Genomic Resources of Three Pulsatilla Species Reveal Evolutionary Hotspots, Species-Specific Sites and Variable Plastid Structure in the Family Ranunculaceae
Abstract
:1. Introduction
2. Results and Discussion
2.1. Size and Structure of Pulsatilla Plastid Genomes
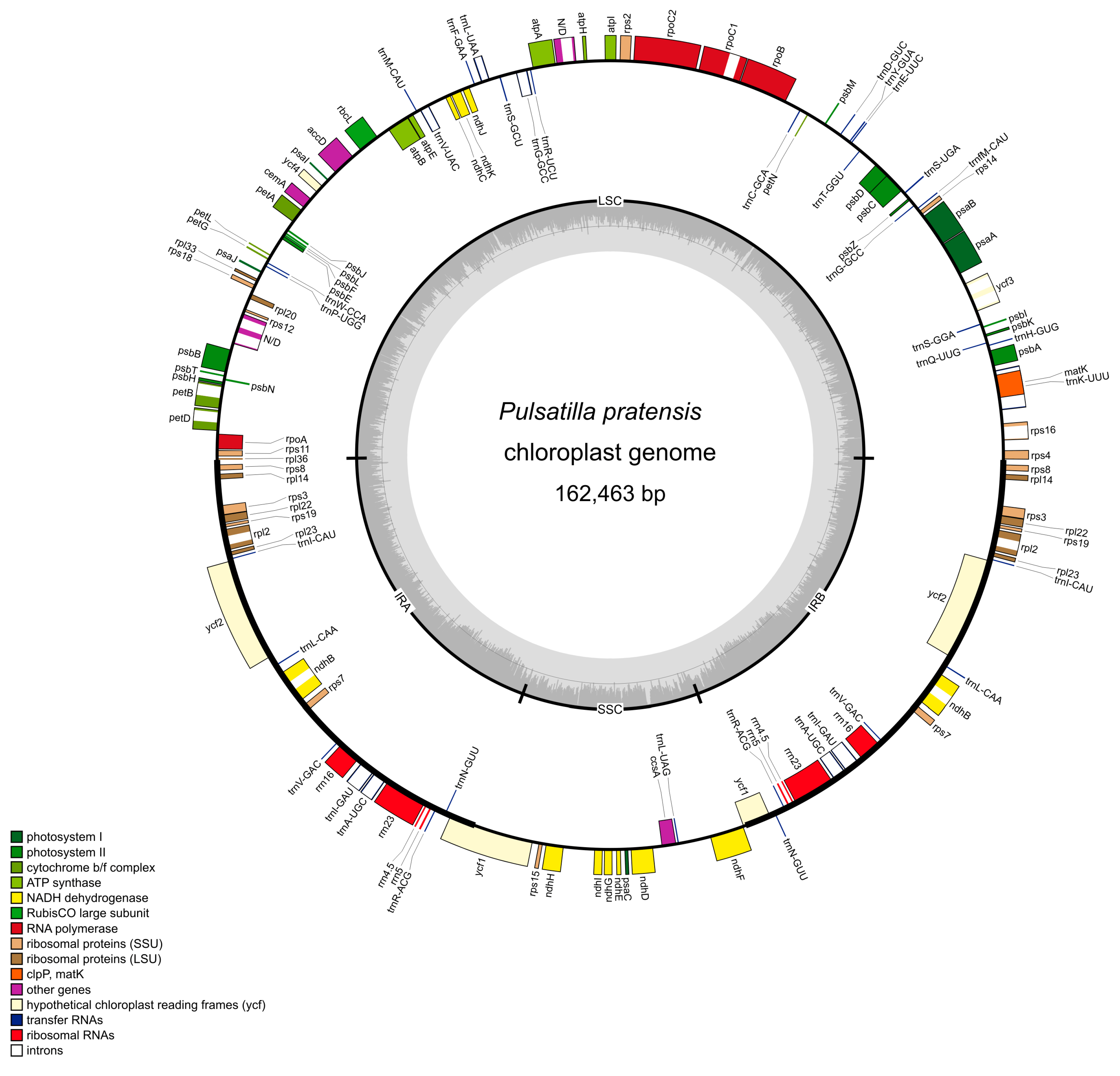
2.2. Variation in the Plastid Genome Structure of the Family Ranunculaceae

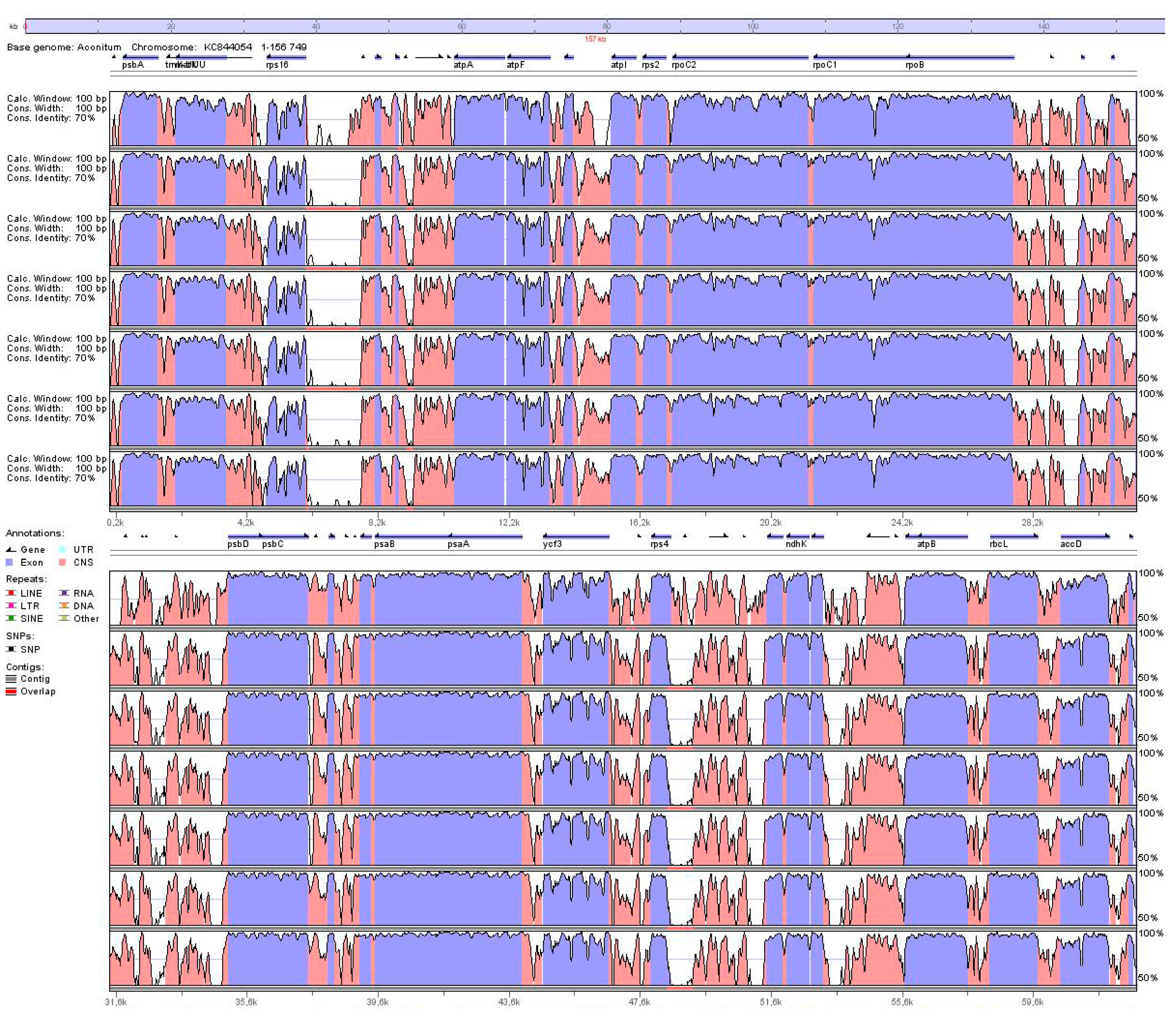

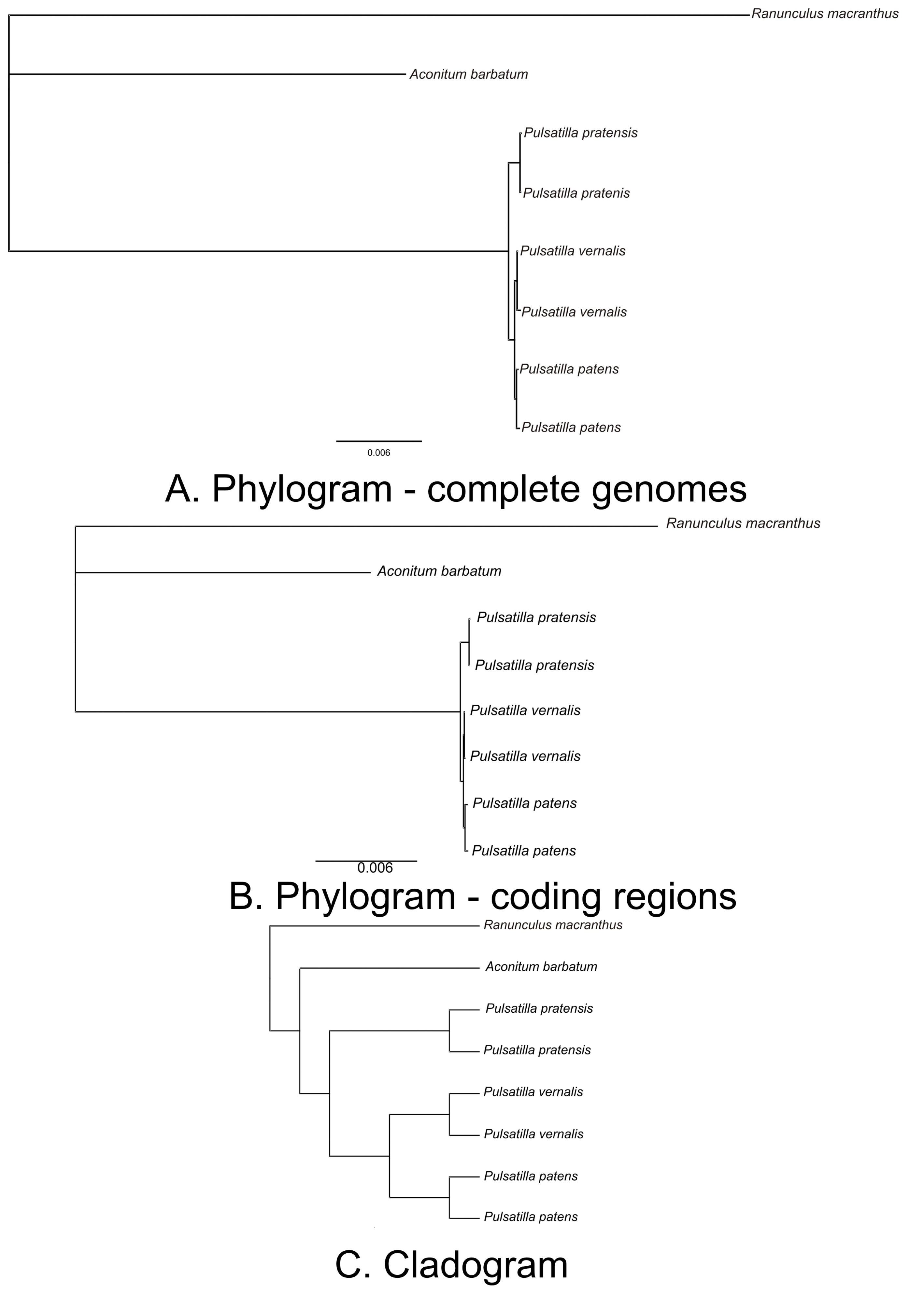
2.3. Identification of the Most Variable Regions of the Pulsatilla Plastid Genome
2.4. Species-Specific Mutations: Barcoding of Closely-Related Pulsatilla Species
2.5. Mutations in Protein Coding Regions (CDS)
2.6. The Characteristics of the Nuclear rRNA Cluster and Its Usefulness in the Studies on Hybridization

3. Experimental Section
3.1. DNA Extraction, Sequencing, Genome Assembly and Annotation
3.2. Selection of Plastid Regions as Markers for Evolutionary Studies of Pulsatilla
3.3. Resolving Phylogenetic Relationships
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Tamura, M. Ranunculaceae. In The Families and Genera of Vascular Plants; Kubitzki, K., Rohwer, J.G., Bittrich, V., Eds.; Springer-Verlag: Berlin, Germany, 1993; Volume 2, pp. 563–583. [Google Scholar]
- Wang, W.C. Ranunculaceae. In Flora of China; Science Press (in Chinese): Beijing, China, 1979; Volume 27, 28. [Google Scholar]
- Xu, Q.; Shu, Z.; He, W.; Chen, L.; Yang, S.; Yang, G. Antitumor activity of Pulsatilla chinensis (Bunge) Regel saponins in human liver tumor 7402 cells in vitro and in vivo. Phytomedicine 2012, 19, 293–300. [Google Scholar] [CrossRef] [PubMed]
- Martin, M.L.; San Roman, L.; Dominguez, A. In vitro activity of protoanemonin on antifungal agent. Planta Med. 1990, 56, 66–69. [Google Scholar] [CrossRef] [PubMed]
- Campbell, W.; Cragg, G.M.L.; Powrie, A.H. Anemonin, protoanemonin and ranunculin from Knowltonia capensis. Phytochemistry 1979, 18, 323–334. [Google Scholar] [CrossRef]
- Akeroyd, J.R. Pulsatilla Miller. Flora Eur. 1993, 1, 264–266. [Google Scholar]
- Uotila, P. Pulsatilla patens × vernalis Suomessa. Memo. Soc. Fauna Flora Fenn. 1980, 56, 111–118. [Google Scholar]
- Uotila, P. Decline of Anemone patens (Ranunculaceae) in Finland. Acta Univ. Ups. Symb. Bot. Ups. 1996, 31, 205–210. [Google Scholar]
- Wójtowicz, W. Pulsatilla patens (L.) Mill. Sasanka otwarta. In Poradnik ochrony siedlisk i Gatunków Natura 2000—Podręcznik Metodyczny, Gatunki roślin (in Polish); Werblan-Jakubiec, H., Sudnik-Wójcikowska, B., Eds.; GIOS: Warszawa, Poland, 2004; pp. 168–171. [Google Scholar]
- Bock, J.H.; Peterson, S.J. Reproductive biology of Pulsatilla patens (Ranunculaceae). Am. Mid. Nat. 1975, 94, 476–478. [Google Scholar] [CrossRef]
- Ordway, E. The phenology and pollination biology of Anemone patens (Ranunculaceae) in western Minesota. In The Prairie: Past, Present and Future: Proceedings of the Ninth North American Prairie Conference; Clambey, G.K., Pemble, R., Eds.; University of Wisconsin–Madison Press: Moorhead, MN, USA, 1986. [Google Scholar]
- Pilt, I.; Kukk, Ü. Pulsatilla patens and Pulsatilla pratensis (Ranunculaceae) in Estonia: Distribution and ecology. Proc. Estonian Acad. Sci. Biol. Ecol. 2002, 51, 242–256. [Google Scholar]
- Kalliovirta, M.; Ryttäri, T.; Heikkinen, R.K. Population structure of a threatened plant, Pulsatilla patens in boreal forests: Modelling relationships to overgrowth and site closure. Biodivers. Conserv. 2006, 15, 3095–3108. [Google Scholar] [CrossRef]
- Grzyl, A.; Kiedrzyński, M.; Zielińska, K.M.; Rewicz, A. The relationschip between climatic conditions and generative reproduction of the lowland population of Pulsatilla vernalis the last breath of a relict plant fluctuating cycle of regeneration. Plant Ecol. 2014, 215, 457–466. [Google Scholar] [CrossRef]
- Priede, G.; Klavina, D. In vitro cultivation and root initiation of the endangered plant Pulsatilla patens. Environ. Exp. Biol. 2011, 9, 71–74. [Google Scholar]
- Sauliene, I.; Brinkyte, E. The impact of phytohormones on pasgueflower (Pulsatilla) regeneration in vitro. Acta Biol. Univ. Daugavp. 2009, 9, 249–254. [Google Scholar]
- Šedivá, J. In vitro seed propagation of some Pulsatilla species (Pulsatilla L.). Acta Pruhaniciana 2002, 73, 48–51. [Google Scholar]
- Šedivá, J.; Žlebčík, J. Summary of findings from a propagation and ex situ conservation of Pulsatilla vernalis, P. pratensis ssp. bohemica, P. patens and P. grandis. Acta Pruhoniciana 2012, 100, 155–160. [Google Scholar]
- Danova, K.; Bertoli, A.; Pistelli, L.; Dimitrov, D.; Pistelli, L. In vitro culture of Balkan endemic and rare Pulsatilla species for conservational purposes and secondary metabolites production. Bot. Serbica 2009, 33, 157–162. [Google Scholar]
- Naumovski, D.; Radič, S.; Pevalek-Kozlina, B. In vitro micropropagation of Pulsatilla pratensis (L.) Miller ssp. nigricans (Störck) Zämelis. Propag. Ornam. Plants 2009, 9, 16–20. [Google Scholar]
- Bang, S.C.; Kim, Y.; Lee, J.H.; Ahn, B.Z. Triterpenoid saponins from the roots of Pulsatilla koreana. J. Nat. Prod. 2005, 68, 268–272. [Google Scholar] [CrossRef] [PubMed]
- Lee, S.H.; Lee, E.; Ko, Y.T. Anti-inflammatory effects of a methanol extract from Pulsatilla koreana in lipopolysaccharide-exposed rats. BMP Rep. 2012, 45, 371–376. [Google Scholar] [CrossRef]
- Hantula, J.; Uotila, P.; Saura, A.; Lokki, J. Chloroplast DNA variation in Anemone s. lato (Ranunculaceae). Plant Syst. Evol. 1989, 163, 81–85. [Google Scholar] [CrossRef]
- Hoot, S.B. Phylogeny of the Ranunculaceae based on epidermal microcharacters and macromorphology. Syst. Bot. 1991, 16, 741–755. [Google Scholar]
- Hoot, S.B. Phylogeny of the Ranunculaceae based on preliminary atpB, rbcL and 18S nuclear ribosomal DNA sequence data. Plant Syst. Evol. 1995, 9, 241–251. [Google Scholar]
- Horandl, E.; Paun, O.; Johanson, J.T.; Lehnebach, C.; Armstrong, T.; Chen, L.; Lockhart, P. Phylogenetic relationships and evolutionary traits in Ranunculus s.l. (Ranunculaceae) inferred from ITS sequence analysis. Mol. Phyl. Evol. 2005, 36, 305–327. [Google Scholar] [CrossRef] [PubMed]
- Tamura, M. Angiospermae ordnung Ranunculales Fam. Ranunculaceae. II. Systematic Part. In Natürliche Pflanzenfamilien. 17aIV; Hiepko, P., Ed.; Duncker & Humblot: Berlin, Germany, 1995; pp. 223–519. [Google Scholar]
- Ehrendorfer, F.; Samuel, R. Contributions to molecular systematic and phylogeny of Anemone and related genera (Ranunculaceae-Anemoninae). Acta Phytotaxon. Sin. 2001, 39, 293–307. [Google Scholar]
- Hoot, S.B.; Meyer, K.M.; Manning, J.C. Phylogeny and reclassification of Anemone (Ranunculaceae), with an emphasis on austral species. Syst. Bot. 2012, 37, 139–152. [Google Scholar]
- Hensen, I.; Obeprieler, C.; Wesche, K. Genetic structure, population size, and seed production of Pulsatilla vulgaris Mill. (Ranunculaceae) in Central Germany. Flora 2005, 200, 3–14. [Google Scholar] [CrossRef]
- Ronikier, M.; Costa, A.; Aguilar, J.F.; Feliner, G.N.; Küpfer, P.; Mirek, Z. Phylogeography of Pulsatilla vernalis (L.) Mill. (Ranunculaceae): Chloroplast DNA reveals two evolutionary lineages across central Europe and Scandinavia. J. Biogeogr. 2008, 35, 1650–1664. [Google Scholar] [CrossRef]
- Hedrick, P.W.; Miller, P.S. Conservation genetics: Techniques and fundamentals. Ecol. Appl. 1992, 2, 30–46. [Google Scholar] [CrossRef]
- Ouborg, N.J.; Piquot, Y.; von Groenendael, M. Population genetic, molecular markers ant the study dispersal in plants. J. Ecol. 1999, 87, 5551–5568. [Google Scholar] [CrossRef]
- Provan, J.; Powell, W.; Hollingsworth, P.M. Chloroplast microsatellites: New tools for studies in plant ecology and evolution. Trends Ecol. Evol. 2001, 16, 142–147. [Google Scholar] [CrossRef]
- Chiang, T-Y.; Chiang, Y-C.; Chen, Y-J.; Chou, C-H.; Havanond, S.; Hong, T-N.; Huang, S. Phylogeography of Kandelia candel in East Asiatic mangroves based on nucleotide variation of chloroplast and mitochondrial DNAs. Mol. Ecol. 2001, 10, 2697–2710. [Google Scholar] [CrossRef] [PubMed]
- Kanno, M.; Yokoyama, J.; Suyama, Y.; Ohyama, M.; Itoh, T.; Suzuki, M. Geographical distribution of two haplotypes of chloroplast DNA in four oak species (Quercus) in Japan. J. Plant Res. 2004, 117, 311–317. [Google Scholar] [CrossRef] [PubMed]
- Birky, C.W. The inheritance of gens in mitochondria and chloroplasts: Laws, mechanisms and models. Annu. Rev. Genet. 2001, 35, 125–148. [Google Scholar] [CrossRef] [PubMed]
- Cruzan, M.B.; Templeton, A.R. Paleoecology and coalescence: Phylogeographic analysis of hypotheses from the fossil record. Trends Ecol. Evol. 2000, 15, 491–496. [Google Scholar] [CrossRef]
- Orive, M.E.; Asmussen, M.A. The effects of pollen and seed migration on nuclear-dicytoplasmic systems. II. A new method for estimating plant gene flow from joint nuclear-cyto plasmic data. Genetics 2000, 155, 833–854. [Google Scholar] [PubMed]
- Petit, R.J.; Pineau, E.; Demeseure, B.; Bacilieri, R.; Ducousso, A.; Kremer, A. Chloroplast DNA footprints of postglacial recolonization by oaks. Proc. Natl. Acad. Sci. USA 1997, 94, 9996–10001. [Google Scholar] [CrossRef] [PubMed]
- Honjo, M.; Ueno, S.; Tsumura, Y.; Washitani, I.; Ohsawa, R. Phylogeographic study based on intraspecific sequence variation of chloroplast DNA for the conservation of genetic diversity in the Japense endangered species Primula sieboldii. Biol. Conserv. 2004, 120, 211–220. [Google Scholar] [CrossRef]
- Yang, J.-B.; Tang, M.; Li, H.-T.; Zhang, Z.-R.; Li, D.-Z. Complete chloroplast genome of the genus Cymbidium: Lights into the species identification, phylogenetic implications and population genetic analyses. BMC Evol. Biol. 2013, 13. [Google Scholar] [CrossRef] [PubMed]
- Demesure, B.; Comps, B.; Petit, R.J. Chloroplast DNA phylogeography of the common beech (Fagus sylvatica L.) in Europe. Evolution 1996, 50, 2515–2520. [Google Scholar] [CrossRef]
- Taberlet, P.; Fumagalli, L.; Wust-Saucy, A.-G.; Cosson, J.-F. Comparative phylogeography and postglacial colonisation routes in Europe. Mol. Ecol. 1998, 7, 453–464. [Google Scholar] [PubMed]
- Hewitt, G.M. Post-glacial re-colonization of European biota. Biol. J. Linn. Soc. 1999, 68, 87–112. [Google Scholar] [CrossRef]
- Newton, A.C.; Allnutt, T.R.; Gillies, A.C.M.; Lowe, A.J.; Ennos, R.A. Molecular phylogeography, intraspecific variation and the conservation of tree species. Trends Ecol. Evol. 1999, 14, 140–145. [Google Scholar] [CrossRef]
- Palme, A.E.; Vendramin, G.G. Chloroplast DNA variation, postglacial recolonisation and hybridisation in hazel, Corylus avellana. Mol. Ecol. 2002, 9, 1769–1780. [Google Scholar] [CrossRef]
- Taberlet, P.; Gielly, L.; Pautou, G.; Bouvet, J. Universal primers for amplification of three noncoding regions of chloroplast DNA. Plant Mol. Biol. 1991, 17, 1105–1109. [Google Scholar] [CrossRef] [PubMed]
- Demesure, B.; Sodzi, N.; Petit, R.J. A set of universal primers for amplification of polymorphic non-coding regions of mitochondrial and chloroplast DNA in plants. Mol. Ecol. 1995, 4, 129–131. [Google Scholar] [CrossRef] [PubMed]
- Shaw, A.J.; Cox, C.J. Variation in “biodiversity value” of peatmoss species in Sphagnum section Acutifolia (Sphagnaceae). Am J. Bot. 2005, 92, 1774–1783. [Google Scholar] [CrossRef] [PubMed]
- Sawicki, J.; Szczecińska, M. A comparison of PCR-based markers for molecular identification of Sphagnum species of the section Acutifolia. Acta Soc. Bot. Pol. 2011, 80, 185–192. [Google Scholar] [CrossRef]
- Gerbi, S.A. Evolution of ribosomal rRNA. In Molecular Evolutionary Genetics; MacIntyre, R.J., Ed.; Plenum: New York, NY, USA, 1985; pp. 419–517. [Google Scholar]
- Wendel, F.F.; Schnabel, A.; Seelanan, T. Bidirectional interlocus concerned evolution following allopolyploid speciation in cotton (Gossypium). Proc. Natl. Acad. Sci. USA 1995, 92, 280–284. [Google Scholar] [CrossRef] [PubMed]
- Karvonen, P.; Szmidt, A.E.; Savolainen, O. Length variation in the internal transcribed spacers of ribosomal DNA in Picea abies and related species. Theor. Appl. Genet. 1994, 89, 969–974. [Google Scholar] [CrossRef] [PubMed]
- Schoch, C.L.; Seifert, K.A.; Huhndorf, S.; Robert, V.; Spouge, J.L.; Levesque, C.A.; Chen, W. Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc. Natl. Acad. Sci. USA 2012, 109, 6241–6246. [Google Scholar] [CrossRef] [PubMed]
- Bednarek-Ochyra, H.; Ochyra, R.; Sawicki, J.; Szczecińska, M. Bucklandiella seppeltii, a new species of Grimmiaceae from Australasia, and its phylogenetic position based on molecular data. Turk. J. Bot. 2014, 38, 1214–1228. [Google Scholar] [CrossRef]
- Denduangboripant, J.; Cronk, Q.C.B. High intraindividual variation in internal transcribed spacer sequences in Aeschynanthus (Gesneriaceae): Implications for phylogenetics. Proc. Biol. Sci. 2000, 267, 1407–1415. [Google Scholar] [CrossRef] [PubMed]
- Won, H.; Renner, S.S. The internal transcribed spacer of ribosomal DNA in the gymnosperm Gnetum. Mol. Phyl. Evol. 2005, 36, 581–597. [Google Scholar] [CrossRef] [PubMed]
- Volkov, R.A.; Komarova, N.Y.; Hemleben, V. Ribosomal DNA in plant hybrids: Inheritance, rearrangement, expression. Syst. Biodivers. 2007, 5, 261–276. [Google Scholar] [CrossRef]
- Small, R.L.; Cronn, R.C.; Wendel, J.F. Use of nuclear genes for phylogeny reconstruction in plants. Aust. Syst. Bot. 2004, 17, 145–170. [Google Scholar] [CrossRef]
- Sun, Y.-L.; Park, W.-G.; Oh, H.-K.; Hong, S.-K. Genetic diversity and hybridization of Pulsatilla tongkangensis based on nrDNA ITS region sequence. Biologia 2014, 69, 24–31. [Google Scholar] [CrossRef]
- Raubeson, L.A.; Peery, R.; Chuley, T.W.; Dziubek, C.; Fourcade, H.M.; Boore, J.L.; Jansen, R.K. Comparative chloroplast genomics: Analyses including new sequences from the angiosperms Nuphar advena and Ranunculus macranthus. BMC Genom. 2007, 8, 174. [Google Scholar] [CrossRef] [PubMed]
- Fajardo, D.; Senalik, D.; Ames, M.; Zhu, H.; Steffan, S.A.; Harbut, R.; Polashock, J.; Vorsa, N.; Gillespie, E.; Kron, K. Complete plastid genome sequence of Vaccinium macrocarpon: Structure, gene content and rearrangements revealed by next generation sequencing. Tree Genet. Genomes 2012, 9, 489–498. [Google Scholar] [CrossRef]
- Szczecińska, M.; Gomolińska, A.; Szkudlarz, P.; Sawicki, J. Plastid and nuclear genomic resources of a relict and endangered plant species—Chamaedaphne calyculata (L.) Moench. (Ericaceae). Turk. J. Bot. 2014, 38, 1229–1238. [Google Scholar] [CrossRef]
- Parks, M.; Cronn, R.; Liston, A. Increasing phylogenetic resolution at a low taxonomic levels using massively parallel sequencing at chloroplast genomes. BMC Biol. 2009, 7. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.-J.; Ma, P.-F.; Li, D.-Z. High-throughput sequencing of six bamboo chloroplast genomes: Phylogenetic implications for temperate woody bamboos (Poaceae: Bambusoideae). PLoS ONE 2011, 6, e20596. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.-S.; Wang, Y.-N.; Hsu, C.-Y.; Lin, C.-P.; Chaw, S.-M. Loss of different inverted repeat copies from the chloroplast genomes of Pinaceae and Cupresophytes and influence of heterotachy on evolution of Gymnosperms phylogeny. Genome Biol. Evol. 2011, 3, 1284–1295. [Google Scholar] [CrossRef] [PubMed]
- Chen, X.; Qiushi, L.; Ying, L.; Qian, J.; Han, J. Chloroplast genome of Aconitum barbatum var. pubreulum (Ranunculaceae) derived from CCS reads using the PacBio RS platform. Front. Plant Sci. 2015, 6. [Google Scholar] [CrossRef]
- Yang, J.B.; Yang, S.X.; Li, H.T.; Yang, J.; Li, D.Z. Comparative chloroplast genomes of Camellia species. PLoS ONE 2013, 8, e73053. [Google Scholar] [CrossRef] [PubMed]
- Yang, M.; Zhang, X.; Liu, G.; Yin, Y.; Chen, K.; Yun, Q.; Zhao, D.; Al-Mssallem, I.S.; Yu, J. The complete chloroplast genome sequence of date Palm (Phoenix dactylifera L.). PLoS ONE 2010, 15. [Google Scholar] [CrossRef] [PubMed]
- Frazer, K.A.; Pachter, L.; Poliakov, A.; Rubin, E.M.; Dubchak, I. VISTA: Computational tools for comparative genomics. Nucleic Acids Res. 2004, 1, W273–W279. [Google Scholar] [CrossRef] [PubMed]
- Korotkova, N.; Nauheimer, L.; Ter-Voskanyan, H.; Allgaier, M.; Borsch, T. Variability among the most rapidly evolving plastid genomic regions is lineage-specific: Implications of pairwise genome comparisons in Pyrus (Rosaceae) and other angiosperms for marker choice. PLoS ONE 2014, 9, e112998. [Google Scholar] [CrossRef] [PubMed]
- Nie, X.; Lv, S.; Zhang, Y.; Du, X.; Wang, L. Complete chloroplast genome sequence of a major invasive species, crofton weed (Ageratina adenophora). PLoS ONE 2012, 7, e36869. [Google Scholar] [CrossRef] [PubMed]
- Dong, W.; Liu, J.; Yu, J.; Wang, L.; Zhou, S. Highly variable chloroplast markers for evaluating plant phylogeny at low taxonomic levels and for DNA barcoding. PLoS ONE 2012, 7, e35071. [Google Scholar] [CrossRef] [PubMed]
- Jukes, T.H.; King, J.L. Deleterious mutations and neutral substitutions. Nature 1971, 231, 114–115. [Google Scholar] [CrossRef] [PubMed]
- Kimura, M. The Neutral Theory of Molecular Evolution; Cambridge University Press: Cambridge, UK, 1983. [Google Scholar]
- Anastasi, G.; Cutroneo, G.; Santoro, G.; Arco, A.; Rizzo, G.; Bramanti, P.; Rinaldi, C.; Sidoti, A.; Amato, A.; Favaloro, A. Costameric proteins in human skeletal muscle during muscular inactivity. J. Anat. 2008, 213, 284–295. [Google Scholar] [CrossRef] [PubMed]
- Ganapathi, M.; Srivastava, P.; Das Sutar, S.K.; Kumar, K.; Dasgupta, D.; Pal Singh, G.; Brahmachari, V.; Brahmachari, S.K. Comparative analysis of chromatin landscape in regulatory regions of human housekeeping and tissue specific genes. BMC Bioinform. 2005, 6, 126. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Rose, A.B. Intron-mediated regulation of gene expression. Curr. Top. Microbiol. Immunol. 2008, 326, 277–290. [Google Scholar] [PubMed]
- Dong, W.; Xu, Ch.; Li, Ch.; Sun, J.; Zuo, Y.; Shi, S. ycf1, the most promising plastid DNA barcode of land plants. Sci. Rep. 2015, 5, 8348. [Google Scholar] [CrossRef] [PubMed]
- Krawczyk, K.; Sawicki, J. The uneven rate of the molecular evolution of gene sequences of DNA-dependent RNA polymerase I of the genus Lamium L. Int. J. Mol. Sci. 2013, 14, 11373–11391. [Google Scholar] [CrossRef] [PubMed]
- Zhang, C.Y.; Wang, F.Y.; Yan, H.F.; Hao, G.; Hu, C.M.; Ge, X.J. Testing DNA barcoding in closely related groups of Lysimachia L. (Myrsinaceae). Mol. Ecol. Res. 2012, 12, 98–108. [Google Scholar] [CrossRef] [PubMed]
- Krawczyk, K.; Szczecińska, M.; Sawicki, J. Evaluation of 11 single-locus and seven multilocus DNA barcodes in Lamium L. (Lamiaceae). Mol. Ecol. Res. 2014, 14, 272–285. [Google Scholar] [CrossRef] [PubMed]
- Sloan, D.B.; Alverson, A.J.; Wu, M.; Palmer, J.D.; Taylor, D.R. Recent acceleration of plastid sequence and structural evolution coincides with extreme mitochondrial divergence in the angiosperm genus Silene. Genome Biol. Evol. 2012, 4, 294–306. [Google Scholar] [CrossRef] [PubMed]
- Erixon, P.; Oxelman, B. Whole-gene positive selection, elevated synonymous substitution rates, duplication, and indel evolution of the chloroplast clpP1 gene. PLoS ONE 2008, 3, e1386. [Google Scholar] [CrossRef] [PubMed]
- Frankham, R.; Briscoe, D.A.; Ballou, J.D. Introduction to Conservation Genetics; Cambridge University Press: New York, NY, USA, 2002. [Google Scholar]
- Kapralov, M.V.; Smith, J.A.C.; Filatov, D.A. Rubisco evolution in C4 eudicots: An analysis of Amaranthaceae sensu lato. PLoS ONE 2012, 7, e52974. [Google Scholar] [CrossRef] [PubMed]
- Hu, S.; Sablok, G.; Wang, B.; Qu, D.; Barbaro, E.; Viola, R.; Li, M.; Varotto, C. Plastome organization and evolution of chloroplast genes in Cardamine species adapted to contrasting habitats. BMC Genom. 2015, 16, 306. [Google Scholar] [CrossRef] [PubMed]
- Gilley, L.; Yong-Ming, Y.; Küpfer, D.; Taberlet, P. Phylogenetic use of noncoding regions in the genus Gentiana L. Chloroplast trnL (UAA) intron versus nuclear ribosomal internal transcribed spacer sequences. Mol. Phyl. Evol. 1996, 3, 460–466. [Google Scholar] [CrossRef] [PubMed]
- Sawicki, J.; Plášek, V.; Szczecińska, M. Molecular evidence do not support the current division of Orthotrichum subgenus Gymnoporus. Plant Syst. Evol. 2009, 279, 125–137. [Google Scholar] [CrossRef]
- Szczecińska, M.; Kwaśniewski, M.; Chwiałkowska, K.; Sawicki, J. Isolation and characterization of microsatellite loci in Pulsatilla patens (L.) Mill. (Ranunculaceae) a rare and endangered plant species in Europe. Conserv. Genet. Resour. 2013, 5, 421–423. [Google Scholar] [CrossRef]
- Wyman, S.K.; Jansen, R.K.; Boore, J.L. Automatic annotation of organellar genomes with DOGMA. Bioinformatics 2004, 20, 3252–3255. [Google Scholar] [CrossRef] [PubMed]
- Darling, A.C.; Mau, B.; Blattner, F.R.; Perna, N.T. Mauve: Multiple alignment of conserved genomic sequence with rearrangements. Genome Res. 2004, 14, 1394–1403. [Google Scholar] [CrossRef] [PubMed]
- Krzywinski, M.; Schein, J.; Bird, I.; Connors, J.; Gascoyne, R.; Horsman, D.; Jones, S.J.; Marra, M.A. Circos: An information aesthetic for comparative genomics. Genome Res. 2009, 19, 1639–1645. [Google Scholar] [CrossRef] [PubMed]
- Faircloth, B.C. MSATCOMMANDER: Detection of microsatellite repeat arrays and automated, locus-specific primer design. Mol. Ecol. Res. 2008, 8, 92–94. [Google Scholar] [CrossRef] [PubMed]
- Edgar, R.C. MUSCLE: Multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res. 2004, 32, 1792–1797. [Google Scholar] [CrossRef] [PubMed]
- Golenberg, E.M.; Clegg, M.T.; Durbin, M.L.; Doebley, J.D.P. Evolution of a noncoding region of the chloroplast genome. Mol. Phylogenet. Evol. 1993, 2, 52–64. [Google Scholar] [CrossRef] [PubMed]
- Morton, B.R.; Clegg, M.T. A chloroplast DNA mutational hotspot and gene conversion in a noncoding region near rbcL in the grass family (Poaceae). Curr. Genet. 1993, 24, 357–365. [Google Scholar] [CrossRef] [PubMed]
- O’Donnell, K. Ribosomal DNA internal transcribed spacers are highly divergent in the phytopathogenic ascomycete Fusarium sambucinum (Gibberella pulicaris). Curr. Genet. 1992, 22, 213–220. [Google Scholar] [CrossRef] [PubMed]
- Akaike, H. A new look at the statistical model identification. IEEE Trans. Autom. Control 1974, 19, 716–723. [Google Scholar] [CrossRef]
- Posada, D.; Crandall, K.A. Modeltest: Testing the model of DNA substitution. Bioinformatics 1998, 14, 817–818. [Google Scholar] [CrossRef] [PubMed]
- Huelsenbeck, J.P.; Ronquist, F.R. MRBAYES: Bayesian inference of phylogeny. Bioinformatics 2001, 17, 754–755. [Google Scholar] [CrossRef] [PubMed]
- Rambaut, A.; Drummond, A.J. Tracer v1.3. 2003. Available online: http://evolve.zoo.ox.ac.uk (accessed on 17 February 2012).
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Szczecińska, M.; Sawicki, J. Genomic Resources of Three Pulsatilla Species Reveal Evolutionary Hotspots, Species-Specific Sites and Variable Plastid Structure in the Family Ranunculaceae. Int. J. Mol. Sci. 2015, 16, 22258-22279. https://doi.org/10.3390/ijms160922258
Szczecińska M, Sawicki J. Genomic Resources of Three Pulsatilla Species Reveal Evolutionary Hotspots, Species-Specific Sites and Variable Plastid Structure in the Family Ranunculaceae. International Journal of Molecular Sciences. 2015; 16(9):22258-22279. https://doi.org/10.3390/ijms160922258
Chicago/Turabian StyleSzczecińska, Monika, and Jakub Sawicki. 2015. "Genomic Resources of Three Pulsatilla Species Reveal Evolutionary Hotspots, Species-Specific Sites and Variable Plastid Structure in the Family Ranunculaceae" International Journal of Molecular Sciences 16, no. 9: 22258-22279. https://doi.org/10.3390/ijms160922258






