Fungi Isolated from Maize (Zea mays L.) Grains and Production of Associated Enzyme Activities
Abstract
:1. Introduction
2. Results and Discussion
2.1. Species Diversity
| Genera and Species | Isolate | Geographic Origin (City, State) | GenBank Accession Number | GenBank or Fusarium-ID (FD) Accession Number of Similar Sequences (Percentage of Identity) |
|---|---|---|---|---|
| Aspergillus flavus | PG 24 | Ponta Grossa, PR | KP691048 | KP036603.1 (100%), KM486551.1 (100%) |
| Aspergillus sp. | BAN 12 | Bandeirantes, PR | - | # |
| Aspergillus sp. | MGA 20 | Maringá, PR | - | # |
| Aspergillus wentii | PG 18 | Ponta Grossa, PR | KP691043 | KM409566.1 (100%), AY373884.1 (100%) |
| Cladosporium cladosporioides | CMA 15 | Clementina, SP | KP691040 | KJ589558.1 (100%), KF986546.1 (100%) |
| C. cladosporioides | CMA 07 | Clementina, SP | KP691034 | KJ589558.1 (100%), JX406506.1 (100%) |
| Curvularia sp. | MGI 04 | Mandaguari, PR | - | # |
| Epicoccum nigrum | PG 16 | Ponta Grossa, PR | KP691041 | JX844158.1 (100%), KJ934366.1 (99%) |
| E. nigrum | PG 23 | Ponta Grossa, PR | KP691047 | KF990155.1 (99%), KC005662.1 (99%) |
| E. nigrum | CMA 14 | Clementina, SP | KP691039 | JX844158.1 (100%), KJ934366.1 (99%) |
| Epicoccum sorghinum | CO 11 | Cruzeiro do Oeste, PR | KP691037 | KM111488.1 (100%), KP050561.1 (99%) |
| Fusarium incarnatum-equiseti * | RV 18-1 | Rio Verde, GO | KP336404 | KF962948.1 (97%), FD_01623 (97.5%) |
| F. incarnatum-equiseti * | CMA 1-2 | Clementina, SP | KP336405 | KF962948.1 (97%), FD_01623 (97.5%) |
| F. incarnatum-equiseti * | CMA 5-1 | Clementina, SP | KP336406 | KF962948.1 (97%), FD_01623 (97.5%) |
| Fusarium subglutinans * | RV 23-2 | Rio Verde, GO | KP336408 | JX867945.1 (100%), KC964122.1 (98%) |
| F. subglutinans * | PG 1-2 | Ponta Grossa, PR | KP336409 | KC964122.1 (99%), JF270302.1 (99%), |
| Fusarium verticillioides | CMA 3-1 | Clementina, SP | KP691051 | KJ125822.1 (100%), KJ125764.1 (100%) |
| F. verticillioides | RV 8-1 | Rio Verde, GO | KP691052 | KJ125822.1 (100%), KJ125764.1 (100%) |
| F. verticillioides | BAN 4-2 | Bandeirantes, PR | KP691053 | KJ125822.1 (100%), KJ125764.1 (100%) |
| F. verticillioides | MGI 1-1 | Mandaguari, PR | KP691054 | KJ125822.1 (100%), KJ125764.1 (100%) |
| F. verticillioides | BAN 2-2 | Bandeirantes, PR | KP691055 | KJ125822.1 (100%), KJ125764.1 (100%) |
| F. verticillioides | MGA 5-1 | Maringá, PR | KP691056 | KJ125822.1 (100%), KJ125764.1 (100%) |
| F. verticillioides | RV 12-2 | Rio Verde, GO | KP691057 | KJ125822.1 (100%), KJ125764.1 (100%) |
| F. verticillioides | MGA 2-2 | Maringá, PR | KP691058 | KJ125822.1 (100%), KJ125764.1 (100%) |
| F. verticillioides | CMA 2-1 | Clementina, SP | KP691059 | KJ125822.1 (100%), KJ125764.1 (100%) |
| F. verticillioides | CPÃ 1-1 | Camapuã, MS | KP691060 | KJ125822.1 (100%), KJ125764.1 (100%) |
| F. verticillioides * | MGI 3-2 | Mandaguari, PR | KP336410 | KF715263.1 (100%), KC599244.1 (99%) |
| F. verticillioides * | RV 12-2 | Rio Verde, GO | KP336411 | KF715263.1 (100%), KC599244.1 (99%) |
| F. verticillioides * | PG 4-1 | Ponta Grossa, PR | KP336412 | KF715263.1 (100%), KC599244.1 (99%) |
| Fusarium andiyazi * | RV 27-1 | Rio Verde, GO | KP336407 | JX974611.1 (99%), KC954401.1 (98%) |
| Fusarium graminearum | MGI 21 | Mandaguari, PR | KP691045 | KM513614.1 (100%), KF624778.1 (100%) |
| Fusarium nygamai * | RV 27-2 | Rio Verde, GO | KP336403 | JF740790.1 (95%), HM243236.1 (95%) |
| Hyphopichia burtonii | CMA 09 | Clementina, SP | KP691036 | KP132302.1 (99%), EU714323.1 (98%) |
| Irpex lacteus | CMA 19 | Clementina, SP | KP691044 | LN714557.1 (100%), KJ831879.1 (100%) |
| Microdochium nivale | PG 22 | Ponta Grossa, PR | KP691046 | JX280606.1 (98%), EF187912.1 (92%), |
| Mucor fragilis | CMA 25 | Clementina, SP | KP691049 | JQ972063.1 (99%), FN650655.1 (99%) |
| Mucor sp. | CO 10 | Cruzeiro do Oeste, PR | - | # |
| Penicillium digitatum | RV 06 | Rio Verde, GO | KP691033 | AY373910.1 (100%), KJ834506.1 (100%) |
| Penicillium sp. | MGI 01 | Mandaguari, PR | - | # |
| Phlebiopsis gigantea | CMA 08 | Clementina, SP | KP691035 | JQ781838.1 (98%), FJ791151.1 (98%) |
| Sarocladium zeae | PG 1-1 | Ponta Grossa, PR | KP691050 | KP132614.1 (100%), KJ188657.1 (98%) |
| Stenocarpella maydis | MGI 03 | Mandaguari, PR | KP691031 | KM030331.1 (100%), KC311732.1 (100%) |
| Talaromyces purpureogenus | MGI 05 | Mandaguari, PR | KP691032 | JX157861.1 (99%), JQ422620.1 (99%) |
| T. purpureogenus | PG 17 | Ponta Grossa, PR | KP691042 | JX157861.1 (99%), JQ422620.1 (99%) |
| T. purpureogenus | MGI 02 | Mandaguari, PR | KP691030 | JX157861.1 (99%), JQ422620.1 (99%) |
| Trichoderma harzianum | CO 13 | Cruzeiro do Oeste, PR | KP691038 | KP050785.1 (100%), KF624792.1 (100%) |
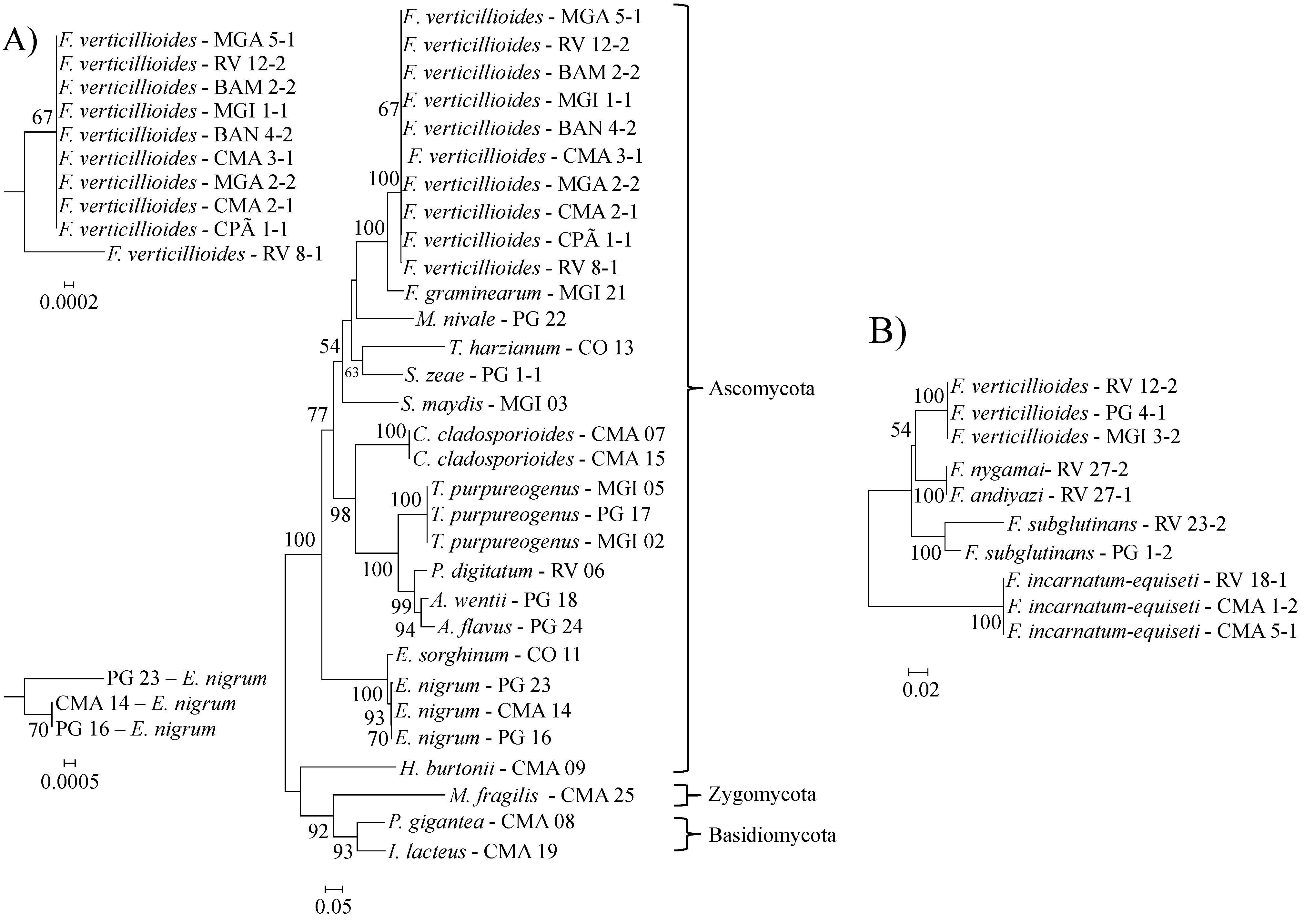
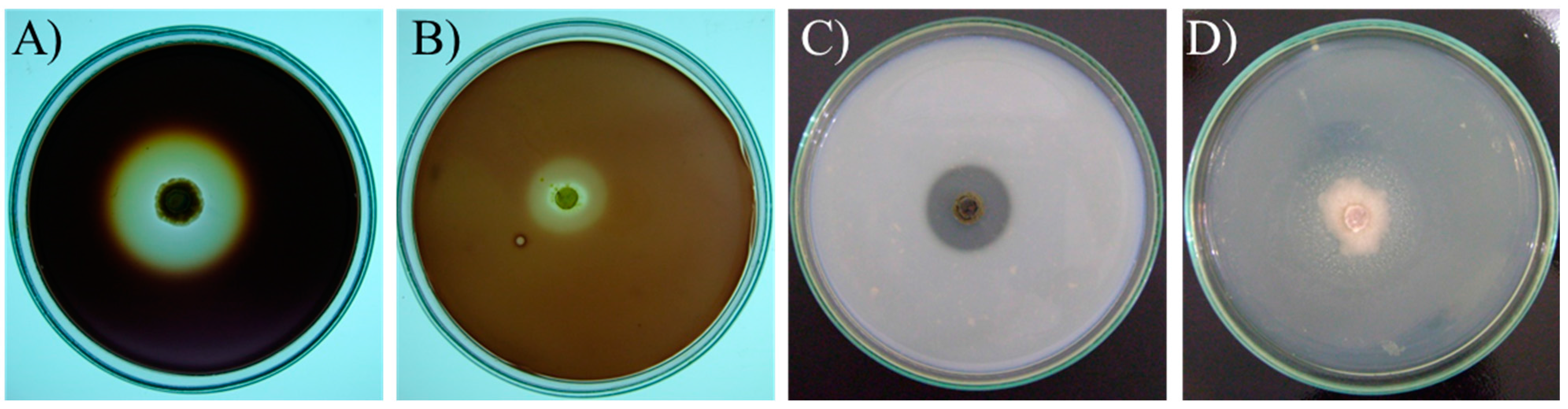
2.2. Extracellular Enzyme Activities
| Fungal Strain | Strain | Amylase * | Cellulase * | Protease * | Lipase * |
|---|---|---|---|---|---|
| A. flavus | PG 24 | 1.24 ± 0.08 b | 1.78 ± 0.03 b | 1.17 ± 0.01 b | 1.24 ± 0.04 f |
| A. wentii | PG 18 | 3.33 ± 0.49 a | 3.92 ± 0.51 a | - | 3.01 ± 0.20 c |
| Aspergillus sp. | BAN 12 | 1.04 ± 0.00 b | 1.84 ± 0.81 b | - | 1.15 ± 0.06 f |
| Aspergillus sp. | MGA 20 | 1.31 ± 0.04 b | 1.57 ± 0.04 b | - | 1.82 ± 0.08 e |
| C. cladosporioides | CMA 07 | - | 4.47 ± 0.50 a | 2.27 ± 0.31 a | 5.60 ± 0.69 a |
| C. cladosporioides | CMA 15 | 2.20 ± 0.40 b | 4.13 ± 0.50 a | 2.33 ± 0.31 a | 5.67 ± 0.31 a |
| Curvularia sp. | MGI 04 | 1.04 ± 0.00 b | 1.36 ± 0.04 b | 1.29 ± 0.03 b | - |
| E. nigrum | CMA 14 | 1.46 ± 0.08 b | 1.62 ± 0.16 b | 1.31 ± 0.06 b | 1.77 ± 0.18 e |
| E. nigrum | PG 16 | 2.25 ± 0.00 b | 2.40 ± 0.36 b | 1.69 ± 0.63 b | 2.97 ± 0.69 c |
| E. nigrum | PG 23 | 1.92 ± 0.62 b | 1.67 ± 0.11 b | 1.09 ± 0.00 b | 2.21 ± 0.29 d |
| E. sorghinum | CO 11 | 1.18 ± 0.03 b | 1.85 ± 0.08 b | 1.04 ± 0.04 b | 1.84 ± 0.28 e |
| F. andiyazi | RV 27-1 | - | - | - | - |
| F. graminearum | MGI 21 | - | - | - | - |
| F. incarnatum-equiseti | RV 18-1 | - | - | - | 2.00 ± 0.00 d |
| F. incarnatum-equiseti | CMA 1-2 | - | - | - | 2.14 ± 0.05 d |
| F. nygamai | RV 27-2 | - | - | - | - |
| F. subglutinans | RV 23-2 | - | - | - | 1.70 ± 0.19 e |
| F. subglutinans | PG 1-2 | - | - | 1.29 ± 0.07 b | 1.64 ± 0.20 e |
| F. verticillioides | MGI 3-2 | - | - | - | - |
| F. verticillioides | RV 12-2 | - | - | - | - |
| F. verticillioides | PG 4-1 | - | - | - | 1.26 ± 0.01 f |
| H. burtonii | CMA 09 | 4.37 ± 1.64 a | 2.10 ± 0.07 b | - | 2.23 ± 0.67 d |
| I. lacteus | CMA 19 | 3.90 ± 0.00 a | - | - | 2.01 ± 0.04 d |
| M. fragilis | CMA 25 | - | - | 1.13 ± 0.02 b | 1.08 ± 0.05 f |
| Mucor sp. | CO 10 | - | - | 1.42 ± 0.03 b | 1.31 ± 0.03 f |
| P. digitatum | RV 06 | - | 3.67 ± 0.31 a | - | 3.87 ± 0.12 b |
| Penicillium sp. | MGI 01 | 1.74 ± 0.04 b | 2.31 ± 0.10 b | 1.53 ± 0.08 b | 1.60 ± 0.05 e |
| S. maydis | MGI 03 | - | 1.36 ± 0.04 b | - | 1.36 ± 0.04 f |
| S. zeae | PG 1-1 | 1.47 ± 0.20 b | 1.84 ± 0.03 b | 1.91 ± 0.09 a | 2.90 ± 0.18 c |
| T. purpureogenus | MGI 02 | - | 1.71 ± 0.09 b | - | 1.31 ± 0.11 f |
| T. harzianum | CO 13 | - | - | 1.27 ± 0.06 b | - |
| T. purpureogenus | MGI 05 | 1.04 ± 0.00 b | 1.42 ± 0.04 b | - | 1.40 ± 0.24 f |
| T. purpureogenus | PG 17 | - | 1.76 ± 0.05 b | - | - |
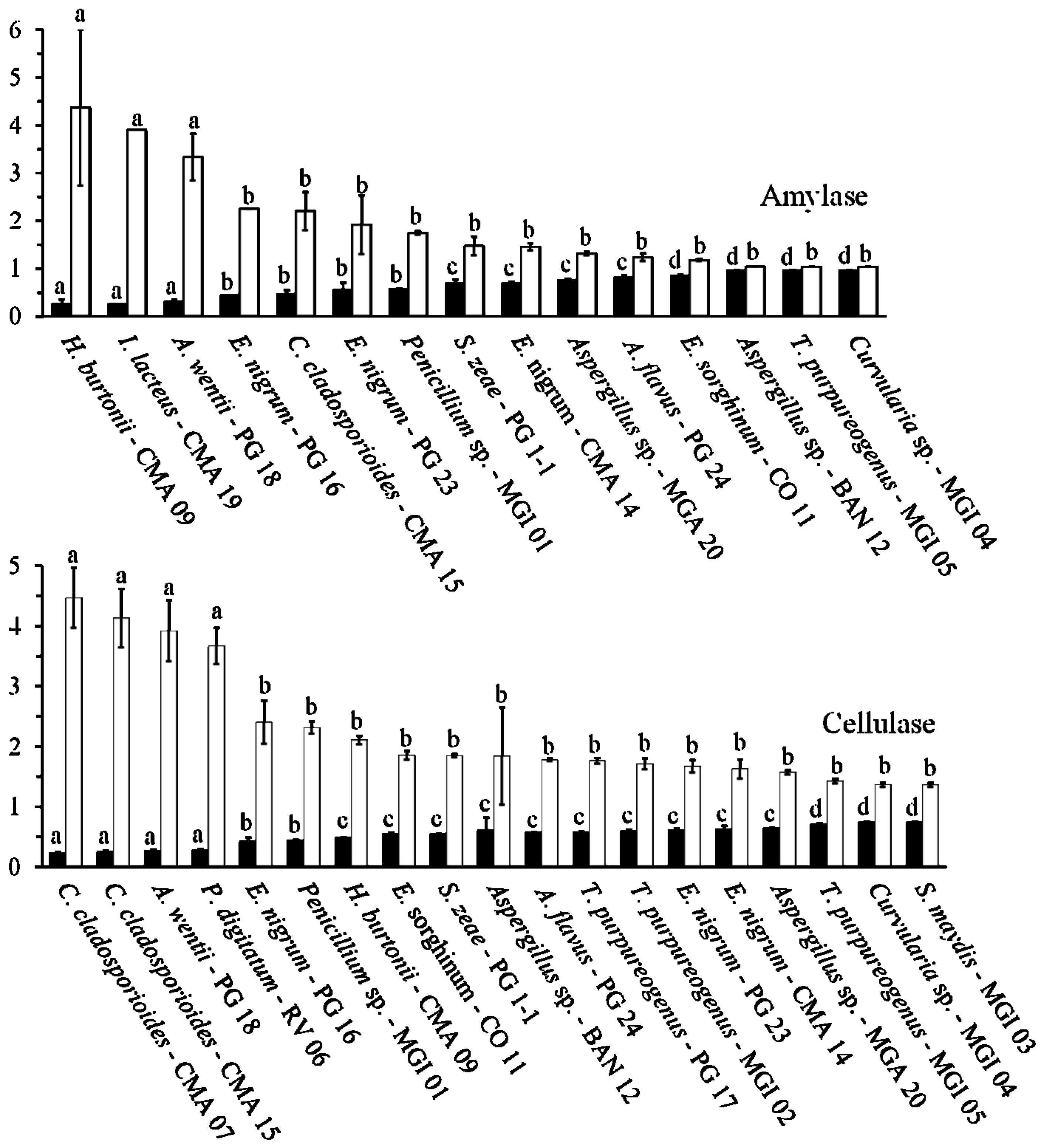
2.3. Amylolytic Activity
2.4. Cellulolytic Activity
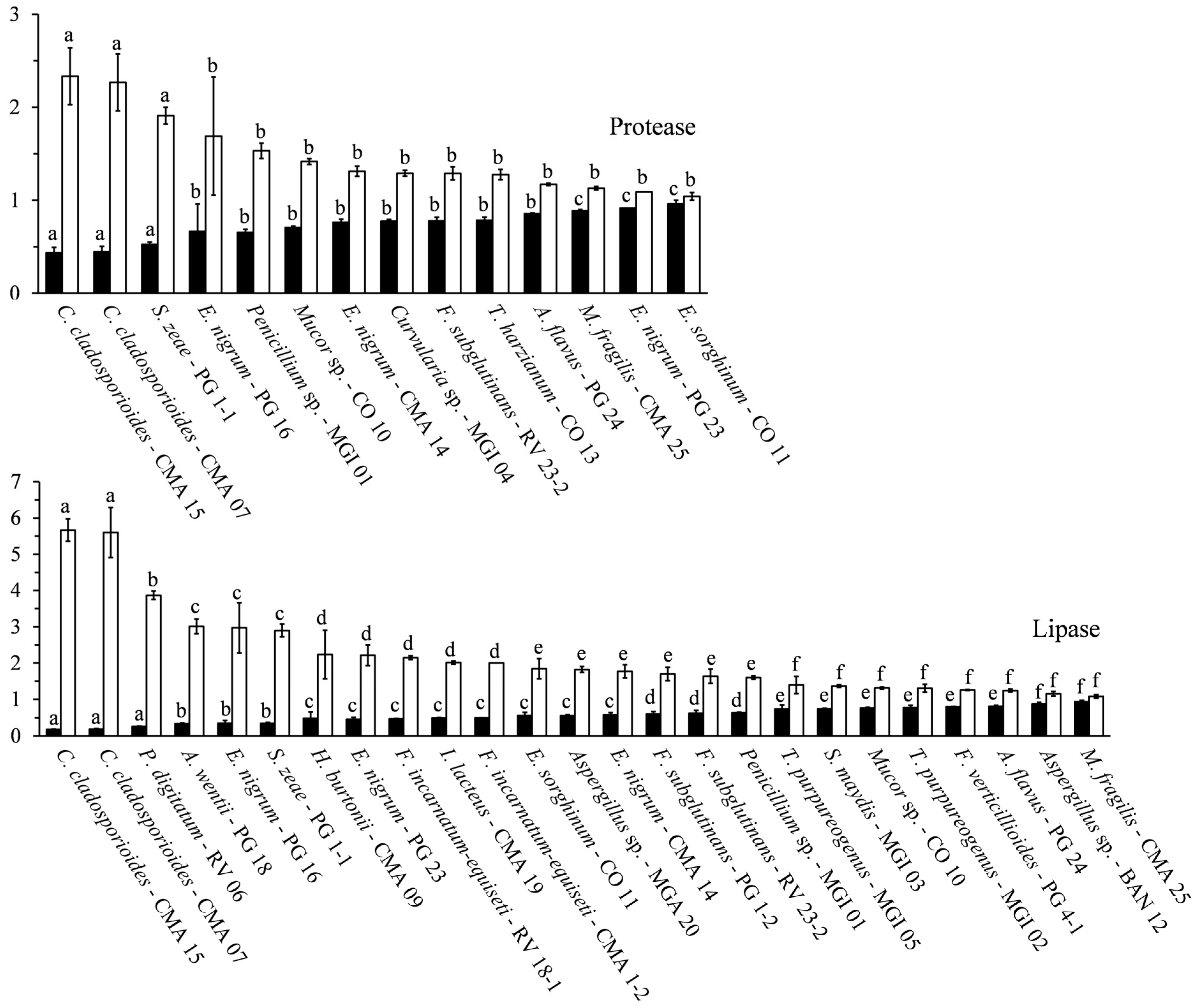
2.5. Proteolytic Activity
2.6. Lipolytic Activity
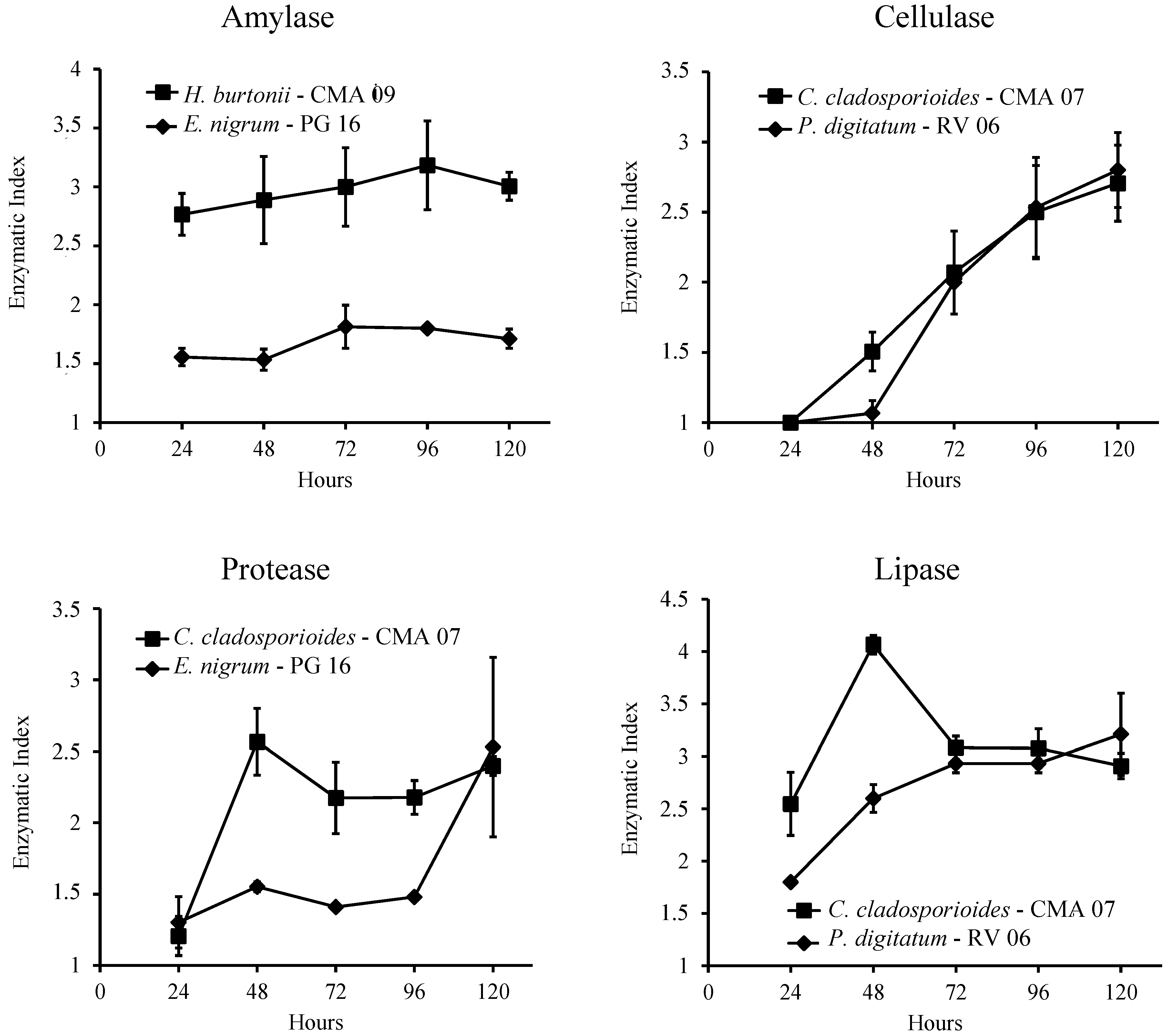
2.7. Quantitative Analysis of Enzyme Production
3. Experimental Section
3.1. Fungal Isolates
3.2. Molecular Identification of the Isolates
3.3. Detection of Extracellular Enzymatic Activity
3.4. Statistical Analysis
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Sarrouh, B.; Santos, T.M.; Miyoshi, A.; Dias, R.; Azevedo, V. Up-to-date insight on industrial enzymes applications and global market. J. Bioprocess Biotechnol. 2012, S4. [Google Scholar] [CrossRef]
- Gurung, N.; Ray, S.; Bose, S.; Rai, V. A broader view: Microbial enzymes and their relevance in industries, medicine, and beyond. BioMed Res. Int. 2013, 2013, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Demain, A.L.; Adrio, J.L. Contributions of microorganisms to industrial biology. Mol. Biotechnol. 2008, 38, 41–55. [Google Scholar] [CrossRef] [PubMed]
- Pandey, A.; Nigam, P.; Soccol, C.R.; Soccol, V.T.; Singh, D.; Mohan, R. Advances in microbial amylases. Biotechnol. Appl. Biochem. 2000, 31, 135–152. [Google Scholar] [CrossRef] [PubMed]
- Gupta, R.; Gigras, P.; Mohapatra, H.; Goswami, V.K.; Chauhan, B. Microbial α-amylases: Biotechnological Perspective. Process Biochem. 2003, 38, 1–18. [Google Scholar] [CrossRef]
- Norouzian, D.; Akbarzadeh, A.; Scharer, J.M.; Moo-Young, M. Fungal glucoamylases. Biotechnol. Adv. 2006, 24, 80–85. [Google Scholar] [CrossRef] [PubMed]
- Sun, H.; Zhao, P.; Ge, X.; Xia, Y.; Hao, Z.; Liu, J.; Peng, M. Recent advances in microbial raw starch degrading enzymes. Appl. Biochem. Biotechnol. 2010, 160, 988–1003. [Google Scholar] [CrossRef] [PubMed]
- Bhat, M.K. Cellulases and related enzymes in biotechnology. Biotechnol. Adv. 2000, 18, 355–383. [Google Scholar] [CrossRef]
- Lynd, L.R.; Weimer, P.J.; van Zyl, W.H.; Pretorius, I.S. Microbial cellulose utilization: Fundamentals and biotechnology. Microbiol. Mol. Biol. Rev. 2002, 66, 506–577. [Google Scholar] [CrossRef] [PubMed]
- Dashtban, M.; Schraft, H.; Qin, W. Fungal bioconversion of lignocellulosic residues; Opportunities & perspectives. Int. J. Biol. Sci. 2009, 5, 578–595. [Google Scholar] [PubMed]
- Zhang, P.; Himmel, M.E.; Mielenz, J.R. Outlook for cellulase improvement: screening and selection strategies. Biotechnol. Adv. 2006, 24, 452–481. [Google Scholar] [CrossRef] [PubMed]
- Kumar, C.G.; Takagi, H. Microbial alkaline proteases: From a bioindustrial viewpoint. Biotechnol. Adv. 1999, 17, 561–594. [Google Scholar] [CrossRef]
- Rao, M.B.; Tanksale, A.M.; Ghatge, M.S.; Deshpande, V.V. Molecular and biotechnological aspects of microbial proteases. Microbiol. Mol. Biol. Rev. 1998, 62, 597–635. [Google Scholar] [PubMed]
- Bornscheuer, U.T. Microbial carboxyl esterases: Classification, properties and application in biocatalysis. FEMS Microbiol. Rev. 2002, 26, 73–81. [Google Scholar] [CrossRef] [PubMed]
- Jaeger, K.-E.; Eggert, T. Lipases for biotechnology. Curr. Opin. Biotechnol. 2002, 13, 390–397. [Google Scholar] [CrossRef]
- Reetz, M.T. Lipases as practical biocatalysts. Curr. Opin. Chem. Biol. 2002, 6, 145–150. [Google Scholar] [CrossRef]
- Pandey, A.; Benjamin, S.; Soccol, C.R.; Nigam, P.; Krieger, N.; Soccol, V.T. The realm of microbial lipases in biotechnology. Biotechnol. Appl. Biochem. 1999, 29, 119–131. [Google Scholar] [PubMed]
- Hasan, F.; Shah, A.A.; Hameed, A. Industrial applications of microbial lipases. Enzym. Microb. Technol. 2006, 39, 235–251. [Google Scholar] [CrossRef]
- Fjerbaek, L.; Christensen, K.V.; Norddahl, B. A review of the current state of biodiesel production using enzymatic transesterification. Biotechnol. Bioeng. 2009, 102, 1297–1315. [Google Scholar] [CrossRef] [PubMed]
- Faria, C.B.; Abe, C.A.L.; Silva, C.N.; Tessmann, D.J.; Barbosa-Tessmann, I.P. New PCR assays for the identification of Fusarium verticillioides, Fusarium subglutinans, and other species of the Gibberella fujikuroi complex. Int. J. Mol. Sci. 2012, 13, 115–132. [Google Scholar] [CrossRef] [PubMed]
- Geiser, D.M.; Jiménez-Gasco, M.M.; Kang, S.; Makalowska, I.; Veeraraghavan, N.; Ward, T.J.; Zhang, N.; Kuldau, G.A.; O’Donnell, K. FUSARIUM-ID v. 1.0: A DNA sequence database for identifying Fusarium. Eur. J. Plant Pathol. 2004, 110, 473–479. [Google Scholar] [CrossRef]
- Bhattacharya, K.; Raha, S. Deteriorative changes of maize, groundnut and soybean seeds by fungi in storage. Mycopathologia 2002, 155, 135–141. [Google Scholar] [CrossRef] [PubMed]
- Niaz, I.; Dawar, S. Detection of seed borne mycoflora in maize (Zea mays L.). Pak. J. Bot. 2009, 41, 443–451. [Google Scholar]
- Rocha, L.O.; Reis, G.M.; Silva, V.N.; Braghini, R.; Teixeira, M.M.G.; Corrêa, B. Molecular characterization and fumonisin production by Fusarium verticillioides isolated from corn grains of different geographic origins in Brazil. Int. J. Food Microbiol. 2011, 145, 9–21. [Google Scholar] [CrossRef] [PubMed]
- Fávaro, L.C.L.; Melo, F.L.; Aguilar-Vildoso, C.I.; Araújo, W.L. Polyphasic analysis of intraspecific diversity in Epicoccum nigrum warrants reclassification into separate species. PLoS ONE 2011, 6, e14828. [Google Scholar] [CrossRef] [PubMed]
- Felsenstein, J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution 1985, 39, 783–791. [Google Scholar] [CrossRef]
- Tamura, K.; Stecher, G.; Peterson, D.; Filipski, A.; Kumar, S. MEGA6: Molecular Evolutionary Genetics Analysis version 6.0. Mol. Biol. Evol. 2013, 30, 2725–2729. [Google Scholar] [CrossRef] [PubMed]
- Takeuchi, A.; Shimizu-Ibuka, A.; Nishiyama, Y.; Mura, K.; Okada, S.; Tokue, C.; Arai, S. Purification and characterization of an α-amylase of Pichia burtonii isolated from the traditional starter “murcha” in Nepal. Biosci. Biotechnol. Biochem. 2006, 70, 3019–3024. [Google Scholar] [CrossRef] [PubMed]
- Wakabayashi, K.; Daicho, H. Purification and characterization of α-amylase from Irpex lacteus (Polyporus tulipiferae). Acad. Bull. Nagano Univ. 1990, 12, 1–19. [Google Scholar]
- Shina, S.; Chakrabarty, S.L. Production of amylase by a submerged culture of Aspergillus wentii. Folia Microbiol. 1978, 23, 6–11. [Google Scholar]
- Saleem, A.; Ebrahim, M.K.H. Production of amylase by fungi isolated from legume seeds collected in Almadinah Almunawwarah, Saudi Arabia. J. Taibah Univ. Sci. 2014, 8, 90–97. [Google Scholar] [CrossRef]
- Chiranjeevi, T.; Rani, G.B.; Chandel, A.K.; Sekhar, P.V.S.; Prakasham, R.S.; Addepally, U. Optimization of holocellulolytic enzymes production by Cladosporium Cladosporioides Using Taguchi-L’16 orthogonal array. J. Biobased Mater. Bioenergy 2012, 6, 148–157. [Google Scholar] [CrossRef]
- Legler, G.; Roeser, K.R.; Illig, H.K. Reaction of β-d-glucosidase A3 from Aspergillus wentii with d-glucal. Eur. J. Biochem. 1979, 101, 85–92. [Google Scholar] [CrossRef] [PubMed]
- Hamada, N.; Ishikawa, K.; Fuse, N.; Kodaira, R.; Shimosaka, M.; Amano, Y.; Kanda, T.; Okazaki, M. Purification, characterization and gene analysis of exo-cellulase II (Ex-2) from the white rot basidiomycete Irpex lacteus. J. Biosci. Bioeng. 1999, 87, 442–451. [Google Scholar] [CrossRef]
- Xu, C.; Ma, F.; Zhang, X. Lignocellulose degradation and enzyme production by Irpex lacteus CD2 during solid-state fermentation of corn stover. J. Biosci. Bioeng. 2009, 108, 372–375. [Google Scholar] [CrossRef] [PubMed]
- Özkan, E.; Ertan, F. Production and determination of some biochemical properties of protease enzyme by Trichothecium roseum under solid state fermentation. Romanian Biotechnol. Lett. 2012, 17, 6903–6912. [Google Scholar]
- Sabotič, J.; Trček, T.; Popovič, T.; Brzin, J. Basidiomycetes harbor a hidden treasure of proteolytic diversity. J. Biotechnol. 2007, 128, 297–307. [Google Scholar] [CrossRef] [PubMed]
- Salvachúa, D.; Martínez, A.T.; Tien, M.; López-Lucendo, M.F.; García, F.; de los Ríos, V.; Martínez, M.J.; Prieto, A. Differential proteomic analysis of the secretome of Irpex lacteus and other white-rot fungi during wheat straw pretreatment. Biotechnol. Biofuels 2013, 6, 115–128. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Chinaglia, S.; Chiarelli, L.R.; Maggi, M.; Rodolfi, M.; Valentini, G.; Picco, A.M. Biochemistry of lipolytic enzymes secreted by Penicillium solitum and Cladosporium cladosporioides. Biosci. Biotechnol. Biochem. 2014, 78, 245–254. [Google Scholar] [CrossRef] [PubMed]
- Chander, H.; Batish, V.K.; Sannabhadti, S.S.; Srinivasan, R.A. Factors affecting lipase production in Aspergillus wentii. J. Food Sci. 1980, 45, 598–600. [Google Scholar] [CrossRef]
- Castellá, G.; Bragulat, M.R.; Rubiales, M.V.; Cabañes, F.J. Malachite green agar, a new selective medium for Fusarium. Mycopathologia 1997, 137, 173–178. [Google Scholar] [CrossRef] [PubMed]
- Leslie, J.F.; Summerell, B.A. The Fusarium Laboratory Manual; Blackwell Publishing: Oxford, UK, 2006. [Google Scholar]
- Nelson, P.E.; Touson, T.A.; Marasas, W.F.O. Fusarium Species, an. Illustrated Manual for Identification; Pennsylvania State University Press: University Park, PA, USA, 1983. [Google Scholar]
- Machado, J.C.; Langerak, C.J.; Jaccoud-Filho, D.S. Seed-Borne Fungi: A Contribution to Routine Seed Health Analysis; ISTA/UFLA: Bassersdorf, Switzerland, 2002. [Google Scholar]
- Pitt, J.I.; Hocking, A.D. Fungi and Food Spoilage, 3rd ed.; Springer: New York, NY, USA, 2009. [Google Scholar]
- Koenig, R.L.; Ploetz, R.C.; Kistler, H.C. Fusarium oxysporum f. sp. cubense consists of a small number of divergent and globally distributed clonal lineages. Phytopathology 1997, 87, 915–923. [Google Scholar]
- White, T.J.; Bruns, T.; Lee, S.; Taylor, J.W. Amplification and direct sequencing of fungal ribosomal RNA genes for phylogenetics. In PCR Protocols, a Guide to Methods and Applications; Innis, M.A., Gelfand, H., Sninsky, J.J., White, T.J., Eds.; Academic Press Inc.: San Diego, CA, USA, 1990; pp. 315–322. [Google Scholar]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Tamura, K.; Nei, M.; Kumar, S. Prospects for inferring very large phylogenies by using the neighbor-joining method. Proc. Natl. Acad. Sci. USA 2004, 101, 11030–11035. [Google Scholar] [CrossRef] [PubMed]
- Ceska, M. Enzyme catalysis of solidified media. Eur. J. Biochem. 1971, 22, 186–192. [Google Scholar] [CrossRef] [PubMed]
- Hankin, L.; Anagnostakis, S.L. The use of solid media for detection of enzyme production by fungi. Mycologia 1975, 67, 597–607. [Google Scholar] [CrossRef]
- Olivares-Bernabeu, C.M.; López-Llorca, L.V. Fungal egg-parasites of plant-parasitic nematodes from Spanish soils. Rev. Iberoam. Micol. 2002, 19, 104–110. [Google Scholar]
- Montenecourt, B.S.; Eveleigh, D.E. Semiquantitative plate assay for determination of cellulase production by Trichoderma viride. Appl. Environ. Microb. 1977, 33, 178–183. [Google Scholar]
- Kasana, R.C.; Salwan, R.; Dhar, H.; Dutt, S.; Gulati, A. A Rapid and easy method for the detection of microbial cellulases on agar plates using Gram’s iodine. Curr. Microbiol. 2008, 57, 503–507. [Google Scholar] [CrossRef] [PubMed]
- Sarath, G.; de La Motte, R.S.; Wagner, F.W. Protease assay methods. In Proteolytics Enzymes: A Pratical Approach; Beynon, R.J., Bonde, J.S., Eds.; University Press: Oxford, UK, 1989; pp. 25–54. [Google Scholar]
- Sierra, G. A simple method for the detection of lipolytic activity of microorganisms and some observations on the influence of the contact between cells and fatty substrates. Anton. Leeuw. 1957, 23, 15–22. [Google Scholar] [CrossRef]
© 2015 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abe, C.A.L.; Faria, C.B.; De Castro, F.F.; De Souza, S.R.; Santos, F.C.d.; Da Silva, C.N.; Tessmann, D.J.; Barbosa-Tessmann, I.P. Fungi Isolated from Maize (Zea mays L.) Grains and Production of Associated Enzyme Activities. Int. J. Mol. Sci. 2015, 16, 15328-15346. https://doi.org/10.3390/ijms160715328
Abe CAL, Faria CB, De Castro FF, De Souza SR, Santos FCd, Da Silva CN, Tessmann DJ, Barbosa-Tessmann IP. Fungi Isolated from Maize (Zea mays L.) Grains and Production of Associated Enzyme Activities. International Journal of Molecular Sciences. 2015; 16(7):15328-15346. https://doi.org/10.3390/ijms160715328
Chicago/Turabian StyleAbe, Camila Agnes Lumi, Carla Bertechini Faria, Fausto Fernandes De Castro, Sandra Regina De Souza, Fabiane Cristina dos Santos, Cleiltan Novais Da Silva, Dauri José Tessmann, and Ione Parra Barbosa-Tessmann. 2015. "Fungi Isolated from Maize (Zea mays L.) Grains and Production of Associated Enzyme Activities" International Journal of Molecular Sciences 16, no. 7: 15328-15346. https://doi.org/10.3390/ijms160715328





