Homology Modeling Study of Bovine μ-Calpain Inhibitor-Binding Domains
Abstract
:1. Introduction
2. Results
2.1. Structural Comparison between Active and Inactive Bovine CAPN1 Models
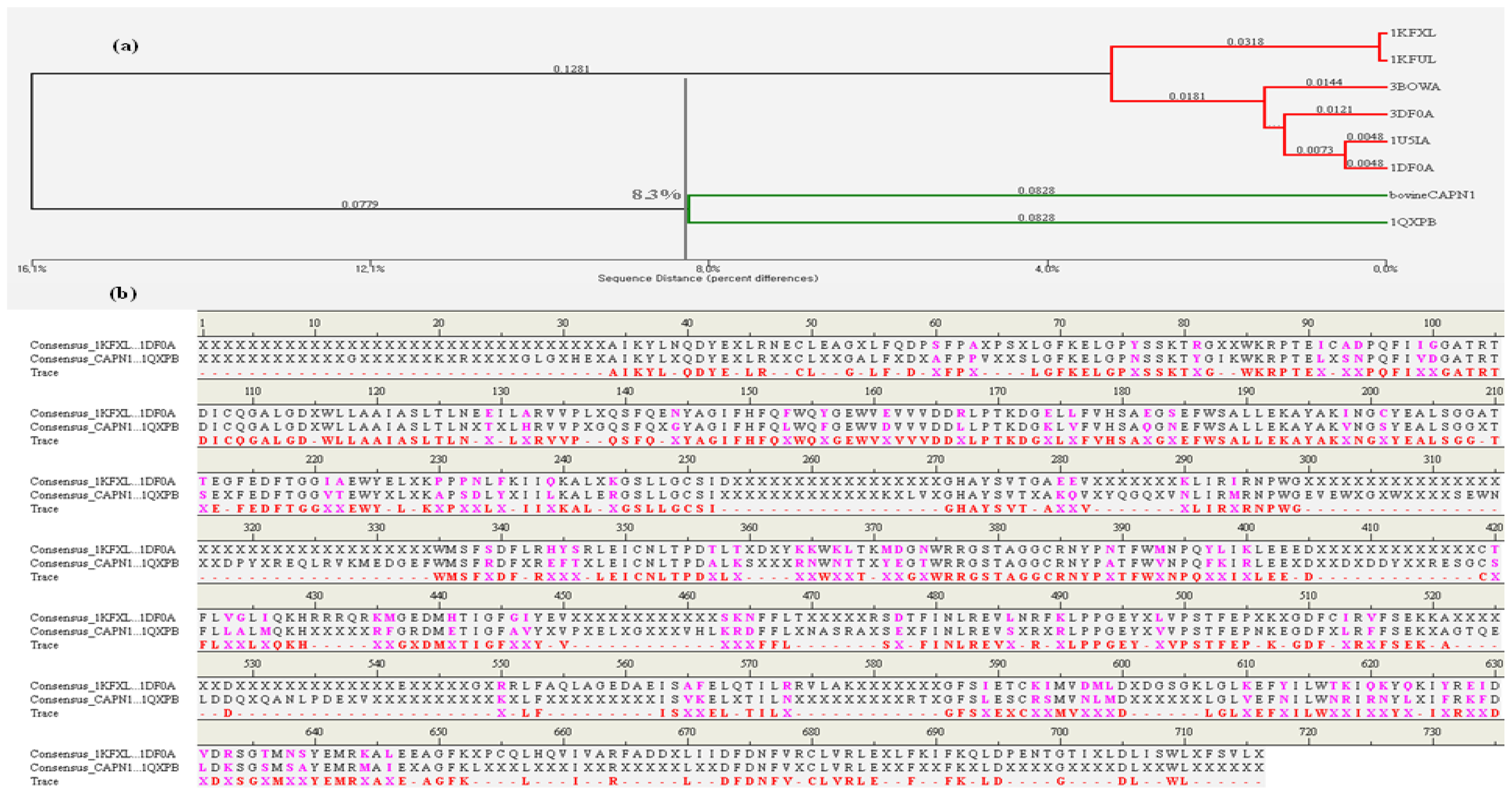
2.1.1. Comparison of the Large and Small Subunits in CAPN1 and CAPN2 Molecules
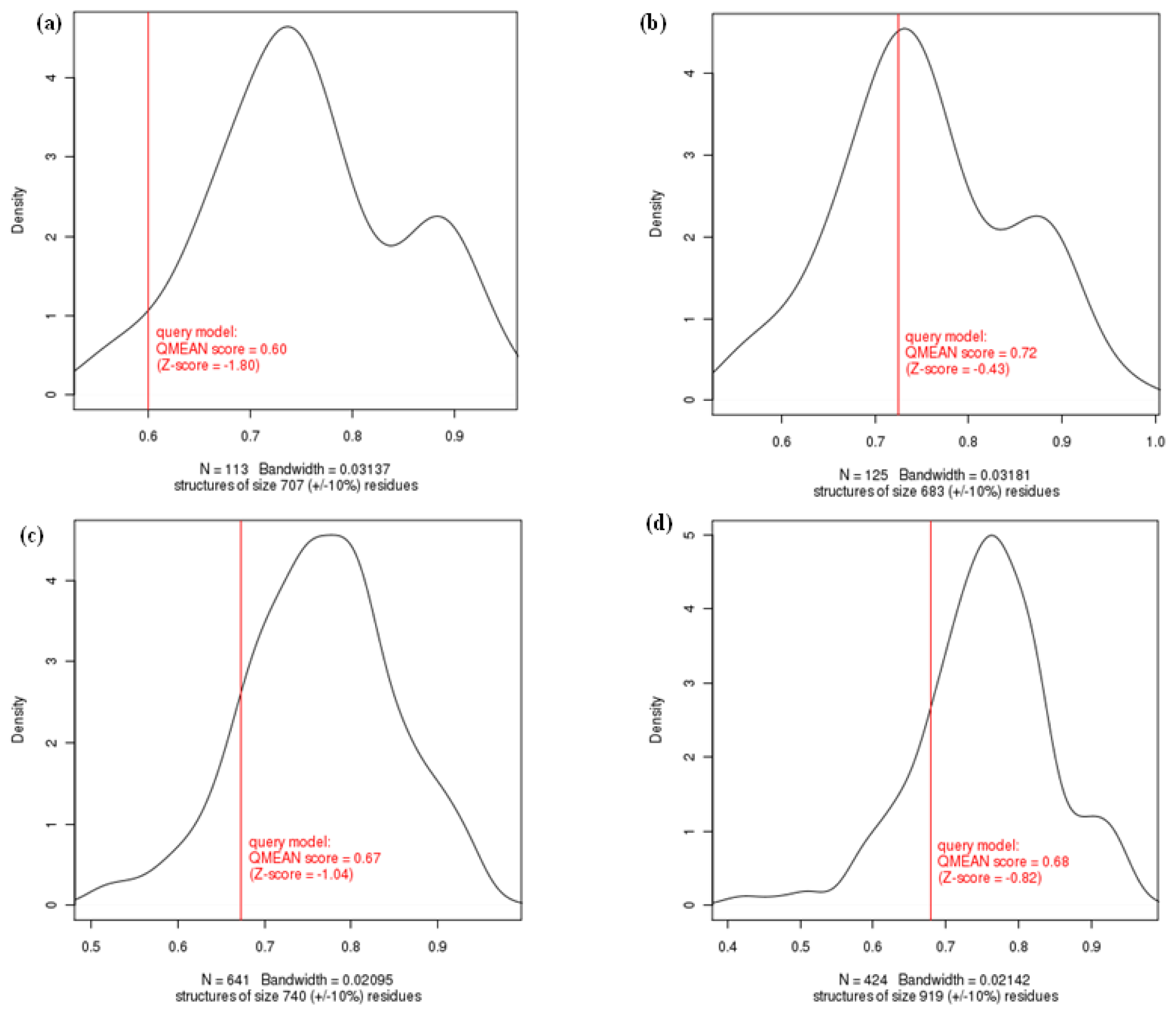
2.1.2. Evaluation of Homology Modeled Proteins
2.1.3. Overlay Structure Similarity Using Field-Based Alignment
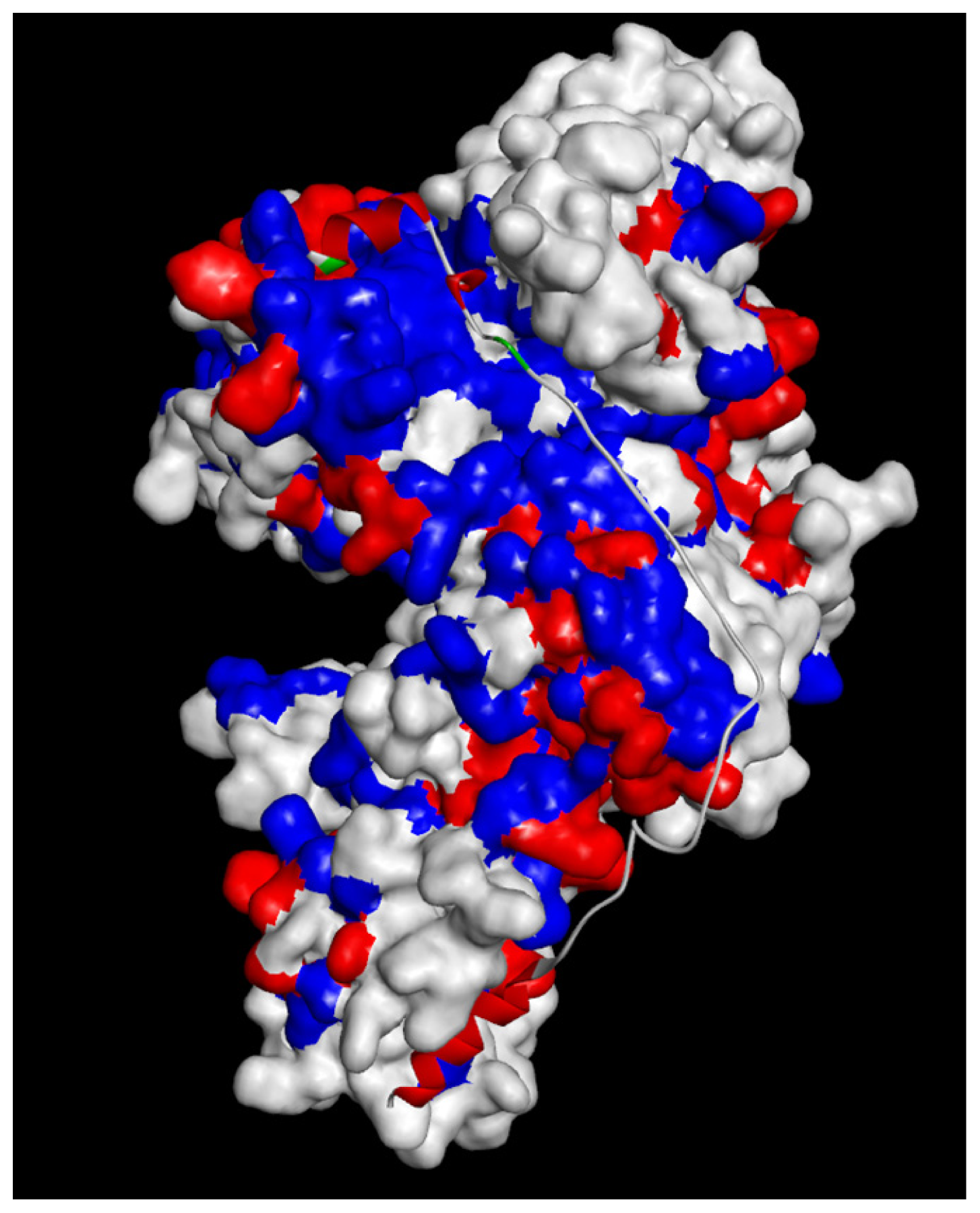
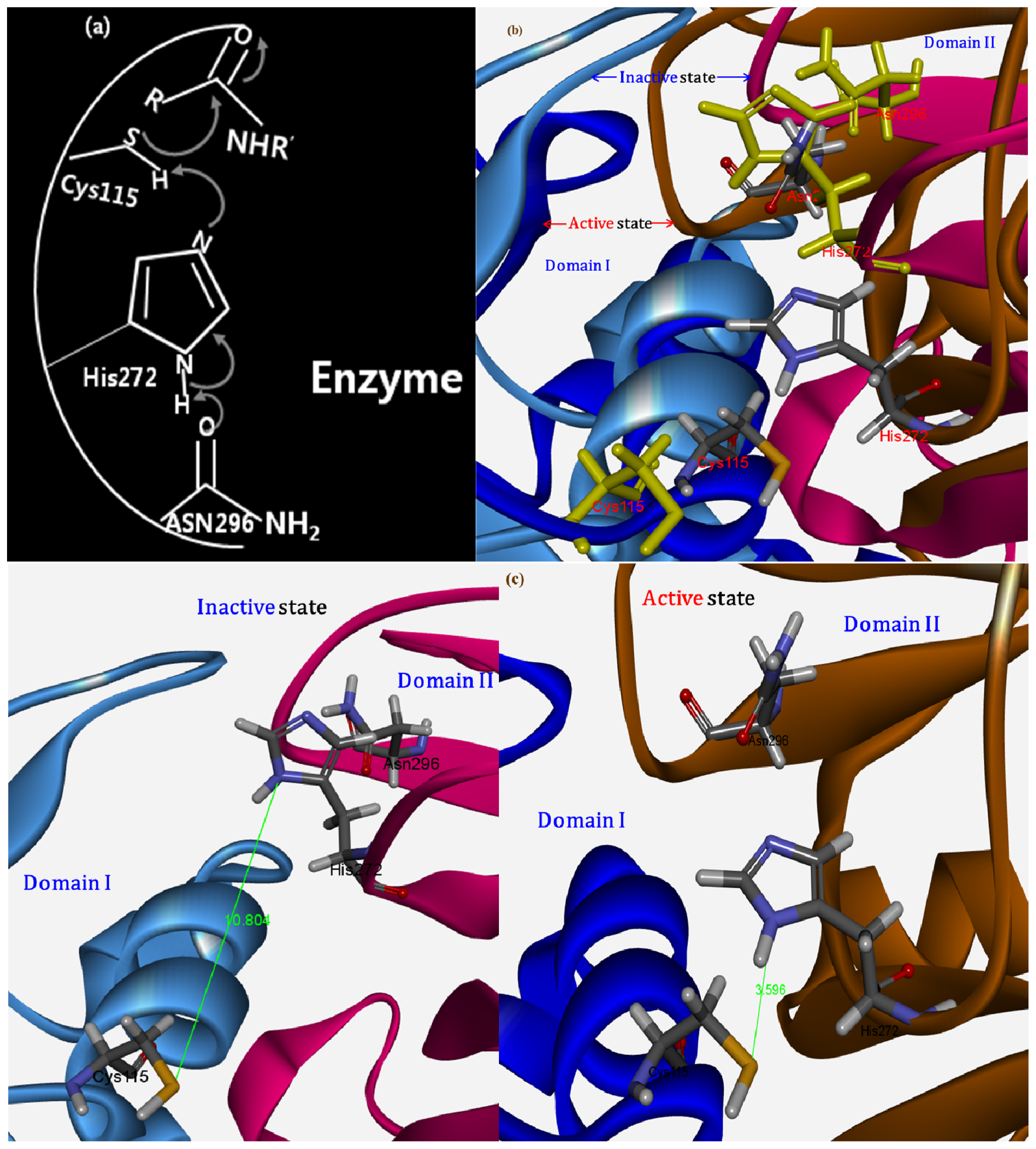
2.1.4. Potential Functional Residues and Interaction Interfaces
2.1.5. Comparison of Ca2+-Binding Patterns in CAPN Subgroups
2.1.6. The Role of Structural Water Molecules for Their Ca2+-Binding Patterns
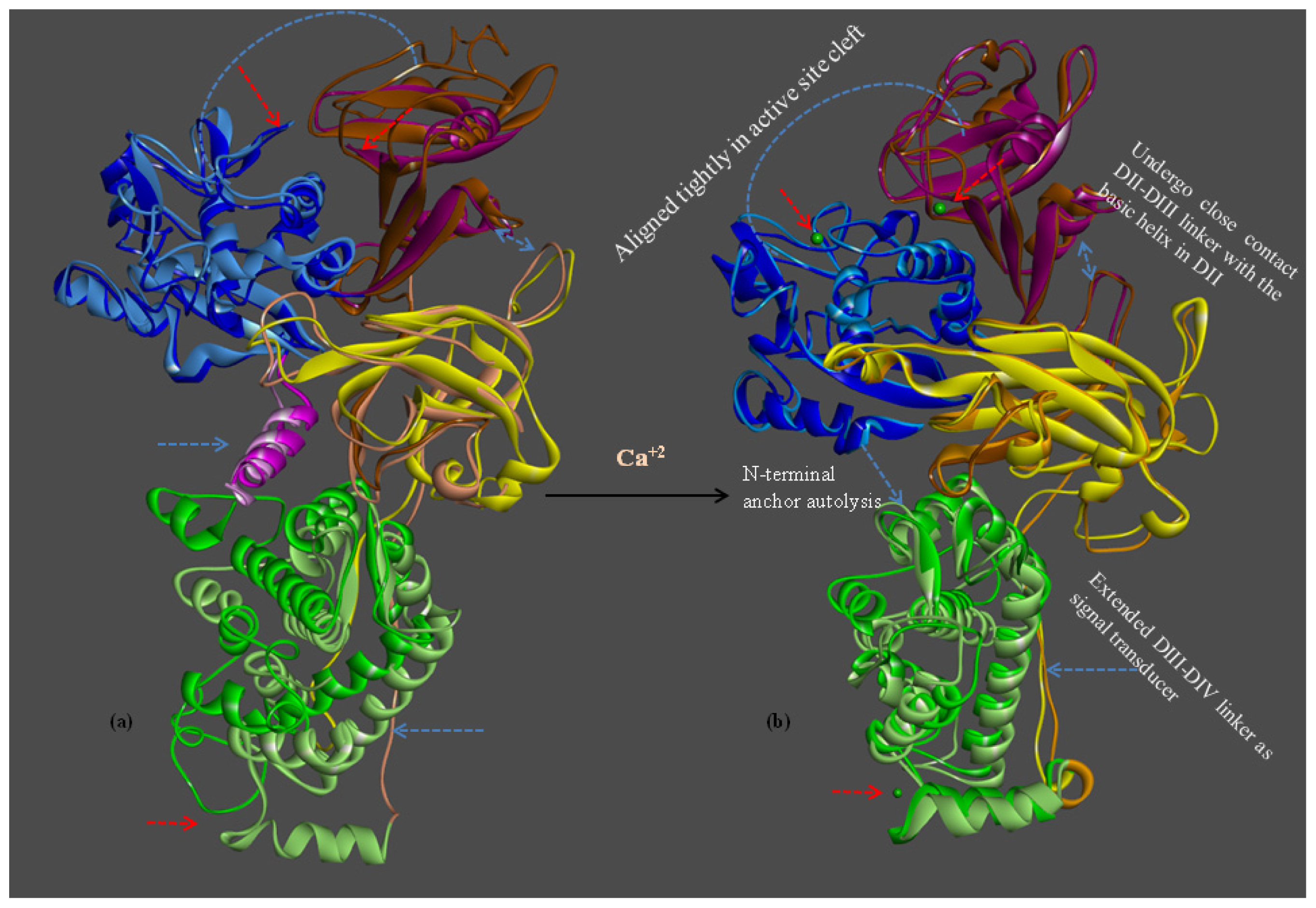
2.1.7. Structure Flexibility and Changes on CAPN Activation
2.2. A Structural Model for the Inhibition of CAPN1 by CAST4 from Both Bovines
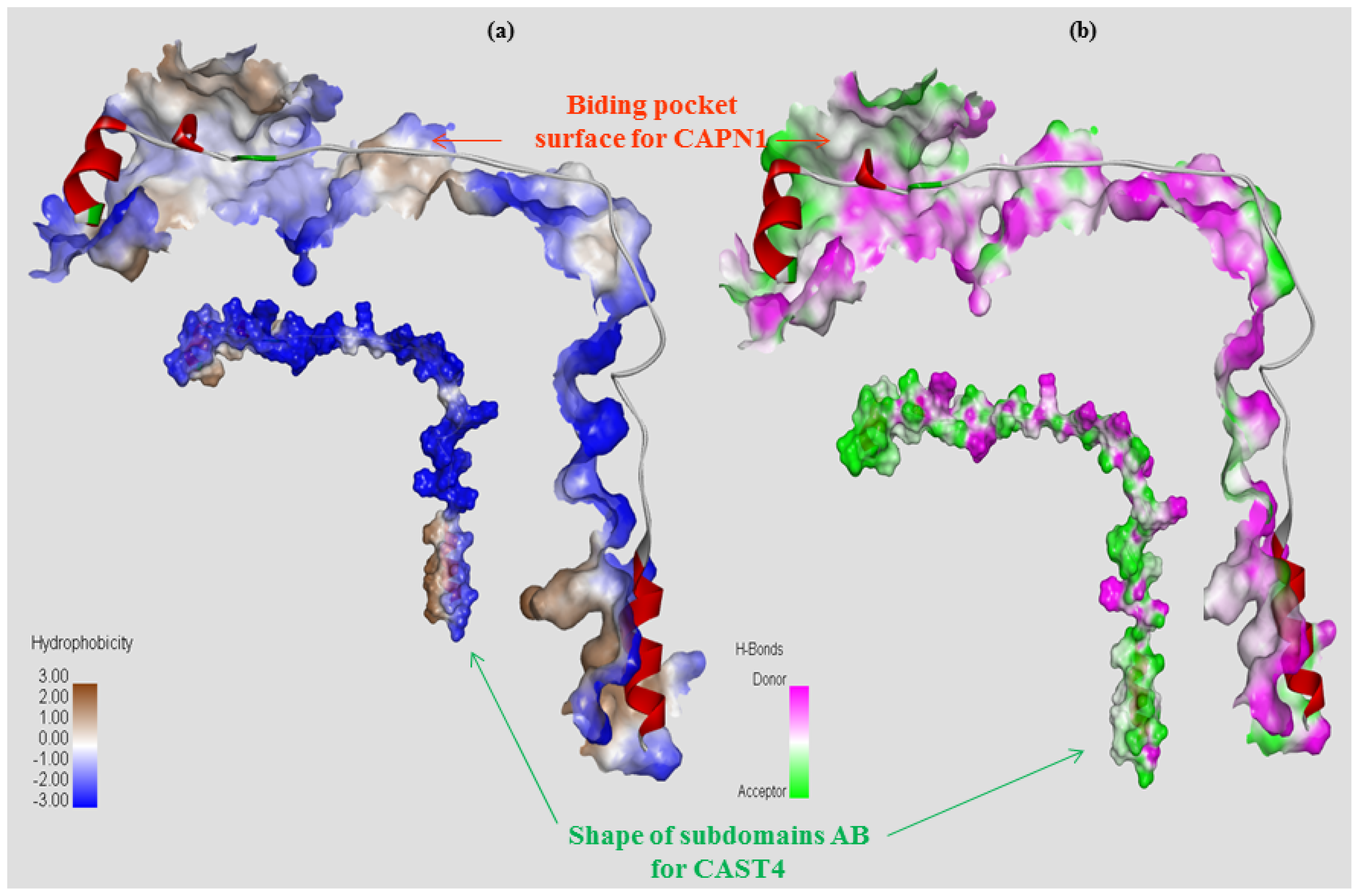
2.2.1. Structure Modeling and Evaluation of the Bovine CAPN1/CAST4 Complex
2.2.2. Functional Characterization of the Inhibitory Motif of CAST4 Subdomain B
3. Discussion
4. Materials and Methods
4.1. A Structural Model of the Bovine CAPN1 Protein in the Absence of Calcium
4.2. Predicting the Complex Structure between Calcium Bound, Bovine CAPN1 and CAST
5. Conclusions
Acknowledgments
Conflicts of Interest
References
- Goll, D.E.; Thompson, V.F.; Li, H.; Wei, W.; Cong, J. The capain system. Physiol. Rev 2003, 83, 731–801. [Google Scholar]
- Davis, T.L.; Walker, J.R.; Finerty, P.J., Jr.; Mackenzie, F.; Newman, E.M.; Dhe-Paganon, S. The crystal structures of human calpains 1 and 9 imply diverse mechanisms of action and auto-inhibition. J. Mol. Biol 2007, 366, 216–229. [Google Scholar]
- Macqueen, D.J.; Delbridge, M.L.; Manthri, S.; Johnston, I.A. A newly classified vertebrate calpain protease, directly ancestral to CAPN1 and 2, episodically evolved a restricted physiological function in placental mammals. Mol. Biol. Evol 2010, 27, 1886–1902. [Google Scholar]
- Preziosa, E.; Liu, S.; Terova, G.; Gao, X.; Liu, H.; Kucuktas, H.; Terhune, J.; Liu, Z. Effect of nutrient restriction and re-feeding on calpain family genes in skeletal muscle of channel catfish (lctalurus punctatus). PLoS One 2013, 8, e59404. [Google Scholar]
- Sorimachi, H.; Ono, Y. Regulation and physiological roles of the calpain system in muscular disorders. Cardiovasc. Res 2010, 96, 11–22. [Google Scholar]
- Suzuki, K.; Hata, S.; Kawabata, Y.; Sorimachi, H. Structure, activation and biology of calpain. Diabetes 2004, 53, S12–S18. [Google Scholar]
- Crawford, C.; Brown, N.R.; Willis, A.C. Studies of the active site of m-calpain and the interaction with calpastatin. Biochem. J 1993, 296, 135–142. [Google Scholar]
- Hanna, R.A.; Campbell, R.L.; Davies, P.L. Calcium-bound structure of calpain and its mechanism of inhibition by calpastatin. Nature 2008, 456, 409–412. [Google Scholar]
- Moldoveanu, T.; Gehring, K.; Green, D.R. Concerted multi-pronged attack by calpastatin to occlude the catalytic cleft of heterodimeric calpains. Nature 2008, 456, 404–408. [Google Scholar]
- Suzuki, K. The structure of calpains and the calpain gene. In Intracelluar Calcium-Dependent Proteolysis; Mellgren, R.L., Murachi, T., Eds.; CRC: Boca Raton, FL, USA, 1990; pp. 25–35. [Google Scholar]
- Pal, G.P.; de Veyra, T.; Elce, J.S.; Jia, Z. Crystal structure of a μ-like calpain reveals a partially activated conformation with low Ca2+ requirement. Structure 2003, 11, 1521–1526. [Google Scholar]
- Dutt, P.; Spriggs, C.N.; Davies, P.L.; Jia, Z.; Elce, J.S. Origins of the difference in Ca2+ requirement for activation of μ-and m-calpain. Biochem. J 2002, 367, 263–269. [Google Scholar]
- Cuerrier, D.; Moldoveanu, T.; Davies, P.L. Determination of peptide substrate specificity for μ-calapain by a peptide library-based approach. J. Biol. Chem 2005, 280, 40632–40641. [Google Scholar]
- DuVerle, D.A.; Ono, Y.; Sorimachi, H.; Mamitsuka, H. Calpain cleavage prediction using multiple kernel learning. PLoS One 2011, 6, e19035. [Google Scholar]
- Tompa, P.; Buzder-Lantos, P.; Tantos, A.; Farkas, A.; Szilágyi, A.; Bánóczi, Z.; Hudecz, F.; Friedrich, P. On the sequential determinants of calpain cleavage. J. Biol. Chem 2004, 279, 20775–20785. [Google Scholar]
- Yoshizawa, T.; Sorimachi, H.; Tomioka, S.; Ishiura, S.; Suzuki, K. A catalytic subunit of calpain possess full proteolytic activity. FEBS Lett 1995, 358, 101–103. [Google Scholar]
- Yoshizawa, T.; Sorimachi, H.; Tomioka, S.; Ishiura, S.; Suzuki, K. Calpain dissociate into subunits in the presence of calcium ions. Biochem. Biophys. Res. Commun 1995, 208, 376–383. [Google Scholar]
- Todd, B.; Moore, D.; Deivanayagam, C.C.; Lin, G.D.; Chattopadhyay, D.; Maki, M.; Wang, K.K.; Narayana, S.V. A structural model for the inhibition of calpain by calpastatin: crystal structures of the native domain VI of calpain and its complexes with calpastatin peptide a small molecule inhibitor. J. Mol. Biol 2003, 328, 131–146. [Google Scholar]
- Zimmerman, U.J.P.; Schlaepfer, W.W. Two stage autolysis of the catalytic subunit initiates activation of calpain I. Biochim. Biophys. Acta 1991, 1078, 192–198. [Google Scholar]
- Hosfield, C.M.; Elce, J.S.; David, P.L.; Jia, Z.C. Crystal structure of calpain reveals the structural basis for Ca2+-dependent protease activity and a novel mode of enzyme activation. EMBO J 1999, 18, 6880–6889. [Google Scholar]
- Strobl, S.; Fernandez-Catalan, C.; Braun, M.; Huber, R.; Masumoto, H.; Nakagawa, K. The crystal structure of calcium-free human m-calpain suggests an electrostatic switch mechanism for activation by calcium. Proc. Natl. Acad. Sci. USA 2000, 97, 588–592. [Google Scholar]
- Lin, G.D.; Chattopadhyay, D.; Maki, M.; Wang, K.K.; Carson, M.; Jin, L.; Yuen, P.W.; Takano, E.; Hatanaka, M.; DeLucas, L.J.; et al. Crystal structure of calcium bound domain VI of calpain at 1.9 Å resolution and its role in enzyme assembly, regulation, and inhibitor binding. Nat. Struct. Biol 1997, 4, 539–547. [Google Scholar]
- Blanchard, H.; Grochulski, P.; Li, Y.; Arthur, J.S.; Davies, P.L.; Elce, J.S.; Cygler, M. Structure of a calpain Ca2+-binding domain reveals a novel EF-hand and Ca2+-induced conformational changes. Nat. Struct. Biol 1997, 4, 532–538. [Google Scholar]
- Schechter, I.; Berger, A. On the size of the active site in proteases. I. Papain. Biochem. Biophys. Res. Commun 1967, 27, 157–162. [Google Scholar]
- Sorimachi, H.; Hata, S.; Ono, Y. Impact of genetic insights into calpain biology. J. Biochem 2011, 150, 23–37. [Google Scholar]
- Croall, A.D.; DeMartino, G.N. Calcium-activated neutral protease (calpain) systems structure, function, and regulation. Physiol. Rev 1991, 71, 813–847. [Google Scholar]
- Lee, M.S.; Kwon, Y.T.; Li, M.; Peng, J.; Friedlander, R.M.; Tsai, L.H. Neurotoxicity induces cleavage of p35 to p25 by calpain. Nature 2000, 405, 360–364. [Google Scholar]
- Gregoriou, M.; Willis, A.C.; Pearson, M.A.; Crawford, C. The calpain cleavage sites in the epidermal growth factor receptor kinase domain. Eur. J. Biochem 1994, 223, 455–464. [Google Scholar]
- Hayashi, M.; Saito, Y.; Kawashima, S. Hydrolysis of protamine by calcium-activated neutral protease (CANP). J. Biochem 1985, 97, 1363–1370. [Google Scholar]
- Lovell, S.C.; Davis, I.W.; Arendale, W.B.; de Bakker, P.I.; Word, J.M.; Prisant, M.G.; Richardson, J.S.; Richardson, D.C. Structure Validation by C-alpha geometry: phi, psi and C-beta deviation. Proteins 2003, 50, 437–450. [Google Scholar]
- Benkert, P.; Biasini, M.; Schwede, T. Toward the estimation of absolute quality of individual protein structure models. Bioinformatics 2011, 27, 343–350. [Google Scholar]
- Kearsly, S.K.; Smith, G.M. An alternative method for the alignment of molecular structures: Maximizing electrostatic and steric overlap. Tetrahedron Comput. Methodol 1990, 13, 615–633. [Google Scholar]
- Hartigan, J.A. Clustering Algorithms; John Wiley & Sons: New York, NY, USA, 1975. [Google Scholar]
- Moldoveanu, T.; Jia, Z.; Davies, P.L. Calpain activation by cooperate Ca2+ binding at two non-EF-hand sites. J. Biol. Chem 2004, 279, 6106–6114. [Google Scholar]
- Tudor, M.; Christopher, M.H.; Daniel, L.; John, S.E.; Zongchao, J.; Peter, L.D. A Ca2+ switch aligns the active site of calpain. Cell 2002, 108, 649–660. [Google Scholar]
- Zenon, G. Structure of a trapped intermediate of calmodulin: Calcium regulation of EF-hand Proteins from a new perspective. J. Mol. Biol 2005, 346, 1351–1366. [Google Scholar]
- Zenon, G. Insights into modulation of calcium signaling by magnesium in calmodulin, troponin C and regulated EF-hand proteins. Biochim. Biophys. Acta 2011, 1813, 913–921. [Google Scholar]
- Pietsch, M.; Chua, K.C.; Abell, A.D. Calpains: Attractive targets for the development of synthetic inhibitors. Curr. Top Med. Chem 2010, 10, 270–293. [Google Scholar]
- Moldoveanu, T.; Campbell, R.L. Crystal structure of calpain-E64 and -Leupeptin inhibitor complexes reveal mobile loops gating the active site. J. Mol. Biol 2004, 343, 1313–1326. [Google Scholar]
- Jia, Z.; Petrounevitch, V.; Wong, A.; Moldoveanu, T.; Davies, P.L.; Elce, J.S.; Beckmann, J.S. Mutations in calpain 3 associated with limb girdle muscular dystrophy: Analysis by molecular modeling and by mutation in m-calcpain. Biophys. J 2001, 80, 2590–2596. [Google Scholar]
- Saez, M.E.; Ramirez-Lorca, R.; Moron, F.J.; Ruiz, A. The therapeutic potential of the calpain family: new aspects. Drug Discov. Today 2006, 11, 917–923. [Google Scholar]
- Hosfield, C.M.; Moldoveanu, T.; Davies, P.L.; Elce, J.S.; Jia, Z. Calpain mutants with increased Ca2+ sensitivity and implications for the role of the C2-like domain. J. Biol. Chem 2001, 276, 7404–7407. [Google Scholar]
- Hosfield, C.M.; Elce, J.S.; Jia, Z. Activation of calpain by Ca2+: Roles of the large subunit N-terminal and domain III-IV linker peptides. J. Mol. Biol 2004, 343, 1049–1053. [Google Scholar]
- Tompa, P.; Emori, Y.; Sorimachi, H.; Suzuki, K.; Friedrich, P. Domain III of calpain is a Ca2+-regulated phospholipid-binding domain. Biochem. Biophys. Res. Commun 2001, 280, 1333–1339. [Google Scholar]
- Shiraha, H.; Glading, A.; Chou, J.; Jia, Z.; Wells, A. Activation of m-calpain (calpain II) by epidermal growth factor is limited by protein kinase A phosphorylation of m-calapin. Mol. Cell. Biol 2002, 22, 2716–2727. [Google Scholar]
- Turk, B. Targeting protease: Successes, failures and future prospects. Nat. Rev. Drug Discov 2006, 5, 785–799. [Google Scholar]
- Betts, R.; Weinsheimer, S.; Blouse, G.E.; Anagli, J. Structural determinants of the calpain inhibitory activity of calpastatin peptide B27-WT. J. Biol. Chem 2003, 278, 7800–7809. [Google Scholar]
- Seung-Hwan, L.; Seung-Chang, K.; Han-ha, C.; Soo-Hyun, C.; Hyeong-Cheol, K.; Dajeong, L.; Bong-Hwan, C.; Chang-Gwan, D.; Cedric, G.; Boh-suk, Y.; et al. Mutations in calpastatin and μ-calpain are associated with meat tenderness, flavor, and juiciness of Hanwoo (Korean cattle): Molecular modeling the effects of substitutions in the calpastatin/μ-calpain complex. Meat Sci 2014, 96, 1501–1508. [Google Scholar]
- Cong, M.; Thompson, V.F.; Goll, D.E.; Antin, P.B. The bovine calpastatin gene promoter and a new N-terminal region of the protein are targets for cAMP-dependent protein kinase activity. J. Biol. Chem 1998, 273, 660–666. [Google Scholar]
- Killefer, J.; Koohmaraie, M. Bovine skeletal muscle calpastatin: cloning sequence analysis, and steady-state mRNA expression. J. Anim. Sci 1994, 72, 604–614. [Google Scholar]
- Maki, M.; Ma, H.; Takano, E.; Adachi, Y.; Lee, W.J.; Hatanaka, M.; Murachi, T. Calpastatins: Biochemical and molecular biological studies. Biomed. Biochem. Acta 1991, 50, 509–516. [Google Scholar]
- Hanna, R.A.; Garcia-Diaz, B.E.; Davis, P.L. Calpastatin simultaneously binds four calpains with different kinetics constants. FEBS Lett 2007, 581, 2894–2898. [Google Scholar]
- Hata, S.; Abe, M.; Suzuki, H.; Kitamura, F.; Toyama-Sorimachi, N.; Abe, K.; Sakimura, K.; Sorimachi, H. Calpain 8/nCL-2 and calpain 9/nCL-4 constitute an active protease complex, G-calpain, involved in gastric mucosal defense. PLoS Genet 2010, 6, e1001040. [Google Scholar]
- Kawasaki, H.; Emori, Y.; Imajoh-Ohmi, S.; Minami, Y.; Suzuki, K. Identification and characterization of inhibitory sequences in four repeating domains of the endogenous inhibitor of calcium-dependent protease. J. Biochem 1989, 106, 274–281. [Google Scholar]
- Singh, N.; Shrivastav, A.; Olson, D.; Lakshmikuttyamma, A.; Ross, A.; Parr, T.; Bardsley, R.G.; Sharma, R.K. Cardiac high molecular weight calmodulin-binding protein is homologous to calpastatin I and calpastatin II. Biochem. Biophys. Res. Commun 2008, 373, 387–391. [Google Scholar]
- Skoog, B.; Wichman, A. Calculation of the isoelectric points of polypeptides from the amino acid composition. Trends Anal. Chem 1986, 5, 82–83. [Google Scholar]
- Lee, W.J.; Ma, H.; Takano, E.; Yang, H.Q.; Hatanaka, M.; Maki, M. Molecular diversity in amino-terminal domains of human calpastatin by exon skipping. J. Biol. Chem 1992, 267, 8437–8442. [Google Scholar]
- Nishimura, T.; Goll, D.E. Binding of calpain fragments to calpastatin. J. Biol. Chem 1991, 266, 11842–11850. [Google Scholar]
- Ma, H.; Yang, H.Q.; Takano, E.; Hatanaka, M.; Maki, M. Amino-terminal conserved region in proteinase inhibitor domain of calpastatin potentiates its calpain inhibitory activity by interesting with calmodulin-like domain of proteinase. J. Biol. Chem 1994, 269, 24430–24436. [Google Scholar]
- Maki, M.; Bagci, H.; Hamaguchi, K.; Ueda, M.; Murachi, T.; Hatanaka, M. Inhibition of calpain by a synthetic oligopeptide corresponding to an exon of the human calpastatin gene. J. Biol. Chem 1989, 264, 18866–18869. [Google Scholar]
- Ono, Y.; Sorimachi, H. Calpains: An elaborate proteolytic system. Biochim. Biophys. Acta 2012, 1824, 224–236. [Google Scholar]
- Sali, A.; Pottertone, L.; Yuan, F.; Vlijmen, H.; Karplus, M. Evaluation of comparative protein modeling by MODELLER. Prot. Struct. Funct. Genet 1995, 23, 318–326. [Google Scholar]
- Shen, M.Y.; Sali, A. Statistical potential for assessment and prediction of protein structures. Protein Sci 2006, 15, 2507–2524. [Google Scholar]
- Sali, A.; Overington, J.P. Derivation of rules for comparative protein modeling from a database of protein structure alignments. Protein Sci 1994, 31, 582–1596. [Google Scholar]
- Discovery Studio. Version 3.1; Available online: http://accelrys.com/products/discovery-studio/ accessed on 7 March 2013.
- Castrignanó, T.; de Meo, P.D.; Cozzetto, D.; Talamo, I.G.; Tramontano, A. The PMDB protein model database. Nucleic Acids Res 2006, 34, D306–D309. [Google Scholar]





| CAPN1 vs. Bovine | Sequence identity (%) | ||||
|---|---|---|---|---|---|
| Domain I (DI) | Domain II (DII) | Domain III (DIII) | Domain IV (DIV) | Overall * | |
| House mouse | 91.6 | 85.5 | 87.3 | 93.6 | 89.5 |
| Norway rat | 91.6 | 84.9 | 87.3 | 93.6 | 89.7 |
| Dog | 95.3 | 91.0 | 94.3 | 97.7 | 94.7 |
| Rhesus monkey | 94.8 | 93.4 | 93.6 | 98.3 | 95.1 |
| Human | 95.3 | 91.6 | 94.3 | 98.3 | 94.7 |
| Chimpanzee | 95.3 | 92.2 | 94.3 | 98.3 | 94.8 |
| Bovine CAPN2 | 77.0 | 65.7 | 59.9 | 51.7 | 62.5 |
| QMEAN6 score | Inactive template structure (1QXP:B) | Inactive bovine CAPN1 structural model | Activated template structure (1KXR:B) | Activated bovine CAPN1 structural model | CAST4 bound to CAPN2 of template structure (3BOW) | CAST4 bound to CAPN1 structural model | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Raw score | Z-score | Raw score | Z-score | Raw score | Z-score | Raw score | Z-score | Raw score | Z-score | Raw score | Z-score | |
| C-beta interaction energy | −170.90 | −0.82 | −124.35 | −1.00 | −73.49 | −1.20 | −160.43 | −0.63 | −314.42 | −0.01 | −213.98 | −0.30 |
| All-atom pairwise energy | −16,945.12 | −0.41 | −14,017.57 | −0.85 | −7440.68 | −0.89 | −17,049.35 | −0.18 | −30,398.27 | 0.40 | −19,083.34 | −0.46 |
| Solvation energy | −31.92 | −1.62 | −27.92 | −1.65 | −11.59 | −2.08 | −36.17 | −1.21 | −110.00 | 0.60 | −52.80 | −1.30 |
| Torsion angle energy | −69.46 | −2.81 | −79.90 | −2.55 | −111.55 | 1.03 | −179.04 | −0.24 | −230.02 | −0.34 | −190.68 | −0.27 |
| Secondary structure agreement | 78.2% | −0.12 | 80.2% | 0.28 | 74.8% | −1.40 | 80.5% | 0.35 | 80.4% | −0.19 | 78.1% | −0.76 |
| Solvent accessibility agreement | 74.7% | −1.16 | 74.9% | −1.13 | 89.4% | 1.64 | 78.2% | −0.54 | 76.6% | −0.58 | 75.9% | −0.88 |
| QMEAN6 score | 0.592 | −1.90 | 0.606 | −1.74 | 0.915 | 1.58 | 0.724 | −0.43 | 0.678 | −0.82 | 0.673 | −1.04 |
| Residues | 788 | 703 | 321 | 683 | 914 | 742 | ||||||
| PDFs total energy | - | 5790.3301 | - | 1348.0272 | - | 108,223 | ||||||
| DOPE score | - | −74,339.4609 | - | −80,511.2891 | - | −86,723.3 | ||||||
| PDB ID | Molecular description | The Ca2+ binding site | The residue conservation pattern * |
|---|---|---|---|
| 1KXR | Ca2+ bound protease core of CAPN1(rat) | Site 1: Val99, Gly101, Asp106, Glu185 Site 2: Glu302, Asp309, Met 329, Asp331, Glu333 | Site 1: VGDE Site 2: EDMDE Site 3: ADEE |
| 1ZCM | CAPN1 protease core inhibited by ZLLYCH2F(human) | Site 1: Val99, Gly101, Asp106, Glu185 Site 2: Glu302, ASP309, Met329, Asp331, Glu333 | |
| Activated CAPN1 structure model (bovine) | Site 1: Val99, Asp100, Gly101, Thr103, Asp106, Glu185 Site 2: Glu302, ASP309, Met329, Asp331, Glu333 Site 3: Ala557, Asp560, Glu562, Glu567 | ||
| 3BOW | Complex of CAPN2 and CAST(rat) | Site 1: Ile89, Gly90, Gly91, Asp96, Glu175 Site2: Glu292, Asp299, Gln319, Asp321, Glu323 Site3: Ala542, Asp545, Glu547, Glu552 Site 4: Asp585, Asp587, Ser589, Lys591, Glu596 Site 5: Asp615, Asp617, Ser618, Thr621, Glu626 Site 6: Asp658, Asn661 | Site 1: IGGDE Site 2: EDQDE Site 3: ADEE Site 4: DDSLE Site 5:DDRTE Site 6:DDDN |
| 3DF0 | Complex of CAPN2 and CAST(rat) | Site 1: Ile89, Gly90, Gly91, Asp96, Glu175 Site2: Glu292, Asp299, Gln319, Asp321, Glu323 Site 3: Ala542, Asp545, Ala546, Glu547, Glu552 Site 4: Glu547, Asp585, Asp587, Ser589, Lys591, Glu596 Site 5: Asp615, Asp617, Ser618, Thr621, Glu626 Site 6: Asp570, Asp658, Asp660, Asn661 |
| PDB ID | Molecular description | Experimental details | Length | Sequence identity (%) | Sequence similarity (%) | E-value | Template |
|---|---|---|---|---|---|---|---|
| 1QXP:B | Like CAPN1 (rat) | X-ray Diffraction (2.80 Å) | 788 | 74.6 | 82.4 | 0.0 | T |
| 1KXR:B | Ca2+ bound protease core of CAPN1 (rat) | X-ray Diffraction (2.07 Å) | 321 | 80.2 | 85.8 | 1 × 10−17 | T |
| 3BOW:A | CAPN2 catalytic subunit (rat) | X-ray Diffraction (2.40 Å) | 680 | 59.4 | 77 | 0.0 | T |
| 3BOW:C | CAST 4 domain (rat) | X-ray Diffraction (2.40 Å) | 65 | 72.9 | 86.4 | 3 × 10−10 | T |
| 1KFX:L | Form I of CAPN2 (human) | X-ray Diffraction (3.15 Å) | 640 | 57.1 | 75.7 | 0.0 | - |
| 1KFU:L | Form II of CAPN2 (human) | X-ray Diffraction (2.50 Å) | 699 | 61.4 | 81.3 | 0.0 | - |
| 3DF0:A | CAPN2 catalytic subunit (rat) | X-ray Diffraction (2.95 Å) | 676 | 59.5 | 77.5 | 0.0 | - |
| 1U5I:A | CAPN2 mutant Lys10Thr (rat) | X-ray Diffraction (2.86 Å) | 625 | 56.1 | 73.4 | 0.0 | - |
| The regulation system | Interaction regions | Calpain residues * |
|---|---|---|
| The catalytic subunit of CAPN1/subdomains AB of CAST4 complex from both bovines (model structure) | CAPN1 and CAST4 subdomain B | Leu73, Ser78, Lys79, Ile83, Cys108, Gln109, Gly110, Ala111, Leu112, Gly113, Cys115, Trp116, Lys171, Lys174, Leu175, Val176, His179, Ser209, Thr210, Glu212, Ser251, Asp253, Ile254, Ser255, Ser256, Asp259, Met260, Ala262, Val263, Val269, Lys270, Gly271, His272, Ala273, Trp298, Glu300, Arg347, Glu349, Cys384, Arg385, Asn386, Pro388, Trp392, Asp439, Met440, Thr442, Tyr448, Lys463, Asp465, Phe467, Leu468, Ser469, Asn470, Ala471, Ser472, Arg475, Gln478, Phe479, Ile480, Asn481, Leu482, Arg483, Phe503 |
| CAPN1 and CAST4 subdomain A | Asn548, Leu552, Gln555, Leu556, Ile571, Arg574, Ile575, Lys578, His579, Asp581, Phe612, Trp616, Ile619, Arg620, Leu623, Arg627, Asp630, Leu631, Lys633, Gly635 | |
| The overall CAPN2/CAST4 complex from both rats (PDB ID: 3BOW) | CAP2 and CAST4 subdomain B | Leu63, Gly100, Ala101, Leu102, Lys161, Leu169, Gly198, Ile244, Thr245, Asp249, Gly261, His262, Ala263, Trp288, Arg375, Asn376, Leu454, Thr455, Arg457, Ala458, Arg461, Phe465, Asn467, Phe489 |
| CAP2 and CAST4 subdomain A | Leu537, Gln540, Leu541, Ile556, Val560, Arg564, Trp601, Gln605, Gln608, Arg612, Asp615 | |
| CAP2 and CAST4 subdomain C | Leu106, Leu110, Ile125, Val129, Arg132, His133, Trp170 | |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Chai, H.-H.; Lim, D.; Lee, S.-H.; Chai, H.-Y.; Jung, E. Homology Modeling Study of Bovine μ-Calpain Inhibitor-Binding Domains. Int. J. Mol. Sci. 2014, 15, 7897-7938. https://doi.org/10.3390/ijms15057897
Chai H-H, Lim D, Lee S-H, Chai H-Y, Jung E. Homology Modeling Study of Bovine μ-Calpain Inhibitor-Binding Domains. International Journal of Molecular Sciences. 2014; 15(5):7897-7938. https://doi.org/10.3390/ijms15057897
Chicago/Turabian StyleChai, Han-Ha, Dajeong Lim, Seung-Hwan Lee, Hee-Yeoul Chai, and Eunkyoung Jung. 2014. "Homology Modeling Study of Bovine μ-Calpain Inhibitor-Binding Domains" International Journal of Molecular Sciences 15, no. 5: 7897-7938. https://doi.org/10.3390/ijms15057897





