Network-Driven Proteogenomics Unveils an Aging-Related Imbalance in the Olfactory IκBα-NFκB p65 Complex Functionality in Tg2576 Alzheimer’s Disease Mouse Model
Abstract
:1. Introduction
2. Results and Discussion
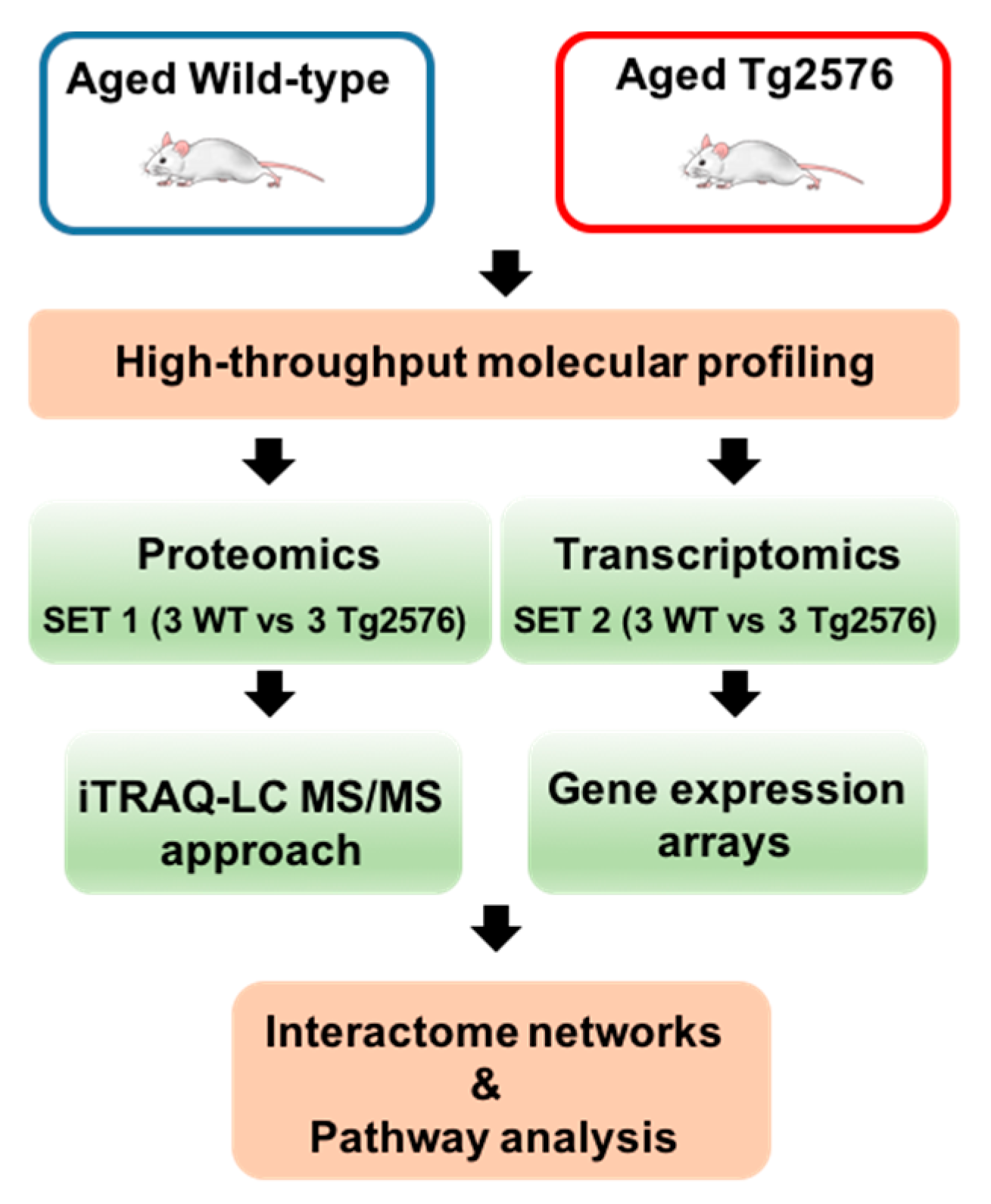
2.1. Molecular Alterations Detected in the Olfactory Bulb (OB) of 18-Month-Old Tg2576 Mice
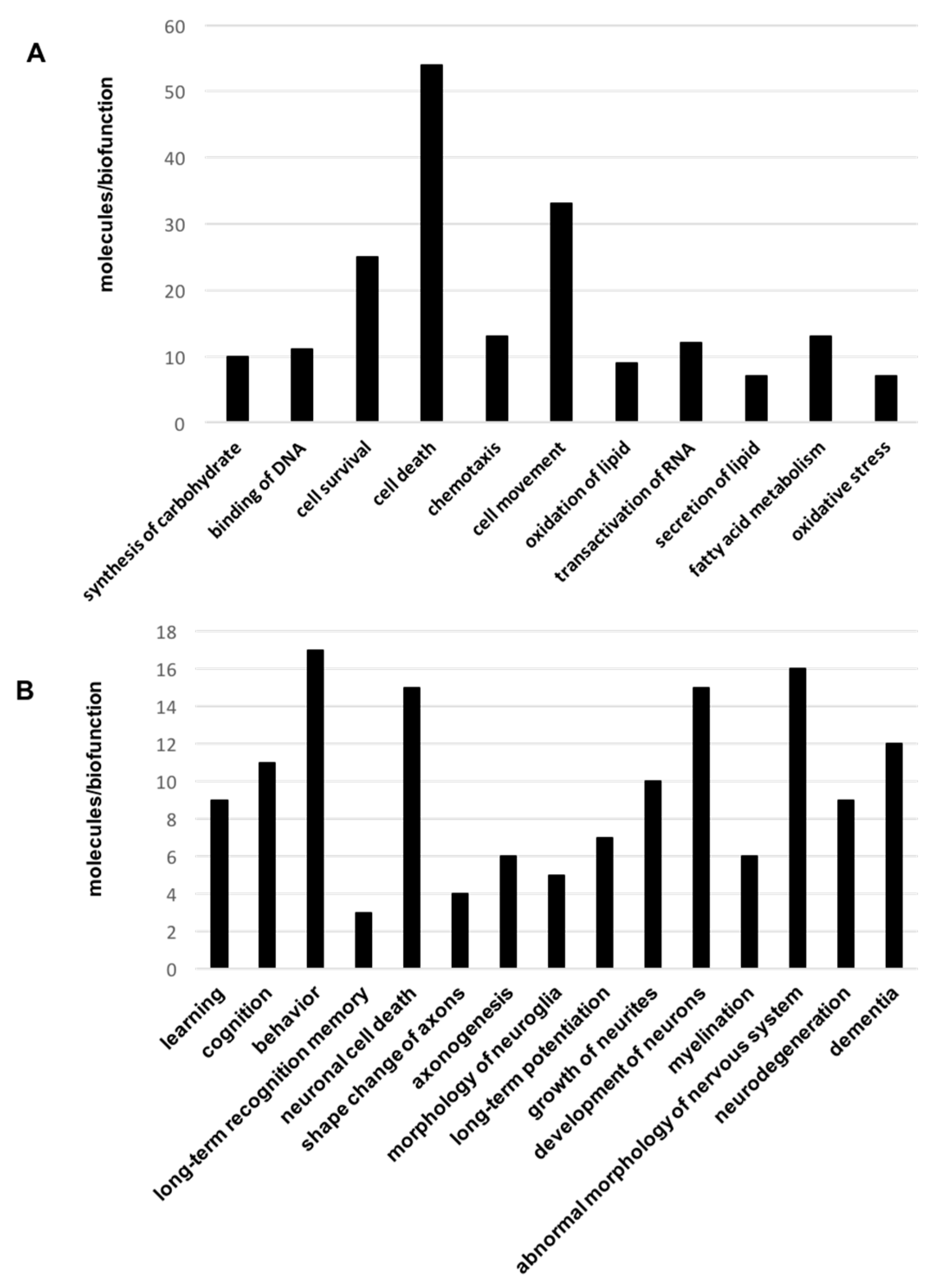
2.2. Biological Functions and Neuronal-Specific Processes Altered in the OB of Aged Tg2576 Mice
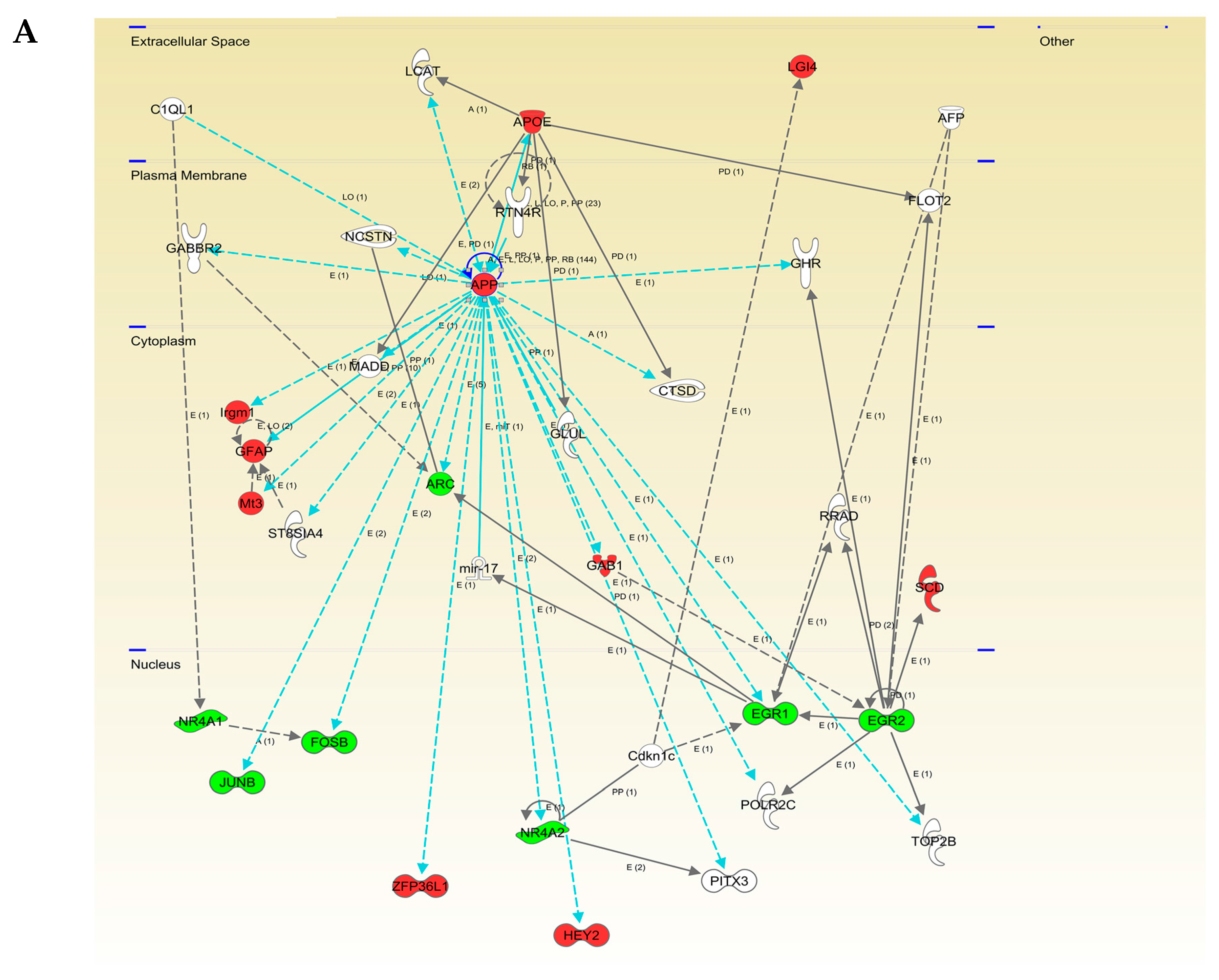
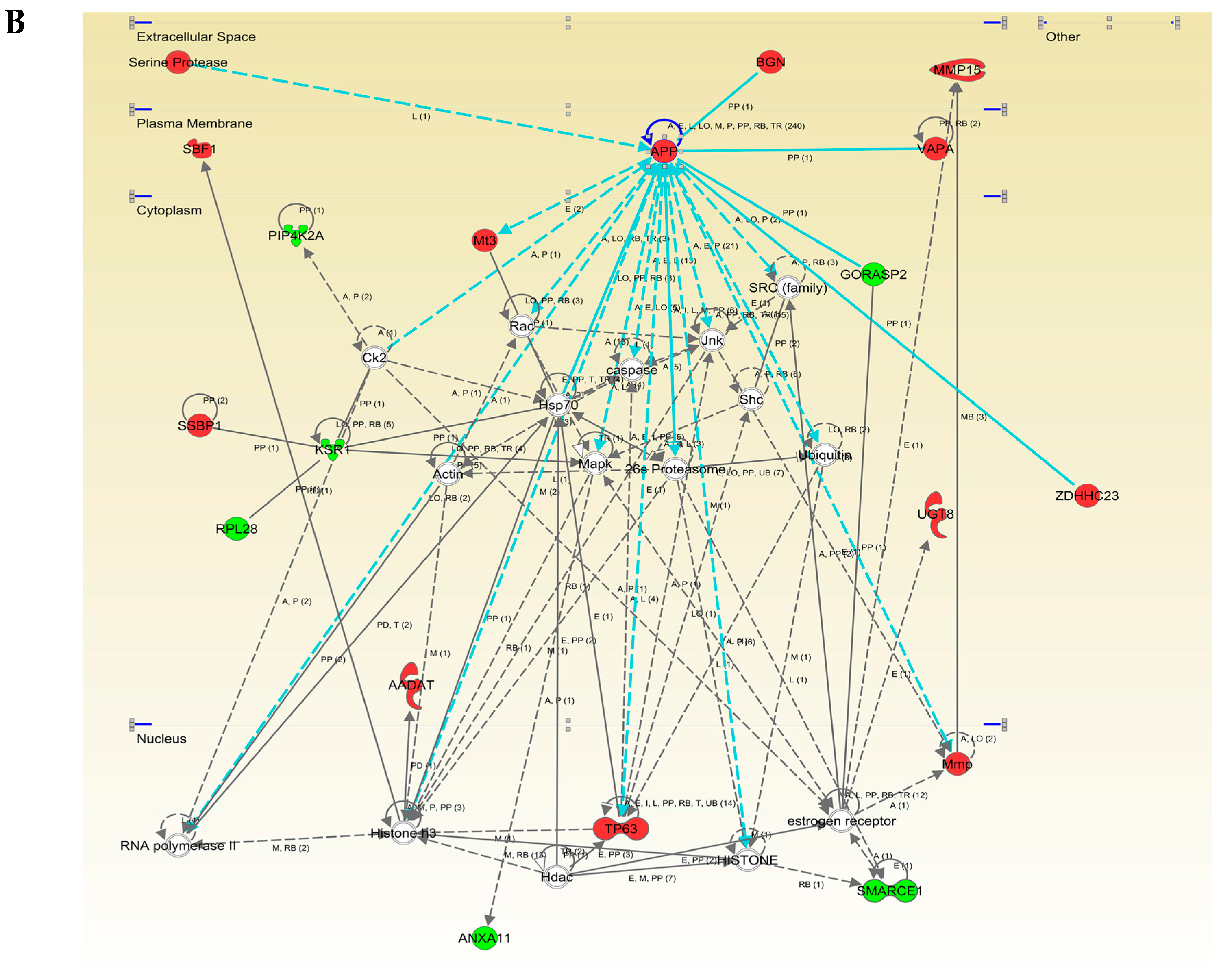
2.3. Functional Interactome of the hAPPSw Isoform at the Olfactory Level: Characterization of Potential Hubs by Network-Driven Proteogenomics
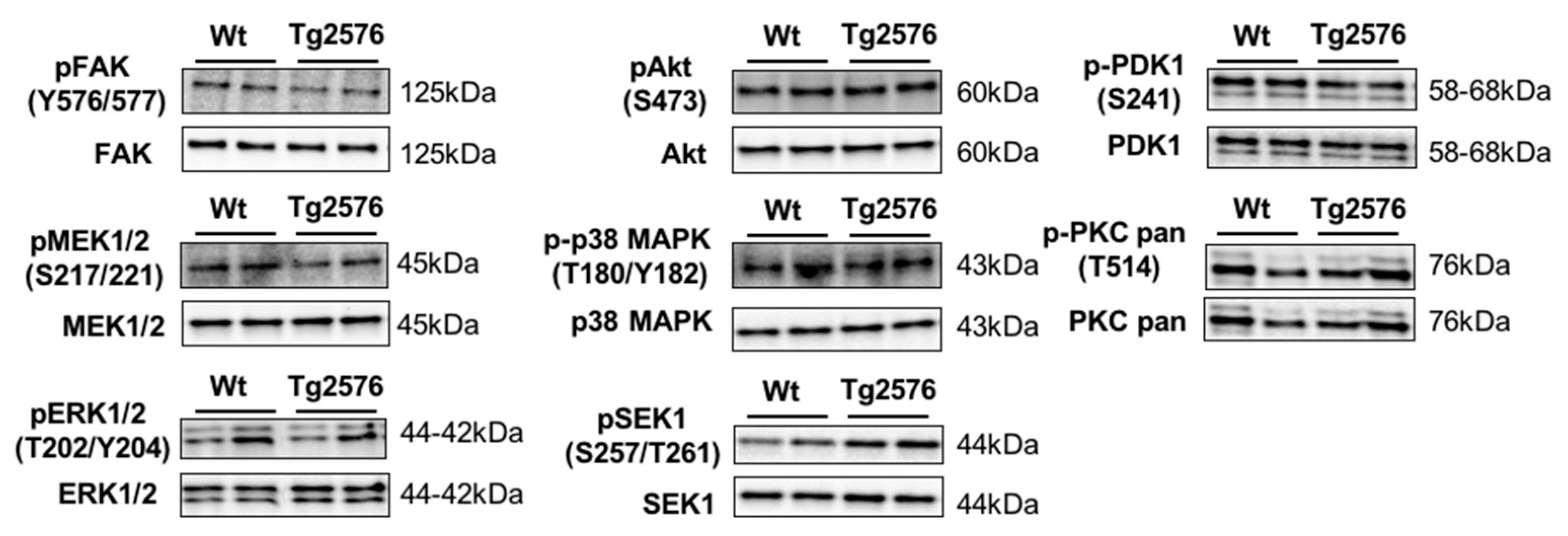
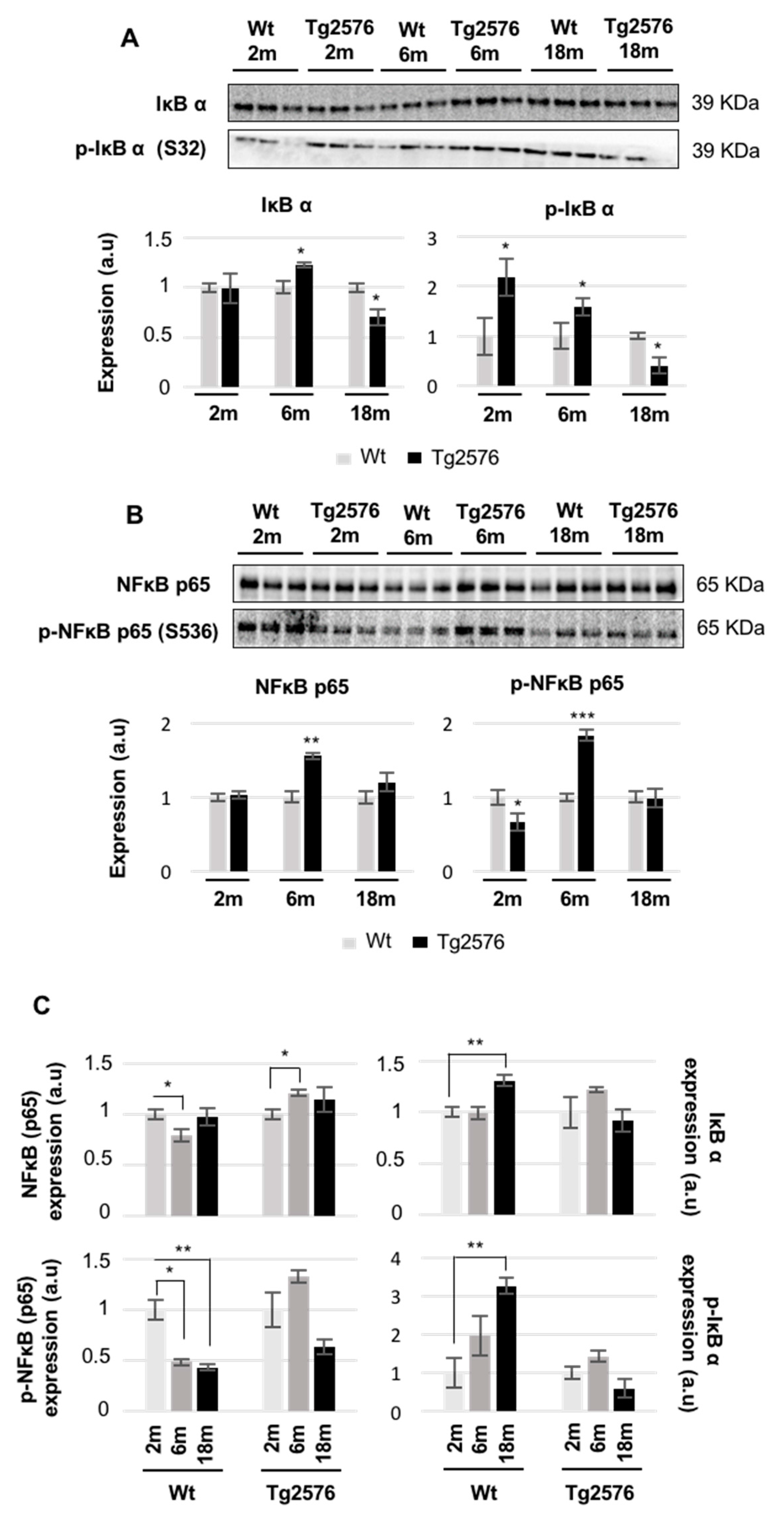
2.4. An Impairment in the Olfactory IκBα-NFκB p65 Complex Functionality during the Progression of Alzheimer’s Disease (AD) in Tg2576 Mice
3. Materials and Methods
3.1. Materials
3.2. Animals
3.3. Olfactory Proteomics
3.4. OB Transcriptomics
3.5. Bioinformatics
3.6. Immunoblotting Analysis
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
Abbreviations
| AD | Alzheimer’s disease |
| APP | Amyloid precursor protein |
| Aβ | β-amyloid |
| ERK | Extracellular signal-regulated kinase |
| FAK | Focal adhesion kinase |
| OB | Olfactory bulb |
| MS/MS | Tandem mass-spectrometry |
| PDK1 | Phosphoinositide-dependent protein kinase 1 |
| PKC | Protein kinase C |
| p38 MAPK | p38 Mitogen-activated protein kinase |
| WT | Wild-type |
References
- Bahar-Fuchs, A.; Chetelat, G.; Villemagne, V.L.; Moss, S.; Pike, K.; Masters, C.L.; Rowe, C.; Savage, G. Olfactory deficits and amyloid-β burden in Alzheimer’s disease, mild cognitive impairment, and healthy aging: A PIB PET study. J. Alzheimers Dis. 2010, 22, 1081–1087. [Google Scholar] [CrossRef] [PubMed]
- Attems, J.; Walker, L.; Jellinger, K.A. Olfactory bulb involvement in neurodegenerative diseases. Acta Neuropathol. 2014, 127, 459–475. [Google Scholar] [CrossRef] [PubMed]
- Hsiao, K.; Chapman, P.; Nilsen, S.; Eckman, C.; Harigaya, Y.; Younkin, S.; Yang, F.; Cole, G. Correlative memory deficits, Aβ elevation, and amyloid plaques in transgenic mice. Science 1996, 274, 99–102. [Google Scholar] [CrossRef] [PubMed]
- Puzzo, D.; Gulisano, W.; Palmeri, A.; Arancio, O. Rodent models for Alzheimer’s disease drug discovery. Expert Opin. Drug Discov. 2015, 10, 703–711. [Google Scholar] [CrossRef] [PubMed]
- Westerman, M.A.; Cooper-Blacketer, D.; Mariash, A.; Kotilinek, L.; Kawarabayashi, T.; Younkin, L.H.; Carlson, G.A.; Younkin, S.G.; Ashe, K.H. The relationship between Aβ and memory in the Tg2576 mouse model of Alzheimer’s disease. J. Neurosci. 2002, 22, 1858–1867. [Google Scholar] [PubMed]
- Janus, C.; Pearson, J.; McLaurin, J.; Mathews, P.M.; Jiang, Y.; Schmidt, S.D.; Chishti, M.A.; Horne, P.; Heslin, D.; French, J.; et al. Aβ peptide immunization reduces behavioural impairment and plaques in a model of Alzheimer’s disease. Nature 2000, 408, 979–982. [Google Scholar] [CrossRef] [PubMed]
- Chen, G.; Chen, K.S.; Knox, J.; Inglis, J.; Bernard, A.; Martin, S.J.; Justice, A.; McConlogue, L.; Games, D.; Freedman, S.B.; et al. A learning deficit related to age and β-amyloid plaques in a mouse model of Alzheimer’s disease. Nature 2000, 408, 975–979. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Jacobsen, J.S.; Wu, C.C.; Redwine, J.M.; Comery, T.A.; Arias, R.; Bowlby, M.; Martone, R.; Morrison, J.H.; Pangalos, M.N.; Reinhart, P.H.; et al. Early-onset behavioral and synaptic deficits in a mouse model of Alzheimer’s disease. Proc. Natl. Acad. Sci. USA 2006, 103, 5161–5166. [Google Scholar] [CrossRef] [PubMed]
- Reddy, P.H.; McWeeney, S.; Park, B.S.; Manczak, M.; Gutala, R.V.; Partovi, D.; Jung, Y.; Yau, V.; Searles, R.; Mori, M.; et al. Gene expression profiles of transcripts in amyloid precursor protein transgenic mice: Up-regulation of mitochondrial metabolism and apoptotic genes is an early cellular change in Alzheimer’s disease. Hum. Mol. Genet. 2004, 13, 1225–1240. [Google Scholar] [CrossRef] [PubMed]
- Lalande, J.; Halley, H.; Balayssac, S.; Gilard, V.; Dejean, S.; Martino, R.; Frances, B.; Lassalle, J.M.; Malet-Martino, M. 1H NMR metabolomic signatures in five brain regions of the AβPPswe Tg2576 mouse model of Alzheimer’s disease at four ages. J. Alzheimers Dis. 2014, 39, 121–143. [Google Scholar] [PubMed]
- Nistico, R.; Ferraina, C.; Marconi, V.; Blandini, F.; Negri, L.; Egebjerg, J.; Feligioni, M. Age-related changes of protein sumoylation balance in the AβPP Tg2576 mouse model of Alzheimer’s disease. Front. Pharmacol. 2014, 5, 63. [Google Scholar] [PubMed]
- Fodero, L.R.; Saez-Valero, J.; McLean, C.A.; Martins, R.N.; Beyreuther, K.; Masters, C.L.; Robertson, T.A.; Small, D.H. Altered glycosylation of acetylcholinesterase in APP (SW) Tg2576 transgenic mice occurs prior to amyloid plaque deposition. J. Neurochem. 2002, 81, 441–448. [Google Scholar] [CrossRef] [PubMed]
- King, D.L.; Arendash, G.W. Behavioral characterization of the Tg2576 transgenic model of Alzheimer’s disease through 19 months. Physiol. Behav. 2002, 75, 627–642. [Google Scholar] [CrossRef]
- Doty, R.L. The olfactory vector hypothesis of neurodegenerative disease: Is it viable? Ann. Neurol. 2008, 63, 7–15. [Google Scholar] [CrossRef] [PubMed]
- Lehman, E.J.; Kulnane, L.S.; Lamb, B.T. Alterations in β-amyloid production and deposition in brain regions of two transgenic models. Neurobiol. Aging 2003, 24, 645–653. [Google Scholar] [CrossRef]
- Guerin, D.; Sacquet, J.; Mandairon, N.; Jourdan, F.; Didier, A. Early locus coeruleus degeneration and olfactory dysfunctions in Tg2576 mice. Neurobiol. Aging 2009, 30, 272–283. [Google Scholar] [CrossRef] [PubMed]
- Young, J.W.; Sharkey, J.; Finlayson, K. Progressive impairment in olfactory working memory in a mouse model of mild cognitive impairment. Neurobiol. Aging 2009, 30, 1430–1443. [Google Scholar] [CrossRef] [PubMed]
- Wesson, D.W.; Levy, E.; Nixon, R.A.; Wilson, D.A. Olfactory dysfunction correlates with amyloid-β burden in an Alzheimer’s disease mouse model. J. Neurosci. 2010, 30, 505–514. [Google Scholar] [CrossRef] [PubMed]
- Wesson, D.W.; Borkowski, A.H.; Landreth, G.E.; Nixon, R.A.; Levy, E.; Wilson, D.A. Sensory network dysfunction, behavioral impairments, and their reversibility in an Alzheimer’s β-amyloidosis mouse model. J. Neurosci. 2011, 31, 15962–15971. [Google Scholar] [CrossRef] [PubMed]
- Fernandez-Irigoyen, J.; Corrales, F.J.; Santamaria, E. Proteomic atlas of the human olfactory bulb. J. Proteom. 2012, 75, 4005–4016. [Google Scholar] [CrossRef] [PubMed]
- Zelaya, M.V.; Perez-Valderrama, E.; de Morentin, X.M.; Tunon, T.; Ferrer, I.; Luquin, M.R.; Fernandez-Irigoyen, J.; Santamaria, E. Olfactory bulb proteome dynamics during the progression of sporadic Alzheimer’s disease: Identification of common and distinct olfactory targets across alzheimer-related co-pathologies. Oncotarget 2015, 6, 39437–39456. [Google Scholar] [CrossRef] [PubMed]
- Lachen-Montes, M.; Gonzalez-Morales, A.; Zelaya, M.V.; Perez-Valderrama, E.; Ausin, K.; Ferrer, I.; Fernandez-Irigoyen, J.; Santamaria, E. Olfactory bulb neuroproteomics reveals a chronological perturbation of survival routes and a disruption of prohibitin complex during Alzheimer’s disease progression. Sci. Rep. 2017, 7, 9115. [Google Scholar] [CrossRef] [PubMed]
- Lachen-Montes, M.; Gonzalez-Morales, A.; de Morentin, X.M.; Perez-Valderrama, E.; Ausin, K.; Zelaya, M.V.; Serna, A.; Aso, E.; Ferrer, I.; Fernandez-Irigoyen, J.; et al. An early dysregulation of FAK and MEK/ERK signaling pathways precedes the β-amyloid deposition in the olfactory bulb of APP/PS1 mouse model of Alzheimer’s disease. J. Proteom. 2016, 148, 149–158. [Google Scholar] [CrossRef] [PubMed]
- Stein, T.D.; Johnson, J.A. Lack of neurodegeneration in transgenic mice overexpressing mutant amyloid precursor protein is associated with increased levels of transthyretin and the activation of cell survival pathways. J. Neurosci. 2002, 22, 7380–7388. [Google Scholar] [PubMed]
- George, A.J.; Gordon, L.; Beissbarth, T.; Koukoulas, I.; Holsinger, R.M.; Perreau, V.; Cappai, R.; Tan, S.S.; Masters, C.L.; Scott, H.S.; et al. A serial analysis of gene expression profile of the Alzheimer’s disease Tg2576 mouse model. Neurotox. Res. 2010, 17, 360–379. [Google Scholar] [CrossRef] [PubMed]
- Tan, L.; Wang, X.; Ni, Z.F.; Zhu, X.; Wu, W.; Zhu, L.Q.; Liu, D. A systematic analysis of genomic changes in Tg2576 mice. Mol. Neurobiol. 2013, 47, 883–891. [Google Scholar] [CrossRef] [PubMed]
- Shin, S.J.; Lee, S.E.; Boo, J.H.; Kim, M.; Yoon, Y.D.; Kim, S.I.; Mook-Jung, I. Profiling proteins related to amyloid deposited brain of Tg2576 mice. Proteomics 2004, 4, 3359–3368. [Google Scholar] [CrossRef] [PubMed]
- Gillardon, F.; Rist, W.; Kussmaul, L.; Vogel, J.; Berg, M.; Danzer, K.; Kraut, N.; Hengerer, B. Proteomic and functional alterations in brain mitochondria from Tg2576 mice occur before amyloid plaque deposition. Proteomics 2007, 7, 605–616. [Google Scholar] [CrossRef] [PubMed]
- Shevchenko, G.; Wetterhall, M.; Bergquist, J.; Hoglund, K.; Andersson, L.I.; Kultima, K. Longitudinal characterization of the brain proteomes for the Tg2576 amyloid mouse model using shotgun based mass spectrometry. J. Proteome Res. 2012, 11, 6159–6174. [Google Scholar] [CrossRef] [PubMed]
- Cuadrado-Tejedor, M.; Cabodevilla, J.F.; Zamarbide, M.; Gomez-Isla, T.; Franco, R.; Perez-Mediavilla, A. Age-related mitochondrial alterations without neuronal loss in the hippocampus of a transgenic model of Alzheimer’s disease. Curr. Alzheimer Res. 2013, 10, 390–405. [Google Scholar] [CrossRef] [PubMed]
- Szklarczyk, D.; Morris, J.H.; Cook, H.; Kuhn, M.; Wyder, S.; Simonovic, M.; Santos, A.; Doncheva, N.T.; Roth, A.; Bork, P.; et al. The string database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 2017, 45, D362–D368. [Google Scholar] [CrossRef] [PubMed]
- Saito, H.; Kubota, M.; Roberts, R.W.; Chi, Q.; Matsunami, H. RTP family members induce functional expression of mammalian odorant receptors. Cell 2004, 119, 679–691. [Google Scholar] [CrossRef] [PubMed]
- Zhang, B.; Gaiteri, C.; Bodea, L.G.; Wang, Z.; McElwee, J.; Podtelezhnikov, A.A.; Zhang, C.; Xie, T.; Tran, L.; Dobrin, R.; et al. Integrated systems approach identifies genetic nodes and networks in late-onset Alzheimer’s disease. Cell 2013, 153, 707–720. [Google Scholar] [CrossRef] [PubMed]
- Ansoleaga, B.; Garcia-Esparcia, P.; Llorens, F.; Moreno, J.; Aso, E.; Ferrer, I. Dysregulation of brain olfactory and taste receptors in AD, PSP and CJD, and AD-related model. Neuroscience 2013, 248, 369–382. [Google Scholar] [CrossRef] [PubMed]
- Lachen-Montes, M.; Zelaya, M.; Segura, V.; Fernández-Irigoyen, J.; Santamaría, E. Progressive modulation of the human olfactory bulb transcriptome during Alzheimer’s disease evolution: Novel insights into the olfactory signaling across proteinopathies. Oncotarget 2017. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Beyer, A.; Aebersold, R. On the dependency of cellular protein levels on mRNA abundance. Cell 2016, 165, 535–550. [Google Scholar] [CrossRef] [PubMed]
- Baker, M.S.; Ahn, S.B.; Mohamedali, A.; Islam, M.T.; Cantor, D.; Verhaert, P.D.; Fanayan, S.; Sharma, S.; Nice, E.C.; Connor, M.; et al. Accelerating the search for the missing proteins in the human proteome. Nat. Commun. 2017, 8, 14271. [Google Scholar] [CrossRef] [PubMed]
- Sakamoto, K.; Karelina, K.; Obrietan, K. CREB: A multifaceted regulator of neuronal plasticity and protection. J. Neurochem. 2011, 116, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Tully, T.; Bourtchouladze, R.; Scott, R.; Tallman, J. Targeting the CREB pathway for memory enhancers. Nat. Rev. Drug Discov. 2003, 2, 267–277. [Google Scholar] [CrossRef] [PubMed]
- Teich, A.F.; Nicholls, R.E.; Puzzo, D.; Fiorito, J.; Purgatorio, R.; Fa, M.; Arancio, O. Synaptic therapy in Alzheimer’s disease: A CREB-centric approach. Neurotherapeutics 2015, 12, 29–41. [Google Scholar] [CrossRef] [PubMed]
- Saura, C.A. CREB-regulated transcription coactivator 1-dependent transcription in Alzheimer’s disease mice. Neurodegener. Dis. 2012, 10, 250–252. [Google Scholar] [CrossRef] [PubMed]
- Saura, C.A.; Valero, J. The role of CREB signaling in Alzheimer’s disease and other cognitive disorders. Rev. Neurosci. 2011, 22, 153–169. [Google Scholar] [CrossRef] [PubMed]
- Kaur, U.; Banerjee, P.; Bir, A.; Sinha, M.; Biswas, A.; Chakrabarti, S. Reactive oxygen species, redox signaling and neuroinflammation in Alzheimer’s disease: The NF-κB connection. Curr. Top. Med. Chem. 2015, 15, 446–457. [Google Scholar] [CrossRef] [PubMed]
- Viatour, P.; Merville, M.P.; Bours, V.; Chariot, A. Phosphorylation of NF-κB and IκB proteins: Implications in cancer and inflammation. Trends Biochem. Sci. 2005, 30, 43–52. [Google Scholar] [CrossRef] [PubMed]
- Kaltschmidt, B.; Uherek, M.; Volk, B.; Baeuerle, P.A.; Kaltschmidt, C. Transcription factor NF-κB is activated in primary neurons by amyloid β peptides and in neurons surrounding early plaques from patients with alzheimer disease. Proc. Natl. Acad. Sci. USA 1997, 94, 2642–2647. [Google Scholar] [CrossRef] [PubMed]
- Terai, K.; Matsuo, A.; McGeer, P.L. Enhancement of immunoreactivity for NF-kappa B in the hippocampal formation and cerebral cortex of Alzheimer’s disease. Brain Res. 1996, 735, 159–168. [Google Scholar] [CrossRef]
- Ferrer, I.; Marti, E.; Lopez, E.; Tortosa, A. NF-κB immunoreactivity is observed in association with β A4 diffuse plaques in patients with Alzheimer’s disease. Neuropathol. Appl. Neurobiol. 1998, 24, 271–277. [Google Scholar] [CrossRef] [PubMed]
- Paris, D.; Patel, N.; Quadros, A.; Linan, M.; Bakshi, P.; Ait-Ghezala, G.; Mullan, M. Inhibition of Aβ production by NF-κB inhibitors. Neurosci. Lett. 2007, 415, 11–16. [Google Scholar] [CrossRef] [PubMed]
- Yoo, S.J.; Lee, J.H.; Kim, S.Y.; Son, G.; Kim, J.Y.; Cho, B.; Yu, S.W.; Chang, K.A.; Suh, Y.H.; Moon, C. Differential spatial expression of peripheral olfactory neuron-derived BACE1 induces olfactory impairment by region-specific accumulation of β-amyloid oligomer. Cell Death Dis. 2017, 8, e2977. [Google Scholar] [CrossRef] [PubMed]
- Nagayama, S.; Homma, R.; Imamura, F. Neuronal organization of olfactory bulb circuits. Front. Neural Circuits 2014, 8, 98. [Google Scholar] [CrossRef] [PubMed]
- Cuadrado-Tejedor, M.; Garcia-Osta, A. Current animal models of Alzheimer’s disease: Challenges in translational research. Front. Neurol. 2014, 5, 182. [Google Scholar] [CrossRef] [PubMed]
- Tyanova, S.; Temu, T.; Cox, J. The maxquant computational platform for mass spectrometry-based shotgun proteomics. Nat. Protoc. 2016, 11, 2301–2319. [Google Scholar] [CrossRef] [PubMed]
- Tyanova, S.; Temu, T.; Sinitcyn, P.; Carlson, A.; Hein, M.Y.; Geiger, T.; Mann, M.; Cox, J. The perseus computational platform for comprehensive analysis of (prote)omics data. Nat. Methods 2016, 13, 731–740. [Google Scholar] [CrossRef] [PubMed]
- Vizcaino, J.A.; Deutsch, E.W.; Wang, R.; Csordas, A.; Reisinger, F.; Rios, D.; Dianes, J.A.; Sun, Z.; Farrah, T.; Bandeira, N.; et al. Proteomexchange provides globally coordinated proteomics data submission and dissemination. Nat. Biotechnol. 2014, 32, 223–226. [Google Scholar] [CrossRef] [PubMed]
- Irizarry, R.A.; Bolstad, B.M.; Collin, F.; Cope, L.M.; Hobbs, B.; Speed, T.P. Summaries of Affymetrix GeneChip probe level data. Nucleic Acids Res. 2003, 31, e15. [Google Scholar] [CrossRef] [PubMed]
- Silva, A.R.; Grinberg, L.T.; Farfel, J.M.; Diniz, B.S.; Lima, L.A.; Silva, P.J.; Ferretti, R.E.; Rocha, R.M.; Filho, W.J.; Carraro, D.M.; et al. Transcriptional alterations related to neuropathology and clinical manifestation of Alzheimer’s disease. PLoS ONE 2012, 7, e48751. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Cuadrado-Tejedor, M.; Garcia-Barroso, C.; Sanchez-Arias, J.A.; Rabal, O.; Perez-Gonzalez, M.; Mederos, S.; Ugarte, A.; Franco, R.; Segura, V.; Perea, G.; et al. A first-in-class small-molecule that acts as a dual inhibitor of HDAC and PDE5 and that rescues hippocampal synaptic impairment in Alzheimer’s disease mice. Neuropsychopharmacology 2016, 42, 524–539. [Google Scholar] [CrossRef] [PubMed]



| Upstream Regulator | Molecule Type | p-Value | Target Molecules in OB Omics Dataset |
|---|---|---|---|
| CREB1 | transcription regulator | 0.00000209 | APOE, ARC, EGR1, EGR2, FOSB, GNAL, HLA-A, JUNB, NR4A1, NR4A2, NR4A3, RBP4, SCD, ZDHHC23 |
| NFKBIA | transcription regulator | 0.00916 | BCL2A1, CD82, HLA-A, JUNB, MMP15, NID1, NR4A1 |
| NFKB1 | transcription regulator | 0.0000748 | APOE, APP, BCL2A1, CD82, EGR1, HLA-DMB, NR4A1, TBX21 |
| NFκB (complex) | complex | 0.00174 | APOE, APP, BCL2A1, CD82, EGR1, GFAP, HLA-A, HLA-DMB, JUNB, KLF3 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Palomino-Alonso, M.; Lachén-Montes, M.; González-Morales, A.; Ausín, K.; Pérez-Mediavilla, A.; Fernández-Irigoyen, J.; Santamaría, E. Network-Driven Proteogenomics Unveils an Aging-Related Imbalance in the Olfactory IκBα-NFκB p65 Complex Functionality in Tg2576 Alzheimer’s Disease Mouse Model. Int. J. Mol. Sci. 2017, 18, 2260. https://doi.org/10.3390/ijms18112260
Palomino-Alonso M, Lachén-Montes M, González-Morales A, Ausín K, Pérez-Mediavilla A, Fernández-Irigoyen J, Santamaría E. Network-Driven Proteogenomics Unveils an Aging-Related Imbalance in the Olfactory IκBα-NFκB p65 Complex Functionality in Tg2576 Alzheimer’s Disease Mouse Model. International Journal of Molecular Sciences. 2017; 18(11):2260. https://doi.org/10.3390/ijms18112260
Chicago/Turabian StylePalomino-Alonso, Maialen, Mercedes Lachén-Montes, Andrea González-Morales, Karina Ausín, Alberto Pérez-Mediavilla, Joaquín Fernández-Irigoyen, and Enrique Santamaría. 2017. "Network-Driven Proteogenomics Unveils an Aging-Related Imbalance in the Olfactory IκBα-NFκB p65 Complex Functionality in Tg2576 Alzheimer’s Disease Mouse Model" International Journal of Molecular Sciences 18, no. 11: 2260. https://doi.org/10.3390/ijms18112260







