The Protease Inhibitor CI2c Gene Induced by Bird Cherry-Oat Aphid in Barley Inhibits Green Peach Aphid Fecundity in Transgenic Arabidopsis
Abstract
:1. Introduction
2. Results
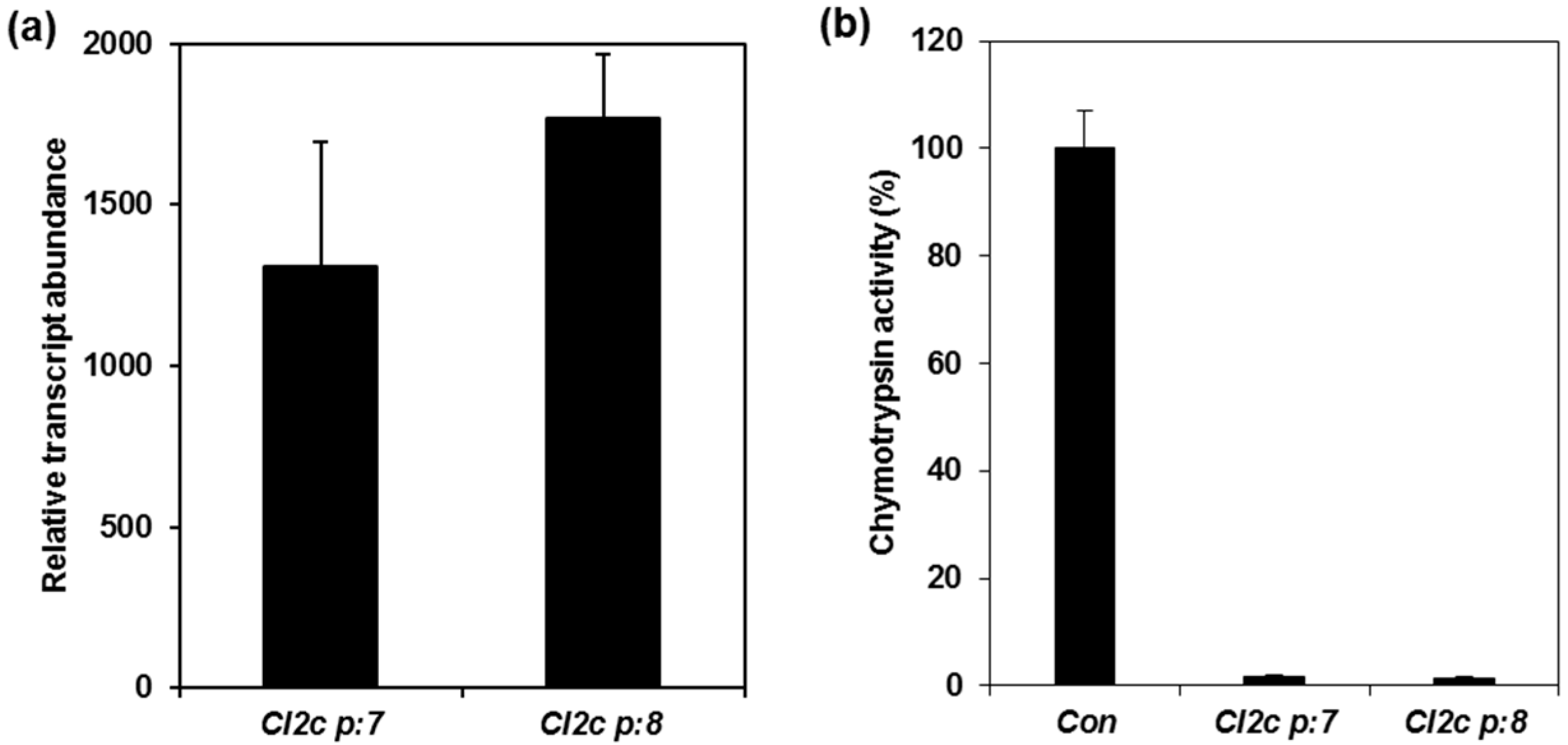
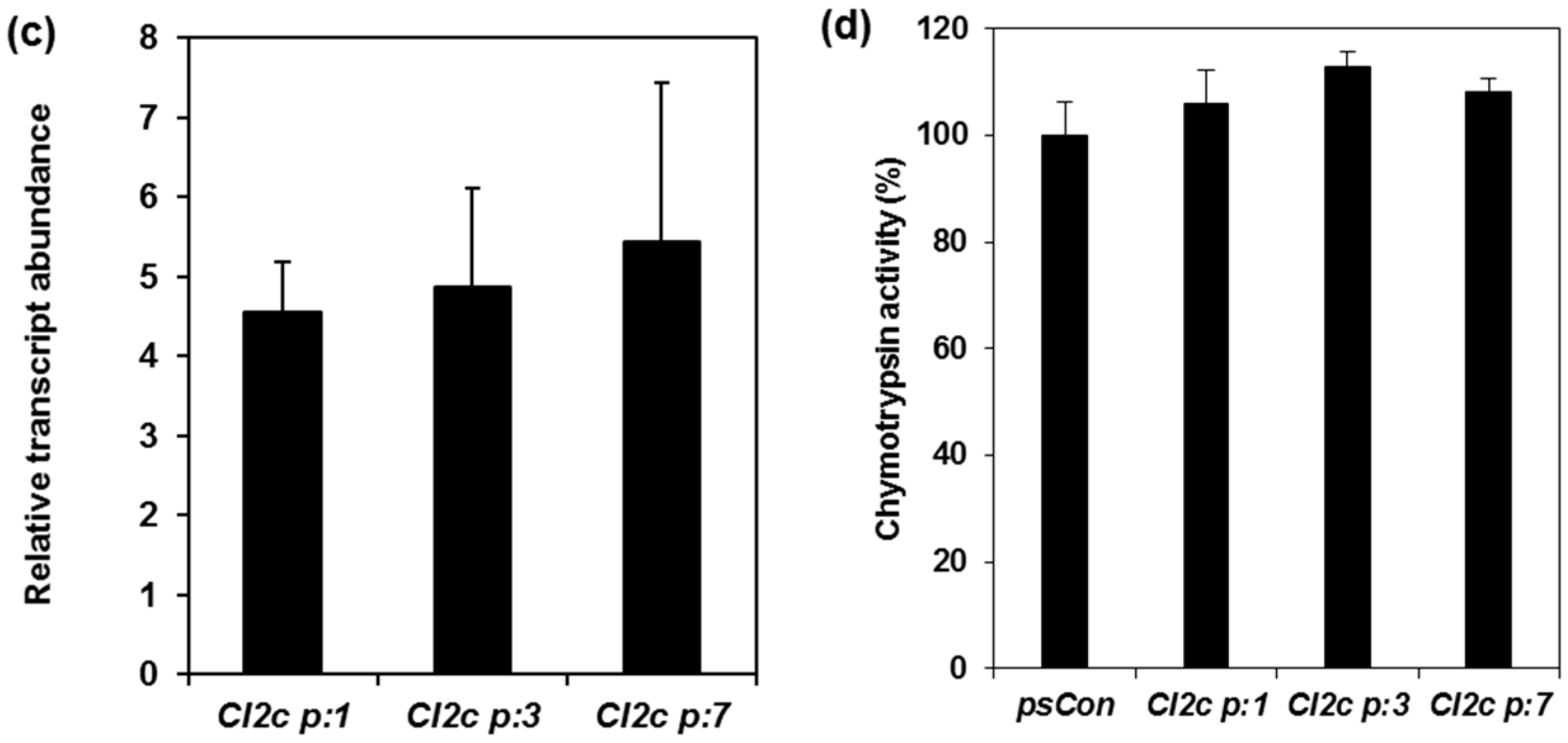
2.1. Confirmation of Transformation and Phenotypes of Arabidopsis Plants Expressing CI2c
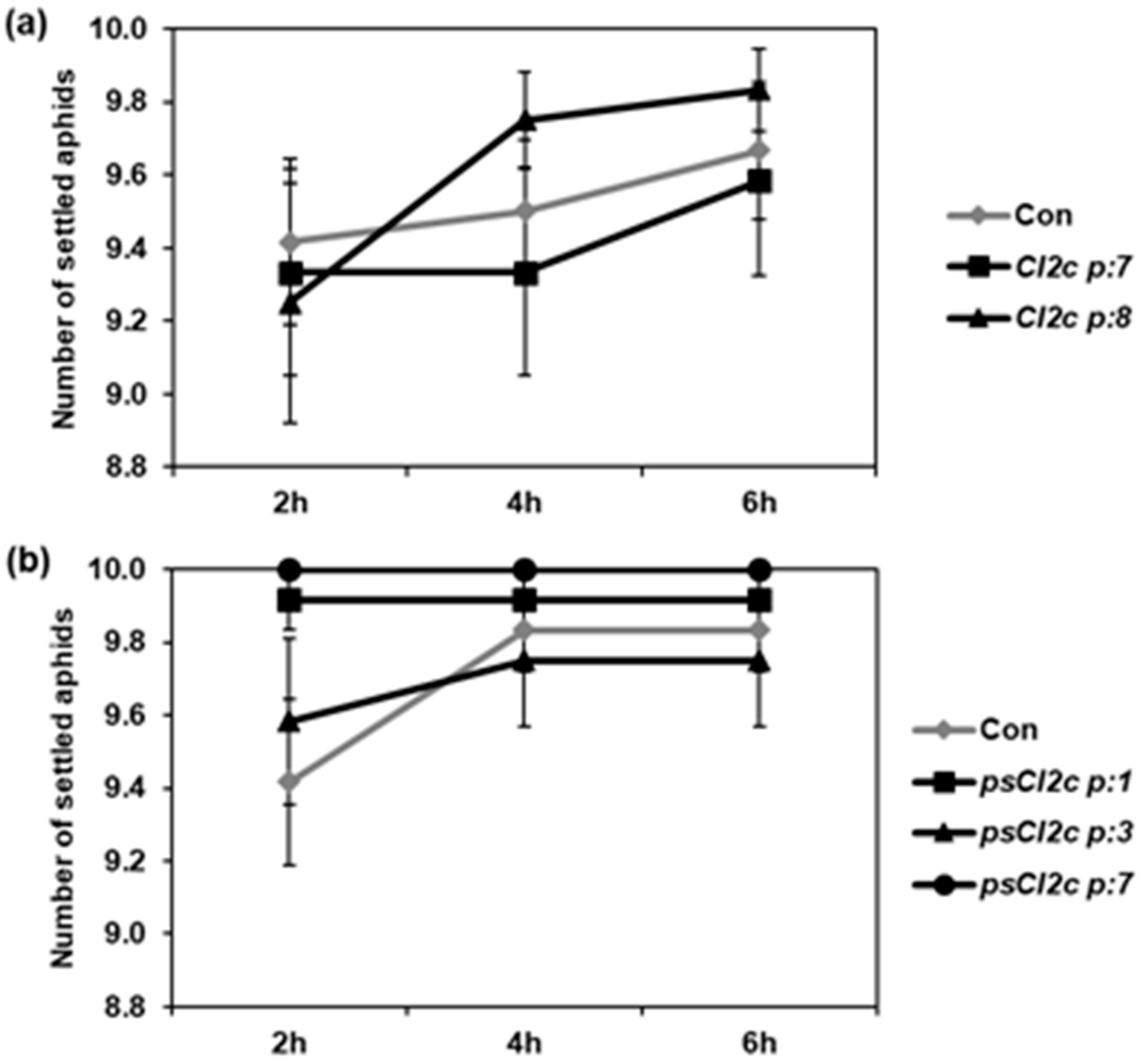
2.2. CI2c Expressed in Arabidopsis Has No or Minor Effect on GPA Settling
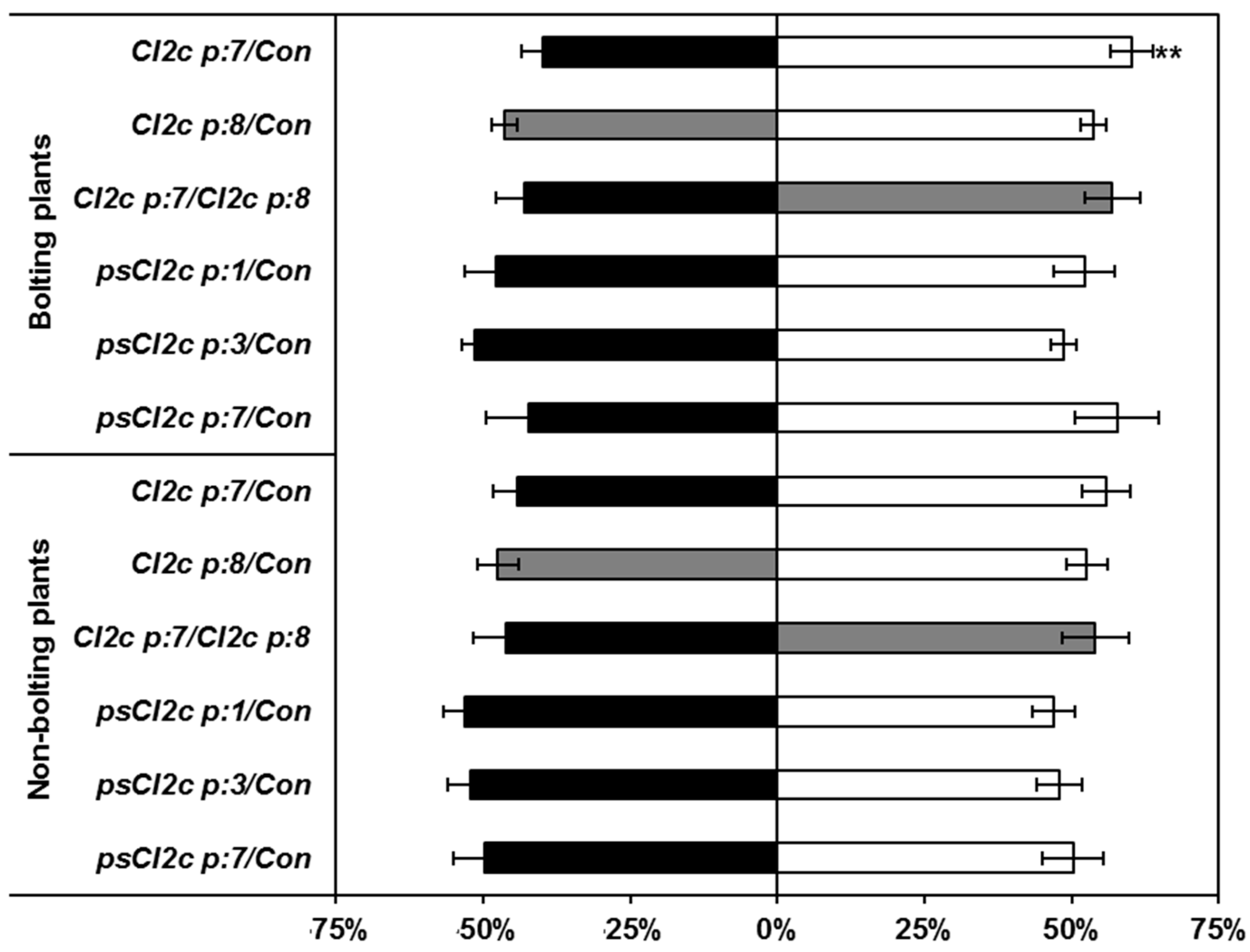
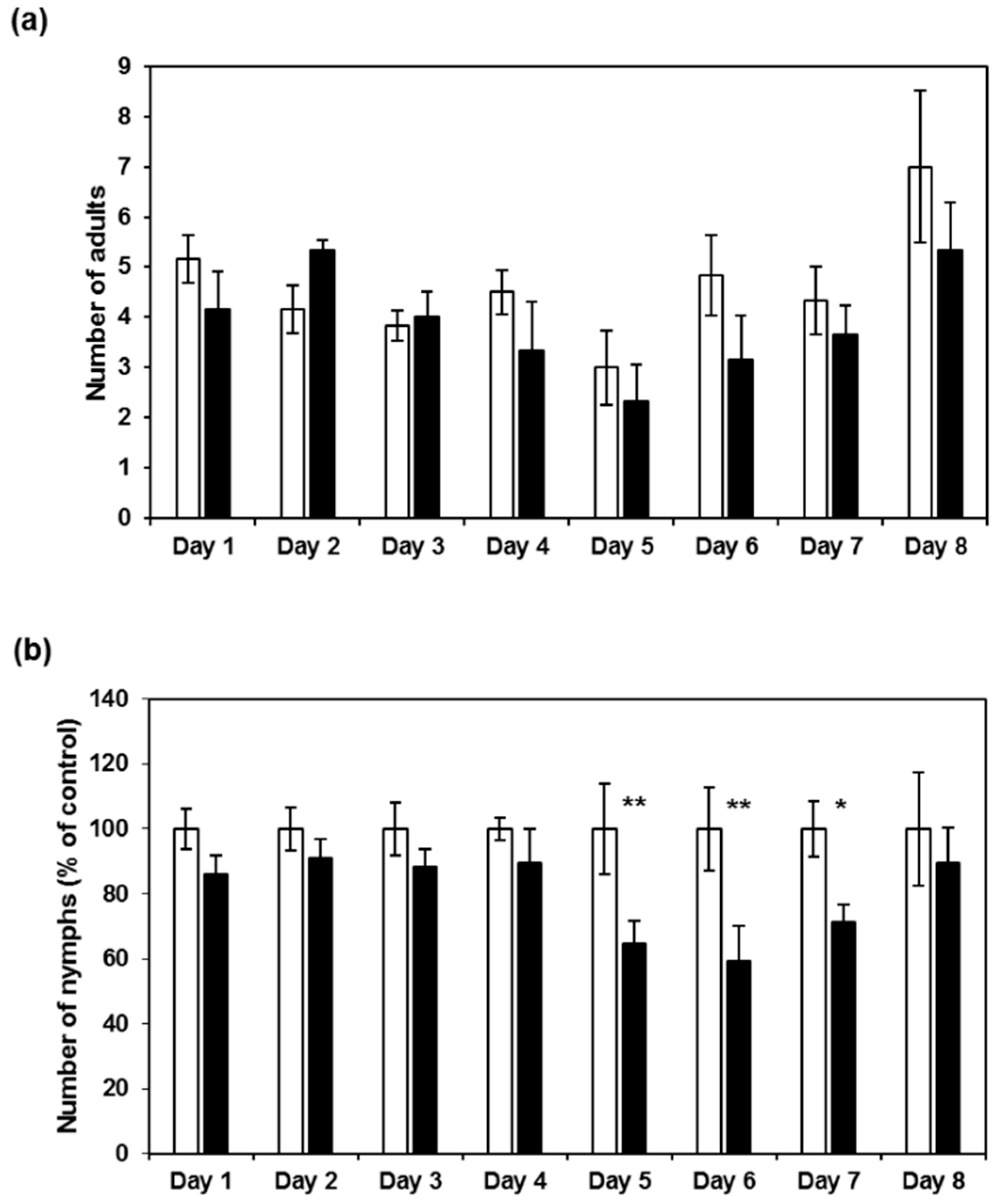
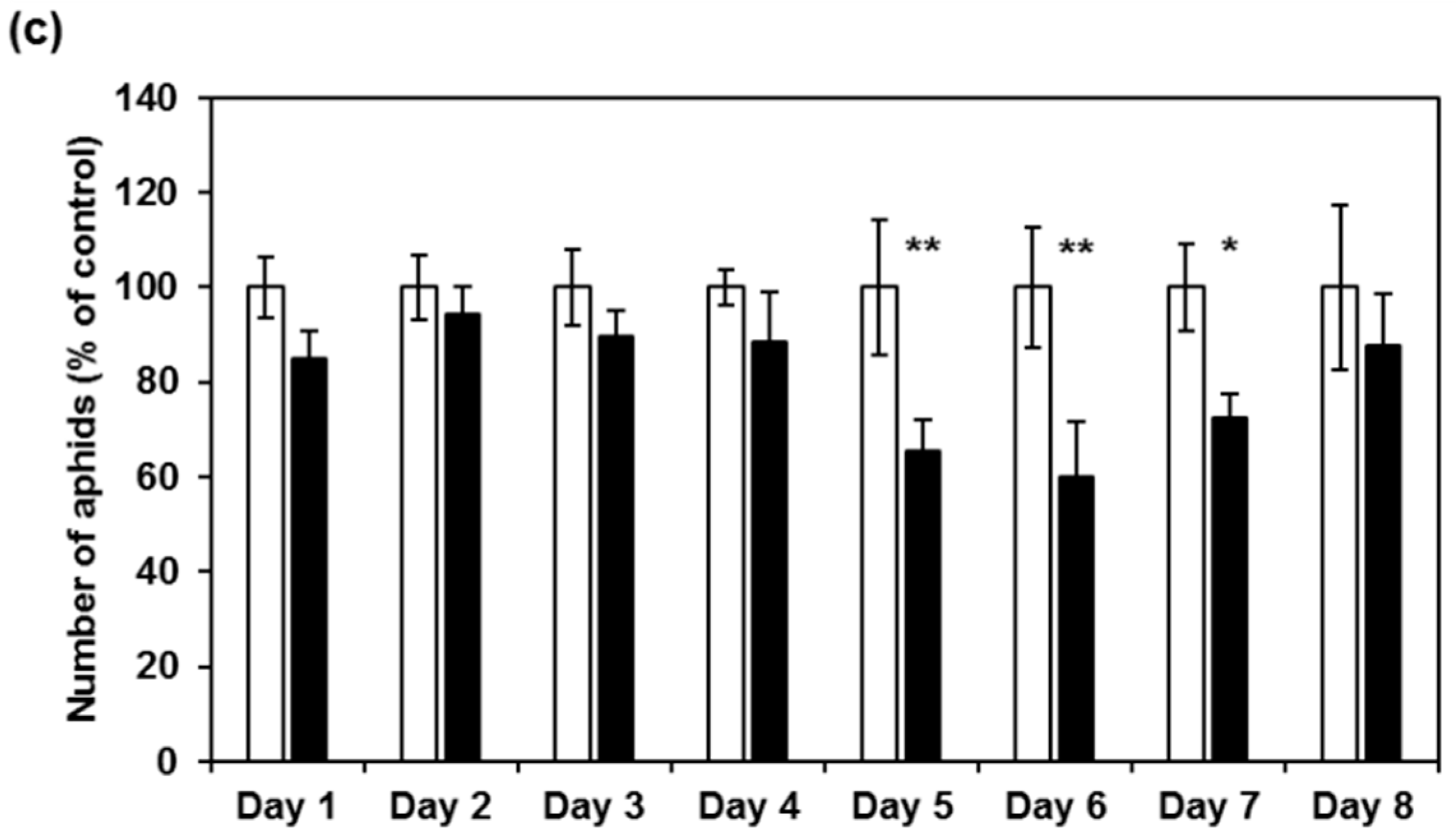
2.3. CI2c Expressed in Arabidopsis Has a Transient Short-Term Effect on GPA Fecundity
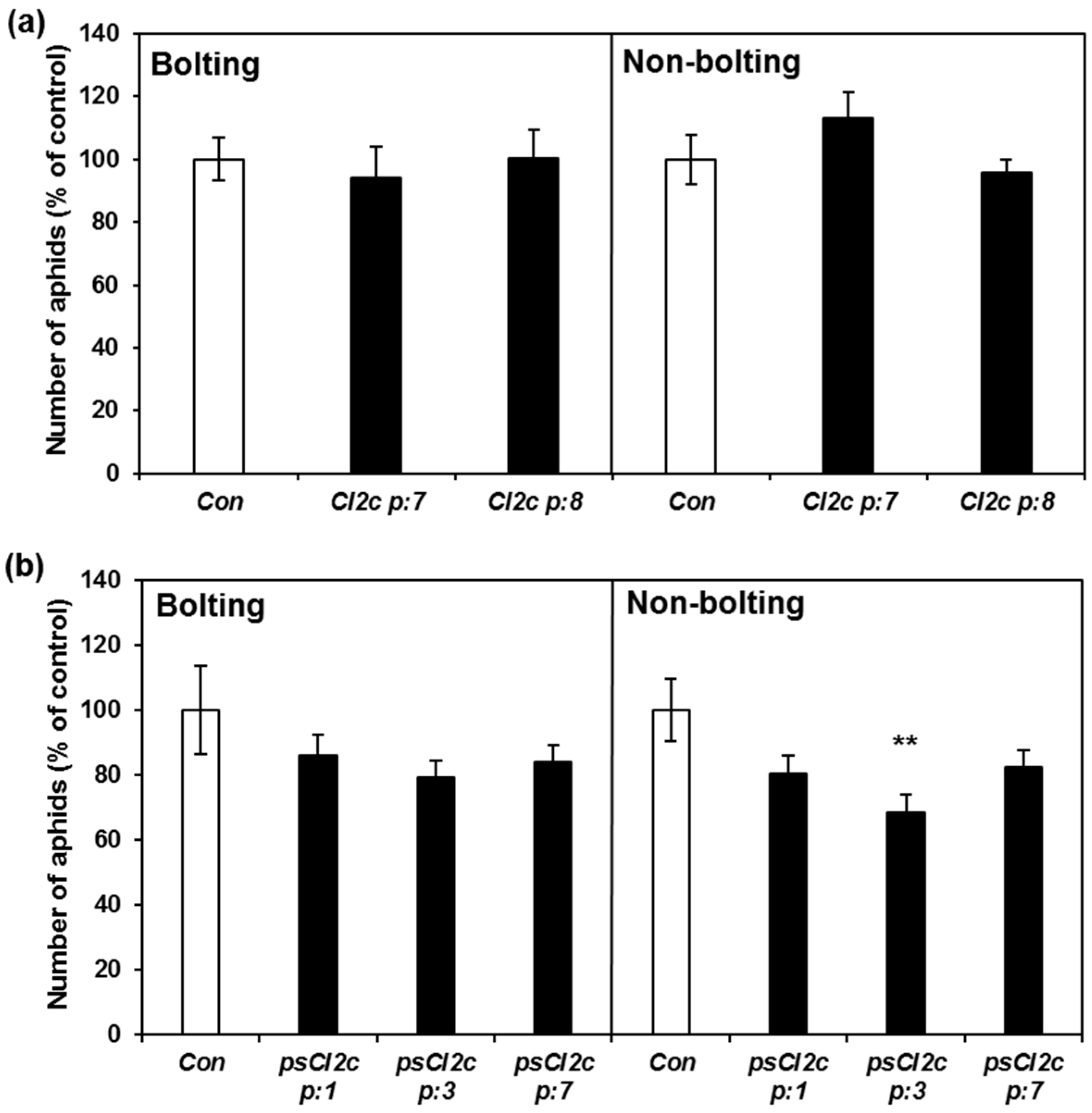
2.4. CI2c Has No Long-Term Effect on GPA Fecundity
3. Discussion
4. Materials and Methods
4.1. Plant Cultivation
4.2. Aphid Rearing
4.3. Plasmid Constructs, Plant Transformation and Selection
4.4. Analyses of Transcript Abundance
4.5. Enzymatic Assay of Protease Inhibitor Activity
4.6. Aphid Settling
4.7. Aphid Fecundity and Life Span
4.8. Statistical and Sequence Analyses
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| BCA | Bird cherry-oat aphid |
| CI2c | Barley chymotrypsin inhibitor 2c |
| GPA | Green peach aphid |
| PI | Proteinase inhibitor |
References
- Quisenberry, S.; Ni, X. Feeding injury. In Aphids as Crop Pests; van Emden, H.F., Harrington, R., Eds.; CABI Publishing: Wallingford, UK, 2007; pp. 331–352. ISBN 9780851998190. [Google Scholar]
- Dewar, A.M. Chemical control. In Aphids as Crop Pests; van Emden, H.F., Harrington, R., Eds.; CABI Publishing: Wallingford, UK, 2007; pp. 391–422. ISBN 9780851998190. [Google Scholar]
- Foster, S.P.; Devine, G.; Devonshire, A.L. Insecticide resistance. In Aphids as Crop Pests; van Emden, H.F., Harrington, R., Eds.; CABI Publishing: Wallingford, UK, 2007; pp. 261–285. ISBN 9780851998190. [Google Scholar]
- Bass, C.; Puinean, A.M.; Zimmer, C.T.; Denholm, I.; Field, L.M.; Foster, S.P.; Gutbrod, O.; Nauen, R.; Slater, R.; Williamson, M.S. The evolution of insecticide resistance in the peach potato aphid, Myzus persicae. Insect Biochem. Mol. Biol. 2014, 51, 41–51. [Google Scholar] [CrossRef] [PubMed]
- Bass, C.; Denholm, I.; Williamson, M.S.; Nauen, R. The global status of insect resistance to neonicotinoid insecticides. Pestic. Biochem. Physiol. 2015, 121, 78–87. [Google Scholar] [CrossRef] [PubMed]
- Hillocks, R.J. Farming with fewer pesticides: EU pesticide review and resulting challenges for UK agriculture. Crop. Prot. 2012, 31, 85–93. [Google Scholar] [CrossRef]
- Schreinemachers, P.; Tipraqsa, P. Agricultural pesticides and land use intensification in high, middle and low income countries. Food Policy 2012, 37, 616–626. [Google Scholar] [CrossRef]
- Dogimont, C.; Bendahmane, A.; Chovelon, V.; Boissot, N. Host plant resistance to aphids in cultivated crops: Genetic and molecular bases, and interactions with aphid populations. C. R. Biol. 2010, 333, 566–573. [Google Scholar] [CrossRef] [PubMed]
- Smith, C.M.; Chuang, W.-P. Plant resistance to aphid feeding: Behavioral, physiological, genetic and molecular cues regulate aphid host selection and feeding. Pest. Manag. Sci. 2014, 70, 528–540. [Google Scholar] [CrossRef] [PubMed]
- Burd, J.D.; Porter, D.R.; Puterka, G.J.; Haley, S.D.; Peairs, F.B. Biotypic variation among North American Russian wheat aphid (Homoptera: Aphididae) populations. J. Econ. Entomol. 2006, 99, 1862–1866. [Google Scholar] [CrossRef] [PubMed]
- Åhman, I.; Ninkovic, V. Volatile interactions between undamaged plants: Effects and potential for breeding resistance to aphids. In Plant Communication from an Ecological Perspective; Baluška, F., Ninkovic, V., Eds.; Springer: Berlin, Germany, 2010; pp. 113–126. [Google Scholar]
- Cheung, W.Y.; Di Giorgio, L.; Åhman, I. Mapping resistance to the bird cherry-oat aphid (Rhopalosiphum. padi) in barley. Plant Breed. 2010, 129, 637–646. [Google Scholar] [CrossRef]
- Louis, J.; Shah, J. Arabidopsis thaliana—Myzus persicae interaction: Shaping the understanding of plant defense against phloem-feeding aphids. Front. Plant Sci. 2013, 4. [Google Scholar] [CrossRef] [PubMed]
- De Vos, M.; Kim, J.H.; Jander, G. Biochemistry and molecular biology of Arabidopsis-aphid interactions. Bioessays 2007, 29, 871–883. [Google Scholar] [CrossRef] [PubMed]
- Truong, D.-H.; Bauwens, J.; Delaplace, P.; Mazzucchelli, G.; Lognay, G.; Francis, F. Proteomic analysis of Arabidopsis thaliana (L.) Heynh responses to a generalist sucking pest (Myzus persicae Sulzer). Plant Biol. 2015, 17, 1210–1217. [Google Scholar] [CrossRef] [PubMed]
- Louis, J.; Lorenc-Kukula, K.; Singh, V.; Reese, J.; Jander, G.; Shah, J. Antibiosis against the green peach aphid requires the Arabidopsis thaliana MYZUS PERSICAE-INDUCED LIPASE1 gene. Plant J. 2010, 64, 800–811. [Google Scholar] [CrossRef] [PubMed]
- Singh, V.; Louis, J.; Ayre, B.G.; Reese, J.C.; Shah, J. TREHALOSE PHOSPHATE SYNTHASE11-dependent trehalose metabolism promotes Arabidopsis thaliana defense against the phloem-feeding insect Myzus persicae. Plant J. 2011, 67, 94–104. [Google Scholar] [CrossRef] [PubMed]
- Avila, C.A.; Arevalo-Soliz, L.M.; Lorence, A.; Goggin, F.L. Expression of α-DIOXYGENASE 1 in tomato and arabidopsis contributes to plant defenses against aphids. Mol. Plant Microbe Interact. 2013, 26, 977–986. [Google Scholar] [CrossRef] [PubMed]
- Beβer, K.; Jarosch, B.; Langen, G.; Kogel, K.-H. Expression analysis of genes induced in barley after chemical activation reveals distinct disease resistance pathways. Mol. Plant Pathol. 2000, 1, 277–286. [Google Scholar]
- Morkunas, I.; Mai, V.C.; Gabryś, B. Phytohormonal signaling in plant responses to aphid feeding. Acta Physiol. Plant 2011, 33, 2057–2073. [Google Scholar] [CrossRef]
- Wei, F.; Wing, R.A.; Wise, R.P. Genome dynamics and evolution of the Mla. (powdery mildew) resistance locus in barley. Plant Cell. 2002, 14, 1903–1917. [Google Scholar] [CrossRef] [PubMed]
- Ryan, C.A. Protease inhibitors in plants: Genes for improving defenses against insects and pathogens. Annu. Rev. Phytopathol. 1990, 28, 425–449. [Google Scholar] [CrossRef]
- Koiwa, H.; Bressan, R.A.; Hasegawa, P.M. Regulation of protease inhibitors and plant defense. Trends Plant Sci. 1997, 2, 379–384. [Google Scholar] [CrossRef]
- Van Loon, L.C.; Rep, M.; Pieterse, C.M.J. Significance of inducible defense-related proteins in infected plants. Annu. Rev. Phytopathol. 2006, 44, 135–162. [Google Scholar] [CrossRef] [PubMed]
- Habib, H.; Fazili, K.M. Plant protease inhibitors: A defense strategy in plants. Biotechnol. Mol. Biol. Rev. 2007, 2, 68–85. [Google Scholar]
- Jashni, M.K.; Mehrabi, R.; Collemare, J.; Mesarich, C.H.; de Wit, P.J.G.M. The battle in the apoplast: Further insights into the roles of proteases and their inhibitors in plant–pathogen interactions. Front. Plant Sci. 2015, 6, 584. [Google Scholar] [CrossRef] [PubMed]
- Rahbé, Y.; Deraison, C.; Bonadé-Bottino, M.; Girard, C.; Nardon, C.; Jouanin, L. Effects of the cysteine protease inhibitor oryzacystatin (OC-I) on different aphids and reduced performance of Myzus persicae on OC-I expressing transgenic oilseed rape. Plant Sci. 2003, 164, 441–450. [Google Scholar] [CrossRef]
- Ribeiro, A.P.O.; Pereira, E.J.G.; Galvan, T.L.; Picanço, M.C.; Picoli, E.A.T.; da Silva, D.J.H.; Fári, M.G.; Otoni, W.C. Effect of eggplant transformed with oryzacystatin gene on Myzus persicae and Macrosiphum euphorbiae. J. Appl. Entomol. 2006, 130, 84–90. [Google Scholar] [CrossRef]
- Carrillo, L.; Martinez, M.; Álvarez-Alfageme, F.; Castañera, P.; Smagghe, G.; Diaz, I.; Ortego, F. A barley cysteine-proteinase inhibitor reduces the performance of two aphid species in artificial diets and transgenic Arabidopsis plants. Transgenic Res. 2011, 20, 305–319. [Google Scholar] [CrossRef] [PubMed]
- Ashouri, A.; Michaud, D.; Cloutier, C. Unexpected effects of different potato resistance factors to the Colorado potato beetle (Coleoptera:Chrysomelidae) on the potato aphid (Homopthera:Aphididae). Environ. Entomol. 2001, 30, 524–532. [Google Scholar]
- Rahbé, Y.; Ferrasson, E.; Rabesona, H.; Quillien, L. Toxicity to the pea aphid Acyrthosiphon pisum of anti-chymotrypsin isoforms and fragments of Bowman–Birk protease inhibitors from pea seeds. Insect Biochem. Mol. Biol. 2003, 33, 299–306. [Google Scholar] [CrossRef]
- Azzouz, H.; Cherqui, A.; Campan, E.D.M.; Rahbé, Y.; Duport, G.; Jouanin, L.; Kaiser, L.; Giordanengo, P. Effects of plant protease inhibitors, oryzacystatin I and soybean Bowman-Birk inhibitor, on the aphid Macrosiphum euphorbiae (Homoptera, Aphididae) and its parasitoid Aphelinus abdominalis (Hymenoptera, Aphelinidae). J. Insect Physiol. 2005, 51, 75–86. [Google Scholar] [CrossRef] [PubMed]
- Tran, P.; Cheesbrough, T.M.; Keickhefer, R.W. Plant proteinase inhibitors are potential anticereal aphid compounds. J. Econ. Entomol. 1997, 90, 1672–1677. [Google Scholar] [CrossRef]
- Pyati, P.; Bandani, A.R.; Fitches, E.; Gatehouse, J.A. Protein digestion in cereal aphids (Sitobion avenae) as a target for plant defence by endogenous proteinase inhibitors. J. Insect Physiol. 2011, 57, 881–891. [Google Scholar] [CrossRef] [PubMed]
- Casaretto, J.A.; Corcuera, L.J. Proteinase inhibitor accumulation in aphid-infested barley leaves. Phytochemistry. 1998, 49, 2279–2286. [Google Scholar] [CrossRef]
- Delp, G.; Gradin, T.; Åhman, I.; Jonsson, L.M.V. Microarray analysis of the interaction between the aphid Rhopalosiphum padi and host plants reveals both differences and similarities between susceptible and partially resistant barley lines. Mol. Genet. Genom. 2009, 281, 233–248. [Google Scholar] [CrossRef] [PubMed]
- Mehrabi, S.; Åhman, I.; Jonsson, L.M.V. Transcript abundance of resistance- and susceptibility-related genes in a barley breeding pedigree with partial resistance to the bird cherry-oat aphid (Rhopalosiphum. padi L.). Euphytica 2014, 198, 211–222. [Google Scholar] [CrossRef]
- Bhatia, V.; Bhattacharya, R.; Uniyal, P.L.; Singh, R.; Niranjan, R.S. Host generated siRNAs attenuate expression of serine protease gene in Myzus persicae. PLoS ONE 2012, 7, e46343. [Google Scholar] [CrossRef] [PubMed]
- Van Bel, A.J.E..; Will, T. Functional evaluation of proteins in watery and gel saliva of aphids. Front. Plant Sci. 2016, 7, 1840. [Google Scholar] [CrossRef] [PubMed]
- Furch, A.C.U.; van Bel, A.J.E.; Will, T. Aphid salivary proteases are capable of degrading sieve-tube proteins. J. Exp. Bot. 2015, 66, 533–539. [Google Scholar] [CrossRef] [PubMed]
- Giordanengo, P.; Brunissen, L.; Rusterucci, C.; Vincent, C.; van Bel, A.; Dinant, S.; Girousse, C.; Faucher, M.; Bonnemain, J.-L. Compatible plant-aphid interactions: How aphids manipulate plant responses. C. R. Biol. 2010, 333, 516–523. [Google Scholar] [CrossRef] [PubMed]
- Schmülling, T.; Schell, J.; Spena, A. Promoters of the rolA, B, and C genes of Agrobacterium rhizogenes are differentially regulated in transgenic plants. Plant Cell 1989, 1, 665–670. [Google Scholar] [CrossRef] [PubMed]
- Srivastava, A.C.; Ganesan, S.; Ismail, I.O.; Ayre, B.G. Effective carbon partitioning driven by exotic phloem-specific regulatory elements fused to the Arabidopsis thaliana AtSUC2 sucrose-proton symporter gene. BMC Plant Biol. 2009, 9, 7. [Google Scholar] [CrossRef] [PubMed]
- Losvik, A.; Beste, L.; Stephens, J.; Jonsson, L.M.V. Overexpression of the aphid-induced serine protease inhibitor CI2c gene in barley is beneficial to the generalist green peach aphid but does not affect the specialist bird cherry-oat aphid. Int. J. Mol. Sci. 2017. under review. [Google Scholar]
- Booker, J.; Chatfield, S.; Leyser, O. Auxin acts in xylem-associated or medullary cells to mediate apical dominance. Plant Cell 2003, 15, 495–507. [Google Scholar] [CrossRef] [PubMed]
- Pastor, V.; Luna, E.; Mauch-Mani, B.; Ton, J.; Flors, V. Primed plants do not forget. Environ. Exp. Bot. 2013, 94, 46–56. [Google Scholar] [CrossRef]
- Sjölund, R.D. The phloem sieve element: A river runs through it. Plant Cell. 1997, 9, 1137–1146. [Google Scholar] [CrossRef] [PubMed]
- Hunt, D.C.; Chrispeels, M.J. The signal peptide of a vacuolar protein is necessary and sufficient for the efficient secretion of a cytosolic protein. Plant Physiol. 1991, 96, 18–25. [Google Scholar] [CrossRef] [PubMed]
- Williamson, M.S.; Forde, J.; Buxton, B.; Kreis, M. Nucleotide sequence of barley chymotrypsin inhibitor-2 (CI-2) and its expression in normal and high-lysine barley. Eur. J. Biochem. 1987, 165, 99–106. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, U.; Munk, L.; Ullrich, S.E. Immunogold localization of chymotrypsin inhibitor-2, a lysine-rich protein, in developing barley endosperm. Planta 1990, 180, 272–277. [Google Scholar] [CrossRef] [PubMed]
- Powell, G.; Tosh, C.R.; Hardie, J. Host plant selection by aphids: Behavioral, evolutionary, and applied perspectives. Annu. Rev. Entomol. 2006, 51, 309–330. [Google Scholar] [CrossRef] [PubMed]
- Goggin, F.L. Plant-aphid interactions: Molecular and ecological perspectives. Curr. Opin. Plant Biol. 2007, 10, 399–408. [Google Scholar] [CrossRef] [PubMed]
- Will, T.; Steckbauer, K.; Hardt, M.; van Bel, A.J.E. Aphid gel saliva: Sheath structure, protein composition and secretory dependence on stylet-tip milieu. PLoS ONE 2012, 7, e46903. [Google Scholar] [CrossRef] [PubMed]
- Jefferson, R.A.; Kavanagh, T.A.; Bevan, M.W. GUS fusions: β-glucuronidase as a sensitive and versatile gene fusion marker in higher plants. EMBO J. 1987, 6, 3901–3907. [Google Scholar] [PubMed]
- Javaid, S.; Amin, I.; Jander, G.; Mukhtar, Z.; Saeed, N.A.; Mansoor, S. A transgenic approach to control hemipteran insects by expressing insecticidal genes under phloem-specific promoters. Sci. Rep. 2016, 6, 34706. [Google Scholar] [CrossRef] [PubMed]
- Saha, P.; Chakraborti, D.; Sarkar, A.; Dutta, I.; Basu, D.; Das, S. Characterization of vascular-specific RSs1 and rolC promoters for their utilization in engineering plants to develop resistance against hemipteran insect pests. Planta 2007, 226, 429–442. [Google Scholar] [CrossRef] [PubMed]
- Will, T.; Vilcinskas, A. The structural sheath protein of aphids is required for phloem feeding. Insect Biochem. Mol. Biol. 2015, 57, 34–40. [Google Scholar] [CrossRef] [PubMed]
- Byeon, J.H.; Seo, E.S.; Lee, J.B.; Lee, M.J.; Kim, J.K.; Yoo, J.W.; Jung, Y.; Lee, B.L. A specific cathepsin-L-like protease purified from an insect midgut shows antibacterial activity against gut symbiotic bacteria. Dev. Comp. Immunol. 2015, 53, 79–84. [Google Scholar] [CrossRef] [PubMed]
- Jaouannet, M.; Morris, J.A.; Hedley, P.E.; Bos, J.I.B. Characterization of Arabidopsis transcriptional responses to different aphid species reveals genes that contribute to host susceptibility and non-host resistance. PLoS Pathog. 2015, 11, e1004918. [Google Scholar] [CrossRef] [PubMed]
- Karimi, M.; Inzé, D.; Depicker, A. GATEWAYTM vectors for Agrobacterium-mediated plant transformation. Trends Plant Sci. 2002, 7, 193–195. [Google Scholar] [CrossRef]
- Karimi, M.; Bleys, A.; Vanderhaeghen, R.; Hilson, P. Building blocks for plant gene assembly. Plant Physiol. 2007, 145, 1183–1191. [Google Scholar] [CrossRef] [PubMed]
- Clough, S.J.; Bent, A.F. Floral dip: A simplified method for Agrobacterium-mediated transformation of Arabidopsis thaliana. Plant J. 1998, 16, 735–743. [Google Scholar] [CrossRef] [PubMed]
- Czechowski, T.; Stitt, M.; Altmann, T.; Udvardi, M.K.; Scheible, W.-R. Genome-wide identification and testing of superior reference genes for transcript normalization in Arabidopsis. Plant Physiol. 2005, 139, 5–17. [Google Scholar] [CrossRef] [PubMed]
- Wyatt, I.J.; White, P.F. Simple estimation of intrinsic increase rates for aphids and tetranychid mites. J. Appl. Ecol. 1977, 14, 757–766. [Google Scholar] [CrossRef]
- Petersen, T.; Brunak, S.; von Heijne, G.; Nielsen, H. SignalP 4.0: Discriminating signal peptides from transmembrane regions. Nat. Methods 2011, 8, 785–786. [Google Scholar] [CrossRef] [PubMed]

| Control (n = 6) | CI2c p:7 (n = 6) | CI2c p:8 (n = 6) | Control (n = 9) | psCI2c p:3 (n = 9) | |
|---|---|---|---|---|---|
| Days to reproduction | 8.7 ± 0.2 | 8.4 ± 0.3 | 8.7 ± 0.2 | 8.7 ± 0.2 | 9.1 ± 0.3 |
| Reproductive life (days) | 23.6 ± 2.0 | 22.7 ± 1.8 | 24.2 ± 1.4 | 24.4 ±1.7 | 25.3 ±1.5 |
| Life span (days) | 42.0 ± 4.0 | 41.1 ± 3.1 | 42.8 ± 2.7 | 49.1 ±3.4 | 46.6 ± 3.9 |
| Nymphs/individual | 63.9 ± 4.4 | 59.3 ± 2.7 | 59.7 ± 3.8 | 49.0 ± 6.1 | 38.9 ± 2.5 |
| Nymphs/day | 2.8 ± 0.1 | 2.8 ± 0.3 | 2.6 ± 0.3 | 2.0 ± 0.3 | 1.6 ± 0.1 |
| rm | 0.31 ± 0.004 | 0.31 ± 0.008 | 0.30 ± 0.010 | 0.26 ± 0.012 | 0.23 ± 0.014 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Losvik, A.; Beste, L.; Mehrabi, S.; Jonsson, L. The Protease Inhibitor CI2c Gene Induced by Bird Cherry-Oat Aphid in Barley Inhibits Green Peach Aphid Fecundity in Transgenic Arabidopsis. Int. J. Mol. Sci. 2017, 18, 1317. https://doi.org/10.3390/ijms18061317
Losvik A, Beste L, Mehrabi S, Jonsson L. The Protease Inhibitor CI2c Gene Induced by Bird Cherry-Oat Aphid in Barley Inhibits Green Peach Aphid Fecundity in Transgenic Arabidopsis. International Journal of Molecular Sciences. 2017; 18(6):1317. https://doi.org/10.3390/ijms18061317
Chicago/Turabian StyleLosvik, Aleksandra, Lisa Beste, Sara Mehrabi, and Lisbeth Jonsson. 2017. "The Protease Inhibitor CI2c Gene Induced by Bird Cherry-Oat Aphid in Barley Inhibits Green Peach Aphid Fecundity in Transgenic Arabidopsis" International Journal of Molecular Sciences 18, no. 6: 1317. https://doi.org/10.3390/ijms18061317




