Transcriptomics Evidence for Common Pathways in Human Major Depressive Disorder and Glioblastoma
Abstract
:1. Introduction
2. Results
2.1. The Common Co-Occurrence Gene by Text-Mining
2.2. Differentially Expressed Genes
2.3. Functional Enrichment of DEGs
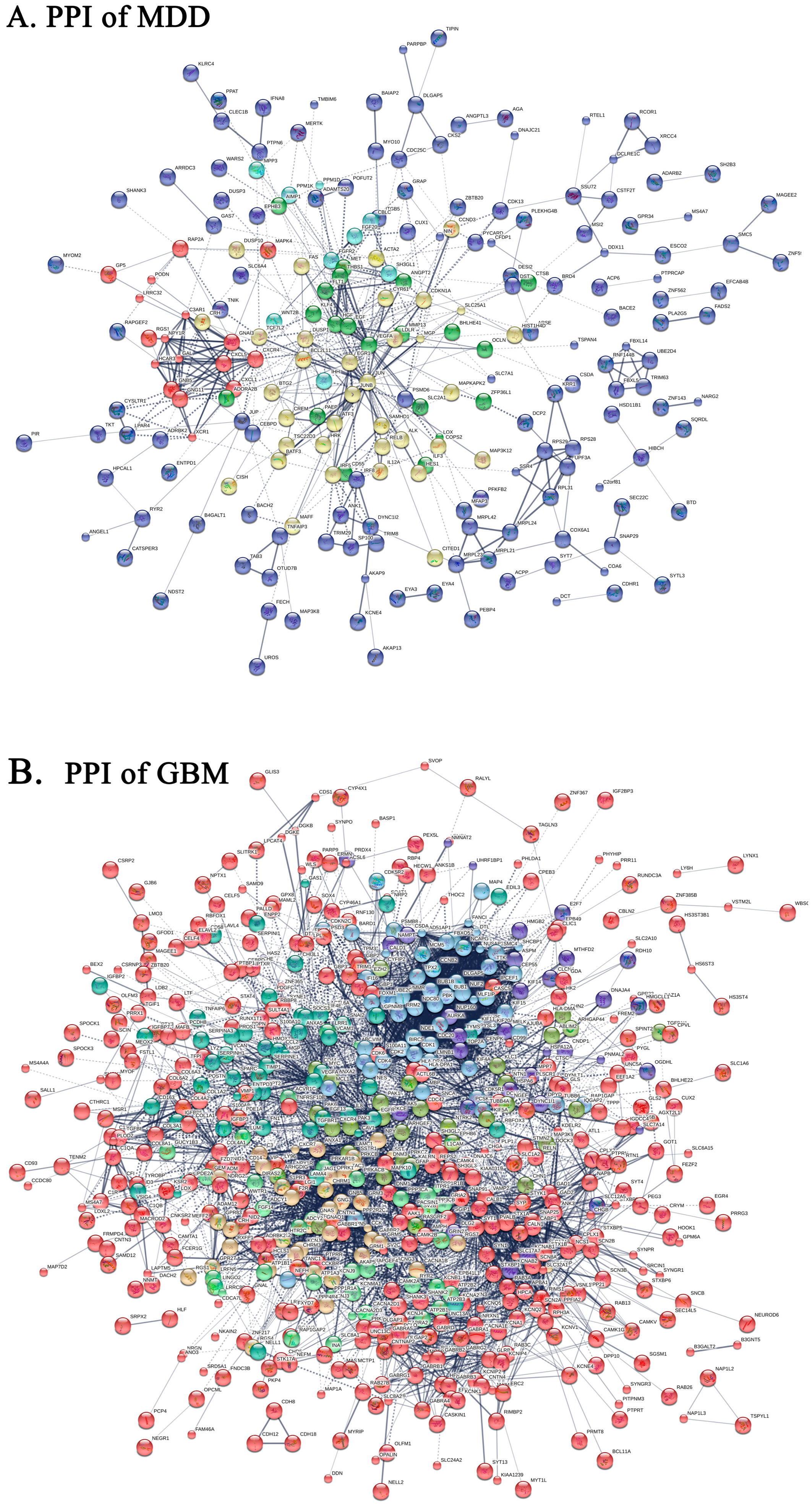
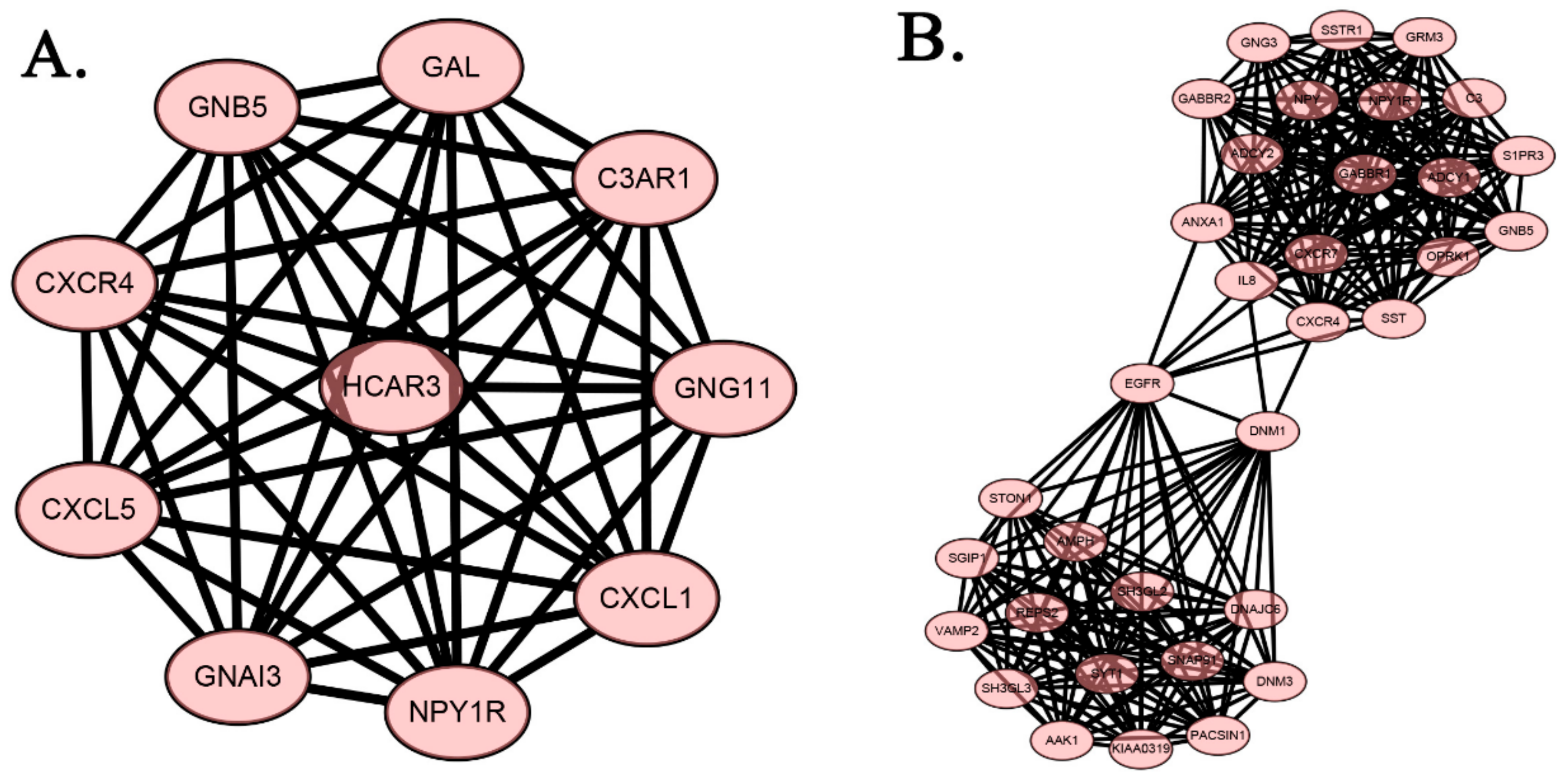
2.4. Protein-Protein Interaction Network of DEGs
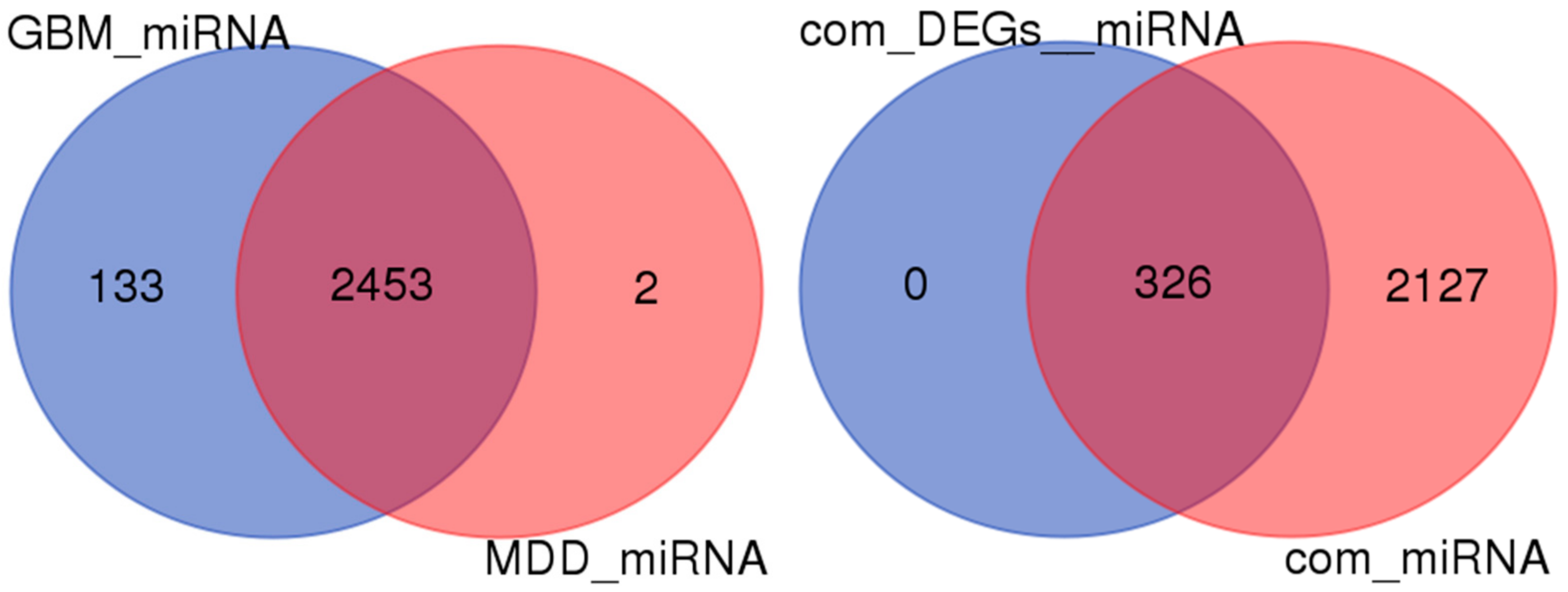
2.5. miRNA
2.6. Functional Enrichment of Common DEGs and Common miRNA
3. Discussion
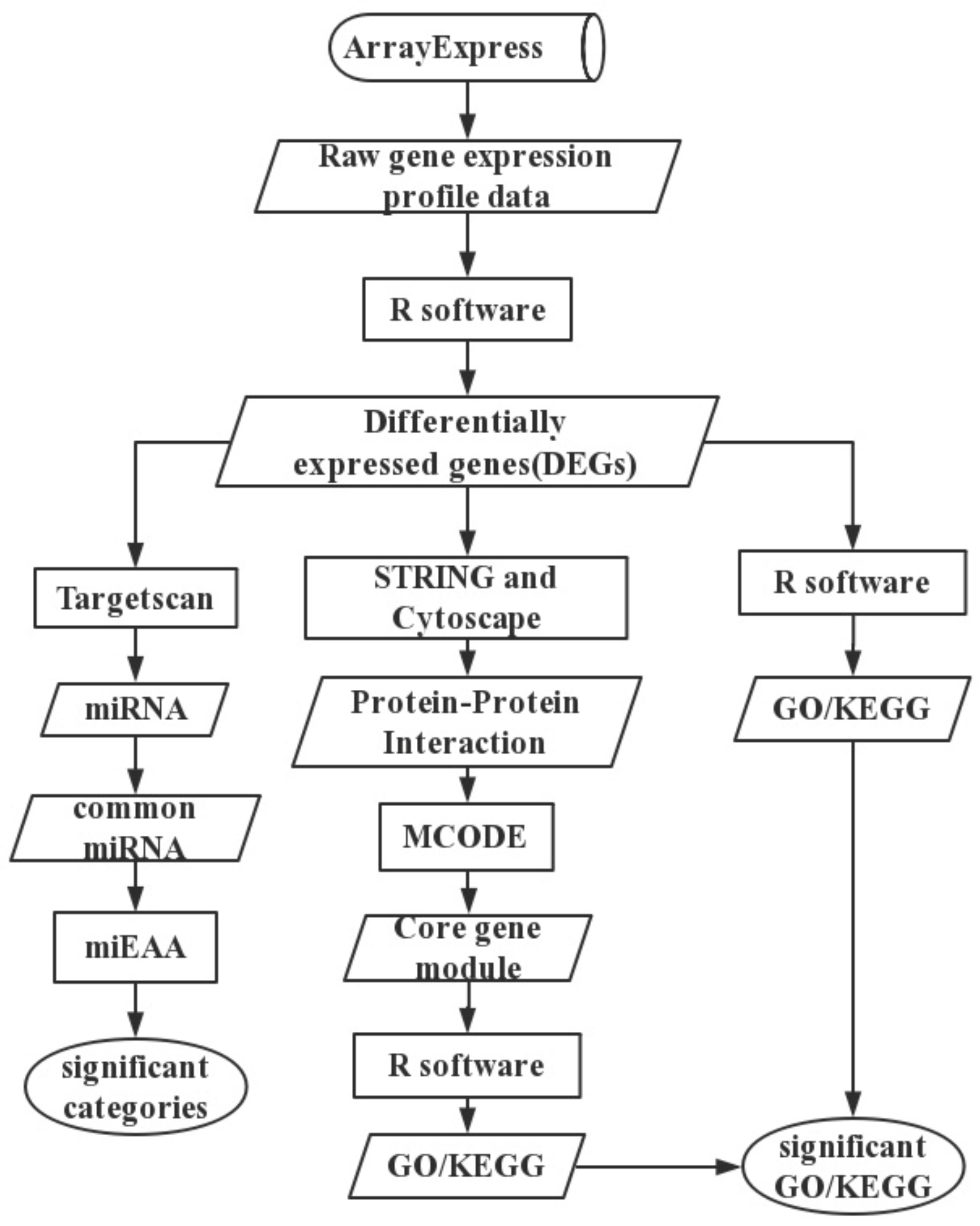
4. Materials and Methods
4.1. Text Mining
4.2. Data and Data Preprocessing
4.3. Differential Expression Analysis
4.4. Functional Enrichment Analysis
4.5. Protein—Protein Interaction Network
4.6. Predicted miRNAs
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Rooney, A.G.; Carson, A.; Grant, R. Depression in cerebral glioma patients: A systematic review of observational studies. J. Natl. Cancer Inst. 2011, 103, 61–76. [Google Scholar] [CrossRef] [PubMed]
- Rooney, A.G.; Brown, P.D.; Reijneveld, J.C.; Grant, R. Depression in glioma: A primer for clinicians and researchers. J. Neurol. Neurosurg. Psychiatry 2014, 85, 230–235. [Google Scholar] [CrossRef] [PubMed]
- Pranckeviciene, A.; Bunevicius, A. Depression screening in patients with brain tumors: A review. CNS Oncol. 2015, 4, 71–78. [Google Scholar] [CrossRef] [PubMed]
- Seddighi, A.; Seddighi, A.S.; Nikouei, A.; Ashrafi, F.; Nohesara, S. Psychological aspects in brain tumor patients: A prospective study. Hell. J. Nucl. Med. 2015, 18 (Suppl. 1), 63–67. [Google Scholar] [PubMed]
- Sah, D.W. Therapeutic potential of rna interference for neurological disorders. Life Sci. 2006, 79, 1773–1780. [Google Scholar] [CrossRef] [PubMed]
- Gautier, L.; Cope, L.; Bolstad, B.M.; Irizarry, R.A. Affy-analysis of affymetrix genechip data at the probe level. Bioinformatics 2004, 20, 307–315. [Google Scholar] [CrossRef] [PubMed]
- Falcon, S.; Gentleman, R. Using gostats to test gene lists for go term association. Bioinformatics 2007, 23, 257–258. [Google Scholar] [CrossRef] [PubMed]
- Ritchie, M.E.; Phipson, B.; Wu, D.; Hu, Y.; Law, C.W.; Shi, W.; Smyth, G.K. Limma powers differential expression analyses for rna-sequencing and microarray studies. Nucleic Acids Res. 2015, 43, e47. [Google Scholar] [CrossRef] [PubMed]
- Carlson, M. KEGG.db: A Set of Annotation Maps for KEGG. Available online: https://bioconductor.org/packages/release/data/annotation/html/KEGG.db.html (accessed on 9 January 2018).
- Franceschini, A.; Szklarczyk, D.; Frankild, S.; Kuhn, M.; Simonovic, M.; Roth, A.; Lin, J.; Minguez, P.; Bork, P.; von Mering, C.; et al. String v9.1: Protein—protein interaction networks, with increased coverage and integration. Nucleic Acids Res. 2013, 41, D808–D815. [Google Scholar] [CrossRef] [PubMed]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef] [PubMed]
- Bader, G.D.; Hogue, C.W. An automated method for finding molecular complexes in large protein interaction networks. BMC Bioinform. 2003, 4, 2. [Google Scholar] [CrossRef] [Green Version]
- Agarwal, V.; Bell, G.W.; Nam, J.W.; Bartel, D.P. Predicting effective microrna target sites in mammalian mrnas. eLife 2015, 4, 1–38. [Google Scholar] [CrossRef] [PubMed]
- Backes, C.; Khaleeq, Q.T.; Meese, E.; Keller, A. Mieaa: Microrna enrichment analysis and annotation. Nucleic Acids Res. 2016, 44, W110–W116. [Google Scholar] [CrossRef] [PubMed]
- Da Silva, R.; Uno, M.; Marie, S.K.; Oba-Shinjo, S.M. Lox expression and functional analysis in astrocytomas and impact of idh1 mutation. PLoS ONE 2015, 10, e0119781. [Google Scholar] [CrossRef] [PubMed]
- Sun, W.; Park, K.W.; Choe, J.; Rhyu, I.J.; Kim, I.H.; Park, S.K.; Choi, B.; Choi, S.H.; Park, S.H.; Kim, H. Identification of novel electroconvulsive shock-induced and activity-dependent genes in the rat brain. Biochem. Biophys. Res. Commun. 2005, 327, 848–856. [Google Scholar] [CrossRef] [PubMed]
- Berent, D.; Macander, M.; Szemraj, J.; Orzechowska, A.; Galecki, P. Vascular endothelial growth factor a gene expression level is higher in patients with major depressive disorder and not affected by cigarette smoking, hyperlipidemia or treatment with statins. Acta Neurobiol. Exp. 2014, 74, 82–90. [Google Scholar]
- Stefano, A.L.D.; Labussiere, M.; Lombardi, G.; Eoli, M.; Bianchessi, D.; Pasqualetti, F.; Farina, P.; Cuzzubbo, S.; Gallego-Perez-Larraya, J.; Boisselier, B. Vegfa snp rs2010963 is associated with vascular toxicity in recurrent glioblastomas and longer response to bevacizumab. J. Neurooncol. 2015, 121, 499–504. [Google Scholar] [CrossRef] [PubMed]
- Lopez-Gines, C.; Cerda-Nicolas, M.; Gil-Benso, R.; Pellin, A.; Lopez-Guerrero, J.A.; Callaghan, R.; Benito, R.; Roldan, P.; Piquer, J.; Llacer, J.; et al. Association of chromosome 7, chromosome 10 and egfr gene amplification in glioblastoma multiforme. Clin. Neuropathol. 2005, 24, 209–218. [Google Scholar] [PubMed]
- Chen, X.; Long, F.; Cai, B.; Chen, X.; Chen, G. A novel relationship for schizophrenia, bipolar and major depressive disorder part 7: A hint from chromosome 7 high density association screen. Behav. Brain Res. 2015, 293, 241–251. [Google Scholar] [CrossRef] [PubMed]
- Fujioka, Y.; Tsuda, M.; Hattori, T.; Sasaki, J.; Sasaki, T.; Miyazaki, T.; Ohba, Y. The ras–pi3k signaling pathway is involved in clathrin-independent endocytosis and the internalization of influenza viruses. PLoS ONE 2011, 6, e16324. [Google Scholar] [CrossRef] [PubMed]
- Li, W.; Li, K.; Zhao, L.; Zou, H. Bioinformatics analysis reveals disturbance mechanism of mapk signaling pathway and cell cycle in glioblastoma multiforme. Gene 2014, 547, 346–350. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Peng, D.; Xie, B.; Jiang, K.; Fang, Y. The extracellular signal-regulated kinase pathway may play an important role in mediating antidepressant-stimulated hippocampus neurogenesis in depression. Med. Hypotheses 2012, 79, 87–91. [Google Scholar] [CrossRef] [PubMed]
- Møller, H.G.; Rasmussen, A.P.; Andersen, H.H.; Johnsen, K.B.; Henriksen, M.; Duroux, M. A systematic review of microrna in glioblastoma multiforme: Micro-modulators in the mesenchymal mode of migration and invasion. Mol. Neurobiol. 2013, 47, 131–144. [Google Scholar] [CrossRef] [PubMed]
- Dwivedi, Y. Micrornas as biomarker in depression pathogenesis. Ann. Psychiatry Ment. Health 2013, 1, 1003. [Google Scholar] [PubMed]
- Miguel-Hidalgo, J.J.; Hall, K.O.; Bonner, H.; Roller, A.M.; Syed, M.; Park, C.J.; Ball, J.P.; Rothenberg, M.E.; Stockmeier, C.A.; Romero, D.G. Microrna-21: Expression in oligodendrocytes and correlation with low myelin mrnas in depression and alcoholism. Prog. Neuropsychopharmacol. Biol. Psychiatry 2017, 79, 503–514. [Google Scholar] [CrossRef] [PubMed]
- Masoudi, M.S.; Mehrabian, E.; Mirzaei, H. Mir-21: A key player in glioblastoma pathogenesis. J. Cell. Biochem. 2017. [Google Scholar] [CrossRef] [PubMed]
- Irshad, K.; Mohapatra, S.K.; Srivastava, C.; Garg, H.; Mishra, S.; Dikshit, B.; Sarkar, C.; Gupta, D.; Chandra, P.S.; Chattopadhyay, P. A combined gene signature of hypoxia and notch pathway in human glioblastoma and its prognostic relevance. PLoS ONE 2015, 10, e0118201. [Google Scholar] [CrossRef] [PubMed]
- Ning, S.; Lei, L.; Wang, Y.; Yang, C.; Liu, Z.; Li, X.; Zhang, K. Preliminary comparison of plasma notch-associated microRNA-34b and -34c levels in drug naive, first episode depressed patients and healthy controls. J. Affect. Disord. 2016, 194, 109–114. [Google Scholar]
- Salvadore, G.; Veen, J.W.V.D.; Zhang, Y.; Marenco, S.; Machadovieira, R.; Baumann, J.; Ibrahim, L.A.; Luckenbaugh, D.A.; Shen, J.; Drevets, W.C. An investigation of amino-acid neurotransmitters as potential predictors of clinical improvement to ketamine in depression. Int. J. Neuropsychopharmacol. 2012, 15, 1063–1072. [Google Scholar] [CrossRef] [PubMed]
- Panosyan, E.H.; Lin, H.J.; Koster, J.; Lasky, J.L. In search of druggable targets for gbm amino acid metabolism. BMC Cancer 2017, 17, 162. [Google Scholar] [CrossRef] [PubMed]
- Wei, B.; Wang, L.; Zhao, X.; Jin, Y.; Kong, D.; Hu, G.; Sun, Z. Co-mutated pathways analysis highlights the coordination mechanism in glioblastoma multiforme. Neoplasma 2014, 61, 424–432. [Google Scholar] [CrossRef] [PubMed]
- Shintaro, O.; Hiroshi, K. Inhibitors of fatty acid amide hydrolase and monoacylglycerol lipase: New targets for future antidepressants. Curr. Neuropharmacol. 2015, 13, 760–775. [Google Scholar]
- Wu, W.J.; Wang, Q.; Zhang, W.; Li, L. Identification and prognostic value of differentially expressed proteins of patients with platinum resistance epithelial ovarian cancer in serum. Zhonghua Fu Chan Ke Za Zhi 2016, 51, 515–523. [Google Scholar] [PubMed]
- Brazma, A.; Parkinson, H.; Sarkans, U.; Shojatalab, M.; Vilo, J.; Abeygunawardena, N.; Holloway, E.; Kapushesky, M.; Kemmeren, P.; Lara, G.G. Arrayexpress—A public repository for microarray gene expression data at the ebi. Nucleic Acids Res. 2003, 31, 68–71. [Google Scholar] [CrossRef] [PubMed]
- Rice, A.M.; McLysaght, A. Dosage-sensitive genes in evolution and disease. BMC Biol. 2017, 15, 78. [Google Scholar] [CrossRef] [PubMed]
- Jeong, H.; Mason, S.P.; Barabasi, A.L.; Oltvai, Z.N. Lethality and centrality in protein networks. Nature 2001, 411, 41–42. [Google Scholar] [CrossRef] [PubMed]
- Hesse, M.; Arenz, C. Microrna maturation and human disease. Methods Mol. Biol. 2014, 1095, 11–25. [Google Scholar] [PubMed]

| Disease | Related Articles | Gene/Protein | Chemical | CC | BP | MF |
|---|---|---|---|---|---|---|
| MDD | 34377 | 1826 | 3511 | 110 | 498 | 104 |
| GBM | 30193 | 4510 | 7779 | 229 | 834 | 244 |
| GBM ∩ MDD | 4 | 1041 | 2248 | 78 | 317 | 52 |
| GBM | MDD |
|---|---|
| HS6ST3; ZNF385B; VSTM2L; EGFR; VEGFA; TOP2A; CDC42; MYC; IL8; FN1; PRKACB; CD44; CDK1; VIM; GFAP; SYP; PPP3CA; SNCA; STX1A; DNM1; GNAO1; CACNA1B; LPL; PCSK2; PRKCB; SYT1; SNAP25; TUBB4A;DLG2; PVALB; CAMK2A; CALB1; CDK2; CAV1; CXCR4; GNB5; VAMP2; NPY; VCAM1; PRKCE; C3; EZH2; CDC20; SST; GAD2; ITPR1; ADCY2; LUM; TAC1; AURKA; CD163; SYN1; SPARC; BIRC5; GABBR2; ANXA1; MGP; GAD1; TYMS; GNG3; SCN2A; MCL1; CNKSR2; NDE1 | JUN; VEGFA; EGF; CXCR4; GNAI3; EGR1; FLT1; CDKN1A; ATF3; CXCL1; GNG11; GNB5; ACTA2; MET; FGFR2; SH3GL1; CXCL5; MGP; THBS1; FAS; IRF5; JUP; RAP2A; TCF7L2; MRPL23; TNFAIP3; CCND3; SLC25A1; MAPK4; BATF3; CD55; CDC25C; MPP3; PPM1D; ILF3; HIST1H4D; CDK13; SSU72; PTPN6; CREM; OCLN; ADORA2B; HIBCH; DYNC1I2; CTSB; MAFF; RYR2; DLGAP5; DCLRE1C; SSR4; ADRBK2; COPS2; COX6A1; LOX; SNAP29; BRD4; DDX11; KRR1; AKAP9; SMC5; ZFP36L1; AIMP1; CFDP1; GAS7; MYO10; GP5; SYT7; ESCO2; MSI2; CLEC1B; FECH; B4GALT1; TKT |
| MDD KEGG | GBM KEGG | Common KEGG |
|---|---|---|
| Alcoholism | Endocytosis | GABAergic synapse |
| Pertussis | Gap junction | Cholinergic synapse |
| Serotonergic synapse | Insulin secretion | Pathways in cancer |
| Dopaminergic synapse | Salivary secretion | Morphine addiction |
| Synaptic vesicle cycle | Circadian entrainment | |
| Gastric acid secretion | Glutamatergic synapse | |
| GnRH signaling pathway | cAMP signaling pathway | |
| Estrogen signaling pathway | Chemokine signaling pathway | |
| Oxytocin signaling pathway | Retrograde endocannabinoid signaling | |
| Neuroactive ligand-receptor interaction | Regulation of lipolysis in adipocytes |
| Category | Subcategory | p-Value | Observed |
|---|---|---|---|
| Pathways | WP411 mRNA processing | 0.060337 | 145 |
| Pathways | hsa00260 Glycine serine and threonine metabolism | 0.060337 | 30 |
| Pathways | hsa00562 Inositol phosphate metabolism | 0.060337 | 99 |
| Pathways | hsa03040 Spliceosome | 0.060337 | 138 |
| Pathways | hsa00330 Arginine and proline metabolism | 0.081737 | 67 |
| Pathways | hsa04330 Notch signaling pathway | 0.081737 | 94 |
| Pathways | P02756 N acetylglucosamine metabolism | 0.095531 | 14 |
| Immune cells | CD3 expressed | 0.008542 | 205 |
| Immune cells | CD19 expressed | 0.029347 | 182 |
| Immune cells | CD14 expressed | 0.040935 | 235 |
| Immune cells | CD56 expressed | 0.055024 | 252 |
| Chromosomal location | Chromosome 7 | 0.036298 | 102 |
| Gene Ontology | GO0042832 defense response to protozoan | 0.0626691 | 11 |
| Gene Ontology | GO0045859 regulation of protein kinase activity | 0.0626691 | 41 |
| Gene Ontology | GO0048304 positive regulation of isotype switching to igg isotypes | 0.0626691 | 11 |
| Gene Ontology | GO0016290 palmitoyl coa hydrolase activity | 0.0665877 | 24 |
| Gene Ontology | GO0044130 negative regulation of growth of symbiont in host | 0.0681013 | 19 |
| Gene Ontology | GO0004439 phosphatidylinositol 4 5 bisphosphate 5 phosphatase activity | 0.0753516 | 16 |
| Gene Ontology | GO0004523 ribonuclease h activity | 0.0753516 | 19 |
| Gene Ontology | GO0031848 protection from non homologous end joining at telomere | 0.0753516 | 16 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xie, Y.; Wang, L.; Xie, Z.; Zeng, C.; Shu, K. Transcriptomics Evidence for Common Pathways in Human Major Depressive Disorder and Glioblastoma. Int. J. Mol. Sci. 2018, 19, 234. https://doi.org/10.3390/ijms19010234
Xie Y, Wang L, Xie Z, Zeng C, Shu K. Transcriptomics Evidence for Common Pathways in Human Major Depressive Disorder and Glioblastoma. International Journal of Molecular Sciences. 2018; 19(1):234. https://doi.org/10.3390/ijms19010234
Chicago/Turabian StyleXie, Yongfang, Ling Wang, Zengyan Xie, Chuisheng Zeng, and Kunxian Shu. 2018. "Transcriptomics Evidence for Common Pathways in Human Major Depressive Disorder and Glioblastoma" International Journal of Molecular Sciences 19, no. 1: 234. https://doi.org/10.3390/ijms19010234





