Cadmium Toxicity Induced Alterations in the Root Proteome of Green Gram in Contrasting Response towards Iron Supplement
Abstract
:1. Introduction
2. Results
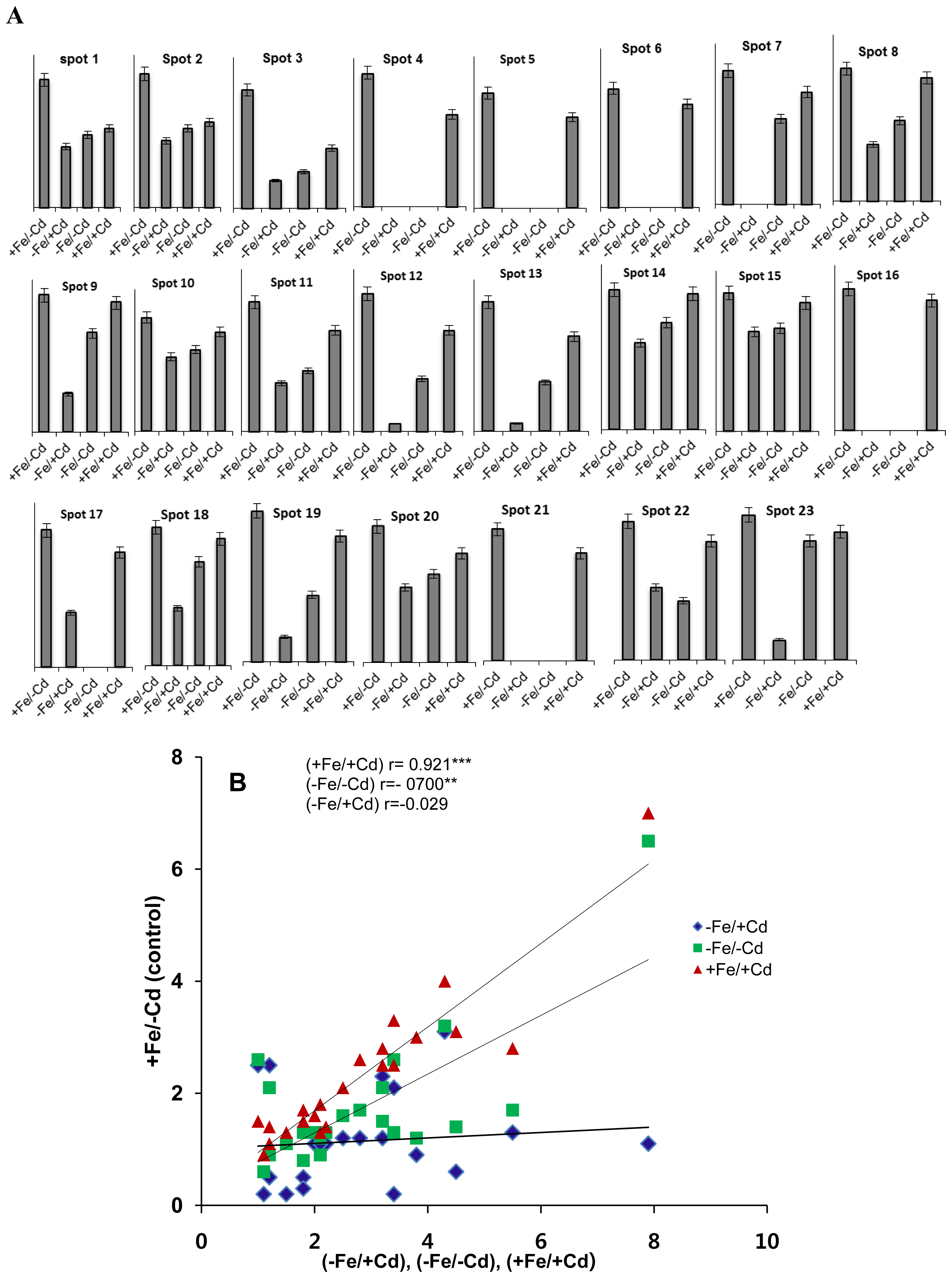
2.1. Differentially Expressed Proteins in Roots by 2DE
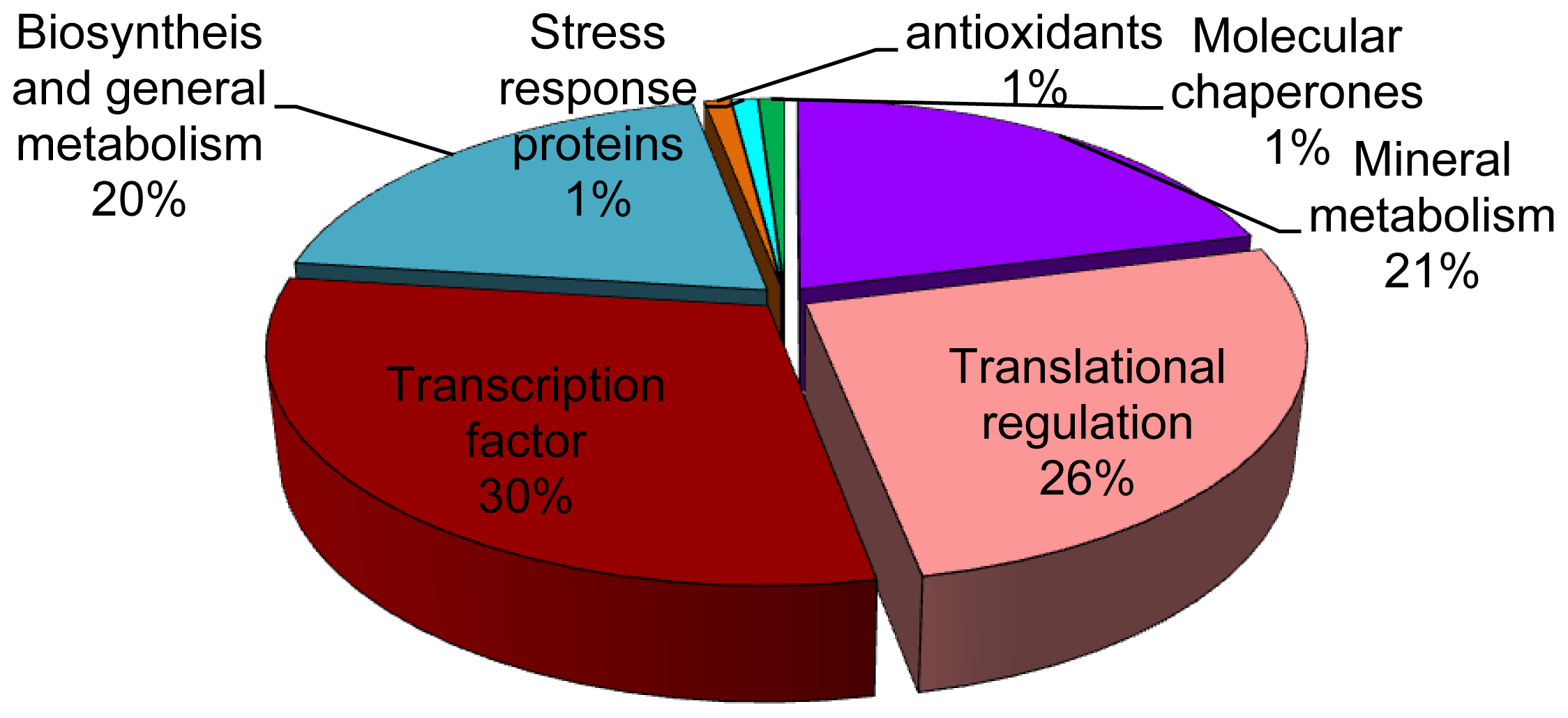
2.2. Identification of Differentially Expressed Proteins in Roots
2.3. Comparison of Diffrentailly Expressed Proteins among Four Treatments
3. Discussion
4. Methods
4.1. Plant Culture and Treatment
4.2. Protein Extraction, 2-D SDS-PAGE and Image Analysis
5. Conclusions
Acknowledgments
Conflicts of Interest
Abbreviations
| 2DE | two dimensional gel electrophoresis |
| SDS-PAGE | sodium dodecyl sulfate poly acrylamide gel electrophoresis) |
| MALDI-TOF | matrix assisted laser desorption/ionization-time of flight |
| IPG | immobilized pH gradient |
| IAA | Iodoacetamide |
| DTT | dithiothriotol |
- Author ContributionsSowbiya Muneer designed and performed the experiments, Khalid Rehman Hakeem analyzed the data, Rozi Mohamed contributed in analytical softwares, Jeon Hyun Lee contributed in statistical analysis of the data and helpful discussions, Sowbiya Muneer and Khalid Rehman Hakeem wrote the manuscript.
References
- Wu, H.; Chen, C.; Du, J.; Liu, H.; Yan, C.; Zhang, Y.; He, Y.; Wang, Y.; Chu, C.; Feng, Z.; Li, J.; Ling, H.Q. Co-over expression FIT with AtbHLH38 or AtbHLH39 in Arabidopsis-Enhanced Cadmium Tolerance via Increased Cadmium Sequestration in Roots and Improved Iron Homeostasis of Shoots. Plant Physiol 2012, 158, 790–800. [Google Scholar]
- Meda, A.R.; Scheuermann, E.B.; Prechsl, U.E.; Erenoglu, B.; Schaaf, G.; Hayen, H.; Weber, G.; Wiren, N.V. Iron Acquisition by phytosiderophores contributes to cadmium tolerance. Plant Physiol 2007, 143, 1761–1773. [Google Scholar]
- Santos, C.S.; Silva, A.I.; Serrao, I.; Carvalho, A.L.; Vasconcelos, M.W. Transcriptomics analysis of iron deficiency related genes the legumes. Food Res. Int 2013, 54, 1162–1171. [Google Scholar]
- Vasconcelos, M.; Eckert, H.; Arahana, V.; Graef, G.; Grusak, M.A.; Clemente, T. Molecular and phenotypic characterization of transgenic soybean expressing the Arabidopsis ferric chelate reductase geneFRO2. Planta 2006, 224, 1116–1128. [Google Scholar]
- Muneer, S.; Kim, T.H.; Qureshi, M.I. Fe Modulates Cd-induced Oxidative stress and the expression of Stress Responsive Proteins in the Nodules ofVigna radiata. Plant Growth Regul 2012, 68, 421–433. [Google Scholar]
- Muneer, S.; Ahmad, J.; Qureshi, M.I. Involvement of Fe nutrition in modulating oxidative stress and the expression of stress responsive proteins in leaves of Vigna radiata L. Aust. J. Crop Sci 2013, 7, 1333–1342. [Google Scholar]
- Sanita di Toppi, L.; Gabbrielli, R. Response to cadmium in higher plants. Environ. Exp. Bot 1999, 4, 105–130. [Google Scholar]
- Halliwell, B.; Gutteridge, J.M.C. Free Radicals in Biology and Medicine, 3rd ed; Oxford University Press: New York, NY, USA, 1999. [Google Scholar]
- Siedlecka, A. Some aspects of interaction between heavy metals and plant mineral nutrients. Acta Soc. Bot. Pol 1995, 64, 265–272. [Google Scholar]
- Siedlecka, A.; Krupa, Z. Cd/Fe interaction in higher plants-its consequences for the photosynthetic apparatus. Photosynthetica 1999, 36, 321–331. [Google Scholar]
- Astolfi, S.; Zuchi, Z.; Chiani, A.; Passera, C. In vivo and in vitro effects of cadmium on H+ ATPase activity of plasmamembrane vesicles from oat (Avena sativa L.) roots. J. Plant Physiol 2003, 160, 387–393. [Google Scholar]
- Wallace, A.; Wallace, G.A.; Cha, J.W. Some modifications in trace metal toxicities and deficiencies in plants resulting from interactions with other elements and chelating agents—The special case of iron. J. Plant Nutr 1992, 15, 1589–1598. [Google Scholar]
- Hossain, Z.; Hajika, M.; Komatsu, S. Comparative proteome analysis of high and low cadmium accumulating soybeans under cadmium stress. Amino Acids 2012, 43, 2393–2416. [Google Scholar]
- Bah, A.M.; Sun, H.; Chen, F.; Zhou, J.; Dai, H.; Zhang, G.; Wu, F. Comparative proteomic analysis of Typha angustifolia leaf under chromium, cadmium and lead stress. J. Hazard. Mater 2010, 184, 191–203. [Google Scholar]
- Eide, D.; Broderius, M.; Fett, J.; Guerinot, M.L. A novel iron-regulated metal transporter from plants identified by functional expression in yeast. Proc. Natl. Acad. Sci. USA 1996, 93, 5624–5628. [Google Scholar]
- Cohen, C.K.; Fox, T.C.; Gravin, D.F.; Kochian, L.V. The role of iron deficiency stress responses in stimulating heavy metal transporter in plants. Plant Physiol 1998, 116, 1063–1072. [Google Scholar]
- Korshunova, Y.O.; Eide, D.; Clark, W.G.; Guerinot, M.L.; Pakrasi, H.B. The IRT1 protein from Arabidopsis thaliana is a metal transporter with a broad substrate range. Plant Mol. Biol 1999, 40, 37–44. [Google Scholar]
- Vert, G.; Natasha, G.; Dedaldechamp, F.; Gaymard, F.; Guerinot, M.L.; Briat, J.F.; Curie, C. IRT1, an Arabidopsis transporter essential for iron uptake from soil and for plant growth. Plant Cell 2002, 14, 1223–1233. [Google Scholar]
- Yoshihara, T.; Hodoshima, H.; Miyanao, Y.; Shoji, K.; Shimada, H.; Goto, F. Cadmium inducible iron deficiency response observed from macro and molecular views in Tobacco plants. Plant Cell Rep 2006, 25, 365–373. [Google Scholar]
- Beven, M.; Bancroft, I.; Bent, E.; Love, K.; Goodman, H.; Dean, C.; Bergkamp, R.; Dirkse, W.; van Staveren, M.; Stiekema, W.; et al. Analysis of 1.9 Mb contiguous sequence from chromosome 4 of Arabidopsis thaliana. Nature 1998, 391, 485–488. [Google Scholar]
- Hart, J.J.; Welch, R.; Norvell, W.A.; Sullivan, L.A.; Kochian, L.V. Characterization of cadmium binding, uptake, and translocation in intact seedlings of bread and durum wheat cultivars. Plant Physiol 1998, 116, 1413–1420. [Google Scholar]
- Pandey, A.; Choudhary, M.K.; Bhushan, D.; Chattopadhyay, A.; Chakraborty, S.; Datta, A.; Chakroborty, N. The Nuclear proteome of chickpea (Cicer arietinum L.) reveals predicted and un-expected proteins. J. Proteome Res 2006, 5, 3301–3311. [Google Scholar]
- Choudhary, M.K.; Basu, D.; Datta, A.; Chakraborty, N.; Chakroborty, S. Dehydration-responsive Nuclear Proteome of Rice (Oryza sativa L.) Illustrates Protein Network, Novel Regulators of Cellular Adaptation, and Evolutionary. Mol. Cell. Proteomics 2009, 8, 1579–1598. [Google Scholar]
- Luo, S.; Ishida, H.; Makino, A.; Mae, T. Fe2+-catalyzed site specific cleavage of the large subunit of ribulose 1,5-bisphosphatecarboxylase close to the active site. J. Biol. Chem 2002, 277, 12382–12387. [Google Scholar]
- Muneer, S.; Lee, B.R.; Bae, D.W.; Kim, T.H. Changes in expression of proteins involved in alleviation of Fe-deficiency by sulfur nutrition in Brassica napus L. Acta Physiol. Plant 2013, 35, 3037–3045. [Google Scholar]
- Hell, R.; Stephan, U.W. Iron uptake, trafficking and homeostasis in plants. Planta 2003, 216, 541–551. [Google Scholar]
- Schmidt, W. Iron solutions: Acquisition strategies and signaling pathways in plants. Trends Plant Sci 2003, 8, 188–193. [Google Scholar]
- Mukherjee, I.; Campbell, N.H.; Ash, J.S.; Connolly, E.L. Expression profiling of the Arabidopsis ferric chelate reductase (FRO) gene family reveals differential regulation by iron and copper. Planta 2006, 223, 1178–1190. [Google Scholar]
- Connolly, E.L.; Fett, J.P.; Guerinot, M.L. Expression of the IRT1 metal transporter is controlled by metals at the levels of transcript and protein accumulation. Plant Cell 2002, 14, 1347–1357. [Google Scholar]
- Cohen, C.K.; Gravin, D.F.; Kochian, L.V. Kinetic properties of a micronutrient transporter from Pisum sativum indicate a primary function in Fe uptake from the soil. Planta 2004, 218, 784–792. [Google Scholar]
- Shanmugam, V.; Lo, J.C.; Wu, C.L.; Wang, S.L.; Lai, C.C.; Connolly, E.L.; Huang, J.L.; Yeh, K.C. Differential expression and regulation of iron-regulated metal transporters in Arabidopsis halleri and Arabidopsis thaliana—The role in zinc tolerance. New Phytol 2011, 190, 125–137. [Google Scholar]
- Qureshi, M.I.; Muneer, S.; Bashir, H.; Ahmad, J.; Iqbal, M. Nodule physiology and proteomics of stressed legumes. Adv. Bot. Res 2010, 56, 1–38. [Google Scholar]


| Spot | Acession number | Homology | % coverage | Matched peptides | Mascot score | Mr value | Ther.pI/Exp.pI | Species |
|---|---|---|---|---|---|---|---|---|
| 1 | gi/35172183 | beta-ketoacyl-acyl carrier protein synthase III | 20 | 82 | 53 | 41,844 | 6.6/6.7 | Glycine max |
| 2 | gi/89257673 | C-terminal processing protease, putative | 14 | 70 | 39 | 54,888 | 8.9/7.0 | Brassica olereaca |
| 3 | gi/42571085 | DVL family protein | 25 | 28 | 38 | 6234 | 8.9/7.0 | Arabidopsis thaliana |
| 4 | gi/255548021 | Polygalacturonase, putative | 25 | 61 | 38 | 26,835 | 5.4/5.5 | Ricinus communis |
| 5 | gi/187473813 | Cysteine proteinase inhibitor | 72 | 44 | 39 | 6888 | 9.3/6.9 | Sonneratia griffithii |
| 6 | gi/187473851 | Cysteine proteinase inhibitor | 72 | 44 | 39 | 6830 | 9.5/7.0 | Sonneratia griffithii |
| 7 | gi/11935116 | Ethylene receptor | 57 | 63 | 60 | 12,773 | 7.5/7.0 | Populus trichocarpa |
| 8 | gi/145333373 | (S)-2-hydroxy-acid oxidase | 18 | 60 | 38 | 34,382 | 7.7/7.0 | Arabidopsis thaliana |
| 9 | gi/224112805 | AP2/ERF domain-containing transcription factor | 21 | 50 | 39 | 26,024 | 5.0/4.5 | Populus trichocarpa |
| 10 | gi/110740110 | Putative cysteinyl-tRNA synthetase | 32 | 46 | 38 | 15,724 | 7.8/6.3 | Arabidopsis thaliana |
| 11 | gi/255587821 | Small nuclear ribonucleoprotein f, putative | 30 | 28 | 49 | 9802 | 9.0/4.5 | Ricinus communis |
| 12 | gi/379068466 | Nucleotide-binding site leucine-rich repeat protein | 27 | 73 | 68 | 30,937 | 5.0/5.0 | Rhododendron formosanum |
| 13 | gi/77553861 | Retrotransposon protein | 42 | 64 | 40 | 16,417 | 8.7/6.8 | Oryza sativa |
| 14 | gi/224178035 | Magnesium chelatase | 23 | 79 | 40 | 37,402 | 4.0/4.7 | Pyramimonas parkeae |
| 15 | gi/15239904 | Calcium binding protein | 38 | 61 | 39 | 17,200 | 4.2/6.0 | Arabidopsis thaliana |
| 16 | gi/55742654 | Heat shock protein 70 | 20 | 44 | 80 | 75,480 | 5.1/5.0 | Galus gallus |
| 17 | gi/34395061 | Pathogenesis related protein1 | 39 | 61 | 88 | 16,417 | 4.6/4.5 | Oryza sativa |
| 18 | gi/105990543 | Pathogenesis related protein 2 | 30 | 48 | 55 | 16,518 | 4.8/4.5 | Zea mays |
| 19 | gi/124301261 | Glutathione S transferase omega like protein | 32 | 106 | 42 | 37,448 | 5.7/5.0 | Medicago trancatula |
| 20 | gi/125987561 | Ribosomal protein L23 | 54 | 52 | 39 | 11,192 | 10/6.5 | Sorghum bicolor |
| 21 | gi/302398837 | HD domain class transcription factor | 14 | 80 | 40 | 646,677 | 6/4.6 | Malus domestica |
| 22 | gi/89257673 | C-terminal processing protease, putative | 14 | 88 | 39 | 54,888 | 8.9/7.0 | Brassica olerecea |
| 23 | gi/357460053 | (+) neomenthol dehydrogenase | 23 | 70 | 66 | 32,472 | 6.3/4.2 | Medicago trancatula |
© 2014 by the authors; licensee MDPI, Basel, Switzerland This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Muneer, S.; Hakeem, K.R.; Mohamed, R.; Lee, J.H. Cadmium Toxicity Induced Alterations in the Root Proteome of Green Gram in Contrasting Response towards Iron Supplement. Int. J. Mol. Sci. 2014, 15, 6343-6355. https://doi.org/10.3390/ijms15046343
Muneer S, Hakeem KR, Mohamed R, Lee JH. Cadmium Toxicity Induced Alterations in the Root Proteome of Green Gram in Contrasting Response towards Iron Supplement. International Journal of Molecular Sciences. 2014; 15(4):6343-6355. https://doi.org/10.3390/ijms15046343
Chicago/Turabian StyleMuneer, Sowbiya, Khalid Rehman Hakeem, Rozi Mohamed, and Jeong Hyun Lee. 2014. "Cadmium Toxicity Induced Alterations in the Root Proteome of Green Gram in Contrasting Response towards Iron Supplement" International Journal of Molecular Sciences 15, no. 4: 6343-6355. https://doi.org/10.3390/ijms15046343





