Efficient Generation of Somatic Cell Nuclear Transfer-Competent Porcine Cells with Mutated Alleles at Multiple Target Loci by Using CRISPR/Cas9 Combined with Targeted Toxin-Based Selection System
Abstract
:1. Introduction
2. Results
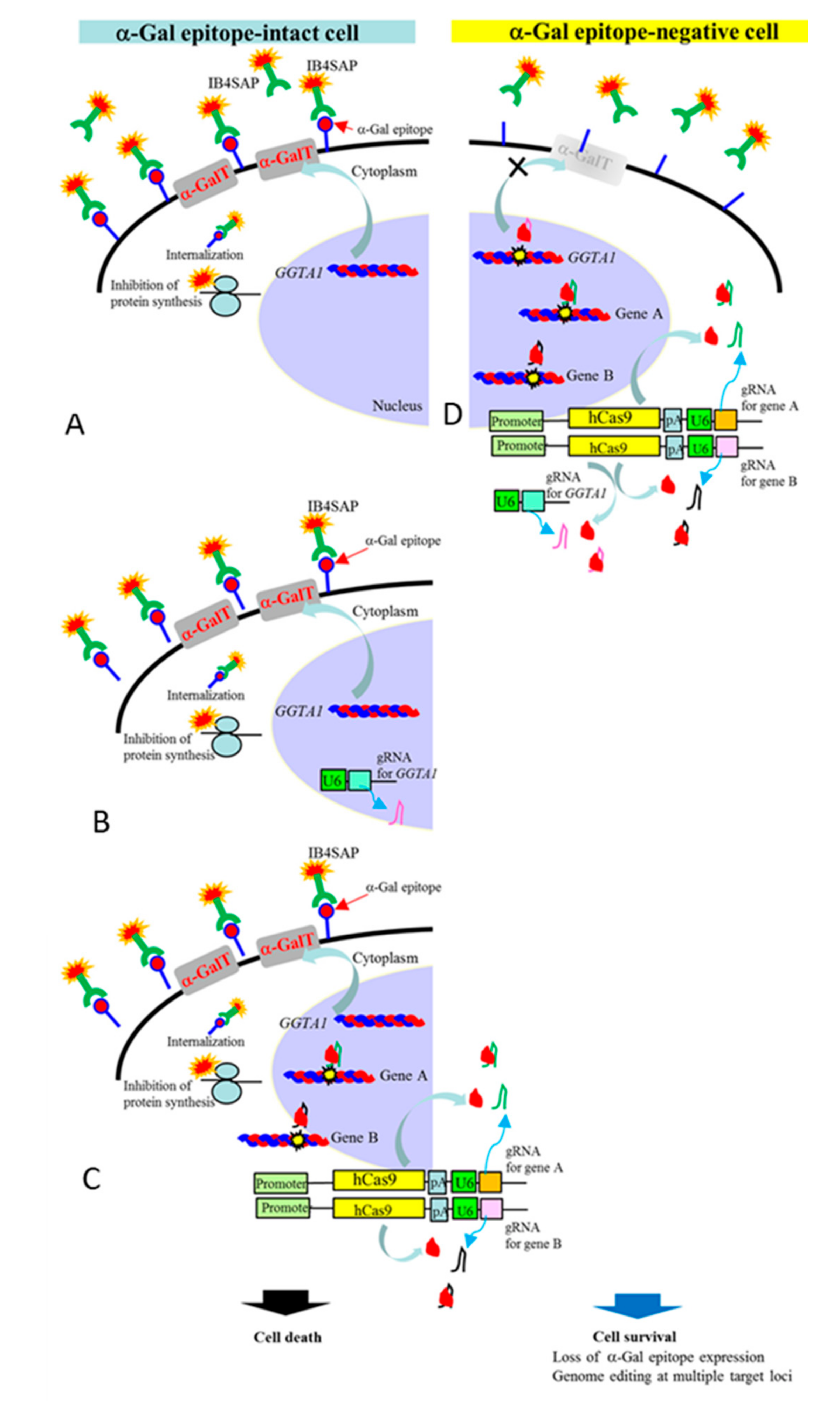
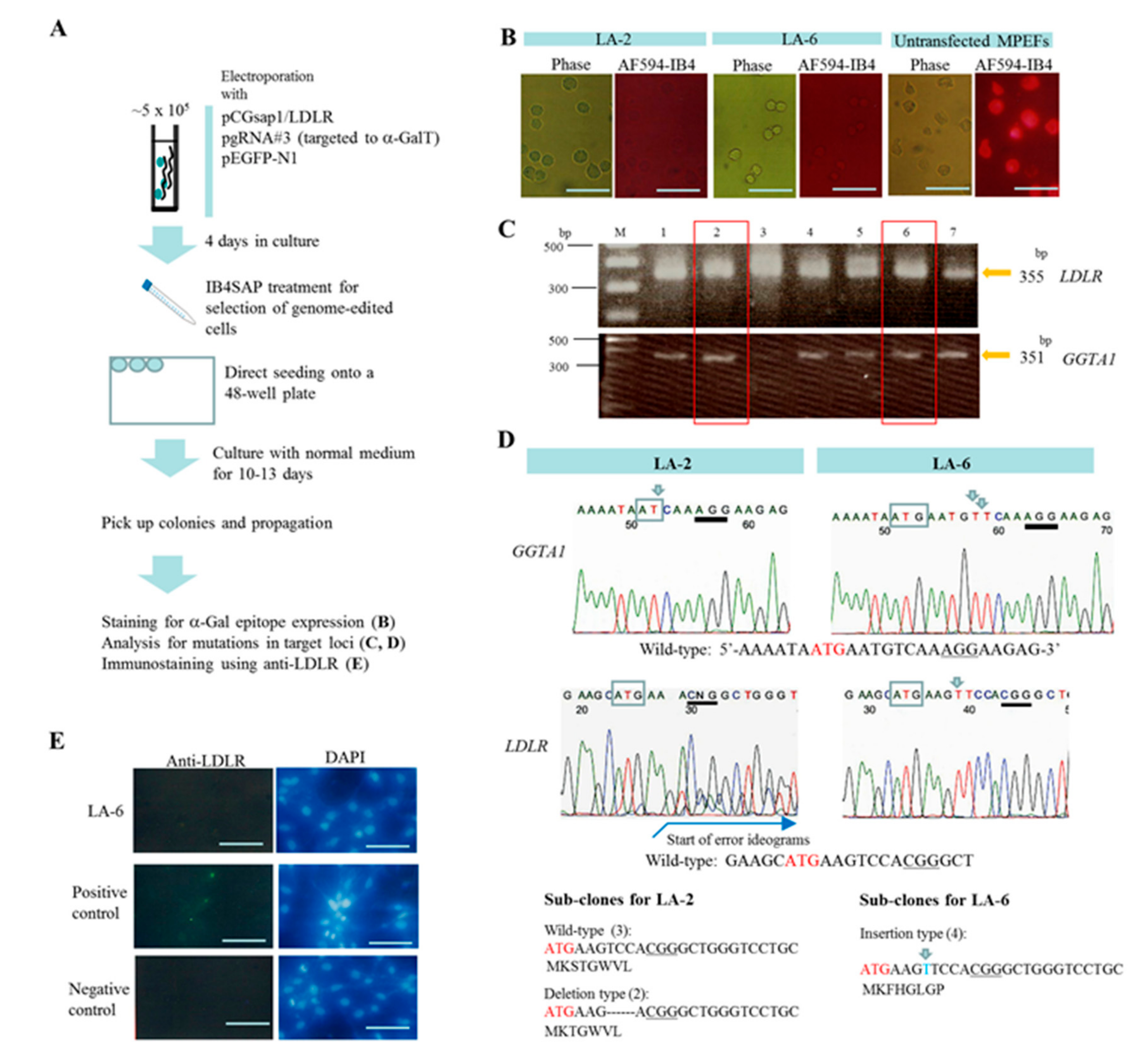
2.1. Experiment 1: Genome Editing of Two Target Loci through One-Shot Transfection and Subsequent Selection with IB4SAP
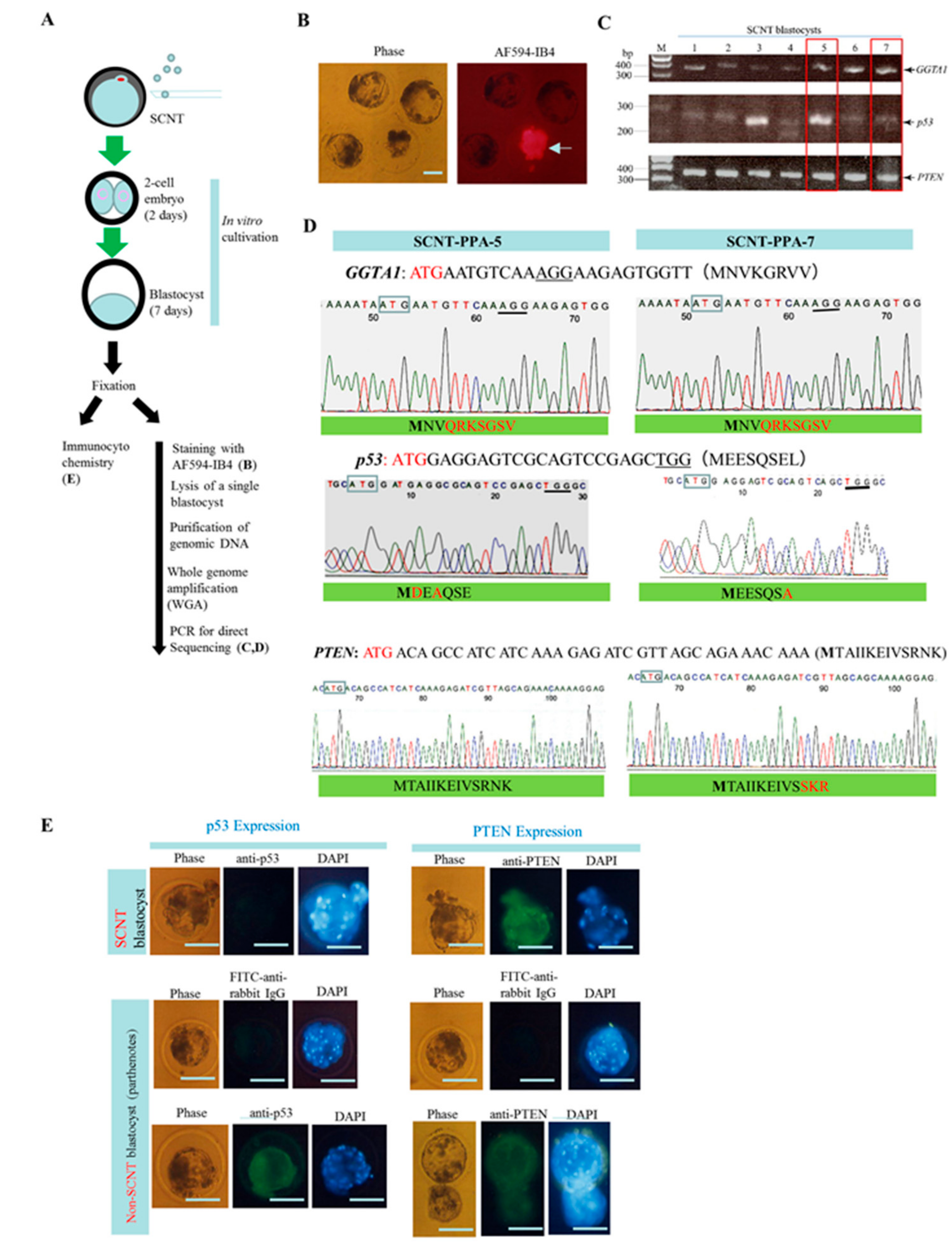
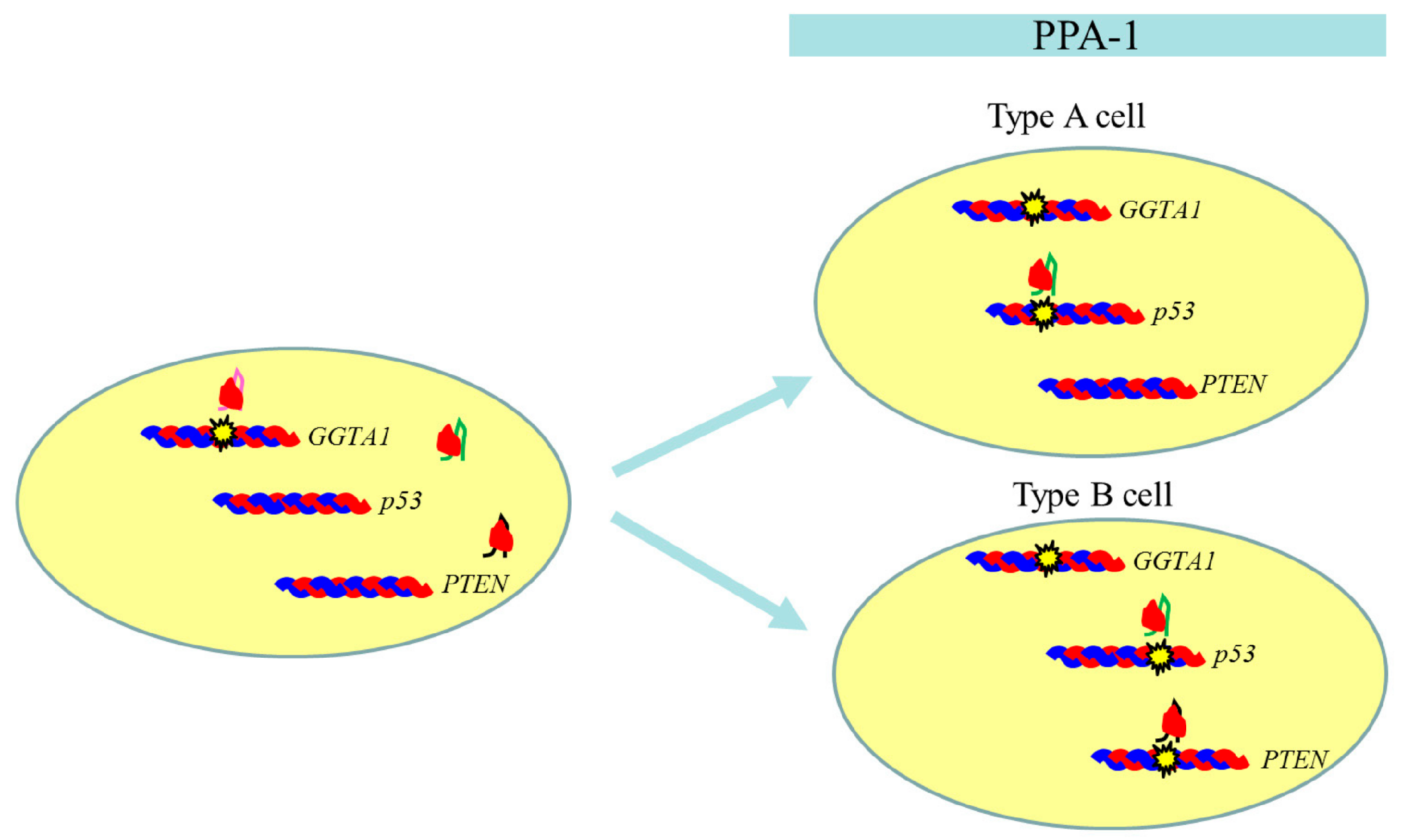
2.2. Experiment 2: Genome Editing of Two Target Loci through One-Shot Transfection and Subsequent Selection with IB4SAP
2.3. Experiment 3: Detecting Possible Off-Target Mutations in the Target Sites of Genome-Edited Cells
3. Discussion
4. Materials and Methods
4.1. Cells
4.2. Construction of pCGsap1-Based Vectors for Simultaneous Expression of hCas9 and sgRNA
4.3. Transfection and Selection of Genome-Edited Clones after Targeted Toxin-Based Selection
4.4. SCNT
4.5. Genomic DNA Isolation, PCR, and Sanger Sequencing of Mutated Sites
4.6. WGA Using A Single Blastocyst
4.7. Screening and Detection of Off-Target Sites
4.8. T7E1-Based Assay and Sequencing
4.9. Staining with AF594-IB4 and Detection of Fluorescence
4.10. Immunocytochemistry
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| AF594-IB4 | Alexa Fluor 594-labeled BS-I-B4 isolectin |
| α-GalT | α-1,3-galactosyltransferase |
| DAPI | 4′,6-Diamidino-2-phenylindole, dihydrochloride |
| D-PBS | Dulbecco’s modified phosphate-buffered saline without Ca2+ and Mg2+, pH 7.4 |
| EGFP | Enhanced green fluorescent protein |
| EndoGalC | Endo-β-galactosidase C |
| FACS | Fluorescence-activated cell sorting |
| FBS | Fetal bovine serum |
| GFP | Green fluorescent protein |
| GM | Genetically modified |
| h | Humanized |
| IB4SAP | Saporin toxin-labeled BS-I-B4 isolectin |
| KO | Knockout |
| LDLR | Low density lipoprotein receptor |
| MPEF | Microminipig embryonic fibroblastic cell |
| NGS | Normal goat serum |
| NHEJ | Non-homologous end joining |
| PEF | Porcine embryonic fibroblastic cell |
| PAM | Protospacer adjacent motif |
| PCR | Polymerase chain reaction |
| PFA | Paraformaldehyde |
| PTEN | Deletion of phosphatase and tensin homolog from chromosome 10 |
| SCNT | Somatic cell nuclear transfer |
| sg | Single guide |
| VPA | Valproic acid |
| WGA | Whole genome amplification |
References
- Hsu, P.D.; Scott, D.A.; Weinstein, J.A.; Ran, F.A.; Konermann, S.; Agarwala, V.; Li, Y.; Fine, E.J.; Wu, X.; Shalem, O.; et al. DNA targeting specificity of RNA-guided Cas9 nucleases. Nat. Biotechnol. 2013, 31, 827–832. [Google Scholar] [CrossRef] [PubMed]
- Harrison, M.M.; Jenkins, B.V.; O’Connor-Giles, K.M.; Wildonger, J. A CRISPR view of development. Genes Dev. 2014, 28, 1859–1872. [Google Scholar] [CrossRef] [PubMed]
- Mali, P.; Yang, L.; Esvelt, K.M.; Aach, J. RNA-guided human genome engineering via Cas9. Science 2013, 339, 823–826. [Google Scholar] [CrossRef] [PubMed]
- Cong, L.; Ran, F.A.; Cox, D.; Lin, S.; Barretto, R.; Habib, N.; Hsu, P.D.; Wu, X.; Jiang, W.; Marraffini, L.A.; et al. Multiplex genome engineering using CRISPR/Cas systems. Science 2013, 339, 819–823. [Google Scholar] [CrossRef] [PubMed]
- Ran, F.A.; Hsu, P.D.; Wright, J.; Agarwala, V.; Scott, D.A.; Zhang, F. Genome engineering using the CRISPR-Cas9 system. Nat. Protoc. 2013, 8, 2281–2308. [Google Scholar] [CrossRef] [PubMed]
- Li, K.; Wang, G.; Andersen, T.; Zhou, P.; Pu, W.T. Optimization of genome engineering approaches with the CRISPR/Cas9 system. PLoS ONE 2014, 9, e105779. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Heo, Y.T.; Quan, X.; Xu, Y.N.; Baek, S.; Choi, H.; Kim, N.H.; Kim, J. CRISPR/Cas9 nuclease-mediated gene knock-in in bovine-induced pluripotent cells. Stem Cells Dev. 2015, 24, 393–402. [Google Scholar] [CrossRef] [PubMed]
- Bauer, D.E.; Canver, M.C.; Orkin, S.H. Generation of genomic deletions in mammalian cell lines via CRISPR/Cas9. J. Vis. Exp. 2015, 95, e52118. [Google Scholar] [CrossRef] [PubMed]
- Ho, T.T.; Zhou, N.; Huang, J.; Koirala, P.; Xu, M.; Fung, R.; Wu, F.; Mo, Y.Y. Targeting non-coding RNAs with the CRISPR/Cas9 system in human cell lines. Nucleic Acids Res. 2015, 43, e17. [Google Scholar] [CrossRef] [PubMed]
- Zhu, Z.; Verma, N.; Gonzalez, F.; Shi, Z.D.; Huangfu, D. CRISPR/Cas-mediated selection-free knockin strategy in human embryonic stem cells. Stem Cell Rep. 2015, 4, 1103–1111. [Google Scholar] [CrossRef] [PubMed]
- Akasaka, E.; Watanabe, S.; Himaki, H.; Ohtsuka, M.; Yoshida, M.; Miyoshi, K.; Sato, M. Enrichment of xenograft-competent genetically modified pig cells using a targeted toxin, isolectin BS-I-B4 conjugate. Xenotransplantation 2010, 17, 81–89. [Google Scholar] [CrossRef] [PubMed]
- Sato, M.; Akasaka, E.; Saitoh, I.; Ohtsuka, M.; Nakamura, S.; Sakurai, T.; Watanabe, S. Targeted toxin-based selectable drug-free enrichment of mammalian cells with high transgene expression. Biology 2013, 2, 341–355. [Google Scholar] [CrossRef] [PubMed]
- Galili, U. The α-gal epitope (Gal α1–3Gal β1–4GlcNAc-R) in xenotransplantation. Biochimie 2001, 83, 557–563. [Google Scholar] [CrossRef]
- Cooper, D.K.; Gollackner, B.; Sachs, D.H. Will the pig solve the transplantation backlog? Ann. Rev. Med. 2002, 53, 133–147. [Google Scholar] [CrossRef] [PubMed]
- Vulchanova, L.; Olson, T.H.; Stone, L.S.; Riedl, M.S.; Elde, R.; Honda, C.N. Cytotoxic targeting of isolectin IB4-binding sensory neurons. Neuroscience 2001, 108, 143–155. [Google Scholar] [CrossRef]
- Ogawa, H.; Muramatsu, H.; Kobayashi, T.; Morozumi, K.; Yokoyama, I.; Kurosawa, N.; Nakao, A.; Muramatsu, T. Molecular cloning of endo-β-galactosidase C and its application in removing α-galactosyl xenoantigen from blood vessels in the pig. J. Biol. Chem. 2000, 275, 19368–19374. [Google Scholar] [CrossRef] [PubMed]
- Yazaki, S.; Iwamoto, M.; Onishi, A.; Miwa, Y.; Suzuki, S.; Fuchimoto, D.; Sembon, S.; Furusawa, T.; Hashimoto, M.; Oishi, T.; et al. Successful cross-breeding of cloned pigs expressing endo-beta-galactosidase C and human decay accelerating factor. Xenotransplantation 2009, 16, 511–521. [Google Scholar] [CrossRef] [PubMed]
- Vaughan, H.A.; Loveland, B.E.; Sandrin, M.S. Gal(α1-3)Gal is the major xenoepitope expressed on pig endothelial cells recognized by naturally occurring cytotoxic human antibodies. Transplantation 1994, 58, 879–882. [Google Scholar] [CrossRef] [PubMed]
- Sato, M.; Miyoshi, K.; Nagao, Y.; Nishi, Y.; Ohtsuka, M.; Nakamura, S.; Sakurai, T.; Watanabe, S. The combinational use of CRISPR/Cas9-based gene editing and targeted toxin technology enables efficient biallelic knockout of the α-1,3-galactosyltransferase gene in porcine embryonic fibroblasts. Xenotransplantation 2014, 21, 291–300. [Google Scholar] [CrossRef] [PubMed]
- Lai, L.; Kolber-Simonds, D.; Park, K.W.; Cheong, H.T.; Greenstein, J.L.; Im, G.S.; Samuel, M.; Bonk, A.; Rieke, A.; Day, B.N.; et al. Production of α-1,3-galactosyltransferase knockout pigs by nuclear transfer cloning. Science 2002, 295, 1089–1092. [Google Scholar] [CrossRef] [PubMed]
- Dai, Y.; Vaught, T.D.; Boone, J.; Chen, S.H.; Phelps, C.J.; Ball, S.; Monahan, J.A.; Jobst, P.M.; McCreath, K.J.; Lamborn, A.E.; et al. Targeted disruption of the α1,3-galactosyltransferase gene in cloned pigs. Nat. Biotechnol. 2002, 20, 251–255. [Google Scholar] [CrossRef] [PubMed]
- Phelps, C.J.; Koike, C.; Vaught, T.D.; Boone, J.; Wells, K.D.; Chen, S.H.; Ball, S.; Specht, S.M.; Polejaeva, I.A.; Monahan, J.A.; et al. Production of α1,3-galactosyltransferase-deficient pigs. Science 2003, 299, 411–414. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Kolber-Simonds, D.; Lai, L.; Watt, S.R.; Denaro, M.; Arn, S.; Augenstein, M.L.; Betthauser, J.; Carter, D.B.; Greenstein, J.L.; Hao, Y.; et al. Production of α-1,3-galactosyltransferase null pigs by means of nuclear transfer with fibroblasts bearing loss of heterozygosity mutations. Proc. Natl. Acad. Sci. USA 2004, 101, 7335–7340. [Google Scholar] [CrossRef] [PubMed]
- Hauschild, J.; Petersen, B.; Santiago, Y.; Queisser, A.L.; Carnwath, J.W.; Lucas-Hahn, A.; Zhang, L.; Meng, X.; Gregory, P.D.; Schwinzer, R.; et al. Efficient generation of a biallelic knockout in pigs using zinc-finger nucleases. Proc. Natl. Acad. Sci. USA 2011, 108, 12013–12017. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.T.; Kwon, D.K.; Park, A.R.; Lee, E.J.; Yun, Y.J.; Ji, D.Y.; Lee, K.; Park, K.W. Production of α1,3-galactosyltransferase targeted pigs using transcription activator-like effector nuclease-mediated genome editing technology. J. Vet. Sci. 2016, 17, 89–96. [Google Scholar] [CrossRef] [PubMed]
- Leren, T.P. Sorting an LDL receptor with bound PCSK9 to intracellular degradation. Atherosclerosis 2014, 237, 76–81. [Google Scholar] [CrossRef] [PubMed]
- Al-Allaf, F.A.; Coutelle, C.; Waddington, S.N.; David, A.L.; Harbottle, R.; Themis, M. LDLR-Gene therapy for familial hypercholesterolaemia: Problems, progress, and perspectives. Int. Arch. Med. 2010, 3, 36. [Google Scholar] [CrossRef] [PubMed]
- Davis, B.T.; Wang, X.J.; Rohret, J.A.; Struzynski, J.T.; Merricks, E.P.; Bellinger, D.A.; Rohret, F.A.; Nichols, T.C.; Rogers, C.S. Targeted disruption of LDLR causes hypercholesterolemia and atherosclerosis in Yucatan miniature pigs. PLoS ONE 2014, 9, e93457. [Google Scholar] [CrossRef] [PubMed]
- Matlashewski, G.; Lamb, P.; Pim, D.; Peacock, J.; Crawford, L.; Benchimol, S. Isolation and characterization of a human p53 cDNA clone: Expression of the human p53 gene. EMBO J. 1984, 3, 3257–3262. [Google Scholar] [PubMed]
- Li, J.; Yen, C.; Liaw, D.; Podsypanina, K.; Bose, S.; Wang, S.I.; Puc, J.; Miliaresis, C.; Rodgers, L.; McCombie, R.; et al. PTEN, a putative protein tyrosine phosphatase gene mutated in human brain, breast, and prostate cancer. Science 1997, 275, 1943–1947. [Google Scholar] [CrossRef] [PubMed]
- Couto, S.S.; Cao, M.; Duarte, P.C.; Banach-Petrosky, W.; Wang, S.; Romanienko, P.; Wu, H.; Cardiff, R.D.; Abate-Shen, C.; Cunha, G.R. Simultaneous haploinsufficiency of Pten and Trp53 tumor suppressor genes accelerates tumorigenesis in a mouse model of prostate cancer. Differentiation 2009, 77, 103–111. [Google Scholar] [CrossRef] [PubMed]
- Carnero, A.; Paramio, J.M. The PTEN/PI3K/AKT Pathway in vivo, Cancer Mouse Models. Front. Oncol. 2014, 4, 252. [Google Scholar] [CrossRef] [PubMed]
- Sakurai, T.; Watanabe, S.; Kamiyoshi, A.; Sato, M.; Shindo, T. A single blastocyst assay optimized for detecting CRISPR/Cas9 system-induced indel mutations in mice. BMC Biotechnol. 2014, 14, 69. [Google Scholar] [CrossRef] [PubMed]
- Akasaka, E.; Ozawa, A.; Mori, H.; Mizobe, Y.; Yoshida, M.; Miyoshi, K.; Sato, M. Whole-genome amplification-based GenomiPhi for multiple genomic analysis of individual early porcine embryos. Theriogenology 2011, 75, 1543–1549. [Google Scholar] [CrossRef] [PubMed]
- Fu, Y.; Foden, J.A.; Khayter, C.; Maeder, M.L.; Reyon, D.; Joung, J.K.; Sander, J.D. High-frequency off-target mutagenesis induced by CRISPR-Cas nucleases in human cells. Nat. Biotechnol. 2013, 31, 822–826. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Pattanayak, V.; Lin, S.; Guilinger, J.P.; Ma, E.; Doudna, J.A.; Liu, D.R. High-throughput profiling of off-target DNA cleavage reveals RNA-programmed Cas9 nuclease specificity. Nat. Biotechnol. 2013, 31, 839–843. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Fujimura, T.; Takahagi, Y.; Shigehisa, T.; Nagashima, H.; Miyagawa, S.; Shirakura, R.; Murakami, H. Production of α1,3-galactosyltransferase gene-deficient pigs by somatic cell nuclear transfer: A novel selection method for gal α1,3-Gal antigen-deficient cells. Mol. Reprod. Dev. 2008, 75, 1372–1378. [Google Scholar] [CrossRef] [PubMed]
- Reyes, L.M.; Estrada, J.L.; Ivary, B.; Sidner, R.A.; Paris, L.L.; Tector, A.J. Efficient selection of Gal-knockout pig cells for somatic cell nuclear transfer. J. Surg. Res. 2013, 184, e37–e42. [Google Scholar] [CrossRef] [PubMed]
- Xin, J.; Yang, H.; Fan, N.; Zhao, B.; Ouyang, Z.; Liu, Z.; Zhao, Y.; Li, X.; Song, J.; Yang, Y.; et al. Highly efficient generation of GGTA1 biallelic knockout inbred mini-pigs with TALENs. PLoS ONE 2013, 8, e84250. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Estrada, J.L.; Burlak, C.; Montgomery, J.; Butler, J.R.; Santos, R.M.; Wang, Z.Y.; Paris, L.L.; Blankenship, R.L.; Downey, S.M.; et al. Efficient generation of genetically distinct pigs in a single pregnancy using multiplexed single-guide RNA and carbohydrate selection. Xenotransplantation 2015, 22, 20–31. [Google Scholar] [CrossRef] [PubMed]
- Cheng, W.; Zhao, H.; Yu, H.; Xin, J.; Wang, J.; Zeng, L.; Yuan, Z.; Qing, Y.; Li, H.; Jia, B.; et al. Efficient generation of GGTA1-null Diannan miniature pigs using TALENs combined with somatic cell nuclear transfer. Reprod. Biol. Endocrinol. 2016, 14, 77. [Google Scholar] [CrossRef] [PubMed]
- Feng, C.; Li, X.; Cui, H.; Long, C.; Liu, X.; Tian, X.; Pan, D.; Luo, Y. Highly efficient generation of GGTA1 knockout pigs using a combination of TALEN mRNA and magnetic beads with somatic cell nuclear transfer. J. Integr. Agric. 2016, 15, 1540–1549. [Google Scholar] [CrossRef]
- Sato, M.; Koriyama, M.; Watanabe, S.; Ohtsuka, M.; Sakurai, T.; Inada, E.; Saitoh, I.; Nakamura, S.; Miyoshi, K. Direct injection of CRISPR/Cas9-related mRNA into cytoplasm of parthenogenetically activated porcine oocytes causes frequent mosaicism for indel mutations. Int. J. Mol. Sci. 2015, 16, 17838–17856. [Google Scholar] [CrossRef] [PubMed]
- Bao, L.; Chen, H.; Jong, U.; Rim, C.; Li, W.; Lin, X.; Zhang, D.; Luo, Q.; Cui, C.; Huang, H.; et al. Generation of GGTA1 biallelic knockout pigs via zinc-finger nucleases and somatic cell nuclear transfer. Sci. China Life Sci. 2014, 57, 263–268. [Google Scholar] [CrossRef] [PubMed]
- Miyoshi, K.; Kawaguchi, H.; Maeda, K.; Sato, M.; Akioka, K.; Noguchi, M.; Horiuchi, M.; Tanimoto, A. Birth of cloned microminipigs derived from somatic cell nuclear transfer embryos that have been transiently treated with valproic acid. Cell. Reprogram. 2016, 18, 390–400. [Google Scholar] [CrossRef] [PubMed]
- Nakayama, A.; Sato, M.; Shinohara, M.; Matsubara, S.; Yokomine, T.; Akasaka, E.; Yoshida, M.; Takao, S. Efficient transfection of primarily cultured porcine embryonic fibroblasts using the Amaxa nucleofection systemTM. Cloning Stem Cells 2007, 9, 523–534. [Google Scholar] [CrossRef] [PubMed]
- Sato, M.; Ohtsuka, M.; Miura, H.; Miyoshi, K.; Watanabe, S. Determination of the optimal concentration of several selective drugs useful for generating multi-transgenic porcine embryonic fibroblasts. Reprod. Domest. Anim. 2012, 47, 759–765. [Google Scholar] [CrossRef] [PubMed]
- Sakurai, T.; Kamiyoshi, A.; Kawate, H.; Mori, C.; Watanabe, S.; Tanaka, M.; Uetake, R.; Sato, M.; Shindo, T. A non-inheritable maternal Cas9-based multiple-gene editing system in mice. Sci. Rep. 2016, 6, 20011. [Google Scholar] [CrossRef] [PubMed]
- Sato, K.; Yoshida, M.; Miyoshi, K. Utility of ultrasound stimulation for activation of pig oocytes matured in vitro. Mol. Reprod. Dev. 2005, 72, 396–403. [Google Scholar] [CrossRef] [PubMed]
- Miyoshi, K.; Sato, K.; Yoshida, M. In vitro development of cloned embryos derived from miniature pig somatic cells after activation by ultrasound stimulation. Cloning Stem Cells 2006, 8, 159–165. [Google Scholar] [CrossRef] [PubMed]
- Miyoshi, K.; Fujimoto, Y.; Mori, H.; Yoshida, M. Activation and parthenogenetic development of pig oocytes exposed to ultrasound in media containing different concentrations of Ca2+. J. Reprod. Dev. 2008, 54, 42–45. [Google Scholar] [CrossRef] [PubMed]
- Himaki, T.; Mizobe, Y.; Tsuda, K.; Suetomo, M.; Yamakuchi, H.; Miyoshi, K.; Takao, S.; Yoshida, M. Effect of postactivation treatment with latrunculin a on in vitro and in vivo development of cloned embryos derived from kidney fibroblasts of an aged Clawn miniature boar. J. Reprod. Dev. 2012, 58, 398–403. [Google Scholar] [CrossRef] [PubMed]

| Oligonucleotides (Oligo) Used for Cloning into SapI Site in pCGSap1 | Target Gene | Reference |
|---|---|---|
| LDLR-oligo: 5′-ACC GGCTG GAAGC ATGAA GTCCA G-3′ 3′-CCGAC CTTCG TACTT CAGGT CAAA-5′ | Porcine LDLR | Sus scrofa breed mixed chromosome 2, Sscrofa10.2 Sequence ID: ref (NC_010444.3) |
| p53-oligo: 5′-ACC GGAGG AGTCG CAGTC CGAGC G-3′ 3′-CCTCC TCAGC GTCAG GCTCG CAAA-5′ | Porcine p53 | Sus scrofa breed mixed chromosome 12, Sscrofa10.2 Sequence ID: ref (NC_010454.3) |
| PTEN-oligo: 5′-ACC AGATC GTTAG CAGAA ACAAA G-3′ 3′-TCTAG CAATC GTCTT TGTTT CAAA-5′ | Porcine PTEN | Sus scrofa breed mixed chromosome 14, Sscrofa10.2 Sequence ID: ref (NC_010456.4) |
| Property | Loci Targeted | LA-1 | LA-2 | LA-3 | Clones LA-4 | LA-5 | LA-6 | LA-7 |
|---|---|---|---|---|---|---|---|---|
| Direct sequencing of PCR products 1 | GGTA1 LDLR | Normal Normal | Bi-allelic Mono-allelic | Normal Normal | Bi-allelic Bi-allelic | Bi-allelic Mono-allelic | Bi-allelic Bi-allelic | Bi-allelic Mono-allelic |
| Expression of α-Gal epitope, as evaluated by cytochemical staining with AF594-IB4 2 | GGTA1 | ++ | − | ++ | − | − | − | − |
| Expression of LDLR, as evaluated by immunocytochemical staining using anti-LDLR 2 | LDLR | + | NT | NT | NT | NT | − | NT |
| Clones | Locus | Genotyping of Clones by Direct Sequencing of PCR Products and Inserts Sub-Cloned into TA Cloning Vector | Expression of α-Gal Epitope, as Evaluated by Cytochemical Staining with AF594-IB4 | Expression of Protein, as Evaluated by Immunocytochemical Staining Using Antibodies |
|---|---|---|---|---|
| PPA-1 | GGTA1 | Deletion type (bi-allelic) | − | |
| PTEN | Mixture of indel type (bi-allelic) and wild-type | No | Yes | |
| p53 | Mixture of deletion (bi-allelic) and replacement (bi-allelic) types | No | ||
| PPA-2 | GGTA1 | Deletion type (bi-allelic) | − | |
| PTEN | Deletion type (mono-allelic) | No | Yes | |
| p53 | Insertion type (bi-allelic) | No |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Sato, M.; Miyoshi, K.; Nakamura, S.; Ohtsuka, M.; Sakurai, T.; Watanabe, S.; Kawaguchi, H.; Tanimoto, A. Efficient Generation of Somatic Cell Nuclear Transfer-Competent Porcine Cells with Mutated Alleles at Multiple Target Loci by Using CRISPR/Cas9 Combined with Targeted Toxin-Based Selection System. Int. J. Mol. Sci. 2017, 18, 2610. https://doi.org/10.3390/ijms18122610
Sato M, Miyoshi K, Nakamura S, Ohtsuka M, Sakurai T, Watanabe S, Kawaguchi H, Tanimoto A. Efficient Generation of Somatic Cell Nuclear Transfer-Competent Porcine Cells with Mutated Alleles at Multiple Target Loci by Using CRISPR/Cas9 Combined with Targeted Toxin-Based Selection System. International Journal of Molecular Sciences. 2017; 18(12):2610. https://doi.org/10.3390/ijms18122610
Chicago/Turabian StyleSato, Masahiro, Kazuchika Miyoshi, Shingo Nakamura, Masato Ohtsuka, Takayuki Sakurai, Satoshi Watanabe, Hiroaki Kawaguchi, and Akihide Tanimoto. 2017. "Efficient Generation of Somatic Cell Nuclear Transfer-Competent Porcine Cells with Mutated Alleles at Multiple Target Loci by Using CRISPR/Cas9 Combined with Targeted Toxin-Based Selection System" International Journal of Molecular Sciences 18, no. 12: 2610. https://doi.org/10.3390/ijms18122610







