1. Introduction
Autoimmune thyroid diseases (AITDs) are a group of organ-specific and polygenic inherited autoimmune diseases, with an estimated prevalence of up to 1%–5% of the general population [
1]. AITDs mainly consist of two clinical subtypes of Graves’ disease (GD) and Hashimoto’s thyroiditis (HT). GD is predominantly characterized by a variable combination of hyperthyroidism, diffused goiter and high level of thyroid stimulating hormone receptor antibody (TRAb). Meanwhile, some GD patients may present extrathyroidal manifestations, including ophthalmopathy, pretibial myxedema and clubbed fingers. Clinical features of HT include the presence of antibody against thyroid peroxidase (TPOAb) or thyroglobulin (TgAb). Additionally, some patients with HT harbor extensive apoptosis of thyrocytes leading to hypothyroidism. Although there are some common characteristics in GD and HT, such as destruction of thyroid tissue and the existence of circulating thyroid autoantibodies including TRAb, TPOAb and TgAb, the clinical presentations and mechanisms of the two subtypes are different from each other to some extent; for example, our previous studies found that GD, HT or even Graves’ ophthalmopathy (GO) have specific genetic backgrounds [
2,
3]. The pathogenesis of AITDs remains unclear, although there is much evidence demonstrating that the interaction between genetic factors and environmental components may be involved in their etiology [
4,
5].
More recently, an increasing body of research has confirmed that several specific genes are associated with multiple autoimmune diseases [
6,
7], implicating that many autoimmune diseases may share some genetic risk factors. For instance, TNFAIP3 has been identified to be related to the genetic etiology of systemic lupus erythematosus (SLE) [
8], rheumatoid arthritis (RA) [
9], systemic sclerosis (SSc) [
10]. Additionally, we also found the relationship between this gene and GD [
11]. All these data documented that variants in several genes probably contribute to dysregulation of common immune pathways, and then are involved in the pathological procedure of diverse autoimmune diseases.
The tumor necrosis factor superfamily member 4 (
TNFSF4) gene encodes a cytokine (OX40L), which is expressed on antigen-presenting cells (APCs) to provide co-stimulatory signals to T cells. In recent years, the
TNFSF4 gene polymorphisms have been reported to be an important predisposition factor to SLE [
12], RA [
6], SSc [
13] and primary Sjogren’s syndrome (pSS) [
14]. However, to date, whether
TNFSF4 gene variations are associated with AITDs has not been investigated.
In the present study, we evaluated whether mutations in TNFSF4 gene are genetically predisposed in Han Chinese populations to AITDs via a case-control study. Single nucleotide polymorphisms (SNPs) tagging four independent susceptibility loci were genotyped in a large cohort of AITDs patients and normal healthy controls. We also analyzed the association between each polymorphism locus and the predisposition to different subtypes of AITDs, including GD, HT and ophthalmopathy.
3. Discussion
The
TNFSF4 gene, also known as the OX40 ligand (OX40L), encodes the OX40L protein which is a co-stimulatory cytokine and belongs to the TNF ligand family. The protein mainly participates in the interaction of T-cell and antigen-presenting cell (APC), T-cell activation and B-cell differentiation, providing CD28-independent co-stimulatory signals for activated CD4
+ T cells [
15].
TNFSF4, located in chromosome 1 (1q25), contains three exons and two introns (in NCBI database). Previous studies have shown that polymorphisms of
TNFSF4 can confer risk to diverse autoimmune diseases, such as SLE, RA, SSc and pSS, but it remains unknown whether genetic mutations of
TNFSF4 region may induce occurrence of AITDs, which attracts our interest.
AITDs are also regarded as autoimmune diseases targeting the thyroid with a complex genetic and environmental etiology, manifesting mainly as GD and HT. It is notable that genetic factors play a prominent role in the occurrence and persistence of AITDs. Given that autoimmune diseases may share a common genetic predisposition, and that immune dysregulation plays a vital role in AITDs [
16,
17], we hypothesized that variants within the
TNFSF4 gene, which is a crucial immune regulator, could also elicit abnormal OX40L expression and dysfunction, thus affecting T-cell activation and leading to unbalanced immune regulation and its resultant occurrence of AITDs.
In the present work, we observed the association between four loci of
TNFSF4 gene and AITDs patients in the Han Chinese population. We found that the frequency of genotype GG in rs3850641 was slightly lower in AITDs patients, probably suggesting it could decrease susceptibility to AITDs. In addition, frequencies of GG genotype in rs3850641 and TT genotype in rs7514229 also decreased in AITDs subjects, confirming that variant genotype GG in rs3860541 was indeed a factor protecting people from AITDs, as was variant genotype TT in rs7514229. Our results suggested polymorphisms in the
TNFSF4 gene region, one SNP in 3′UTR (rs7514229) and two intronic SNPs (rs3860541 and rs1234313), may be associated with AITDs susceptibility. To our knowledge, variants in the intron of a gene may influence its expression and regulate its function [
18], 3′UTR polymorphisms in the gene region are of important regulation function. We therefore speculated that the molecular action underlying genetic pathology of AITDs is that
TNFSF4 SNPs may affect the expression of
TNFSF4 gene and down-regulate T-cell activation, which requires further in-depth research to confirm.
Further, to investigate the association between genotype and clinical manifestations, we carried out the clinical sub-phenotype analysis. The AITDs occurrence in teenagers (≤18 young patients) may be due to their genetic family history of this disease [
19,
20], which corresponded with our results showing the frequency of family history was much higher in AITDs patients with disease-onset age ≤18 years old. Meanwhile, marginally significant differences in frequencies of rs7514229 genotype TT and disease-onset age were found between AITDs patients with disease-onset age ≤18 years old and AITDs patients with disease-onset age ≥19 years old. Similar correlations between gene mutations and disease-onset age were reported in RA [
21], type 1 diabetes [
22] and multiple sclerosis [
23]. Furthermore, we revealed that frequency of GG genotype of rs3850641 declined slightly in GD subgroup of AITDs, although without significance. Nevertheless, we observed that
TNFSF4 SNPs were not associated with AITDs patients with ophthalmopathy or family history. Several studies provided clues that thyroid-associated ophthalmopathy (TAO) was correlated with the impact of environmental elements, especially current smoking history [
24,
25]. Recent studies are suggesting that genetic markers also affect the susceptibility of TAO [
26], including genetic variants in the STAT3 [
27], TSHR [
28] and HLA-DR3 [
29] regions. However, our results cannot add the
TNFSF4 gene to the list of the predisposition of thyroid-associated ophthalmopathy (TAO). Moreover, allele A from rs3850641 was associated with the decreased risk for the HT subgroup of hypothyroidism by 41.5%. In HT, hypothyroidism is more associated with a family history of thyroid dysfunction [
20]. Our study showed HT hypothyroidism patients with higher ratio of family history, which was consistent with the previous research [
20]. To our best knowledge, we were the first to find that genetic factors are also involved in etiology of hypothyroidism in HT. Why do these SNPs not show their susceptibility to GD or TAO? It is possible that thyroid eye disease or TAO is a different disease than Graves’ disease and Hashimoto’s thyroiditis. In addition, a recent paper found that polymorphisms in calsequestrin (CASQ1) are correlated with HT and GO, but not Graves’ hyperthyroiditis (GH) [
30]. Interestingly, our study found SNPs in
TNFSF4 are associated with hypothyroidism of Hashimoto’s thyroiditis, but not thyroid orbitopathy or GD. These two studies do not show contradictory results, and illustrate the complexity of the diseases, GD, HT and TAO or GO. For instance, our previous studies indeed found
UBE2L3 and
CLEC16A gene polymorphisms to be associated with susceptibility to HT rather than GD and TAO or GO [
2,
3]. Obviously, the genetic mechanisms of these diseases are still unclear, so more research is needed to reveal the pathomechanism of thyroid ophthalmopathy.
Overall, we provided the first evidence for genetic association between four susceptibility loci in the TNFSF4 gene in Chinese AITDs patients, with samples exclusively from the Han Chinese population. Nevertheless, considering the validation of a convincing association and discovery of population differences, the importance of replication studies in some different populations should not be overlooked. The statistical power calculated in this research was very strong (larger than 0.8) to detect the association, and it has adequately reached a significant result. Simultaneously, the sample size in this study was large enough with 1,048 cases and 909 controls to effectively reduce the type of errors (type 1 error and type 2 error).
4. Materials and Methods
4.1. Subjects
A total of 1,048 Chinese patients with AITDs (693 GD and 355 HT) and 909 healthy Chinese controls were recruited. All AITDs patients were enrolled from the Out-Patient Department of Endocrinology of Jinshan Hospital of Fudan University. Ethnically and geographically matched and unrelated healthy controls were recruited from the Healthy Check-Up Center of the same hospital.
All AITDs patients were diagnosed as previously described [
2,
27]. GD patients were diagnosed based on their clinical manifestations and biochemical assessments of hyperthyroidism and the positive circulating TRAb, with or without positive TPOAb or TgAb and diffusive goiter of the thyroid. HT was defined based on the high level of either TPOAb or TgAb, with or without clinical and biochemical hypothyroidism and the presence of an enlarged thyroid. A minority of HT patients were further confirmed by fine needle aspiration biopsies. All the control subjects showed negative thyroid antibodies against TPO. In the current study, TPOAb, TgAb and TRAb were detected with highly specific and sensitive immunochemiluminescence kits from Roche Company (Shanghai, China).
All the subjects, including AITDs patients and controls, were ethnic Han Chinese. Written informed consent was obtained from all participants and the research was approved by the Ethics Committee of Jinshan Hospital of Fudan University (JYLL-2014-06, 2014/2/21), respectively.
4.2. DNA Sample Preparation
Genomic DNA were extracted from 2 mL of peripheral venous blood from each subject using RelaxGene Blood DNA System (Tiangen Biotech Company, Beijing, China), according to the manufacturer’s protocol. The concentration and A260/A280 ratio of all DNA samples were measured by NANO DROP 2000 Spectrophotometer (Thermo Scientific Company, Waltham, MA, USA). Finally, the DNA samples with great purity and concentration were used for next genotyping.
4.3. Single Nucleotide Polymorphism (SNP) Selection and Genotyping
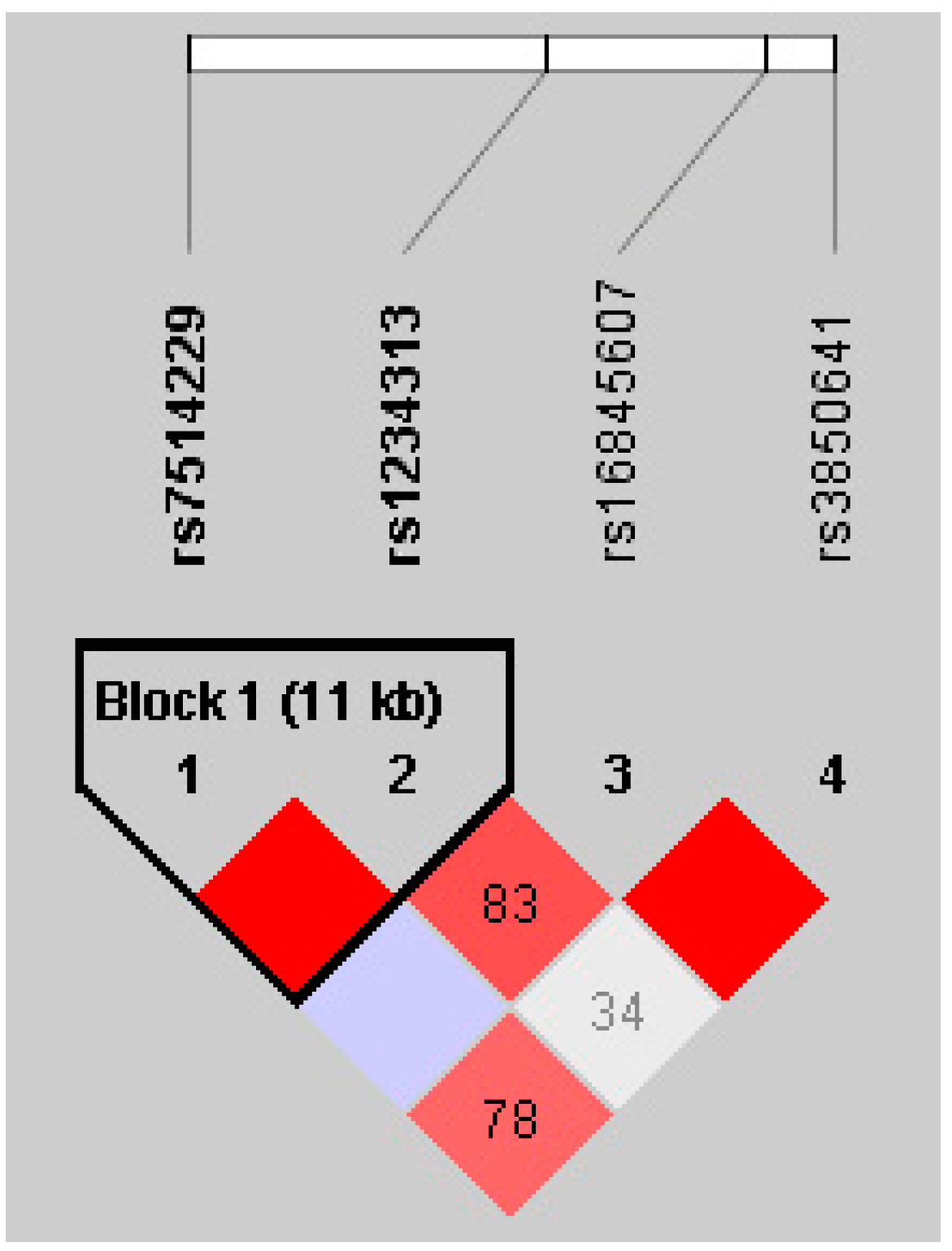
Marker-tagging SNPs were chosen from the Hapmap CHB data using the Tagger programme of Haploview software (Whitehead Institute for Biomedical Research, MIT Media Lab, and Broad Institute of Harvard and MIT) to satisfy the following criteria: minor allele frequency (MAF) >0.1, Hardy-Weinberg equilibrium (HWE) with p > 0.001 and logarithm of odds (LOD) >3.0. For the TNFSF4 gene of 23 kb with 42 SNPs in Hapmap CHB population, we selected four loci covering the whole region of the TNFSF4 gene to capture all the most common variants. Four tag SNPs were selected including rs7514229 located in the 3′ untranslated region (UTR), as well as rs1234313, rs16845607 and rs3850641 in intron 1 of the TNFSF4 region.
Genotyping of the four SNPs was undertaken using the ligase detection reaction (LDR) platform according to the manufacturer’s instructions. Moreover, to ensure detection quality, each reaction was performed in duplicate, and blank samples without DNA were used as negative controls. Furthermore, only SNPs and samples that passed the 95% quality control threshold were subjected to further statistical analysis and SNPs with allele frequencies not meeting Hardy-Weinberg equilibrium (HWE) were removed from the next analysis. The primers specific to the four SNPs at the TNFSF4 loci are “rs7514229” forward-GATAACACAGAATCATCCAG and reverse-TTGTAGCACATGTTTCCCTG; “rs1234313” forward-ATCTAACACTGGCTCTAGTC and reverse-GCCATTCTGACTAGAATAGG; “rs16845607” forward-AGATATAGCTACCAAGCTCC and reverse-GATGAGAAAACAGAGGCTAC; “rs3850641” forward-GCTGTCACTTTGAAGCTTTG and reverse-TGCCTGATCAAACACATTAC.
4.4. Clinical Sub-Phenotype Analysis
Clinical sub-phenotype stratification analysis was conducted using a case-only approach, in which basic allelic and genotypic examination was performed by comparing minor allele and genotype frequency of cases with a specific sub-phenotype to the whole case group. The clinical sub-phenotypes include: (1) the age of disease onset (≤18 years old versus ≥19 years old); (2) presence or absence of ophthalmopathy which was defined as a distinctive disorder characterized by inflammation and swelling of the extraocular muscles, eyelid retraction, periorbital edema, episcleral vascular injection, conjunctive swelling and proptosis; (3) presence or absence of hypothyroidism in HT patients; and (4) presence or absence of AITDs family history, which was defined as the subjects’ first-degree relatives including parents, children and siblings or second-degree relatives such as grandparents, uncles and aunts who had AITDs.
4.5. Statistical Analysis
Clinical data were described as M ± SD (mean ± standard deviation). Hardy-Weinberg equilibrium (HWE) concordance test in the controls and patient samples, linkage disequilibrium (LD) test and haplotype frequency calculation were performed using HaploView 4.2 (Whitehead Institute for Biomedical Research, MIT Media Lab, and Broad Institute of Harvard and MIT). In order to analyze whether the four predisposing loci are associated with AITDs, allele and genotype frequencies were compared between AITDs cases and healthy controls using the Chi-square test (χ2-test) or Fisher’s exact test. LD among the selected SNPs was measured using the pairwise LD measures D’ and r2. All data were statistically calculated with the SPSS 18.0 software (International Business Machines Corporation, Armonk, NY, USA). A p value of less than 0.05 was considered statistically significant. Odds ratio (OR) and 95% confidence interval (95% CI) were applied to assess the association between each genotype and AITDs.
4.6. Power Calculation
Power calculations for AITDs in this research considered allele frequency of SNPs from 0.05 to 0.5, a population prevalence of 1%–5% for AITDs, and OR of 0.2–0.5 at a 0.05 significant level. As a result, this study had sufficient power (larger than 0.8) to detect the association of OR of 0.2 or above with 1,048 cases and 909 controls.