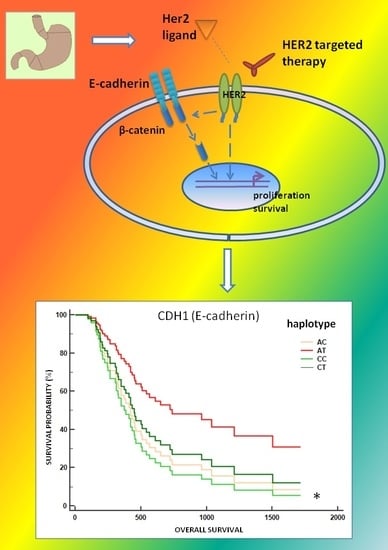
Characterizing Metastatic HER2-Positive Gastric Cancer at the CDH1 Haplotype
Abstract
:1. Introduction
2. Results
2.1. Patient Characteristics
2.2. CDH1 Mutations
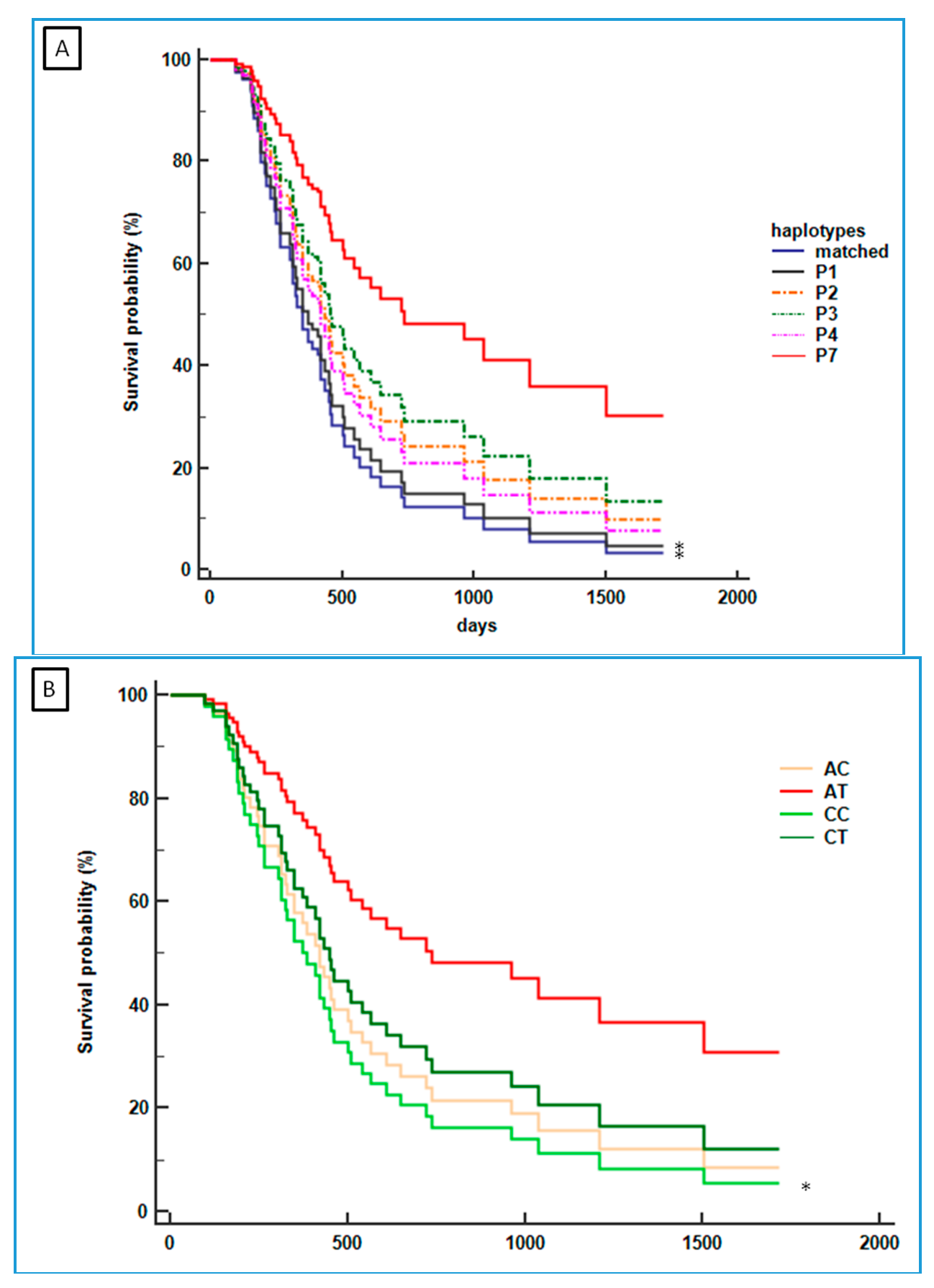
2.3. Association between CDH1 P7-Haplotype and mGC-HER2
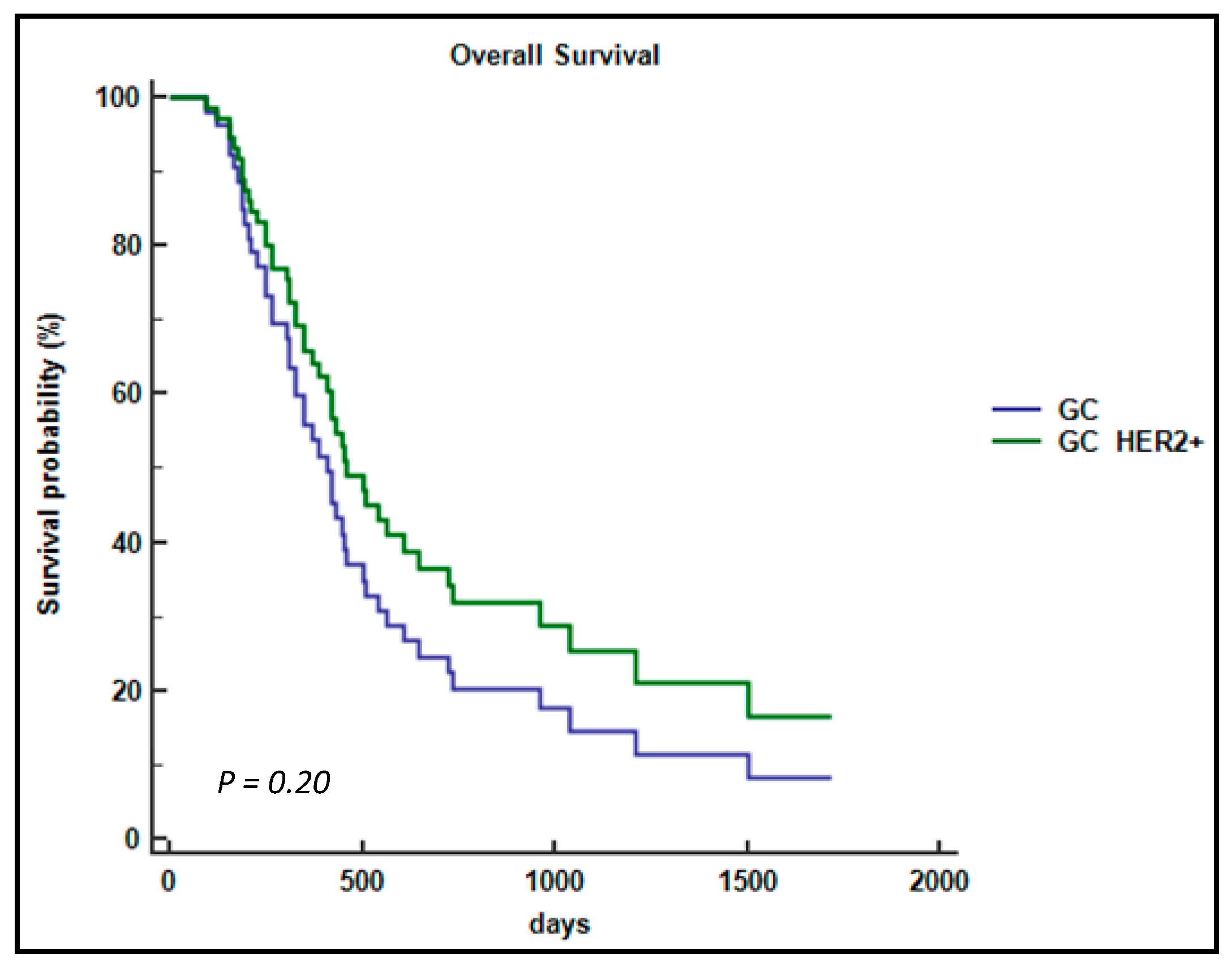
2.4. Association between the CDH1 P7 Haplotype and the Survival of mGC-HER2 Patients
3. Discussion
4. Materials and Methods
4.1. Study Population
4.2. Genotyping Analysis
4.3. Immunohistochemistry
4.4. Statistical Analysis
5. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| GC | Gastric cancer |
| mGC | Metastatic gastric cancer |
| HER2 | Human epidermal growth factor receptor 2 |
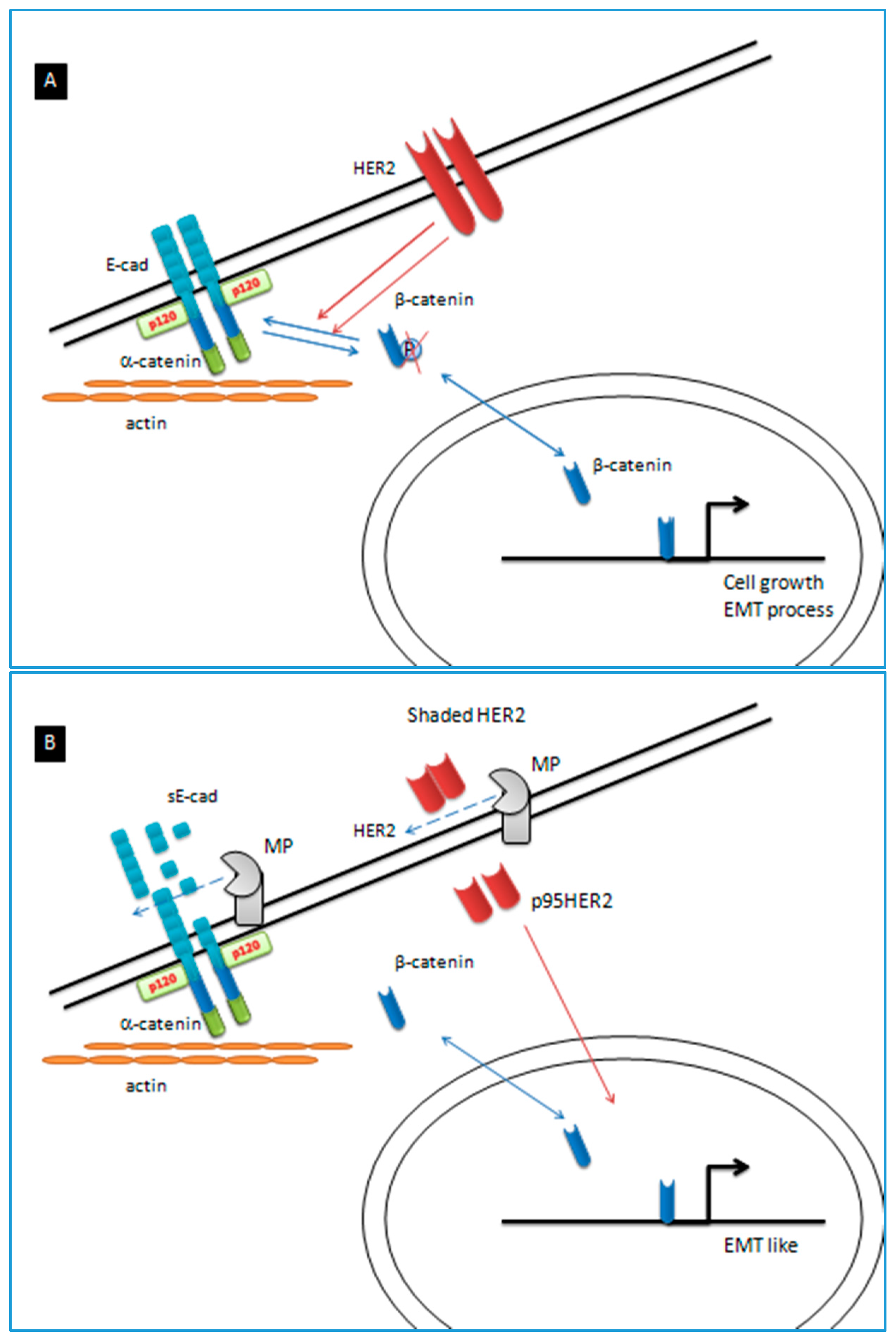
| E-cad | E-cadherin |
| IHC | Immunohistochemical |
| EBV | Epstein-barr virus |
| EMT | Epithelial-to-mesenchymal transition |
| MP | Metalloproteinase |
References
- Siegel, R.L.; Miller, K.D.; Jemal, A. Cancer statistics, 2017. CA Cancer J. Clin. 2017, 67, 7–30. [Google Scholar] [CrossRef] [PubMed]
- Santeufemia, D.A.; Lumachi, F.; Fadda, G.M.; Lo Re, G.; Miolo, G.; Basso, S.M.M.; Chiara, G.B.; Tumolo, S. Comment on “Repetitive transarterial chemoembolization (TACE) of liver metastases from gastric cancer: Local control and survival results”: Will there be clinical implications in the future? Eur. J. Radiol. 2013, 82, 1591–1592. [Google Scholar] [CrossRef] [PubMed]
- Romano, F.; Garancini, M.; Uggeri, F.; Degrate, L.; Nespoli, L.; Gianotti, L.; Nespoli, A.; Uggeri, F. Surgical treatment of liver metastases of gastric cancer: State of the art. World J. Surg. Oncol. 2012, 10, 157. [Google Scholar] [CrossRef] [PubMed]
- Catalano, V.; Graziano, F.; Santini, D.; D’Emidio, S.; Baldelli, A.M.; Rossi, D.; Vincenzi, B.; Giordani, P.; Alessandroni, P.; Testa, E.; et al. Second-line chemotherapy for patients with advanced gastric cancer: Who may benefit? Br. J. Cancer 2008, 99, 1402–1407. [Google Scholar] [CrossRef] [PubMed]
- Orditura, M.; Galizia, G.; Sforza, V.; Gambardella, V.; Fabozzi, A.; Laterza, M.M.; Andreozzi, F.; Ventriglia, J.; Savastano, B.; Mabilia, A.; et al. Treatment of gastric cancer. World J. Gastroenterol. 2014, 20, 1635–1649. [Google Scholar] [CrossRef] [PubMed]
- Bang, Y.-J.; Van Cutsem, E.; Feyereislova, A.; Chung, H.C.; Shen, L.; Sawaki, A.; Lordick, F.; Ohtsu, A.; Omuro, Y.; Satoh, T.; et al. Trastuzumab in combination with chemotherapy versus chemotherapy alone for treatment of HER2-positive advanced gastric or gastro-oesophageal junction cancer (ToGA): A phase 3, open-label, randomised controlled trial. Lancet 2010, 376, 687–697. [Google Scholar] [CrossRef]
- Garattini, S.K.; Basile, D.; Cattaneo, M.; Fanotto, V.; Ongaro, E.; Bonotto, M.; Negri, F.V.; Berenato, R.; Ermacora, P.; Cardellino, G.G.; et al. Molecular classifications of gastric cancers: Novel insights and possible future applications. World J. Gastrointest. Oncol. 2017, 9, 194–208. [Google Scholar] [CrossRef] [PubMed]
- Bonotto, M.; Garattini, S.K.; Basile, D.; Ongaro, E.; Fanotto, V.; Cattaneo, M.; Cortiula, F.; Iacono, D.; Cardellino, G.G.; Pella, N.; et al. Immunotherapy for gastric cancers: Emerging role and future perspectives. Expert Rev. Clin. Pharmacol. 2017, 10, 609–619. [Google Scholar] [CrossRef] [PubMed]
- Lubarsch, O.; Henke, F.; Rössle, R. Handbuch der Speziellen Pathologischen Anatomie und Histologie. Springer 1937, 9(part 3). Available online: http://www.springer.com/series/206 (accessed on 30 November 2017).
- Siewert, J.R.; Stein, H.J. Carcinoma of the gastroesophageal junction—Classification, pathology and extent of resection. Dis. Esophagus 1996, 9, 173–182. [Google Scholar] [CrossRef]
- WHO Classification of Tumours of the Digestive System, Fourth Edition. Available online: http://apps.who.int/bookorders/WHP/detart1.jsp?sesslan=1&codlan=1&codcol=70&codcch=4003 (accessed on 14 November 2017).
- Lauren, P. The two histological main types of gastric carcinoma: Diffuse and so-called intestinal-type carcinoma. An attempt at a histo-clinical classification. Acta Pathol. Microbiol. Scand. 1965, 64, 31–49. [Google Scholar] [CrossRef] [PubMed]
- Shan, L.; Ying, J.; Lu, N. HER2 expression and relevant clinicopathological features in gastric and gastroesophageal junction adenocarcinoma in a Chinese population. Diagn. Pathol. 2013, 8, 76. [Google Scholar] [CrossRef] [PubMed]
- The Cancer Genome Atlas Research Network. Comprehensive molecular characterization of gastric adenocarcinoma. Nature 2014, 513, 202–209. [Google Scholar] [CrossRef] [Green Version]
- Ye, X.S.; Yu, C.; Aggarwal, A.; Reinhard, C. Genomic alterations and molecular subtypes of gastric cancers in Asians. Chin. J. Cancer 2016, 35, 42. [Google Scholar] [CrossRef] [PubMed]
- Chu, C.-M.; Chen, C.-J.; Chan, D.-C.; Wu, H.-S.; Liu, Y.-C.; Shen, C.-Y.; Chang, T.-M.; Yu, J.; Harn, H.-J.; Yu, C.-P.; et al. CDH1 polymorphisms and haplotypes in sporadic diffuse and intestinal gastric cancer: A case–control study based on direct sequencing analysis. World J. Surg. Oncol. 2014, 12, 80. [Google Scholar] [CrossRef] [PubMed]
- Dang, H.-Z.; Yu, Y.; Jiao, S.-C. Prognosis of HER2 over-expressing gastric cancer patients with liver metastasis. World J. Gastroenterol. 2012, 18, 2402–2407. [Google Scholar] [CrossRef] [PubMed]
- Nami, B.; Wang, Z. HER2 in breast cancer stemness: A negative feedback loop towards trastuzumab resistance. Cancers 2017, 9, 40. [Google Scholar] [CrossRef] [PubMed]
- Namikawa, T.; Munekage, E.; Munekage, M.; Maeda, H.; Yatabe, T.; Kitagawa, H.; Sakamoto, K.; Obatake, M.; Kobayashi, M.; Hanazaki, K. Evaluation of a trastuzumab-containing treatment regimen for patients with unresectable advanced or recurrent gastric cancer. Mol. Clin. Oncol. 2016, 5, 74–78. [Google Scholar] [CrossRef] [PubMed]
- Caggiari, L.; Miolo, G.; Canzonieri, V.; De Zorzi, M.; Alessandrini, L.; Corona, G.; Cannizzaro, R.; Santeufemia, D.A.; Cossu, A.; Buonadonna, A.; et al. A new mutation of the CDH1 gene in a patient with an aggressive signet-ring cell carcinoma of the stomach. Cancer Biol. Ther. 2017, 1–6. [Google Scholar] [CrossRef] [PubMed]
- Pisignano, G.; Napoli, S.; Magistri, M.; Mapelli, S.N.; Pastori, C.; Marco, S.D.; Civenni, G.; Albino, D.; Enriquez, C.; Allegrini, S.; et al. A promoter-proximal transcript targeted by genetic polymorphism controls E-cadherin silencing in human cancers. Nat. Commun. 2017, 8, 15622. [Google Scholar] [CrossRef] [PubMed]
- Zhan, Z.; Wu, J.; Zhang, J.-F.; Yang, Y.-P.; Tong, S.; Zhang, C.-B.; Li, J.; Yang, X.-W.; Dong, W. CDH1 gene polymorphisms, plasma CDH1 levels and risk of gastric cancer in a Chinese population. Mol. Biol. Rep. 2012, 39, 8107–8113. [Google Scholar] [CrossRef] [PubMed]
- Jiang, B.; Zhu, K.; Shao, H.; Bao, C.; Ou, J.; Sun, W. Lack of association between the CDH1 polymorphism and gastric cancer susceptibility: A meta-analysis. Sci. Rep. 2015, 5, 7891. [Google Scholar] [CrossRef] [PubMed]
- Chen, B.; Zhou, Y.; Yang, P.; Liu, L.; Qin, X.-P.; Wu, X.-T. CDH1 -160C>A gene polymorphism is an ethnicity-dependent risk factor for gastric cancer. Cytokine 2011, 55, 266–273. [Google Scholar] [CrossRef] [PubMed]
- Memni, H.; Macherki, Y.; Klayech, Z.; Ben-Haj-Ayed, A.; Farhat, K.; Remadi, Y.; Gabbouj, S.; Mahfoudh, W.; Bouzid, N.; Bouaouina, N.; et al. E-cadherin genetic variants predict survival outcome in breast cancer patients. J. Transl. Med. 2016, 14, 320. [Google Scholar] [CrossRef] [PubMed]
- Matsusaka, S.; Zhang, W.; Cao, S.; Hanna, D.L.; Sunakawa, Y.; Sebio, A.; Ueno, M.; Yang, D.; Ning, Y.; Parekh, A.; et al. TWIST1 polymorphisms predict survival in patients with metastatic colorectal cancer receiving first-line bevacizumab plus oxaliplatin-based chemotherapy. Mol. Cancer Ther. 2016, 15, 1405–1411. [Google Scholar] [CrossRef] [PubMed]
- Song, H.; Wang, X.; Yan, J.; Mi, N.; Jiao, X.; Hao, Y.; Zhang, W.; Gao, Y. Association of single-nucleotide polymorphisms of CDH1 with nonsyndromic cleft lip with or without cleft palate in a northern Chinese Han population. Medicine (Baltimore) 2017, 96, e5574. [Google Scholar] [CrossRef] [PubMed]
- Kluijt, I.; Siemerink, E.J.M.; Ausems, M.G.E.M.; van Os, T.A.M.; de Jong, D.; Simões-Correia, J.; van Krieken, J.H.; Ligtenberg, M.J.; Figueiredo, J.; van Riel, E.; et al. Dutch working group on hereditary gastric cancer CDH1-related hereditary diffuse gastric cancer syndrome: Clinical variations and implications for counseling. Int. J. Cancer 2012, 131, 367–376. [Google Scholar] [CrossRef] [PubMed]
- Benusiglio, P.R.; Malka, D.; Rouleau, E.; De Pauw, A.; Buecher, B.; Noguès, C.; Fourme, E.; Colas, C.; Coulet, F.; Warcoin, M.; et al. CDH1 germline mutations and the hereditary diffuse gastric and lobular breast cancer syndrome: A multicentre study. J. Med. Genet. 2013, 50, 486–489. [Google Scholar] [CrossRef] [PubMed]
- Frebourg, T.; Oliveira, C.; Hochain, P.; Karam, R.; Manouvrier, S.; Graziadio, C.; Vekemans, M.; Hartmann, A.; Baert-Desurmont, S.; Alexandre, C.; et al. Cleft lip/palate and CDH1/E-cadherin mutations in families with hereditary diffuse gastric cancer. J. Med. Genet. 2006, 43, 138–142. [Google Scholar] [CrossRef] [PubMed]
- Mossey, P.A.; Little, J.; Munger, R.G.; Dixon, M.J.; Shaw, W.C. Cleft lip and palate. Lancet 2009, 374, 1773–1785. [Google Scholar] [CrossRef]
- Pinheiro, H.; Bordeira-Carrico, R.; Seixas, S.; Carvalho, J.; Senz, J.; Oliveira, P.; Inacio, P.; Gusmao, L.; Rocha, J.; Huntsman, D.; et al. Allele-specific CDH1 downregulation and hereditary diffuse gastric cancer. Hum. Mol. Genet. 2010, 19, 943–952. [Google Scholar] [CrossRef] [PubMed]
- Repetto, O.; De Paoli, P.; De Re, V.; Canzonieri, V.; Cannizzaro, R. Levels of Soluble E-Cadherin in Breast, Gastric, and Colorectal Cancers. Available online: https://www.hindawi.com/journals/bmri/2014/408047/ (accessed on 17 November 2017).
- D’souza, B.; Taylor-Papadimitriou, J. Overexpression of ERBB2 in human mammary epithelial cells signals inhibition of transcription of the E-cadherin gene. Proc. Natl. Acad. Sci. USA 1994, 91, 7202–7206. [Google Scholar] [CrossRef] [PubMed]
- Liu, J.; Chen, X.; Mao, Y.; Qu, Q.; Shen, K. Association of epithelial-mesenchymal transition with lapatinib resistance through multipe pathways activation in HER2-positive breast cancer. J. Clin. Oncol. 2014, 32, e11579. [Google Scholar] [CrossRef]
- Cheng, J.-C.; Qiu, X.; Chang, H.-M.; Leung, P.C.K. HER2 mediates epidermal growth factor-induced down-regulation of E-cadherin in human ovarian cancer cells. Biochem. Biophys. Res. Commun. 2013, 434, 81–86. [Google Scholar] [CrossRef] [PubMed]
- Garziera, M.; Canzonieri, V.; Cannizzaro, R.; Geremia, S.; Caggiari, L.; Zorzi, M.D.; Maiero, S.; Orzes, E.; Perin, T.; Zanussi, S.; et al. Identification and characterization of CDH1 germline variants in sporadic gastric cancer patients and in individuals at risk of gastric cancer. PLoS ONE 2013, 8, e77035. [Google Scholar] [CrossRef] [PubMed]
- Morcillo-Suarez, C.; Alegre, J.; Sangros, R.; Gazave, E.; de Cid, R.; Milne, R.; Amigo, J.; Ferrer-Admetlla, A.; Moreno-Estrada, A.; Gardner, M.; et al. SNP analysis to results (SNPator): A web-based environment oriented to statistical genomics analyses upon SNP data. Bioinformatics 2008, 24, 1643–1644. [Google Scholar] [CrossRef] [PubMed]
- Excoffier, L.; Laval, G.; Schneider, S. Arlequin (version 3.0): An integrated software package for population genetics data analysis. Evol. Bioinform. Online 2007, 1, 47–50. [Google Scholar] [CrossRef] [PubMed]
| CDH1 Region Gene | Reference Polymorphism | cDNA Change | Amino Acid Change | Type of Variant | Genotype | |||
|---|---|---|---|---|---|---|---|---|
| mGC-HER2 (n) | mGC (n) | |||||||
| Promoter | rs5030625 | c.-472delA | Polymorphic variant | G/A (1) | G/A (10) | |||
| Promoter | rs16260 | c.-285C>A | Polymorphic variant | A/A (4) | A/C (6) | A/A (3) | A/C (16) | |
| Promoter | rs34149581 | c.-276T>C | T/C (1) | |||||
| 5′UTR | rs34033771 | c.-71C>G | C/G (1) | |||||
| IV1 | rs3743674 | c.48+6C>T | Polymorphic variant | C/C (1) | T/C (9) | C/C (1) | ||
| EXON3 | rs1801023 | c.345G>A | p.Thr115= | Synonymous variant | G/A (1) | |||
| IV4 | rs33963999 | c.531+10G>C | G/C (2) | |||||
| IV5 | rs189969617 | c.688-14C>T | C/T (1) | |||||
| EXON7 | rs142822590 | c.892G>A | p.Ala298Thr | Missense variant | G/A (1) | |||
| EXON11 | SCV000588228.1 | c.1612delG | p.Asp538Thrfs*19 | Frameshift mutation | delG (1) | |||
| EXON12 | rs35187787 | c.1774G>A | p.Ala592Thr | Missense variant | G/A (1) | |||
| EXON12 | rs33969373 | c.1896C>T | p.HIS632= | Synonymous variant | C/T (2) | |||
| IV12 | rs2276330 | c.1937-13T>C | Polymorphic variant | C/T (2) | C/T (10) | |||
| EXON13 | rs1801552 | c.2076T>C | p.Ala692= | Polymorphic synonymous variant | C/T (5) | T/T (3) | C/T (24) | T/T (5) |
| IV13 | rs35686369 | c.2164+15_2164+16insA | insA (1) | insA (2) | ||||
| EXON14 | rs879026401 | c.2232A>G | p.Pro744= | Synonymous variant | A/G (1) | |||
| EXON14 | rs33964119 | c.2253C>T | p.Asn751= | Synonymous variant | C/T (1) | C/T (2) | ||
| EXON15 | rs587782549 | c.2204G>A | p.Arg796Gln | Missense variant | G/A (1) | |||
| Reference Polymorphism | Allele/Genotype | mGC-HER2 | Frequency | mGC | Frequency | p | OR (95% CI) |
|---|---|---|---|---|---|---|---|
| rs5030625 | |||||||
| Allele | G | 23 | 0.96 | 84 | 0.89 | 0.33 | 2.738 (0.33–22.51) |
| A | 1 | 0.04 | 10 | 0.11 | |||
| Genotype | G/G | 11 | 0.92 | 37 | 0.79 | ||
| G/A | 1 | 0.08 | 10 | 0.21 | |||
| A/A | 0 | 0.00 | 0 | 0.00 | |||
| Dominant model | GG/AA+AG | 11/1 | 0.92/0.08 | 37/10 | 0.79/0.21 | 0.30 | 2.973 (0.34–25.86) |
| Recessive model | AA/AG+GG | 0/12 | 0.00/1.00 | 0/47 | 0.00/1.00 | nv | |
| rs16260 | |||||||
| Allele | A | 14 | 0.58 | 22 | 0.23 | ≤0.001 | 4.582 (1.79–11.75) |
| C | 10 | 0.42 | 72 | 0.77 | |||
| Genotype | A/A | 4 | 0.33 | 3 | 0.06 | ||
| A/C | 6 | 0.50 | 16 | 0.34 | |||
| C/C | 2 | 0.17 | 28 | 0.60 | |||
| Recessive model | CC/AA+AC | 2/10 | 0.17/0.83 | 28/19 | 0.60/0.40 | ≤0.01 | 7.368 (1.45–37.46) |
| Dominant model | AA/AC+CC | 4/8 | 0.33/0.67 | 3/44 | 0.06/0.94 | 0.01 | 7.333 (1.37–39.18) |
| rs3743674 | |||||||
| Allele | T | 22 | 0.92 | 83 | 0.88 | 0.64 | 1.457 (0.30–7.07) |
| C | 2 | 0.08 | 11 | 0.12 | |||
| Genotype | T/T | 11 | 0.92 | 37 | 0.79 | ||
| T/C | 0 | 0.00 | 9 | 0.19 | |||
| C/C | 1 | 0.08 | 1 | 0.02 | |||
| Recessive model | CC/CT+TT | 1/11 | 0.08/0.92 | 1/46 | 0.02/0.98 | 0.29 | 4.182 (0.24–72.21) |
| Dominant model | TT/CC+CT | 11/1 | 0.92/0.08 | 37/10 | 0.79/0.21 | 0.30 | 2.973 (0.34–25.86) |
| rs2276330 | |||||||
| Allele | T | 22 | 0.92 | 84 | 0.90 | 0.74 | 1.309 (0.27–6.42) |
| C | 2 | 0.08 | 10 | 0.11 | |||
| Genotype | T/T | 10 | 0.83 | 37 | 0.79 | ||
| T/C | 2 | 0.17 | 10 | 0.21 | |||
| C/C | 0 | 0.00 | 0 | 0.00 | |||
| Dominant model | TT/CT+CC | 10/2 | 0.83/0.17 | 37/10 | 0.79/0.21 | 0.72 | 1.351 (0.25–7.19) |
| Recessive model | CC/TT+CT | 0/12 | 0.00/1.00 | 0/47 | 0.00/1.00 | nv | |
| rs1801552 | |||||||
| Allele | C | 13 | 0.54 | 60 | 0.64 | 0.39 | 0.670 (0.27–1.66) |
| T | 11 | 0.46 | 34 | 0.36 | |||
| C/C | 4 | 0.33 | 18 | 0.38 | |||
| Genotype | T/C | 5 | 0.42 | 24 | 0.51 | ||
| T/T | 3 | 0.25 | 5 | 0.11 | |||
| Recessive model | TT/CC+CT | 3/9 | 0.25/0.75 | 5/42 | 0.11/0.89 | 0.19 | 2.800 (0.56–13.90) |
| Dominant model | CC/CT+TT | 4/8 | 0.33/0.67 | 18/29 | 0.38/0.62 | 0.75 | 1.241 (0.33–4.72) |
| rs33964119 | |||||||
| C | 23 | 0.96 | 92 | 0.98 | 0.58 | 0.500 (0.04–5.76) | |
| Allele | T | 1 | 0.04 | 2 | 0.02 | ||
| C/C | 11 | 0.92 | 45 | 0.96 | |||
| Genotype | T/C | 1 | 0.08 | 2 | 0.04 | ||
| T/T | 0 | 0.00 | 0 | 0.00 | |||
| Recessive model | CC/CT+TT | 11/1 | 0.92/0.08 | 45/2 | 0.96/0.04 | 0.57 | 2.045 (0.17–24.66) |
| Dominant model | TT/CC+CT | 0/12 | 0.00/1.00 | 0/47 | 0.00/1.00 | nv | |
| Haplotype | mGC (N 94) | Frequency | mGC-HER2 (N 24) | Frequency | p | OR (95% CI) | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| rs5030625 | rs16260 | rs3743674 | rs2276330 | rs1801552 | rs33964119 | |||||||
| P1 | G | C | T | T | C | C | 19 | 0.20 | 2 | 0.08 | 0.24 | 0.359 (0.08–1.67) |
| P2 | G | A | T | T | C | C | 22 | 0.23 | 6 | 0.25 | 1.00 | 1.091 (0.39–3.09) |
| P3 | G | C | T | T | T | C | 31 | 0.33 | 4 | 0.17 | 0.14 | 0.406 (0.13–1.29) |
| P4 | G | C | T | C | C | C | 9 | 0.09 | 2 | 0.08 | 1.00 | 0.859 (0.17–4.26) |
| P5 | A | C | C | T | C | C | 7 | 0.07 | 1 | 0.04 | 0.69 | 0.540 (0.06–4.61) |
| P6 | A | C | C | C | C | C | 1 | 0.01 | 0 | 0.00 | 1.00 | |
| P7 | G | A | T | T | T | C | 0 | 0.00 | 7 | 0.29 | ≤0.001 | |
| P8 | G | C | C | T | T | C | 1 | 0.01 | 0 | 0.00 | 1.00 | |
| P9 | A | C | C | T | T | C | 2 | 0.02 | 0 | 0.00 | 1.00 | |
| P10 | G | C | T | T | C | T | 2 | 0.02 | 1 | 0.04 | 0.50 | 2.00 (0.17–23.03) |
| P11 | G | A | C | T | C | C | 0 | 0.00 | 1 | 0.04 | 0.20 | |
| Patient Identifier | Haplotype | CDH1 Germline Mutation | |||
|---|---|---|---|---|---|
| EXON11 c.1612delG | EXON12 c.1774G>A | IV13 c.2164+15_2164+16insA | EXON14 c.2253C>T | ||
| P287 | P5–P11 | ||||
| P291 | P2–P7 | ||||
| P292 | P4–P7 | ||||
| P296 | P2–P7 | ||||
| P297 | P3–P7 | ||||
| P301 | P2–P3 | ||||
| P303 | P2–P4 | ||||
| P380 | P1–P2 | ||||
| P391 | P2–P7 | ||||
| P486 | P7–P7 | ||||
| P582 | P3–P3 | ||||
| P586 | P1–P10 | ||||
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Caggiari, L.; Miolo, G.; Buonadonna, A.; Basile, D.; Santeufemia, D.A.; Cossu, A.; Palmieri, G.; De Zorzi, M.; Fornasarig, M.; Alessandrini, L.; et al. Characterizing Metastatic HER2-Positive Gastric Cancer at the CDH1 Haplotype. Int. J. Mol. Sci. 2018, 19, 47. https://doi.org/10.3390/ijms19010047
Caggiari L, Miolo G, Buonadonna A, Basile D, Santeufemia DA, Cossu A, Palmieri G, De Zorzi M, Fornasarig M, Alessandrini L, et al. Characterizing Metastatic HER2-Positive Gastric Cancer at the CDH1 Haplotype. International Journal of Molecular Sciences. 2018; 19(1):47. https://doi.org/10.3390/ijms19010047
Chicago/Turabian StyleCaggiari, Laura, Gianmaria Miolo, Angela Buonadonna, Debora Basile, Davide A. Santeufemia, Antonio Cossu, Giuseppe Palmieri, Mariangela De Zorzi, Mara Fornasarig, Lara Alessandrini, and et al. 2018. "Characterizing Metastatic HER2-Positive Gastric Cancer at the CDH1 Haplotype" International Journal of Molecular Sciences 19, no. 1: 47. https://doi.org/10.3390/ijms19010047