J.P.; Schwarz, S.; Hess, D.; Angliker, H.; Fuller-Pace, F.V.; Stahl, H.; Thiry, S.; Siegmann, M. A chicken embryo protein related to the mammalian DEAD-box protein p68 is tightly associated with the highly purified protein-RNA complex of 5-MeC-DNA glycosylase. Nucleic Acids Res. 1999, 27, 3245–3252. [Google Scholar] [CrossRef] [PubMed]
- Yang, L.; Lin, C.; Liu, Z.R. Phosphorylations of DEAD-box p68 RNA helicase are associated with cancer development and cell proliferation. Mol. Cancer Res. 2005, 3, 355–363. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Gao, X.; Huang, Y.; Yang, J.; Liu, Z.R. P68 RNA helicase is a nucleocytoplasmic shuttling protein. Cell Res. 2009, 19, 1388–1400. [Google Scholar] [CrossRef] [PubMed]
- Chen, J.Y.; Chen, W.N.; Poon, K.M.; Zheng, B.J.; Lin, X.; Wang, Y.X.; Wen, Y.M. Interaction between SARS-CoV helicase and a multifunctional cellular protein (DDX5) revealed by yeast and mammalian cell two-hybrid systems. Arch. Virol. 2009, 154, 507–512. [Google Scholar] [CrossRef] [PubMed]
- Zhou, X.; Luo, J.; Mills, L.; Wu, S.; Pan, T.; Geng, G.; Zhang, J.; Luo, H.; Liu, C.; Zhang, H. DDX5 facilitates HIV-1 replication as a cellular co-factor of Rev. PLoS ONE 2013, 8, e65040. [Google Scholar] [CrossRef] [PubMed]
- Goh, P.Y.; Tan, Y.J.; Lim, S.P.; Tan, Y.H.; Lim, S.G.; Fuller-Pace, F.; Hong, W. Cellular RNA helicase p68 relocalization and interaction with the hepatitis C virus (HCV) NS5B protein and the potential role of p68 in HCV RNA replication. J. Virol. 2004, 78, 5288–5398. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Ge, L.L.; Li, P.P.; Wang, Y.; Sun, M.X.; Huang, L.; Ishag, H.; Di, D.D.; Shen, Z.Q.; Fan, W.X.; et al. The DEAD-box RNA helicase DDX5 acts as a positive regulator of Japanese encephalitis virus replication by binding to viral 3′ UTR. Antivir. Res. 2013, 100, 487–499. [Google Scholar] [CrossRef] [PubMed]
- Zhao, S.; Ge, X.; Wang, X.; Liu, A.; Guo, X.; Zhou, L.; Yu, K.; Yang, H. The DEAD-box RNA helicase 5 positively regulates the replication of porcine reproductive and respiratory syndrome virus by interacting with viral Nsp9 in vitro. Virus Res. 2015, 195, 217–224. [Google Scholar] [CrossRef] [PubMed]
- Bortz, E.; Westera, L.; Maamary, J.; Steel, J.; Albrecht, R.A.; Manicassamy, B.; Chase, G.; Martínez-Sobrido, L.; Schwemmle, M.; García-Sastre, A. Host- and strain-specific regulation of influenza virus polymerase activity by interacting cellular proteins. MBio 2011, 2, e00151-11. [Google Scholar] [CrossRef] [PubMed]
- Rahman, M.M.; Bagdassarian, E.; Ali, M.A.M.; McFadden, G. Identification of host DEAD-box RNA helicases that regulate cellular tropism of oncolytic Myxoma virus in human cancer cells. Sci. Rep. 2017, 7, 15710. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Xing, Z.; Mani, S.K.; Bancel, B.; Durantel, D.; Zoulim, F.; Tran, E.J.; Merle, P.; Andrisani, O. RNA helicase DEAD-box protein 5 regulates Polycomb repressive complex 2/Hox transcript antisense intergenic RNA function in hepatitis B virus infection and hepatocarcinogenesis. Hepatology 2016, 64, 1033–1048. [Google Scholar] [CrossRef] [PubMed]
- Ayoubian, H.; Fröhlich, T.; Pogodski, D.; Flatley, A.; Kremmer, E.; Schepers, A.; Feederle, R.; Arnold, G.J.; Grässer, F.A. Antibodies against the mono-methylated arginine-glycine repeat (MMA-RG) of the Epstein-Barr virus nuclear antigen 2 (EBNA2) identify potential cellular proteins targeted in viral transformation. J. Gen. Virol. 2017, 98, 2128–2142. [Google Scholar] [CrossRef] [PubMed]
- Jefferson, M.; Donaszi-Ivanov, A.; Pollen, S.; Dalmay, T.; Saalbach, G.; Powell, P.P. Host factors that interact with the Pestivirus N-terminal protease, Npro, are components of the ribonucleoprotein complex. J. Virol. 2014, 88, 10340–10353. [Google Scholar] [CrossRef] [PubMed]
- Cordin, O.; Banroques, J.; Tanner, N.K.; Linder, P. The DEAD-box protein family of RNA helicases. Gene 2006, 367, 17–37. [Google Scholar] [CrossRef] [PubMed]
- Linder, P.; Fuller-Pace, F. Happy birthday: 25 years of DEAD-box proteins. Methods Mol. Biol. 2015, 1259, 17–33. [Google Scholar] [PubMed]
- Caruthers, J.M.; McKay, D.B. Helicase structure and mechanism. Curr. Opin. Struct. Biol. 2002, 12, 123–133. [Google Scholar] [CrossRef]
- Tanner, N.K. The newly identified Q motif of DEAD-box helicases is involved in adenine recognition. Cell Cycle 2003, 2, 18–19. [Google Scholar] [CrossRef] [PubMed]
- Rocak, S.; Linder, P. DEAD-box proteins: The driving forces behind RNA metabolism. Nat. Rev. Mol. Cell Biol. 2004, 5, 232–241. [Google Scholar] [CrossRef] [PubMed]
- Cordin, O.; Tanner, N.K.; Doère, M.; Linder, P.; Banroques, J. The newly discovered Q motif of DEAD-box RNA helicases regulates RNA-binding and helicase activity. EMBO J. 2004, 23, 2478–2487. [Google Scholar] [CrossRef] [PubMed]
- Tanner, N.K.; Linder, P. DExD/H box RNA helicases: From generic motors to specific dissociation functions. Mol. Cell 2001, 8, 251–262. [Google Scholar] [CrossRef]
- Schwer, B.; Meszaros, T. RNA helicase dynamics in pre-mRNA splicing. EMBO J. 2000, 19, 6582–6591. [Google Scholar] [CrossRef] [PubMed]
- Schütz, P.; Karlberg, T.; van den Berg, S.; Collins, R.; Lehtiö, L.; Högbom, M.; Holmberg-Schiavone, L.; Tempel, W.; Park, H.W.; Hammarström, M.; et al. Comparative structural analysis of human DEAD-box RNA helicases. PLoS ONE 2010, 5, e12791. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.J.; Lee, S.G. The DEAD-box RNA helicase DDX3 interacts with DDX5, co-localizes with it in the cytoplasm during the G2/M phase of the cycle, and affects its shuttling during mRNP export. J. Cell Biochem. 2012, 113, 985–996. [Google Scholar] [CrossRef] [PubMed]
- Decroly, E.; Debarnot, C.; Ferron, F.; Bouvet, M.; Coutard, B.; Imbert, I.; Gluais, L.; Papageorgiou, N.; Sharff, A.; Bricogne, G.; et al. Crystal structure and functional analysis of the SARS-coronavirus RNA cap 2′-O-methyltransferase nsp10/nsp16 complex. PLoS Pathog. 2011, 7, e1002059. [Google Scholar] [CrossRef] [PubMed]
- Bouvet, M.; Imbert, I.; Subissi, L.; Gluais, L.; Canard, B.; Decroly, E. RNA 3′-end mismatch excision by the severe acute respiratory syndrome coronavirus nonstructural protein nsp10/nsp14 exoribonuclease complex. Proc. Natl. Acad. Sci. USA 2012, 109, 9372–9377. [Google Scholar] [CrossRef] [PubMed]
- Thiel, V.; Ivanov, K.A.; Putics, A.; Hertzig, T.; Schelle, B.; Bayer, S.; Weissbrich, B.; Snijder, E.J.; Rabenau, H.; Doerr, H.W.; et al. Mechanisms and enzymes involved in SARS coronavirus genome expression. J. Gen. Virol. 2003, 84, 2305–2315. [Google Scholar] [CrossRef] [PubMed]
- Adedeji, A.O.; Marchand, B.; Te Velthuis, A.J.; Snijder, E.J.; Weiss, S.; Eoff, R.L.; Singh, K.; Sarafianos, S.G. Mechanism of nucleic acid unwinding by SARS-CoV helicase. PLoS ONE 2012, 7, e36521. [Google Scholar] [CrossRef] [PubMed]
- Lee, N.R.; Kwon, H.M.; Park, K.; Oh, S.; Jeong, Y.J.; Kim, D.E. Cooperative translocation enhances the unwinding of duplex DNA by SARS coronavirus helicase nsp13. Nucleic Acids Res. 2010, 38, 7626–7636. [Google Scholar] [CrossRef] [PubMed]
- Subissi, L.; Imbert, I.; Ferron, F.; Collet, A.; Coutard, B.; Decroly, E.; Canard, B. SARS-CoV ORF1b-encoded nonstructural proteins 12–16: Replicative enzymes as antiviral targets. Antivir. Res. 2014, 101, 122–130. [Google Scholar] [CrossRef] [PubMed]
- Kwong, A.D.; Rao, B.G.; Jeang, K.T. Viral and cellular RNA helicases as antiviral targets. Nat. Rev. Drug Discov. 2005, 4, 845–853. [Google Scholar] [CrossRef] [PubMed]
- Xu, L.; Khadijah, S.; Fang, S.; Wang, L.; Tay, F.P.; Liu, D.X. The cellular RNA helicase DDX1 interacts with coronavirus nonstructural protein 14 and enhances viral replication. J. Virol. 2010, 84, 8571–8583. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.H.; Chen, P.J.; Yeh, S.H. Nucleocapsid phosphorylation and RNA helicase DDX1 recruitment enables coronavirus transition from discontinuous to continuous transcription. Cell Host Microbe 2014, 16, 462–472. [Google Scholar] [CrossRef] [PubMed]
- Emmott, E.; Munday, D.; Bickerton, E.; Britton, P.; Rodgers, M.A.; Whitehouse, A.; Zhou, E.M.; Hiscox, J.A. The cellular interactome of the coronavirus infectious bronchitis virus nucleocapsid protein and functional implications for virus biology. J. Virol. 2013, 87, 9486–9500. [Google Scholar] [CrossRef] [PubMed]
- Carter, C.A.; Ehrlich, L.S. Cell biology of HIV-1 infection of macrophages. Annu. Rev. Microbiol. 2008, 62, 425–443. [Google Scholar] [CrossRef] [PubMed]
- Faust, T.B.; Binning, J.M.; Gross, J.D.; Frankel, A.D. Making sense of multifunctional proteins: Human immunodeficiency virus type 1 accessory and regulatory proteins and connections to transcription. Annu. Rev. Virol. 2017, 4, 241–260. [Google Scholar] [CrossRef] [PubMed]
- Fellay, J.; Shianna, K.V.; Ge, D.; Colombo, S.; Ledergerber, B.; Weale, M.; Zhang, K.; Gumbs, C.; Castagna, A.; Cossarizza, A.; et al. A whole-genome association study of major determinants for host control of HIV-1. Science 2007, 317, 944–947. [Google Scholar] [CrossRef] [PubMed]
- Brass, A.L.; Dykxhoorn, D.M.; Benita, Y.; Yan, N.; Engelman, A.; Xavier, R.J.; Lieberman, J.; Elledge, S.J. Identification of host proteins required for HIV infection through a functional genomic screen. Science 2008, 319, 921–926. [Google Scholar] [CrossRef] [PubMed]
- Bushman, F.D.; Malani, N.; Fernandes, J.; D’Orso, I.; Cagney, G.; Diamond, T.L.; Zhou, H.; Hazuda, D.J.; Espeseth, A.S.; König, R.; et al. Host cell factors in HIV replication: Meta-analysis of genome-wide studies. PLoS Pathog. 2009, 5, e1000437. [Google Scholar] [CrossRef] [PubMed]
- Fang, J.; Kubota, S.; Yang, B.; Zhou, N.; Zhang, H.; Godbout, R.; Pomerantz, R.J. A DEAD-box protein facilitates HIV-1 replication as a cellular co-factor of Rev. Virology 2004, 330, 471–480. [Google Scholar] [CrossRef] [PubMed]
- Jeang, K.T.; Yedavalli, V. Role of RNA helicases in HIV-1 replication. Nucleic Acids Res. 2006, 34, 4198–4205. [Google Scholar] [CrossRef] [PubMed]
- Yasuda-Inoue, M.; Kuroki, M.; Ariumi, Y. Distinct DDX DEAD-box RNA helicases cooperate to modulate the HIV-1 Rev function. Biochem. Biophys. Res. Commun. 2013, 434, 803–808. [Google Scholar] [CrossRef] [PubMed]
- Ariumi, Y. Multiple functions of DDX3 RNA helicase in gene regulation, tumorigenesis, and viral infection. Front. Genet. 2014, 5, 423. [Google Scholar] [CrossRef] [PubMed]
- Williams, C.A.; Abbink, T.E.; Jeang, K.T.; Lever, A.M. Identification of RNA helicases in human immunodeficiency virus 1 (HIV-1) replication—A targeted small interfering RNA library screen using pseudotyped and WT HIV-1. J. Gen. Virol. 2015, 96, 1484–1489. [Google Scholar] [CrossRef] [PubMed]
- Fujii, R.; Okamoto, M.; Aratani, S.; Oishi, T.; Ohshima, T.; Taira, K.; Baba, M.; Fukamizu, A.; Nakajima, T. A role of RNA helicase A in cis-acting transactivation response element-mediated transcriptional regulation of human immunodeficiency virus type 1. J. Biol. Chem. 2001, 276, 5445–5451. [Google Scholar] [CrossRef] [PubMed]
- Lai, M.C.; Wang, S.W.; Cheng, L.; Tarn, W.Y.; Tsai, S.J.; Sun, H.S. Human DDX3 interacts with the HIV-1 Tat protein to facilitate viral mRNA translation. PLoS ONE 2013, 8, e68665. [Google Scholar] [CrossRef] [PubMed]
- Yasuda-Inoue, M.; Kuroki, M.; Ariumi, Y. DDX3 RNA helicase is required for HIV-1 Tat function. Biochem. Biophys. Res. Commun. 2013, 441, 607–611. [Google Scholar] [CrossRef] [PubMed]
- Rausch, J.W.; Grice, S.F. HIV rev assembly on the rev response element (RRE): A structural perspective. Viruses 2015, 7, 3053–3075. [Google Scholar] [CrossRef] [PubMed]
- Sherpa, C.; Rausch, J.W.; Le Grice, S.F.; Hammarskjold, M.L.; Rekosh, D. The HIV-1 Rev response element (RRE) adopts alternative conformations that promote different rates of virus replication. Nucleic Acids Res. 2015, 43, 4676–4686. [Google Scholar] [CrossRef] [PubMed]
- Sithole, N.; Williams, C.; Vaughan, A.; Lever, A. The roles of DEAD-box helicases in the life cycle of HIV-1. Lancet 2015, 385, S89. [Google Scholar] [CrossRef]
- Moss, B. Poxvirus DNA replication. Cold Spring Harb. Perspect. Biol. 2013, 5, a010199. [Google Scholar] [CrossRef] [PubMed]
- Gehring, A.; Bertoletti, A.; Tavis, J.E. Host factor-targeted hepatitis B virus therapies. Intervirology 2014, 57, 158–162. [Google Scholar] [CrossRef] [PubMed]
- König, R.; Stertz, S. Recent strategies and progress in identifying host factors involved in virus replication. Curr. Opin. Microbiol. 2015, 26, 79–88. [Google Scholar] [CrossRef] [PubMed]
- Radi, M.; Falchi, F.; Garbelli, A.; Samuele, A.; Bernardo, V.; Paolucci, S.; Baldanti, F.; Schenone, S.; Manetti, F.; Maga, G.; et al. Discovery of the first small molecule inhibitor of human DDX3 specifically designed to target the RNA binding site: Towards the next generation HIV-1 inhibitors. Bioorg. Med. Chem. Lett. 2012, 22, 2094–2098. [Google Scholar] [CrossRef] [PubMed]
- Samal, S.K.; Routray, S.; Veeramachaneni, G.K.; Dash, R.; Botlagunta, M. Ketorolac salt is a newly discovered DDX3 inhibitor to treat oral cancer. Sci. Rep. 2015, 5, 9982. [Google Scholar] [CrossRef] [PubMed]
- Bol, G.M.; Vesuna, F.; Xie, M.; Zeng, J.; Aziz, K.; Gandhi, N.; Levine, A.; Irving, A.; Korz, D.; Tantravedi, S.; et al. Targeting DDX3 with a small molecule inhibitor for lung cancer therapy. EMBO Mol. Med. 2015, 7, 648–669. [Google Scholar] [CrossRef] [PubMed]
- McGivern, D.R.; Lemon, S.M. Virus-specific mechanisms of carcinogenesis in hepatitis C virus associated liver cancer. Oncogene 2011, 30, 1969–1983. [Google Scholar] [CrossRef] [PubMed]
- Kao, C.C.; Yi, G.; Huang, H.C. The core of hepatitis C virus pathogenesis. Curr. Opin. Virol. 2016, 17, 66–73. [Google Scholar] [CrossRef] [PubMed]
- Xie, Y.; Ogah, C.A.; Jiang, X.; Li, J.; Shen, J. Nucleoside inhibitors of Hepatitis C virus NS5B polymerase: A systematic review. Curr. Drug Targets 2016, 17, 1560–1576. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, S. Recent advances in the molecular biology of hepatitis C virus. J. Mol. Biol. 2001, 313, 451–464. [Google Scholar] [CrossRef] [PubMed]
- Gao, L.; Tu, H.; Shi, S.T.; Lee, K.J.; Asanaka, M.; Hwang, S.B.; Lai, M.M. Interaction with a ubiquitin-like protein enhances the ubiquitination and degradation of hepatitis C virus RNA-dependent RNA polymerase. J. Virol. 2003, 77, 4149–4159. [Google Scholar] [CrossRef] [PubMed]
- Lim, Y.S.; Tran, H.T.; Park, S.J.; Yim, S.A.; Hwang, S.B. Peptidyl-prolyl isomerase Pin1 is a cellular factor required for hepatitis C virus propagation. J. Virol. 2011, 85, 8777–8788. [Google Scholar] [CrossRef] [PubMed]
- Pham, L.V.; Ngo, H.T.; Lim, Y.S.; Hwang, S.B. Hepatitis C virus non-structural 5B protein interacts with cyclin A2 and regulates viral propagation. J. Hepatol. 2012, 57, 960–966. [Google Scholar] [CrossRef] [PubMed]
- Yoo, Y.H.; Yun, J.; Yoon, C.N.; Lee, J.S. Chemical proteomic identification of T-plastin as a novel host cell response factor in HCV infection. Sci. Rep. 2015, 5, 9773. [Google Scholar] [CrossRef] [PubMed]
- Kuroki, M.; Ariumi, Y.; Hijikata, M.; Ikeda, M.; Dansako, H.; Wakita, T.; Shimotohno, K.; Kato, N. PML tumor suppressor protein is required for HCV production. Biochem. Biophys. Res. Commun. 2013, 430, 592–597. [Google Scholar] [CrossRef] [PubMed]
- Dutta, S.; Gupta, G.; Choi, Y.W.; Kotaka, M.; Fielding, B.C.; Song, J.; Tan, Y.J. The variable N-terminal region of DDX5 contains structural elements and auto-inhibits its interaction with NS5B of hepatitis C virus. Biochem. J. 2012, 446, 37–46. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Harris, D.; Zhang, Z.; Chaubey, B.; Pandey, V.N. Identification of cellular factors associated with the 3′-nontranslated region of the hepatitis C virus genome. Mol. Cell Proteom. 2006, 5, 1006–1018. [Google Scholar] [CrossRef] [PubMed]
- Upadya, M.H.; Aweya, J.J.; Tan, Y.J. Understanding the interaction of hepatitis C virus with host DEAD-box RNA helicases. World J. Gastroenterol. 2014, 20, 2913–2926. [Google Scholar] [CrossRef] [PubMed]
- Ríos-Marco, P.; Romero-López, C.; Berzal-Herranz, A. The cis-acting replication element of the Hepatitis C virus genome recruits host factors that influence viral replication and translation. Sci. Rep. 2016, 6, 25729. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.I.; Lee, Y.M. Japanese encephalitis: The virus and vaccines. Hum. Vaccines Immunother. 2014, 10, 263–279. [Google Scholar] [CrossRef] [PubMed]
- Heffelfinger, J.D.; Li, X.; Batmunkh, N.; Grabovac, V.; Diorditsa, S.; Liyanage, J.B.; Pattamadilok, S.; Bahl, S.; Vannice, K.S.; Hyde, T.B.; et al. Japanese encephalitis surveillance and immunization-Asia and Western Pacific regions, 2016. MMWR Morb. Mortal. Wkly. Rep. 2017, 66, 579–583. [Google Scholar] [CrossRef] [PubMed]
- Selisko, B.; Wang, C.; Harris, E.; Canard, B. Regulation of Flavivirus RNA synthesis and replication. Curr. Opin. Virol. 2014, 9, 74–83. [Google Scholar] [CrossRef] [PubMed]
- Kofler, R.M.; Heinz, F.X.; Mandl, C.W. Capsid protein C of tick-borne encephalitis virus tolerates large internal deletions and is a favorable target for attenuation of virulence. J. Virol. 2002, 76, 3534–3543. [Google Scholar] [CrossRef] [PubMed]
- Katoh, H.; Okamoto, T.; Fukuhara, T.; Kambara, H.; Morita, E.; Mori, Y.; Kamitani, W.; Matsuura, Y. Japanese encephalitis virus core protein inhibits stress granule formation through an interaction with Caprin-1 and facilitates viral propagation. J. Virol. 2013, 87, 489–502. [Google Scholar] [CrossRef] [PubMed]
- Egloff, M.P.; Benarroch, D.; Selisko, B.; Romette, J.L.; Canard, B. An RNA cap (nucleoside-2′-O-)-methyltransferase in the Flavivirus RNA polymerase NS5: Crystal structure and functional characterization. EMBO J. 2002, 21, 2757–2768. [Google Scholar] [CrossRef] [PubMed]
- Uchil, P.D.; Satchidanandam, V. Architecture of the flaviviral replication complex. Protease, nuclease, and detergents reveal encasement within double-layered membrane compartments. J. Biol. Chem. 2003, 278, 24388–24398. [Google Scholar] [CrossRef] [PubMed]
- Li, C.; Ge, L.L.; Li, P.P.; Wang, Y.; Dai, J.J.; Sun, M.X.; Huang, L.; Shen, Z.Q.; Hu, X.C.; Ishag, H.; et al. Cellular DDX3 regulates Japanese encephalitis virus replication by interacting with viral un-translated regions. Virology 2014, 449, 70–81. [Google Scholar] [CrossRef] [PubMed]
- Peterhans, E.; Schweizer, M. Pestiviruses: How to outmaneuver your hosts. Vet. Microbiol. 2010, 142, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Tautz, N.; Tews, B.A.; Meyers, G. The molecular biology of pestiviruses. Adv. Virus Res. 2015, 93, 47–160. [Google Scholar] [PubMed]
- Wang, F.I.; Deng, M.C.; Huang, Y.L.; Chang, C.Y. Structures and functions of Pestivirus glycoproteins: Not simply surface matters. Viruses 2015, 7, 3506–3529. [Google Scholar] [CrossRef] [PubMed]
- Gottipati, K.; Acholi, S.; Ruggli, N.; Choi, K.H. Autocatalytic activity and substrate specificity of the Pestivirus N-terminal protease Npro. Virology 2014, 452–453, 303–309. [Google Scholar] [CrossRef] [PubMed]
- Kappes, M.A.; Faaberg, K.S. PRRSV structure, replication and recombination: Origin of phenotype and genotype diversity. Virology 2015, 479–480, 475–486. [Google Scholar] [CrossRef] [PubMed]
- Yun, S.I.; Lee, Y.M. Overview: Replication of porcine reproductive and respiratory syndrome virus. J. Microbiol. 2013, 51, 711–723. [Google Scholar] [CrossRef] [PubMed]
- Fang, Y.; Snijder, E.J. The PRRSV replicase: Exploring the multifunctionality of an intriguing set of nonstructural proteins. Virus Res. 2010, 154, 61–76. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Zhou, L.; Zhang, J.; Ge, X.; Zhou, R.; Zheng, H.; Geng, G.; Guo, X.; Yang, H. Nsp9 and Nsp10 contribute to the fatal virulence of highly pathogenic porcine reproductive and respiratory syndrome virus emerging in China. PLoS Pathog. 2014, 10, e1004216. [Google Scholar] [CrossRef] [PubMed]
- Xie, J.; Zhou, H.; Cui, J.; Chen, Y.; Zhang, M.; Deng, S.; Zhou, P.; Su, S.; Zhang, G. Inhibition of porcine reproductive and respiratory syndrome virus by specific siRNA targeting Nsp9 gene. Infect. Genet. Evol. 2014, 28, 64–70. [Google Scholar] [CrossRef] [PubMed]
- Harfoot, R.; Webby, R.J. H5 influenza, a global update. J. Microbiol. 2017, 55, 196–203. [Google Scholar] [CrossRef] [PubMed]
- Bui, C.; Bethmont, A.; Chughtai, A.A.; Gardner, L.; Sarkar, S.; Hassan, S.; Seale, H.; MacIntyre, C.R. A systematic review of the comparative epidemiology of avian and human influenza A H5N1 and H7N9—Lessons and unanswered questions. Transbound Emerg. Dis. 2016, 63, 602–620. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Wu, A.; Wang, Y.E.; Quanquin, N.; Li, C.; Wang, J.; Chen, H.W.; Liu, S.; Liu, P.; Zhang, H.; et al. Functional genomics reveals linkers critical for influenza polymerase. J. Virol. 2015, 90, 2938–2947. [Google Scholar] [CrossRef] [PubMed]
- Weigel, T.; Solomaier, T.; Wehmeyer, S.; Peuker, A.; Wolff, M.W.; Reichl, U. A membrane-based purification process for cell culture-derived influenza A virus. J. Biotechnol. 2016, 220, 12–20. [Google Scholar] [CrossRef] [PubMed]
- Zhong, G.; Le, M.Q.; Lopes, T.J.S.; Halfmann, P.; Hatta, M.; Fan, S.; Neumann, G.; Kawaoka, Y. Mutations in the PA protein of avian H5N1 influenza viruses affect polymerase activity and mouse virulence. J. Virol. 2017, 92, e01557-17. [Google Scholar] [CrossRef] [PubMed]
- Fuller-Pace, F.V. The DEAD-box proteins DDX5 (p68) and DDX17 (p72): Multi-tasking transcriptional regulators. Biochim. Biophys. Acta 2013, 1829, 756–763. [Google Scholar] [CrossRef] [PubMed]
- Prockop, S.E.; Vatsayan, A. Epstein-Barr virus lymphoproliferative disease after solid organ transplantation. Cytotherapy 2017, 19, 1270–1283. [Google Scholar] [CrossRef] [PubMed]
- Hau, P.M.; Tsao, S.W. Epstein-Barr virus hijacks DNA damage response transducers to orchestrate its life cycle. Viruses 2017, 9, 341. [Google Scholar] [CrossRef] [PubMed]
- Kempkes, B.; Ling, P.D. EBNA2 and its coactivator EBNA-LP. Curr. Top. Microbiol. Immunol. 2015, 391, 35–59. [Google Scholar] [PubMed]
- Sylvestersen, K.B.; Horn, H.; Jungmichel, S.; Jensen, L.J.; Nielsen, M.L. Proteomic analysis of arginine methylation sites in human cells reveals dynamic regulation during transcriptional arrest. Mol. Cell Proteom. 2014, 13, 2072–2088. [Google Scholar] [CrossRef] [PubMed]
- Kerr, P.J.; Liu, J.; Cattadori, I.; Ghedin, E.; Read, A.F.; Holmes, E.C. Myxoma virus and the Leporipoxviruses: An evolutionary paradigm. Viruses 2015, 7, 1020–1061. [Google Scholar] [CrossRef] [PubMed]
- Chan, W.M.; Rahman, M.M.; McFadden, G. Oncolytic myxoma virus: The path to clinic. Vaccine 2013, 31, 4252–4258. [Google Scholar] [CrossRef] [PubMed]
- Guerra, S.; Nájera, J.L.; González, J.M.; López-Fernández, L.A.; Climent, N.; Gatell, J.M.; Gallart, T.; Esteban, M. Distinct gene expression profiling after infection of immature human monocyte-derived dendritic cells by the attenuated poxvirus vectors MVA and NYVAC. J. Virol. 2007, 81, 8707–8721. [Google Scholar] [CrossRef] [PubMed]
- Moy, R.H.; Cole, B.S.; Yasunaga, A.; Gold, B.; Shankarling, G.; Varble, A.; Molleston, J.M.; ten Oever, B.R.; Lynch, K.W.; Cherry, S. Stem-loop recognition by DDX17 facilitates miRNA processing and antiviral defense. Cell 2014, 158, 764–777. [Google Scholar] [CrossRef] [PubMed]
- Levrero, M.; Testoni, B.; Zoulim, F. HBV cure: Why, how, when? Curr. Opin. Virol. 2016, 18, 135–143. [Google Scholar] [CrossRef] [PubMed]
- Iida-Ueno, A.; Enomoto, M.; Tamori, A.; Kawada, N. Hepatitis B virus infection and alcohol consumption. World J. Gastroenterol. 2017, 23, 2651–2659. [Google Scholar] [CrossRef] [PubMed]
- Seeger, C.; Mason, W.S. Molecular biology of hepatitis B virus infection. Virology 2015, 479–480, 672–686. [Google Scholar] [CrossRef] [PubMed]
- Mani, S.K.K.; Andrisani, O. Hepatitis B virus-associated hepatocellular carcinoma and hepatic cancer stem cells. Genes 2018, 9, 137. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Kim, T.; Bao, M.; Facchinetti, V.; Jung, S.Y.; Ghaffari, A.A.; Qin, J.; Cheng, G.; Liu, Y.J. DDX1, DDX21, and DHX36 helicases form a complex with the adaptor molecule TRIF to sense dsRNA in dendritic cells. Immunity 2011, 34, 866–878. [Google Scholar] [CrossRef] [PubMed]
- Kim, T.; Pazhoor, S.; Bao, M.; Zhang, Z.; Hanabuchi, S.; Facchinetti, V.; Bover, L.; Plumas, J.; Chaperot, L.; Qin, J.; et al. Aspartate-glutamate-alanine-histidine box motif (DEAH)/RNA helicase A helicases sense microbial DNA in human plasmacytoid dendritic cells. Proc. Natl. Acad. Sci. USA 2010, 107, 15181–15186. [Google Scholar] [CrossRef] [PubMed]
- Mitoma, H.; Hanabuchi, S.; Kim, T.; Bao, M.; Zhang, Z.; Sugimoto, N.; Liu, Y.J. The DHX33 RNA helicase senses cytosolic RNA and activates the NLRP3 inflammasome. Immunity 2013, 39, 123–135. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Yuan, B.; Bao, M.; Lu, N.; Kim, T.; Liu, Y.J. The helicase DDX41 senses intracellular DNA mediated by the adaptor STING in dendritic cells. Nat. Immunol. 2011, 12, 959–965. [Google Scholar] [CrossRef] [PubMed]
- Miyashita, M.; Oshiumi, H.; Matsumoto, M.; Seya, T. DDX60, a DEXD/H box helicase, is a novel antiviral factor promoting RIG-I-like receptor-mediated signaling. Mol. Cell Biol. 2011, 31, 3802–3819. [Google Scholar] [CrossRef] [PubMed]
- Nikonov, A.; Mölder, T.; Sikut, R.; Kiiver, K.; Männik, A.; Toots, U.; Lulla, A.; Lulla, V.; Utt, A.; Merits, A.; et al. RIG-I and MDA-5 detection of viral RNA-dependent RNA polymerase activity restricts positive-strand RNA virus replication. PLoS Pathog. 2013, 9, e1003610. [Google Scholar] [CrossRef] [PubMed]
- Ranji, A.; Boris-Lawrie, K. RNA helicases: Emerging roles in viral replication and the host innate response. RNA Biol. 2010, 7, 775–787. [Google Scholar] [CrossRef] [PubMed]
| Virus | Viral Binding Protein | Effect of DDX5 on Viral Replication | Ref. |
|---|---|---|---|
| SARS-CoV | nsp13 | Enhances virus proliferation | [16] |
| IBV-CoV | nsp13 | Enhances virus proliferation | [16] |
| HIV-1 | Rev | Enhances the Rev-dependent RNA export | [17] |
| HCV | NS5B | Required for transcription of viral RNA | [18] |
| JEV | C, NS3 and NS5 | Required for viral RNA replication | [19] |
| PRRSV | nsp9 | Required for viral replication | [20] |
| Influenza virus | NP | Required for viral replication | [21] |
| EBV | EBNA2 | Unknown | [24] |
| Pestivirus | Npro | Unknown | [25] |
| Myxoma virus | Unknown | Inhibits viral replication | [22] |
| HBV | Unknown | Inhibits viral replication | [23] |
| Cell Lines | HCV Strains | Effect on HCV RNA | Effect on HCV Particles | Ref. |
|---|---|---|---|---|
| 293 | HCV-S1 | Reduced | Unknown | [18] |
| Huh7.5 | J6/JFH-1 | Increased | Promotes a late step in viral replication | [81] |
| Huh-7-NS3-3′ | Unknown | Reduced | Unknown | [80] |
| RSc cells | HCV-JFH1 | Reduced | Suppressed | [78] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Cheng, W.; Chen, G.; Jia, H.; He, X.; Jing, Z. DDX5 RNA Helicases: Emerging Roles in Viral Infection. Int. J. Mol. Sci. 2018, 19, 1122. https://doi.org/10.3390/ijms19041122
Cheng W, Chen G, Jia H, He X, Jing Z. DDX5 RNA Helicases: Emerging Roles in Viral Infection. International Journal of Molecular Sciences. 2018; 19(4):1122. https://doi.org/10.3390/ijms19041122
Chicago/Turabian StyleCheng, Wenyu, Guohua Chen, Huaijie Jia, Xiaobing He, and Zhizhong Jing. 2018. "DDX5 RNA Helicases: Emerging Roles in Viral Infection" International Journal of Molecular Sciences 19, no. 4: 1122. https://doi.org/10.3390/ijms19041122





