Structure and Chromosomal Organization of Yeast Genes Regulated by Topoisomerase II
Abstract
:1. Introduction
2. Short Term Transcriptomic Changes after Topoisomerase II Inactivation
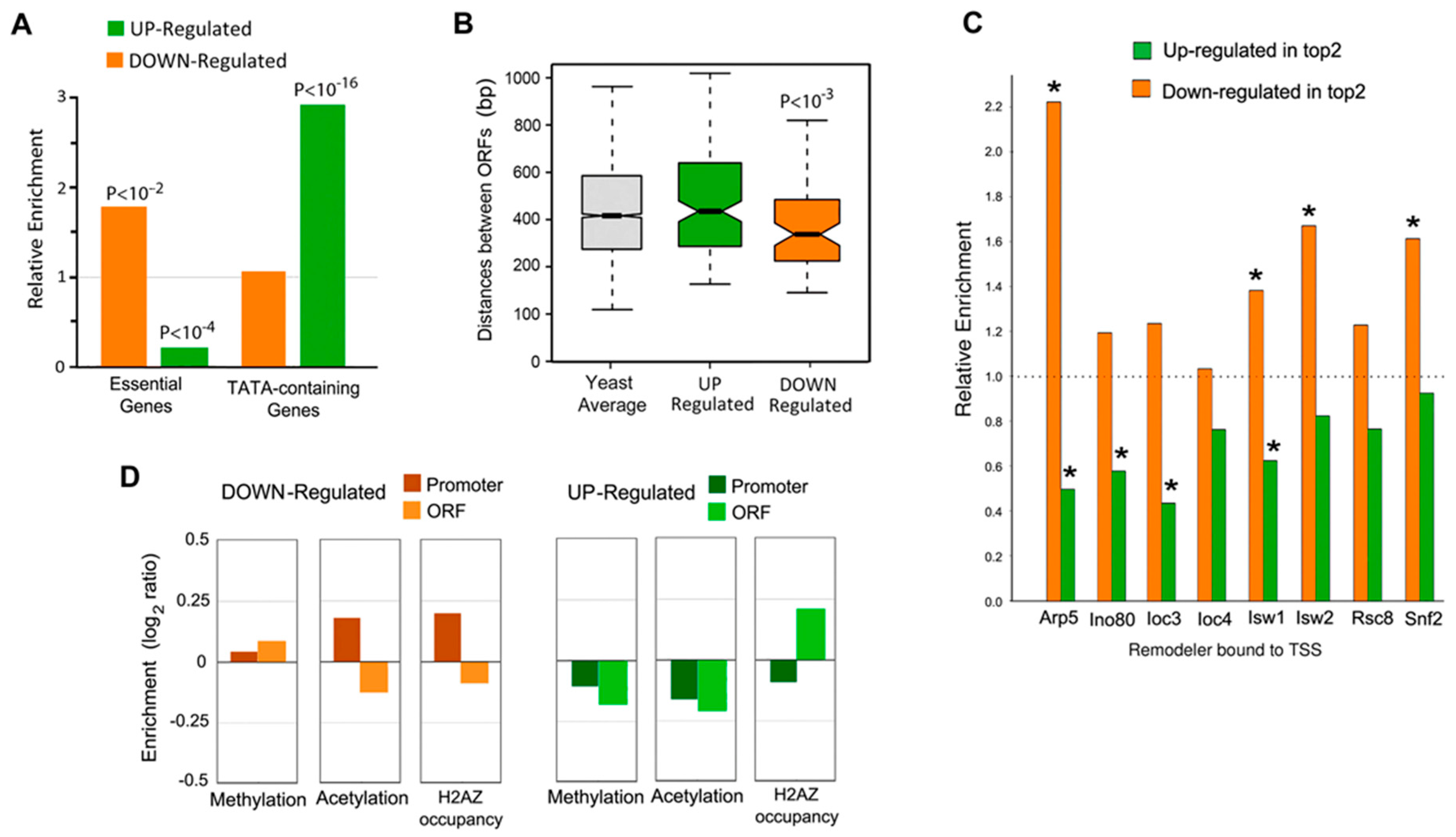
3. Analysis of the General Functional Properties of TOP2-Associated Genes
4. Chromatin Remodeling and Histone Modification Patterns Associated with TOP2 Deregulated Genes
5. The Role of Topo II beyond Single Gene Loci
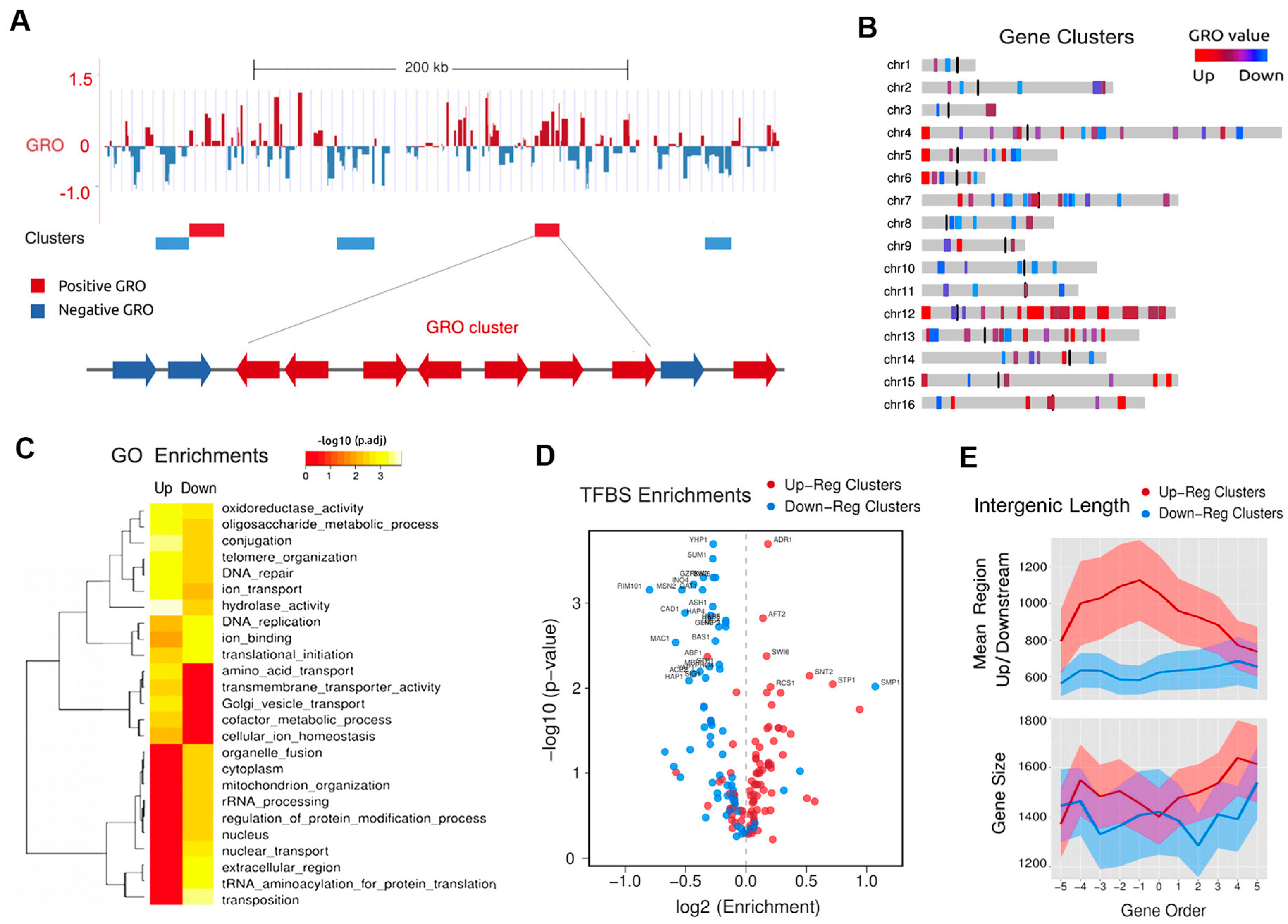
6. Topologically Co-Regulated Gene Clusters in Linear Chromosomes
7. Genic Spacing of TCGCs
8. Long Term Transcriptomic Changes after Topoisomerase II Inactivation
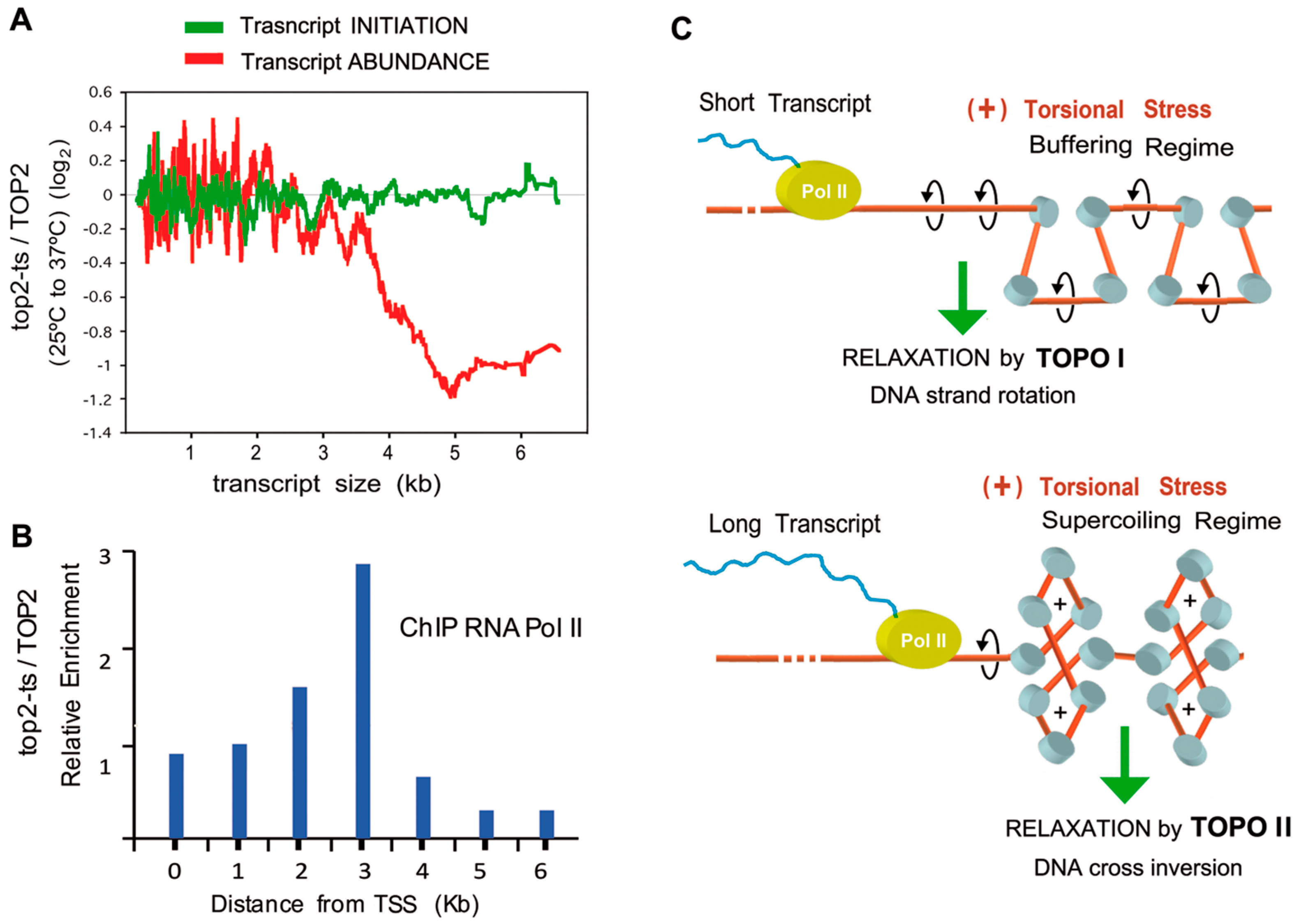
9. Long Term Inactivation of Topo II Diminishes Long Transcripts
10. Conclusions and Perspectives
Supplementary Materials
Supplementary File 1Acknowledgments
Conflicts of Interest
References
- Champoux, J.J. DNA topoisomerases: Structure, function, and mechanism. Annu. Rev. Biochem. 2001, 70, 369–413. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.C. Cellular roles of DNA topoisomerases: A molecular perspective. Nat. Rev. Mol. Cell Biol. 2002, 3, 430–440. [Google Scholar] [CrossRef] [PubMed]
- Pommier, Y.; Leo, E.; Zhang, H.; Marchand, C. DNA topoisomerases and their poisoning by anticancer and antibacterial drugs. Chem. Biol. 2010, 17, 421–433. [Google Scholar] [CrossRef] [PubMed]
- Stewart, L.; Redinbo, M.R.; Qiu, X.; Hol, W.G.; Champoux, J.J. A model for the mechanism of human topoisomerase I. Science 1998, 279, 1534–1541. [Google Scholar] [CrossRef] [PubMed]
- Wang, J.C. Moving one DNA double helix through another by a type II DNA topoisomerase: The story of a simple molecular machine. Q. Rev. Biophys. 1998, 31, 107–144. [Google Scholar] [CrossRef] [PubMed]
- Roca, J. Topoisomerase II: A fitted mechanism for the chromatin landscape. Nucleic Acids Res. 2009, 37, 721–730. [Google Scholar] [CrossRef] [PubMed]
- Nitiss, J.L. DNA topoisomerase II and its growing repertoire of biological functions. Nat. Rev. Cancer 2009, 9, 327–337. [Google Scholar] [CrossRef] [PubMed]
- Piskadlo, E.; Oliveira, R.A. A Topology-Centric View on Mitotic Chromosome Architecture. Int. J. Mol. Sci. 2017, 18, 2751. [Google Scholar] [CrossRef] [PubMed]
- Roca, J. The torsional state of DNA within the chromosome. Chromosoma 2011, 120, 323–334. [Google Scholar] [CrossRef] [PubMed]
- Salceda, J.; Fernández, X.; Roca, J. Topoisomerase II, not topoisomerase I, is the proficient relaxase of nucleosomal DNA. EMBO J. 2006, 25, 2575–2583. [Google Scholar] [CrossRef] [PubMed]
- Joshi, R.S.; Piña, B.; Roca, J. Topoisomerase II is required for the production of long Pol II gene transcripts in yeast. Nucleic Acids Res. 2012, 40, 7907–7915. [Google Scholar] [CrossRef] [PubMed]
- Parvin, J.D.; Sharp, P.A. DNA topology and a minimal set of basal factors for transcription by RNA polymerase II. Cell 1993, 73, 533–540. [Google Scholar] [CrossRef]
- Schultz, M.C.; Brill, S.J.; Ju, Q.; Sternglanz, R.; Reeder, R.H. Topoisomerases and yeast rRNA transcription: Negative supercoiling stimulates initiation and topoisomerase activity is required for elongation. Genes Dev. 1992, 6, 1332–1341. [Google Scholar] [CrossRef] [PubMed]
- Joshi, R.S.; Piña, B.; Roca, J. Positional dependence of transcriptional inhibition by DNA torsional stress in yeast chromosomes. EMBO J. 2010, 29, 740–748. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Roca, J. Transcriptional inhibition by DNA torsional stress. Transcription 2011, 2, 82–85. [Google Scholar] [CrossRef] [PubMed]
- Pedersen, J.M.; Fredsoe, J.; Roedgaard, M.; Andreasen, L.; Mundbjerg, K.; Kruhøffer, M.; Brinch, M.; Schierup, M.H.; Bjergbaek, L.; Andersen, A.H. DNA Topoisomerases Maintain Promoters in a State Competent for Transcriptional Activation in Saccharomyces cerevisiae. PLoS Genet. 2012, 8. [Google Scholar] [CrossRef] [PubMed]
- Sperling, A.S.; Jeong, K.S.; Kitada, T.; Grunstein, M. Topoisomerase II binds nucleosome-free DNA and acts redundantly with topoisomerase I to enhance recruitment of RNA Pol II in budding yeast. Proc. Natl. Acad. Sci. USA 2011, 108, 12693–12698. [Google Scholar] [CrossRef] [PubMed]
- Merino, A.; Madden, K.R.; Lane, W.S.; Champoux, J.J.; Reinberg, D. DNA topoisomerase I is involved in both repression and activation of transcription. Nature 1993, 365, 227–232. [Google Scholar] [CrossRef] [PubMed]
- Mondal, N.; Parvin, J.D. DNA topoisomerase IIalpha is required for RNA polymerase II transcription on chromatin templates. Nature 2001, 413, 435–438. [Google Scholar] [CrossRef] [PubMed]
- Ray, S.; Panova, T.; Miller, G.; Volkov, A.; Porter, A.C.; Russell, J.; Panov, K.I.; Zomerdijk, J.C. Topoisomerase IIα promotes activation of RNA polymerase I transcription by facilitating pre-initiation complex formation. Nat. Commun. 2013, 4, 1598. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bermejo, R.; Capra, T.; Gonzalez-Huici, V.; Fachinetti, D.; Cocito, A.; Natoli, G.; Katou, Y.; Mori, H.; Kurokawa, K.; Shirahige, K.; et al. Genome-Organizing Factors Top2 and Hmo1 Prevent Chromosome Fragility at Sites of S phase Transcription. Cell 2009, 138, 870–884. [Google Scholar] [CrossRef] [PubMed]
- Lotito, L.; Russo, A.; Chillemi, G.; Bueno, S.; Cavalieri, D.; Capranico, G. Global Transcription Regulation by DNA Topoisomerase I in Exponentially Growing Saccharomyces cerevisiae Cells: Activation of Telomere-Proximal Genes by TOP1 Deletion. J. Mol. Biol. 2008, 377, 311–322. [Google Scholar] [CrossRef] [PubMed]
- Durand-Dubief, M.; Persson, J.; Norman, U.; Hartsuiker, E.; Ekwall, K. Topoisomerase I regulates open chromatin and controls gene expression in vivo. EMBO J. 2010, 29, 2126–2134. [Google Scholar] [CrossRef] [PubMed]
- Dykhuizen, E.C.; Hargreaves, D.C.; Miller, E.L.; Cui, K.; Korshunov, A.; Kool, M.; Pfister, S.; Cho, Y.J.; Zhao, K.; Crabtree, G.R. BAF complexes facilitate decatenation of DNA by topoisomerase IIα. Nature 2013, 497, 624–627. [Google Scholar] [CrossRef] [PubMed]
- Thakurela, S.; Garding, A.; Jung, J.; Schübeler, D.; Burger, L.; Tiwari, V.K. Gene regulation and priming by topoisomerase IIα in embryonic stem cells. Nat. Commun. 2013, 4, 2478. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Li, W.; Prescott, E.D.; Burden, S.J.; Wang, J.C. DNA topoisomerase IIbeta and neural development. Science 2000, 287, 131–134. [Google Scholar] [CrossRef] [PubMed]
- Lyu, Y.L.; Wang, J.C. Aberrant lamination in the cerebral cortex of mouse embryos lacking DNA topoisomerase IIbeta. Proc. Natl. Acad. Sci. USA 2003, 100, 7123–7138. [Google Scholar] [CrossRef] [PubMed]
- Nevin, L.M.; Xiao, T.; Staub, W.; Baier, H. Topoisomerase IIbeta is required for lamina-specific targeting of retinal ganglion cell axons and dendrites. Development 2011, 138, 2457–2465. [Google Scholar] [CrossRef] [PubMed]
- Manville, C.M.; Smith, K.; Sondka, Z.; Rance, H.; Cockell, S.; Cowell, I.G.; Lee, K.C.; Morris, N.J.; Padget, K.; Jackson, G.H.; et al. Genome-wide ChIP-seq analysis of human TOP2B occupancy in MCF7 breast cancer epithelial cells. Biol. Open 2015, 4, 1436–1447. [Google Scholar] [CrossRef] [PubMed]
- Ju, B.G.; Lunyak, V.V.; Perissi, V.; Garcia-Bassets, I.; Rose, D.W.; Glass, C.K.; Rosenfeld, M.G. A topoisomerase IIbeta-mediated dsDNA break required for regulated transcription. Science 2006, 312, 1798–1802. [Google Scholar] [CrossRef] [PubMed]
- Haffner, M.C.; Aryee, M.J.; Toubaji, A.; Esopi, D.M.; Albadine, R.; Gurel, B.; Isaacs, W.B.; Bova, G.S.; Liu, W.; Xu, J.; et al. Androgen-induced TOP2B-mediated double-strand breaks and prostate cancer gene rearrangements. Nat Genet. 2010, 42, 668–675. [Google Scholar] [CrossRef] [PubMed]
- Madabhushi, R.; Gao, F.; Pfenning, A.R.; Pan, L.; Yamakawa, S.; Seo, J.; Rueda, R.; Phan, T.X.; Yamakawa, H.; Pao, P.C.; et al. Activity-Induced DNA Breaks Govern the Expression of Neuronal Early-Response Genes. Cell 2015, 161, 1592–1605. [Google Scholar] [CrossRef] [PubMed]
- Uusküla-Reimand, L.; Hou, H.; Samavarchi-Tehrani, P.; Rudan, M.V.; Liang, M.; Medina-Rivera, A.; Mohammed, H.; Schmidt, D.; Schwalie, P.; Young, E.J.; et al. Topoisomerase II beta interacts with cohesin and CTCF at topological domain borders. Genome Biol. 2016, 17. [Google Scholar] [CrossRef]
- Goffeau, A.; Barrell, B.G.; Bussey, H.; Davis, R.W.; Dujon, B.; Feldmann, H.; Galibert, F.; Hoheisel, J.D.; Jacq, C.; Johnston, M.; et al. Life with 6000 Genes. Science 1996, 274, 546–567. [Google Scholar] [CrossRef] [PubMed]
- David, L.; Huber, W.; Granovskaia, M.; Toedling, J.; Palm, C.J.; Bofkin, L.; Jones, T.; Davis, R.W.; Steinmetz, L.M. A high-resolution map of transcription in the yeast genome. Proc. Natl. Acad. Sci. USA 2006, 103, 5320–5325. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Wei, W.; Gagneur, J.; Perocchi, F.; Clauder-Münster, S.; Camblong, J.; Guffanti, E.; Stutz, F.; Huber, W.; Steinmetz, L.M. Bidirectional promoters generate pervasive transcription in yeast. Nature 2009, 457, 1033–1037. [Google Scholar] [CrossRef] [PubMed]
- Tan-Wong, S.M.; Zaugg, J.B.; Camblong, J.; Xu, Z.; Zhang, D.W.; Mischo, H.E.; Ansari, A.Z.; Luscombe, N.M.; Steinmetz, L.M.; Proudfoot, N.J. Gene Loops Enhance Transcriptional Directionality. Science 2012, 338, 671–675. [Google Scholar] [CrossRef] [PubMed]
- Nikolaou, C.; Bermúdez, I.; Manichanh, C.; García-Martinez, J.; Guigó, R.; Pérez-Ortín, J.E.; Roca, J. Topoisomerase II regulates yeast genes with singular chromatin architectures. Nucleic Acids Res. 2013, 41, 9243–9256. [Google Scholar] [CrossRef] [PubMed]
- Tsochatzidou, M.; Malliarou, M.; Papanikolaou, N.; Roca, J.; Nikolaou, C. Genome urbanization: Clusters of topologically co-regulated genes delineate functional compartments in the genome of Saccharomyces cerevisiae. Nucleic Acids Res. 2017, 45, 5818–5828. [Google Scholar] [CrossRef] [PubMed]
- García-Martínez, J.; Aranda, A.; Pérez-Ortín, J.E. Genomic run-on evaluates transcription rates for all yeast genes and identifies gene regulatory mechanisms. Mol. Cell 2004, 15, 303–313. [Google Scholar] [CrossRef] [PubMed]
- Gasch, A.P.; Spellman, P.T.; Kao, C.M.; Carmel-Harel, O.; Eisen, M.B.; Storz, G.; Botstein, D.; Brown, P.O. Genomic Expression Programs in the Response of Yeast Cells to Environmental Changes. Mol. Biol. Cell 2000, 11, 4241–4257. [Google Scholar] [CrossRef] [PubMed]
- Giaever, G.; Chu, A.M.; Ni, L.; Connelly, C.; Riles, L.; Véronneau, S.; Dow, S.; Lucau-Danila, A.; Anderson, K.; André, B.; et al. Functional profiling of the Saccharomyces cerevisiae genome. Nature 2002, 418, 387–391. [Google Scholar] [CrossRef] [PubMed]
- Basehoar, A.D.; Zanton, S.J.; Pugh, B.F. Identification and distinct regulation of yeast TATA box-containing genes. Cell 2004, 116, 699–709. [Google Scholar] [CrossRef]
- Pelechano, V.; Chávez, S.; Pérez-Ortín, J.E. A complete set of nascent transcription rates for yeast genes. PLoS ONE 2010, 5. [Google Scholar] [CrossRef] [PubMed]
- Harbison, C.T.; Gordon, D.B.; Lee, T.I.; Rinaldi, N.J.; Macisaac, K.D.; Danford, T.W.; Hannett, N.M.; Tagne, J.-B.; Reynolds, D.B.; Yoo, J.; et al. Transcriptional regulatory code of a eukaryotic genome. Nature 2004, 431, 99–104. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.; Tillo, D.; Bray, N.; Morse, R.H.; Davis, R.W.; Hughes, T.R.; Nislow, C. A high-resolution atlas of nucleosome occupancy in yeast. Nat. Genet. 2007, 39, 1235–1244. [Google Scholar] [CrossRef] [PubMed]
- Yen, K.; Vinayachandran, V.; Batta, K.; Koerber, R.T.; Pugh, B.F. Genome-wide nucleosome specificity and directionality of chromatin remodelers. Cell 2012, 149, 1461–1473. [Google Scholar] [CrossRef] [PubMed]
- O’Connor, T.R.; Wyrick, J.J. ChromatinDB: A database of genome-wide histone modification patterns for Saccharomyces cerevisiae. Bioinformatics 2007, 23, 1828–1830. [Google Scholar] [CrossRef] [PubMed]
- Tirosh, I.; Barkai, N. Two strategies for gene regulation by promoter nucleosomes. Genome Res. 2008, 18, 1084–1091. [Google Scholar] [CrossRef] [PubMed]
- Zaugg, J.B.; Luscombe, N.M. A genomic model of condition-specific nucleosome behavior explains transcriptional activity in yeast. Genome Res. 2012, 22, 84–94. [Google Scholar] [CrossRef] [PubMed]
- Matthews, H.R. My favourite molecule: Polyamines, chromatin structure and transcription. BioEssays 1993, 15, 561–566. [Google Scholar] [CrossRef] [PubMed]
- Pommier, Y.; Kerrigan, D.; Kohn, K. Topological complexes between DNA and topoisomerase II and effects of polyamines. Biochemistry 1989, 28, 995–1002. [Google Scholar] [CrossRef] [PubMed]
- Bojanowski, K.; Filhol, O.; Cochet, C.; Chambaz, E.M.; Larsen, A.K. DNA topoisomerase II and casein kinase II associate in a molecular complex that is catalytically active. J. Biol. Chem. 1993, 268, 22920–22926. [Google Scholar] [PubMed]
- Alm, K.; Berntsson, P.; Oredsson, S.M. Topoisomerase II is nonfunctional in polyamine-depleted cells. J. Cell. Biochem. 1999, 75, 46–55. [Google Scholar] [CrossRef]
- Ioshikhes, I.P.; Albert, I.; Zanton, S.J.; Pugh, B.F. Nucleosome positions predicted through comparative genomics. Nat. Genet. 2006, 38, 1210–1215. [Google Scholar] [CrossRef] [PubMed]
- Koch, C.M.; Andrews, R.M.; Flicek, P.; Dillon, S.C.; Karaöz, U.; Clelland, G.K.; Wilcox, S.; Beare, D.M.; Fowler, J.C.; Couttet, P.; et al. The landscape of histone modifications across 1% of the human genome in five human cell lines. Genome Res. 2007, 17, 691–707. [Google Scholar] [CrossRef] [PubMed]
- Guenther, M.G.; Levine, S.S.; Boyer, L.A.; Jaenisch, R.; Young, R.A. A Chromatin Landmark and Transcription Initiation at Most Promoters in Human Cells. Cell 2007, 130, 77–88. [Google Scholar] [CrossRef] [PubMed]
- Pokholok, D.K.; Harbison, C.T.; Levine, S.; Cole, M.; Hannett, N.M.; Tong, I.L.; Bell, G.W.; Walker, K.; Rolfe, P.A.; Herbolsheimer, E.; et al. Genome-wide map of nucleosome acetylation and methylation in yeast. Cell 2005, 122, 517–527. [Google Scholar] [CrossRef] [PubMed]
- Shahbazian, M.D.; Grunstein, M. Functions of Site-Specific Histone Acetylation and Deacetylation. Annu. Rev. Biochem. 2007, 76, 75–100. [Google Scholar] [CrossRef] [PubMed]
- Rao, B.; Shibata, Y.; Strahl, B.D.; Lieb, J.D. Dimethylation of Histone H3 at Lysine 36 Demarcates Regulatory and Nonregulatory Chromatin Genome-Wide. Mol. Cell. Biol. 2005, 25, 9447–9459. [Google Scholar] [CrossRef] [PubMed]
- Wagner, E.J.; Carpenter, P.B. Understanding the language of Lys36 methylation at histone H3. Nat. Rev. Mol. Cell Biol. 2012, 13, 115–126. [Google Scholar] [CrossRef] [PubMed]
- Zhang, H.; Roberts, D.N.; Cairns, B.R. Genome-wide dynamics of Htz1, a histone H2A variant that poises repressed/basal promoters for activation through histone loss. Cell 2005, 123, 219–231. [Google Scholar] [CrossRef] [PubMed]
- Talbert, P.B.; Henikoff, S. Histone variants—Ancient wrap artists of the epigenome. Nat. Rev. Mol. Cell Biol. 2010, 11, 264–275. [Google Scholar] [CrossRef] [PubMed]
- Tiwari, V.K.; Burger, L.; Nikoletopoulou, V.; Deogracias, R.; Thakurela, S.; Wirbelauer, C.; Kaut, J.; Terranova, R.; Hoerner, L.; Mielke, C.; et al. Target genes of Topoisomerase II regulate neuronal survival and are defined by their chromatin state. Proc. Natl. Acad. Sci. USA 2012, 109, E934–E943. [Google Scholar] [CrossRef] [PubMed]
- Escargueil, A.E.; Plisov, S.Y.; Skladanowski, A.; Borgne, A.; Meijer, L.; Gorbsky, G.J.; Larsen, A.K. Recruitment of cdc2 kinase by DNA topoisomerase II is coupled to chromatin remodeling. FASEB J. 2001, 15, 2288–2290. [Google Scholar] [CrossRef] [PubMed]
- Hurst, L.D.; Pál, C.; Lercher, M.J. The evolutionary dynamics of eukaryotic gene order. Nat. Rev. Genet. 2004, 5, 299–310. [Google Scholar] [CrossRef] [PubMed]
- Chouvardas, P.; Kollias, G.; Nikolaou, C. Inferring active regulatory networks from gene expression data using a combination of prior knowledge and enrichment analysis. BMC Bioinform. 2016, 17, 181. [Google Scholar] [CrossRef] [PubMed]
- Gehlen, L.R.; Gruenert, G.; Jones, M.B.; Rodley, C.D.; Langowski, J.; O’Sullivan, J.M.; Sullivan, J.M.O. Chromosome positioning and the clustering of functionally related loci in yeast is driven by chromosomal interactions. Nucleus 2012, 3, 370–383. [Google Scholar] [CrossRef] [PubMed]
- Zhu, J.; Zhang, B.; Smith, E.N.; Drees, B.; Brem, R.B.; Kruglyak, L.; Bumgarner, R.E.; Schadt, E.E. Integrating large-scale functional genomic data to dissect the complexity of yeast regulatory networks. Nat. Genet. 2008, 40, 854–861. [Google Scholar] [CrossRef] [PubMed]
- Sugino, R.P.; Innan, H. Natural Selection on Gene Order in the Genome Reorganization Process After Whole-Genome Duplication of Yeast. Mol. Biol. Evol. 2012, 29, 71–79. [Google Scholar] [CrossRef] [PubMed]
- Bajić, D.; Poyatos, J.F. Balancing noise and plasticity in eukaryotic gene expression. BMC Genom. 2012, 13, 343. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brill, S.J.; DiNardo, S.; Voelkel-Meiman, K.; Sternglanz, R. Need for DNA topoisomerase activity as a swivel for DNA replication for transcription of ribosomal RNA. Nature 1987, 326, 414–416. [Google Scholar] [CrossRef] [PubMed]
- Mondal, N.; Zhang, Y.; Jonsson, Z.; Dhar, S.K.; Kannapiran, M.; Parvin, J.D. Elongation by RNA polymerase II on chromatin templates requires topoisomerase activity. Nucleic Acids Res. 2003, 31, 5016–5024. [Google Scholar] [CrossRef] [PubMed]
- García-Rubio, M.L.; Aguilera, A. Topological constraints impair RNA polymerase II transcription and causes instability of plasmid-borne convergent genes. Nucleic Acids Res. 2012, 40, 1050–1064. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Bancaud, A.; Wagner, G.; Conde e Silva, N.; Lavelle, C.; Wong, H.; Mozziconacci, J.; Barbi, M.; Sivolob, A.; Le Cam, E.; Mouawad, L.; et al. Nucleosome Chiral Transition under Positive Torsional Stress in Single Chromatin Fibers. Mol. Cell 2007, 27, 135–147. [Google Scholar] [CrossRef] [PubMed]
- Lavelle, C. DNA torsional stress propagates through chromatin fiber and participates in transcriptional regulation. Nat. Struct. Mol. Biol. 2008, 15, 123–125. [Google Scholar] [CrossRef] [PubMed]
- Lavelle, C.; Victor, J.M.; Zlatanova, J. Chromatin fiber dynamics under tension and torsion. Int. J. Mol. Sci. 2010, 11, 1557–1579. [Google Scholar] [CrossRef] [PubMed]
- Kouzine, F.; Sanford, S.; Elisha-Feil, Z.; Levens, D. The functional response of upstream DNA to dynamic supercoiling in vivo. Nat. Struct. Mol. Biol. 2008, 15, 146–154. [Google Scholar] [CrossRef] [PubMed]

© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Joshi, R.S.; Nikolaou, C.; Roca, J. Structure and Chromosomal Organization of Yeast Genes Regulated by Topoisomerase II. Int. J. Mol. Sci. 2018, 19, 134. https://doi.org/10.3390/ijms19010134
Joshi RS, Nikolaou C, Roca J. Structure and Chromosomal Organization of Yeast Genes Regulated by Topoisomerase II. International Journal of Molecular Sciences. 2018; 19(1):134. https://doi.org/10.3390/ijms19010134
Chicago/Turabian StyleJoshi, Ricky S., Christoforos Nikolaou, and Joaquim Roca. 2018. "Structure and Chromosomal Organization of Yeast Genes Regulated by Topoisomerase II" International Journal of Molecular Sciences 19, no. 1: 134. https://doi.org/10.3390/ijms19010134




