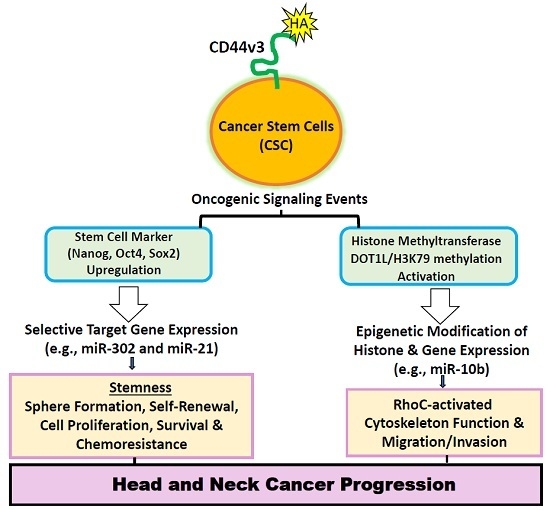
Activation of Matrix Hyaluronan-Mediated CD44 Signaling, Epigenetic Regulation and Chemoresistance in Head and Neck Cancer Stem Cells
Abstract
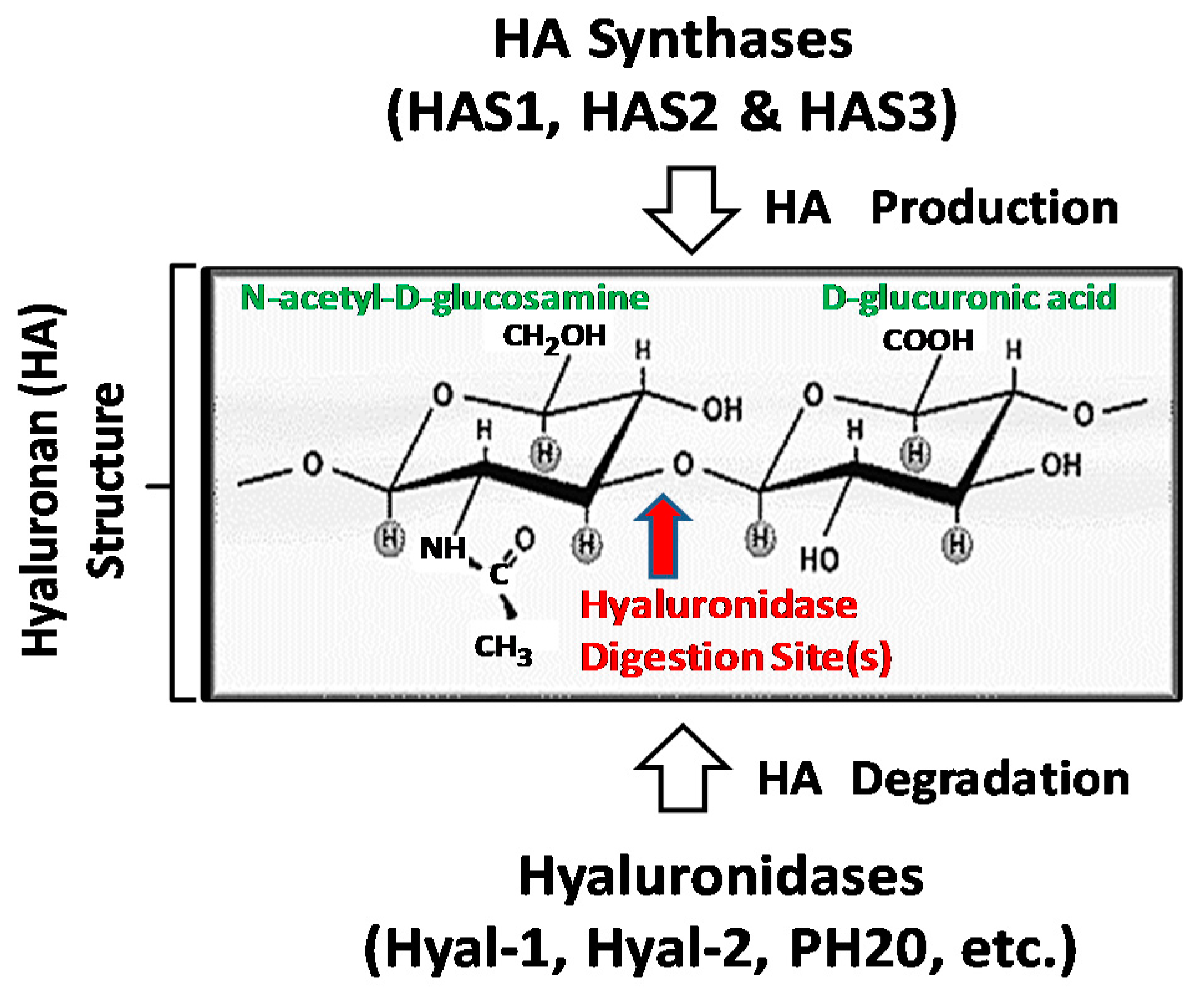
:1. Hyaluronan (HA) Metabolism in Head and Neck Cancer Progression
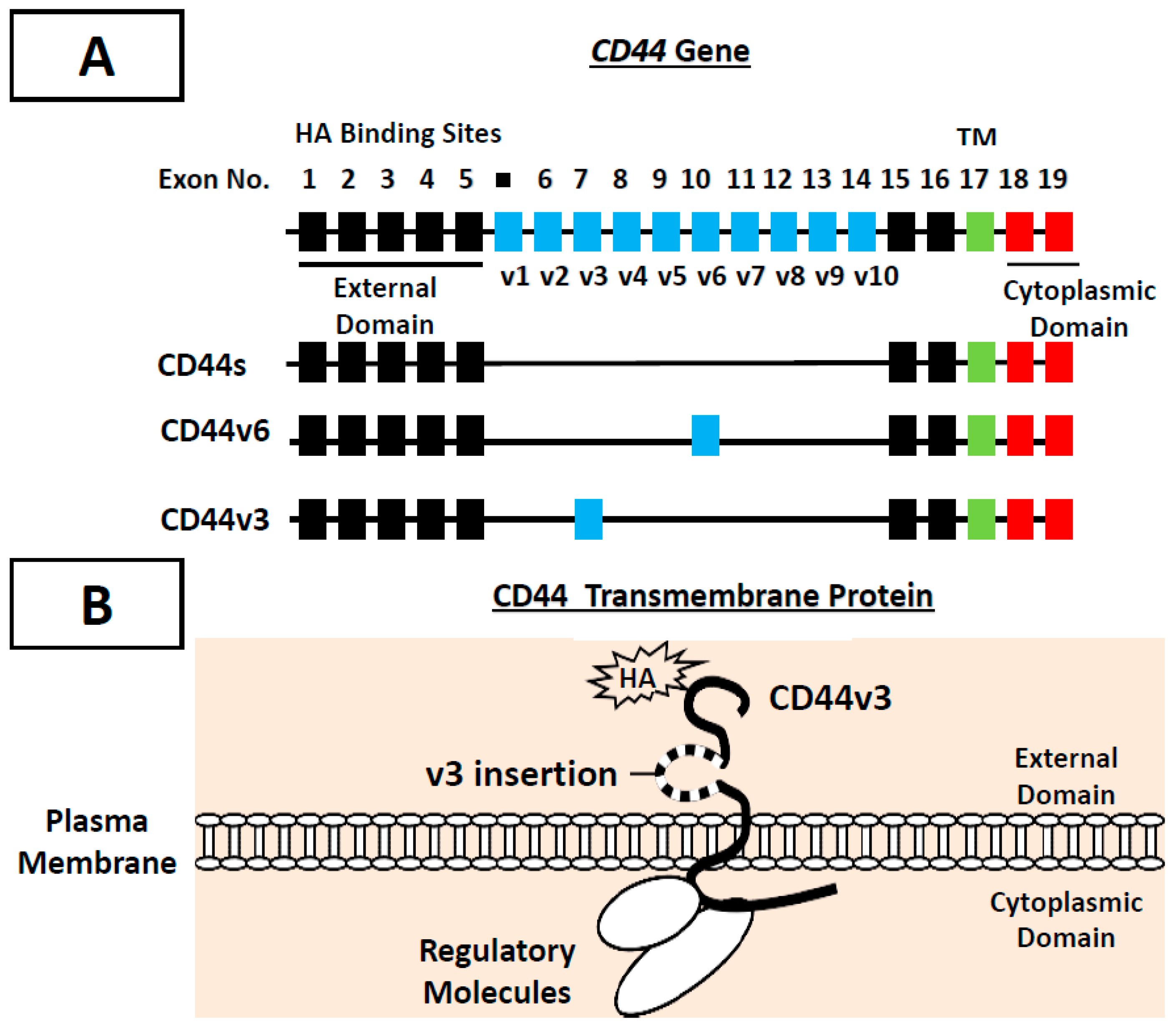
2. CD44 (a HA Receptor) in Cancer Stem Cells (CSCs) and Tumor Progression
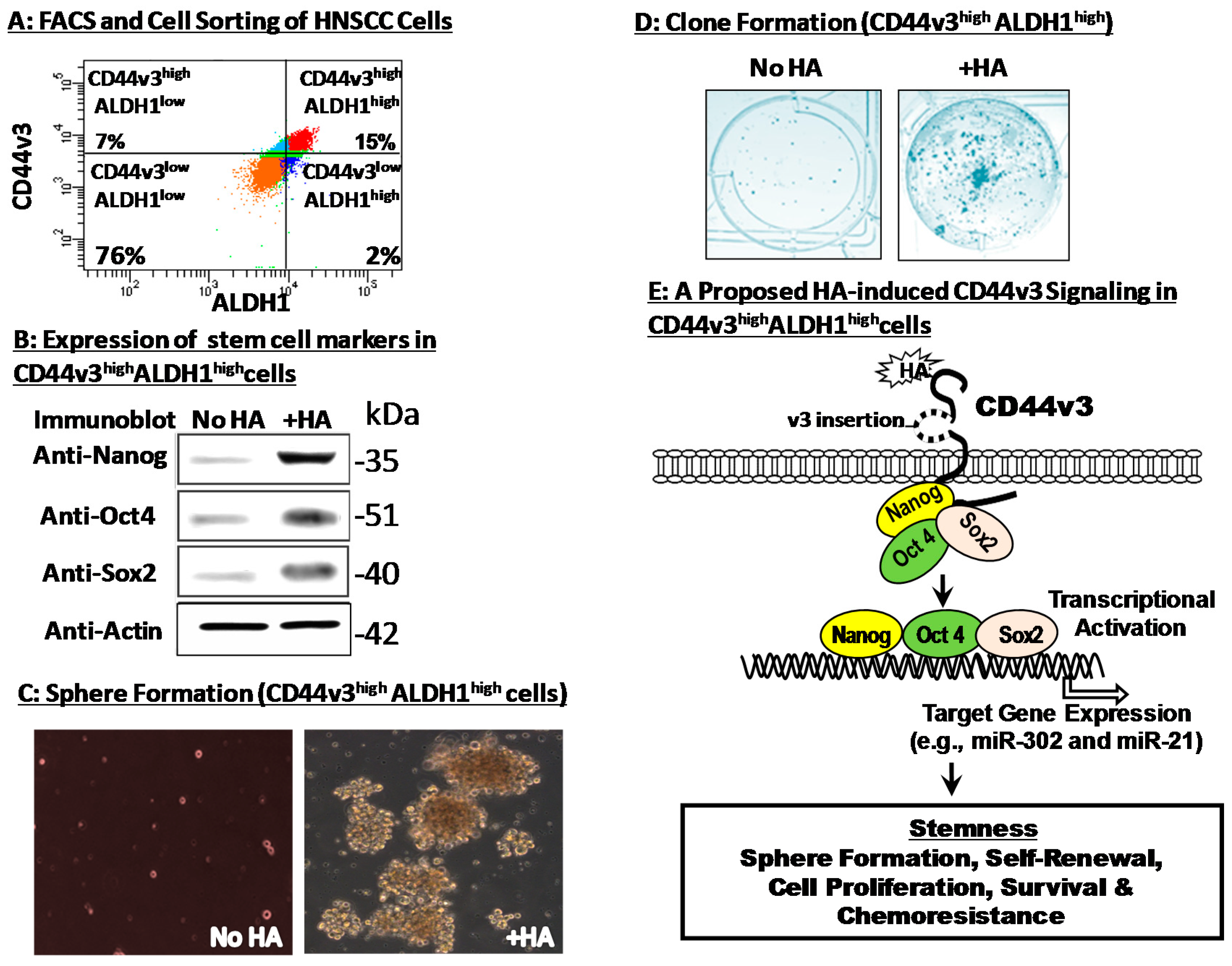
3. Hyaluronan–CD44 Interaction Stimulates Stem Cell Marker Expression, Stemness Properties and Chemoresistance in CD44v3highALDH1high Head and Neck Cancer Stem Cells (CSCs)
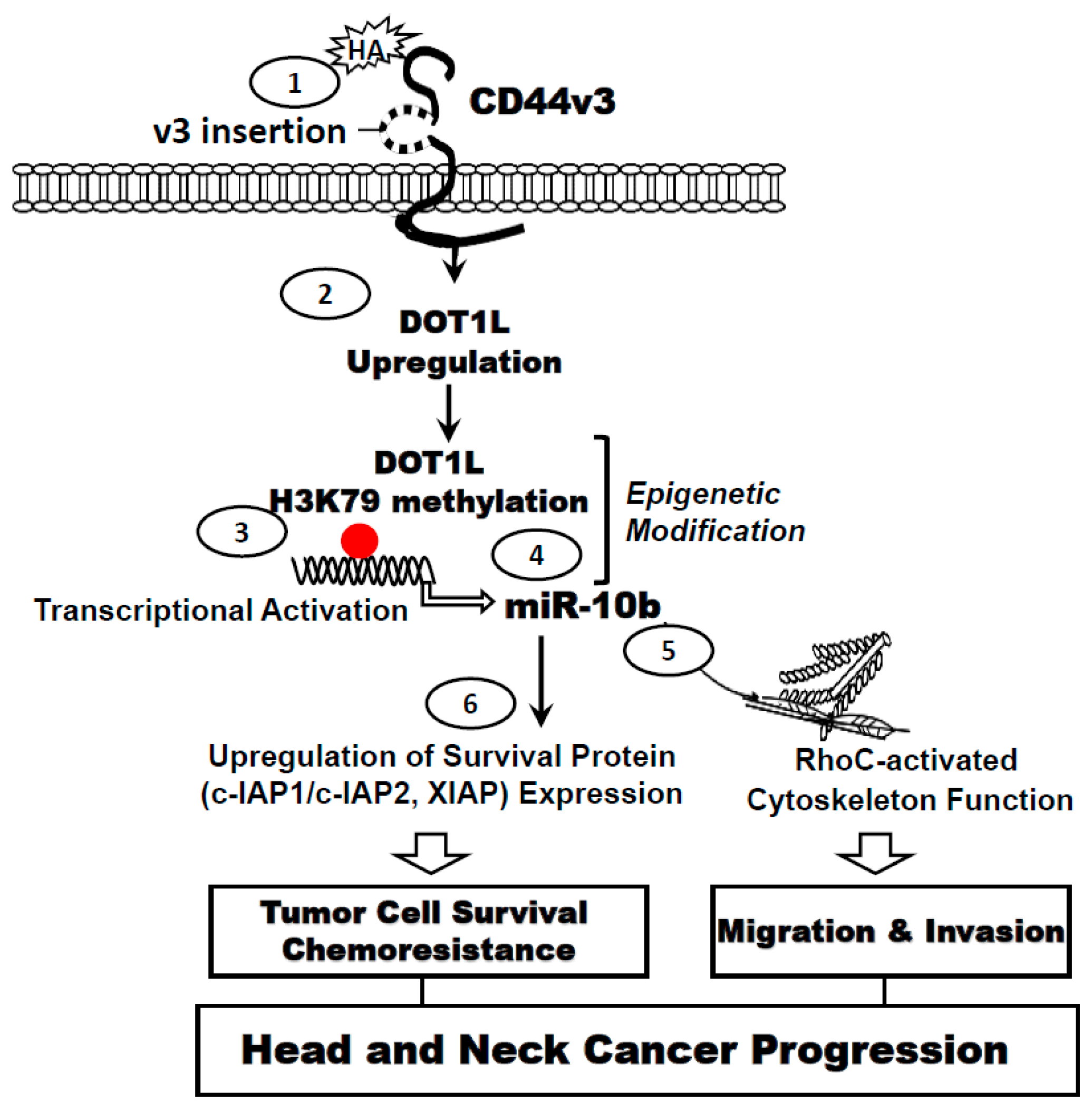
4. Hyaluronan (HA) Promote miRNA-10b Expression, Tumor Cell Invasion and Chemoresistance in Head and Neck Cancer Stem Cells (CSCs)
5. HA–CD44 Interaction Promotes Histone Methyltransferase, DOT1L Expression and Epigenetic Modification in Head and Neck Cancer Stem Cells
6. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Parkin, D.M.; Bray, F.; Ferlay, J.; Pisani, P. Global cancer statistics, 2002. CA Cancer J. Clin. 2002, 55, 74–108. [Google Scholar] [CrossRef]
- Haddad, R.I.; Shin, D.M. Recent advances in head and neck cancer. N. Engl. J. Med. 2008, 359, 1143–1154. [Google Scholar] [CrossRef] [PubMed]
- Leemans, C.R.; Braakhuis, B.J.; Brakenhoff, R.H. The molecular biology of head and neck cancer. Nat. Rev. Cancer 2011, 11, 9–22. [Google Scholar] [CrossRef] [PubMed]
- Pfister, D.G.; Ang, K.K.; Brizel, D.M.; Burtness, B.A.; Cmelak, A.J.; Colevas, A.D.; Dunphy, F.; Eisele, D.W.; Gilbert, J.; Gillison, M.L.; et al. Head and neck cancers. J. Natl. Compr. Cancer Netw. 2011, 9, 596–649. [Google Scholar] [CrossRef]
- Toole, B.P. Proteoglycans and hyaluronan in morphogenesis and differentiation. In Cell Biology of Extracellular Matrix; Hay, E.D., Ed.; Plenum Press: New York, NY, USA, 1991; pp. 305–334. [Google Scholar]
- Chanmee, T.; Ontong, P.; Itano, N. Hyaluronan: A modulator of the tumor microenvironment. Cancer Lett. 2016, 375, 20–30. [Google Scholar] [CrossRef] [PubMed]
- Weigel, P.H.; Hascall, V.C.; Tammi, M. Hyaluronan synthases. J. Biol. Chem. 1997, 272, 13997–14000. [Google Scholar] [CrossRef] [PubMed]
- Itano, N.; Kimata, K. Expression cloning and molecular characterization of HAS protein, a eukaryotic hyaluronan synthase. J. Biol. Chem. 1996, 271, 9875–9878. [Google Scholar] [CrossRef] [PubMed]
- Itano, N.; Kimata, K. Hyaluronan synthase: New directions for hyaluronan research. Trends Glycosci. Glycotechnol. 1998, 10, 23–38. [Google Scholar] [CrossRef]
- Weigel, P.H.; Baggenstoss, B.A.; Washburn, J.L. Hyaluronan synthase assembles hyaluronan on a [GlcNAc(β1,4)]n-GlcNAc(α1→)UDP primer and hyaluronan retains this residual chitin oligomer as a cap at the nonreducing end. Glycobiology 2017, 27, 536–554. [Google Scholar] [CrossRef] [PubMed]
- Spicer, A.P.; McDonald, J.A. Characterization and molecular evolution of a vertebrate hyaluronan synthase gene family. J. Biol. Chem. 1998, 273, 1923–1932. [Google Scholar] [CrossRef] [PubMed]
- Shyjan, A.M.; Heldin, P.; Butcher, E.C.; Yoshino, T.; Briskin, M.J. Functional cloning of the cDNA for a human hyaluronan synthase. J. Biol. Chem. 1996, 271, 23395–23399. [Google Scholar] [CrossRef] [PubMed]
- Tammi, R.H.; Passi, A.G.; Rilla, K.; Karousou, E.; Vigetti, D.; Makkonen, K.; Tammi, M.I. Transcriptional and post-translational regulation of hyaluronan synthesis. FEBS J. 2011, 278, 1419–1428. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.J.; Earle, C.; Wong, G.; Bourguignon, L.Y. Role of hyaluronan synthase 2 to promote CD44-dependent oral cavity squamous cell carcinoma progression. Head Neck 2013, 35, 511–520. [Google Scholar] [CrossRef] [PubMed]
- Stern, R.; Jedrzejas, M.J. Hyaluronidases: Their genomics, structures, and mechanisms of action. Chem. Rev. 2006, 106, 818–839. [Google Scholar] [CrossRef] [PubMed]
- Nykopp, T.K.; Rilla, K.; Tammi, M.I.; Tammi, R.H.; Sironen, R.; Hämäläinen, K.; Kosma, V.M.; Heinonen, S.; Anttila, M. Hyaluronan synthases (HAS1-3) and hyaluronidases (HYAL1-2) in the accumulation of hyaluronan in endometrioid endometrial carcinoma. BMC Cancer 2010, 10, 512–522. [Google Scholar] [CrossRef] [PubMed]
- Csoka, A.B.; Frost, G.I.; Ster, N.R. The six hyaluronidase-like genes in the human and mouse genomes. Matrix Biol. 2001, 20, 499–508. [Google Scholar] [CrossRef]
- Franzmann, E.J.; Schroeder, G.L.; Goodwin, W.J.; Weed, D.T.; Fisher, P.; Lokeshwar, V.B. Expression of tumor markers hyaluronic acid and hyaluronidase (HYAL1) in head and neck tumors. Int. J. Cancer 2003, 106, 438–445. [Google Scholar] [CrossRef] [PubMed]
- Lepperdinger, G.; Müllegger, J.; Kreil, G. Hyal2-less active, but more versatile? Matrix Biol. 2001, 20, 509–514. [Google Scholar] [CrossRef]
- Harada, H.; Takahashi, M. CD44-dependent intracellular and extracellular catabolism of hyaluronic acid by hyaluronidase-1 and -2. J. Biol. Chem. 2007, 282, 5597–5607. [Google Scholar] [CrossRef] [PubMed]
- Wu, M.; Cao, M.; He, Y.; Liu, Y.; Yang, C.; Du, Y.; Wang, W.; Gao, F. A novel role of low molecular weight hyaluronan in breast cancer metastasis. FASEB J. 2015, 29, 1290–1298. [Google Scholar] [CrossRef] [PubMed]
- Flannery, C.R.; Little, C.B.; Hughes, C.E.; Caterson, B. Expression and activity of articular cartilage hyaluronidases. Biochem. Biophys. Res. Commun. 1998, 251, 824–829. [Google Scholar] [CrossRef]
- Csoka, A.B.; Scherer, S.W.; Stern, R. Expression analysis of six paralogous human hyaluronidase genes clustered on chromosomes 3p21 and 7q31. Genomics 1999, 60, 356–361. [Google Scholar] [CrossRef] [PubMed]
- Cherr, G.N.; Yudin, A.I.; Overstreet, J.W. The dual functions of GPI-anchored PH-20: Hyaluronidase and intracellular signaling. Matrix Biol. 2001, 20, 515–525. [Google Scholar] [CrossRef]
- Beech, D.J.; Madan, A.K.; Deng, N. Expression of PH-20 in normal and neoplastic breast tissue. J. Surg. Res. 2002, 103, 203–207. [Google Scholar] [CrossRef] [PubMed]
- Haylock, D.N.; Nilsson, S.K. The role of hyaluronic acid in hemopoietic stem cell biology. Regen. Med. 2006, 1, 437–445. [Google Scholar] [CrossRef] [PubMed]
- Astachov, L.; Vago, R.; Aviv, M.; Nevo, Z. Hyaluronan and mesenchymal stem cells: From germ layer to cartilage and bone. Front. Biosci. 2011, 16, 261–276. [Google Scholar] [CrossRef]
- Franzmann, E.J.; Weed, D.T.; Civantos, F.J.; Goodwin, W.J.; Bourguignon, L.Y. A novel CD44v3 isoform is involved in head and neck squamous cell carcinoma progression. Otolaryngol. Head Neck Surg. 2001, 124, 426–432. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.J.; Wreesmann, V.B.; Bourguignon, L.Y. Association of CD44v3-containing isoforms with tumor cell growth, migration, matrix metalloproteinase expression, and lymph node metastasis in head and neck cancer. Head Neck 2007, 29, 550–558. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.J.; Wong, G.; de Heer, A.M.; Xia, W.; Bourguignon, L.Y. CD44 variant isoforms in head and neck squamous cell carcinoma progression. Laryngoscope 2009, 119, 1518–1530. [Google Scholar] [CrossRef] [PubMed]
- Wang, S.J.; Bourguignon, L.Y. Role of hyaluronan-mediated CD44 signaling in head and neck squamous cell carcinoma progression and chemoresistance. Am. J. Pathol. 2011, 178, 956–963. [Google Scholar] [CrossRef] [PubMed]
- Screaton, G.R.; Bell, M.V.; Jackson, D.G.; Cornelis, F.B.; Gerth, U.; Bell, J.I. Genomic structure of DNA coding the lymphocyte homing receptor CD44 reveals 12 alternatively spliced exons. Proc. Natl. Acad. Sci. USA 1992, 89, 12160–12164. [Google Scholar] [CrossRef] [PubMed]
- Bennett, K.L.; Jackson, D.G.; Simon, J.C.; Tanczos, E.; Peach, R.; Modrell, B.; Stamenkovic, I.; Plowman, G.; Aruffo, A. CD44 isoforms containing exon V3 are responsible for the presentation of heparin-binding growth factor. J. Cell Biol. 1995, 128, 687–698. [Google Scholar] [CrossRef] [PubMed]
- Mack, B.; Gires, O. CD44s and CD44v6 expression in head and neck epithelia. PLoS ONE 2008, 3, e3360. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Peach, R.J.; Hollenbaugh, D.; Stamenkovic, I.; Aruffo, A. Identification of hyaluronic acid binding sites in the extracellular domain of CD44. J. Cell Biol. 1993, 122, 257–264. [Google Scholar] [CrossRef] [PubMed]
- Lokeshwar, V.B.; Iida, N.; Bourguignon, L.Y. The Cell adhesion molecule, GP116, is a new CD44 variant (ex14/v10) involved in hyaluronic acid binding and endothelial cell proliferation. J. Biol. Chem. 2002, 277, 4589–4592. [Google Scholar] [CrossRef]
- Bourguignon, L.Y.; Lokeshwar, V.B.; Chen, X.; Kerrick, W.G. Hyaluronic acid-induced lymphocyte signal transduction and HA receptor (GP85/CD44)-cytoskeleton interaction. J. Immunol. 1993, 151, 6634–6644. [Google Scholar] [PubMed]
- Bourguignon, L.Y. Hyaluronan-mediated CD44 activation of RhoGTPase signaling and cytoskeleton function promotes tumor progression. Semin. Cancer Biol. 2008, 18, 251–259. [Google Scholar] [CrossRef] [PubMed]
- Bourguignon, L.Y.; Wong, G.; Earle, C.; Krueger, K.; Spevak, C.C. Hyaluronan-CD44 interaction promotes c-Src-mediated twist signaling, microRNA-10b expression, and RhoA/RhoC up-regulation, leading to Rho-kinase-associated cytoskeleton activation and breast tumor cell invasion. J. Biol. Chem. 2010, 285, 36721–36735. [Google Scholar] [CrossRef] [PubMed]
- Bourguignon, L.Y.; Shiina, M.; Li, J.J. Hyaluronan-CD44 interaction promotes oncogenic signaling, microRNA functions, chemoresistance, and radiation resistance in cancer stem cells leading to tumor progression. Adv. Cancer Res. 2014, 123, 255–275. [Google Scholar] [PubMed]
- Al-Hajj, M.; Wicha, M.S.; Benito-Hernandez, A.; Morrison, S.J.; Clarke, M.F. Prospective identification of tumorigenic breast cancer cells. Proc. Natl. Acad. Sci. USA 2003, 100, 3983–3988. [Google Scholar] [CrossRef] [PubMed]
- Yeo, T.K.; Nagy, J.A.; Yeo, K.T.; Dvorak, H.F.; Toole, B.P. Increased hyaluronan at sites of attachment to mesentery by CD44-positive mouse ovarian and breast tumor cells. Am. J. Pathol. 1996, 148, 1733–1740. [Google Scholar] [PubMed]
- Dalerba, P.; Cho, R.W.; Clarke, M.F. Cancer stem cells: Models and concepts. Annu. Rev. Med. 2007, 58, 267–284. [Google Scholar] [CrossRef] [PubMed]
- Schulenburg, A.; Ulrich-Pur, H.; Thurnher, D.; Erovic, B.; Florian, S.; Sperr, W.R.; Kalhs, P.; Marian, B.; Wrba, F.; Zielinski, C.C.; et al. Neoplastic stem cells: A novel therapeutic target in clinical oncology. Cancer 2006, 107, 2512–2520. [Google Scholar] [CrossRef] [PubMed]
- Prince, M.E.; Sivanandan, R.; Kaczorowski, A.; Wolf, G.T.; Kaplan, M.J.; Dalerba, P.; Weissman, I.L.; Clarke, M.F.; Ailles, L.E. Identification of a subpopulation of cells with cancer stem cell properties in head and neck squamous cell carcinoma. Proc. Nat. Acad. Sci. USA 2007, 104, 973–978. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.C.; Chen, Y.W.; Hsu, H.S.; Tseng, L.M.; Huang, P.I.; Lu, K.H.; Chen, D.T.; Tai, L.K.; Yung, M.C.; Chang, S.C.; et al. Aldehyde dehydrogenase 1 is a putative marker for cancer stem cells in head and neck squamous cancer. Biochem. Biophys. Res. Commun. 2009, 385, 307–313. [Google Scholar] [CrossRef] [PubMed]
- Clay, M.R.; Tabor, M.; Owen, J.H.; Carey, T.E.; Bradford, C.R.; Wolf, G.T.; Wicha, M.S.; Prince, M.E. Single-marker identification of head and neck squamous cell carcinoma cancer stem cells with aldehyde dehydrogenase. Head Neck 2010, 32, 1195–1201. [Google Scholar] [CrossRef] [PubMed]
- Bourguignon, L.Y.; Wong, G.; Earle, C.; Chen, L. Hyaluronan-CD44v3 interaction with Oct4-Sox2-Nanog promotes miR-302 expression leading to self-renewal, clonal formation, and cisplatin resistance in cancer stem cells from head and neck squamous cell carcinoma. J. Biol. Chem. 2012, 287, 32800–32824. [Google Scholar] [CrossRef] [PubMed]
- Shiina, M.; Bourguignon, L.Y. Selective activation of cancer stem cells by size-specific hyaluronan in head and neck cancer. Int. J. Cell Biol. 2015. [Google Scholar] [CrossRef] [PubMed]
- Bourguignon, L.Y.; Wong, G.; Shiina, M. Up-regulation of histone methyltransferase, DOT1L, by matrix hyaluronan promotes microRNA-10 expression leading to tumor cell invasion and chemoresistance in cancer stem cells from head and neck squamous cell carcinoma. J. Biol. Chem. 2016, 291, 10571–10585. [Google Scholar] [CrossRef] [PubMed]
- Kashyap, V.; Rezende, N.C.; Scotland, K.B.; Shaffer, S.M.; Persson, J.L.; Gudas, L.J.; Mongan, N.P. Regulation of stem cell pluripotency and differentiation involves a mutual regulatory circuit of the NANOG, OCT4, and SOX2 pluripotency transcription factors with polycomb repressive complexes and stem cell microRNAs. Stem Cells Dev. 2009, 18, 1093–1108. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.; He, W.; Lu, C.; Wang, Z.; Wang, J.; Giercksky, K.E.; Nesland, J.M.; Suo, Z. Oct3/4 and Sox2 are significantly associated with an unfavorable clinical outcome in human esophageal squamous cell carcinoma. Anticancer Res. 2009, 29, 1233–1241. [Google Scholar] [PubMed]
- Chiou, S.H.; Yu, C.C.; Huang, C.Y.; Lin, S.C.; Liu, C.J.; Tsai, T.H.; Chou, S.H.; Chien, C.S.; Ku, H.H.; Lo, J.F. Positive correlations of Oct-4 and Nanog in oral cancer stem-like cells and high-grade oral squamous cell carcinoma. Clin. Cancer Res. 2008, 14, 4085–4095. [Google Scholar] [CrossRef] [PubMed]
- Ren, Z.H.; Zhang, C.P.; Ji, T. Expression of SOX2 in oral squamous cell carcinoma and the association with lymph node metastasis. Oncol. Lett. 2016, 11, 1973–1979. [Google Scholar] [CrossRef] [PubMed]
- Habu, N.; Imanishi, Y.; Kameyama, K.; Shimoda, M.; Tokumaru, Y.; Sakamoto, K.; Fujii, R.; Shigetomi, S.; Otsuka, K.; Sato, Y.; et al. Expression of Oct3/4 and Nanog in the head and neck squamous carcinoma cells and its clinical implications for delayed neck metastasis in stage I/II oral tongue squamous cell carcinoma. BMC Cancer 2015, 15, 730–737. [Google Scholar] [CrossRef] [PubMed]
- Islam, F.; Gopalan, V.; Wahab, R.; Smith, R.A.; Lam, A.K. Cancer stem cells in oesophageal squamous cell carcinoma: Identification, prognostic and treatment perspectives. Crit. Rev. Oncol. Hematol. 2015, 96, 9–19. [Google Scholar] [CrossRef] [PubMed]
- Patel, M.; Yang, S. Advances in reprogramming somatic cells to induced pluripotent stem cells. Stem. Cell. Rev. 2010, 6, 367–380. [Google Scholar] [CrossRef] [PubMed]
- Kamachi, Y.; Uchikawa, M.; Kondoh, H. Pairing SOX off: With partners in the regulation of embryonic development. Trends Genet. 2000, 16, 182–187. [Google Scholar] [CrossRef]
- Avilion, A.A.; Nicolis, S.K.; Pevny, L.H.; Perez, L.; Vivian, N.; Lovell-Badge, R. Multipotent cell lineages in early mouse development depend on SOX2 function. Genes Dev. 2003, 17, 126–140. [Google Scholar] [CrossRef] [PubMed]
- Gontan, C.; de Munck, A.; Vermeij, M.; Grosveld, F.; Tibboel, D.; Rottier, R. Sox2 is important for two crucial processes in lung development: Branching morphogenesis and epithelial cell differentiation. Dev. Biol. 2008, 317, 296–309. [Google Scholar] [CrossRef] [PubMed]
- Pesce, M.; Scholer, H.R. Oct-4: Gatekeeper in the beginnings of mammalian development. Stem Cells 2001, 19, 271–278. [Google Scholar] [CrossRef] [PubMed]
- Reers, S.; Pfannerstill, A.C.; Maushagen, R.; Pries, R.; Wollenberg, B. Stem cell profiling in head and neck cancer reveals an Oct-4 expressing subpopulation with properties of chemoresistance. Oral Oncol. 2014, 50, 155–162. [Google Scholar] [CrossRef] [PubMed]
- Herr, W.; Cleary, M.A. The POU domain: Versatility in transcriptional regulation by a flexible two-in-one DNA-binding domain. Gene Dev. 1995, 9, 1679–1693. [Google Scholar] [CrossRef] [PubMed]
- Bourguignon, L.Y.; Earle, C.; Wong, G.; Spevak, C.C.; Krueger, K. Stem cell marker (Nanog) and Stat-3 signaling promote MicroRNA-21 expression and chemoresistance in hyaluronan/CD44-activated head and neck squamous cell carcinoma cells. Oncogene 2012, 31, 149–160. [Google Scholar] [CrossRef] [PubMed]
- Bourguignon, L.Y.; Spevak, C.C.; Wong, G.; Xia, W.; Gilad, E. Hyaluronan-CD44 interaction with protein kinase C(epsilon) promotes oncogenic signaling by the stem cell marker Nanog and the Production of microRNA-21, leading to down-regulation of the tumor suppressor protein PDCD4, anti-apoptosis, and chemotherapy resistance in breast tumor cells. J. Biol. Chem. 2009, 284, 26533–26546. [Google Scholar] [PubMed]
- Nakamura, M.; Nakatani, K.; Uzawa, K.; Ono, K.; Uesugi, H.; Ogawara, K. Establishment and characterization of a cisplatin-resistant oral squamous cell carcinoma cell line, H-1R. Oncol. Rep. 2005, 14, 1281–1286. [Google Scholar] [CrossRef] [PubMed]
- Hunter, A.M.; LaCasse, E.C.; Korneluk, R.G. The inhibitors of apoptosis (IAPs) as cancer targets. Apoptosis 2007, 12, 1543–1568. [Google Scholar] [CrossRef] [PubMed]
- Gyrd-Hansen, M.; Meier, P. IAPs: From caspase inhibitors to modulators of NF-κB, inflammation and cancer. Nat. Rev. Cancer 2010, 10, 561–574. [Google Scholar] [CrossRef] [PubMed]
- Bourguignon, L.Y.; Xia, W.; Wong, G. Hyaluronan-mediated CD44 interaction with p300 and SIRT1 regulates β-catenin signaling and NFκB-specific transcription activity leading to MDR1 and Bcl-xL gene expression and chemoresistance in breast tumor cells. J. Biol. Chem. 2009, 284, 2657–2671. [Google Scholar] [CrossRef] [PubMed]
- Clevers, H. The cancer stem cell: Premises, promises and challenges. Nat. Med. 2011, 17, 313–319. [Google Scholar] [CrossRef] [PubMed]
- Sherida, C.; Kishimoto, H.; Fuchs, R.; Mehrotra, S.; Bhat-Nakshatri, P.; Turner, C.; Goulet, R., Jr.; Badve, S.; Nakshatri, H. CD44+/CD24− breast cancer cells exhibit enhanced invasive properties: An early step necessary for metastasis. Breast Cancer Res. 2006, 8, R59. [Google Scholar] [CrossRef] [PubMed]
- Marhaba, R.; Zoller, M. CD44 in cancer progression: Adhesion, migration and growth regulation. J. Mol. Histol. 2004, 35, 211–231. [Google Scholar] [CrossRef] [PubMed]
- Jiang, W.G.; Puntis, M.C.; Hallett, M.B. Molecular and cellular basis of cancer invasion and metastasis: Implications for treatment. Br. J. Surg. 1994, 81, 1576–1590. [Google Scholar] [CrossRef] [PubMed]
- Vasudevan, S.; Tong, Y.; Steitz, J.A. Switching from repression to activation: MicroRNAs can up-regulate translation. Science 2007, 318, 1931–1934. [Google Scholar] [CrossRef] [PubMed]
- Chang, S.S.; Jiang, W.W.; Smith, I.; Poeta, L.M.; Begum, S.; Glazer, C.; Shan, S.; Westra, W.; Sidransky, D.; Califano, J.A. MicroRNA alterations in head and neck squamous cell carcinoma. Int. J. Cancer 2008, 123, 2791–2797. [Google Scholar] [CrossRef] [PubMed]
- Bourguignon, L.Y.; Wong, G.; Xia, W.; Man, M.Q.; Holleran, W.M.; Elias, P.M. Selective matrix (hyaluronan) interaction with CD44 and RhoGTPase signaling promotes keratinocyte functions and overcomes age-related epidermal dysfunction. J. Dermatol. Sci. 2013, 72, 32–44. [Google Scholar] [CrossRef] [PubMed]
- Bourguignon, L.Y. Matrix hyaluronan-activated CD44 signaling promotes keratinocyte activities and improves abnormal epidermal functions. Am. J. Pathol. 2014, 184, 1912–1919. [Google Scholar] [CrossRef] [PubMed]
- Sasayama, T.; Nishihara, M.; Kondoh, T.; Hosoda, K.; Kohmura, E. MicroRNA-10b is overexpressed in malignant glioma and associated with tumor invasive factors, uPAR and RhoC. Int. J. Cancer 2009, 125, 1407–1413. [Google Scholar] [CrossRef] [PubMed]
- Shilatifard, A. Chromatin modifications by methylation and ubiquitination: Implications in the regulation of gene expression. Annu. Rev. Biochem. 2006, 75, 243–269. [Google Scholar] [CrossRef] [PubMed]
- Ng, H.H.; Feng, Q.; Wang, H.; Erdjument-Bromage, H.; Tempst, P.; Zhang, Y.; Struhl, K. Lysine methylation within the globular domain of histone H3 by Dot1 is important for telomeric silencing and Sir protein association. Genes Dev. 2002, 16, 1518–1527. [Google Scholar] [CrossRef] [PubMed]
- Van Leeuwen, F.; Gafken, P.R.; Gottschling, D.E. Dot1p modulates silencing in yeast by methylation of the nucleosome core. Cell 2002, 109, 745–756. [Google Scholar] [CrossRef]
- Feng, Q.; Wang, H.; Ng, H.H.; Erdjument-Bromage, H.; Tempst, P.; Struhl, K.; Zhang, Y. Methylation of H3-lysine 79 is mediated by a new family of HMTases without a SET domain. Curr. Biol. 2002, 12, 1052–1058. [Google Scholar] [CrossRef]
- San-Segundo, P.A.; Roeder, G.S. Role for the silencing protein Dot1 in meiotic checkpoint control. Mol. Boil. Cell 2000, 11, 3601–3615. [Google Scholar] [CrossRef]
- Janzen, C.J.; Hake, S.B.; Lowell, J.E.; Cross, G.A. Selective di- or trimethylation of histone H3 Lysine 76 by two DOT1 homologs is important for cell cycle regulation in Trypanosoma brucei. Mol. Cell 2006, 23, 497–507. [Google Scholar] [CrossRef] [PubMed]
- Bernt, K.M.; Zhu, N.; Sinha, A.U.; Vempati, S.; Faber, J.; Krivtsov, A.V.; Feng, Z.; Punt, N.; Daigle, A.; Bullinger, L.; et al. MLL-rearranged leukemia is dependent on aberrant H3K79 methylation by DOT1L. Cancer Cell 2011, 20, 66–78. [Google Scholar] [CrossRef] [PubMed]
- Daigle, S.R.; Olhava, E.J.; Therkelsen, C.A.; Majer, C.R.; Sneeringer, C.J.; Song, J.; Johnston, L.D.; Scott, M.P.; Smith, J.J.; Xiao, Y.; et al. Selective killing of mixed lineage leukemia cells by a potent small-molecule DOT1L inhibitor. Cancer Cell 2011, 20, 53–65. [Google Scholar] [CrossRef] [PubMed]
- Daigle, S.R.; Olhava, E.J.; Therkelsen, C.A.; Basavapathruni, A.; Jin, L.; Boriack-Sjodin, P.A.; Allain, C.J.; Klaus, C.R.; Raimondi, A.; Scott, M.P.; et al. Potent inhibition of DOT1L as treatment of MLL-fusion leukemia. Blood 2013, 122, 1017–1025. [Google Scholar] [CrossRef] [PubMed]
- Kim, W.; Kim, R.; Park, G.; Park, J.W.; Kim, J.E. Deficiency of H3K79 histone methyltransferase Dot1-like protein (DOT1L) inhibits cell proliferation. J. Biol. Chem. 2012, 287, 5588–5599. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.; Teruya-Feldstein, J.; Weinberg, R.A. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 2007, 449, 682–688. [Google Scholar] [CrossRef] [PubMed]
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bourguignon, L.Y.W.; Earle, C.; Shiina, M. Activation of Matrix Hyaluronan-Mediated CD44 Signaling, Epigenetic Regulation and Chemoresistance in Head and Neck Cancer Stem Cells. Int. J. Mol. Sci. 2017, 18, 1849. https://doi.org/10.3390/ijms18091849
Bourguignon LYW, Earle C, Shiina M. Activation of Matrix Hyaluronan-Mediated CD44 Signaling, Epigenetic Regulation and Chemoresistance in Head and Neck Cancer Stem Cells. International Journal of Molecular Sciences. 2017; 18(9):1849. https://doi.org/10.3390/ijms18091849
Chicago/Turabian StyleBourguignon, Lilly Y. W., Christine Earle, and Marisa Shiina. 2017. "Activation of Matrix Hyaluronan-Mediated CD44 Signaling, Epigenetic Regulation and Chemoresistance in Head and Neck Cancer Stem Cells" International Journal of Molecular Sciences 18, no. 9: 1849. https://doi.org/10.3390/ijms18091849
APA StyleBourguignon, L. Y. W., Earle, C., & Shiina, M. (2017). Activation of Matrix Hyaluronan-Mediated CD44 Signaling, Epigenetic Regulation and Chemoresistance in Head and Neck Cancer Stem Cells. International Journal of Molecular Sciences, 18(9), 1849. https://doi.org/10.3390/ijms18091849